Abstract
Cells often use multiple pathways to repair the same DNA lesion, and the choice of pathway has substantial implications for the fidelity of genome maintenance. DNA interstrand crosslinks covalently link the two strands of DNA, and thereby block replication and transcription; the cytotoxicity of these crosslinks is exploited for chemotherapy. In Xenopus egg extracts, the collision of replication forks with interstrand crosslinks initiates two distinct repair pathways. NEIL3 glycosylase can cleave the crosslink1; however, if this fails, Fanconi anaemia proteins incise the phosphodiester backbone that surrounds the interstrand crosslink, generating a double-strand-break intermediate that is repaired by homologous recombination2. It is not known how the simpler NEIL3 pathway is prioritized over the Fanconi anaemia pathway, which can cause genomic rearrangements. Here we show that the E3 ubiquitin ligase TRAIP is required for both pathways. When two replisomes converge at an interstrand crosslink, TRAIP ubiquitylates the replicative DNA helicase CMG (the complex of CDC45, MCM2–7 and GINS). Short ubiquitin chains recruit NEIL3 through direct binding, whereas longer chains are required for the unloading of CMG by the p97 ATPase, which enables the Fanconi anaemia pathway. Thus, TRAIP controls the choice between the two known pathways of replication-coupled interstrand-crosslink repair. These results, together with our other recent findings3,4 establish TRAIP as a master regulator of CMG unloading and the response of the replisome to obstacles.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All relevant data are available from the authors and/or are included with this Letter. Source images are available in Supplementary Fig. 1.
References
Semlow, D. R., Zhang, J., Budzowska, M., Drohat, A. C. & Walter, J. C. Replication-dependent unhooking of DNA interstrand cross-links by the NEIL3 glycosylase. Cell 167, 498–511.e14 (2016).
Räschle, M. et al. Mechanism of replication-coupled DNA interstrand crosslink repair. Cell 134, 969–980 (2008).
Larsen, N. B. et al. Mechanism of replication-coupled DNA-protein crosslink proteolysis by SPRTN and the proteasome. Mol. Cell 73, 574–588 (2019).
Deng, L. et al. Mitotic CDK promotes replisome disassembly, fork breakage, and complex DNA rearrangements. Mol. Cell (in the press).
Price, N. E., Catalano, M. J., Liu, S., Wang, Y. & Gates, K. S. Chemical and structural characterization of interstrand cross-links formed between abasic sites and adenine residues in duplex DNA. Nucleic Acids Res. 43, 3434–3441 (2015).
Kottemann, M. C. & Smogorzewska, A. Fanconi anaemia and the repair of Watson and Crick DNA crosslinks. Nature 493, 356–363 (2013).
Zhang, J. et al. DNA interstrand cross-link repair requires replication-fork convergence. Nat. Struct. Mol. Biol. 22, 242–247 (2015).
Fullbright, G., Rycenga, H. B., Gruber, J. D. & Long, D. T. p97 promotes a conserved mechanism of helicase unloading during DNA cross-link repair. Mol. Cell. Biol. 36, 2983–2994 (2016).
Amunugama, R. et al. Replication fork reversal during DNA interstrand crosslink repair requires CMG unloading. Cell Rep.s 23, 3419–3428 (2018).
Knipscheer, P. et al. The Fanconi anemia pathway promotes replication-dependent DNA interstrand cross-link repair. Science 326, 1698–1701 (2009).
Massaad, M. J. et al. Deficiency of base excision repair enzyme NEIL3 drives increased predisposition to autoimmunity. J. Clin. Invest. 126, 4219–4236 (2016).
Parmar, K., D’Andrea, A. & Niedernhofer, L. J. Mouse models of Fanconi anemia. Mutat. Res. 668, 133–140 (2009).
Sejersted, Y. et al. Endonuclease VIII-like 3 (Neil3) DNA glycosylase promotes neurogenesis induced by hypoxia-ischemia. Proc. Natl Acad. Sci. USA 108, 18802–18807 (2011).
Torisu, K., Tsuchimoto, D., Ohnishi, Y. & Nakabeppu, Y. Hematopoietic tissue-specific expression of mouse Neil3 for endonuclease VIII-like protein. J. Biochem. 138, 763–772 (2005).
Chapard, C., Hohl, D. & Huber, M. The role of the TRAF-interacting protein in proliferation and differentiation. Exp. Dermatol. 21, 321–326 (2012).
Harley, M. E. et al. TRAIP promotes DNA damage response during genome replication and is mutated in primordial dwarfism. Nat. Genet. 48, 36–43 (2016).
Räschle, M. et al. Proteomics reveals dynamic assembly of repair complexes during bypass of DNA cross-links. Science 348, 1253671 (2015).
Hoffmann, S. et al. TRAIP is a PCNA-binding ubiquitin ligase that protects genome stability after replication stress. J. Cell Biol. 212, 63–75 (2016).
Huang, J. et al. The DNA translocase FANCM/MHF promotes replication traverse of DNA interstrand crosslinks. Mol. Cell 52, 434–446 (2013).
Mutreja, K. et al. ATR-mediated global fork slowing and reversal assist fork traverse and prevent chromosomal breakage at DNA interstrand cross-links. Cell Rep. 24, 2629–2642 (2018).
Long, D. T., Joukov, V., Budzowska, M. & Walter, J. C. BRCA1 promotes unloading of the CMG helicase from a stalled DNA replication fork. Mol. Cell 56, 174–185 (2014).
Priego Moreno, S., Bailey, R., Campion, N., Herron, S. & Gambus, A. Polyubiquitylation drives replisome disassembly at the termination of DNA replication. Science 346, 477–481 (2014).
Dewar, J. M., Low, E., Mann, M., Räschle, M. & Walter, J. C. CRL2Lrr1 promotes unloading of the vertebrate replisome from chromatin during replication termination. Genes Dev. 31, 275–290 (2017).
Liu, M. et al. Expression and purification of active mouse and human NEIL3 proteins. Protein Expr. Purif. 84, 130–139 (2012).
Wang, B. et al. Structure and ubiquitin interactions of the conserved zinc finger domain of Npl4. J. Biol. Chem. 278, 20225–20234 (2003).
Wallace, B. D. et al. APE2 Zf-GRF facilitates 3′-5′ resection of DNA damage following oxidative stress. Proc. Natl Acad. Sci. USA 114, 304–309 (2017).
Dutrillaux, B., Aurias, A., Dutrillaux, A. M., Buriot, D. & Prieur, M. The cell cycle of lymphocytes in Fanconi anemia. Hum. Genet. 62, 327–332 (1982).
Sparks, J. L. et al. The CMG helicase bypasses DNA-protein cross-links to facilitate their repair. Cell 176, 167–181.e21 (2019).
Lebofsky, R., Takahashi, T. & Walter, J. C. DNA replication in nucleus-free Xenopus egg extracts. Methods Mol. Biol. 521, 229–252 (2009).
Dewar, J. M., Budzowska, M. & Walter, J. C. The mechanism of DNA replication termination in vertebrates. Nature 525, 345–350 (2015).
Joukov, V., Chen, J., Fox, E. A., Green, J. B. & Livingston, D. M. Functional communication between endogenous BRCA1 and its partner, BARD1, during Xenopus laevis development. Proc. Natl Acad. Sci. USA 98, 12078–12083 (2001).
Walter, J. & Newport, J. Initiation of eukaryotic DNA replication: origin unwinding and sequential chromatin association of Cdc45, RPA, and DNA polymerase α. Mol. Cell 5, 617–627 (2000).
Byun, T. S., Pacek, M., Yee, M. C., Walter, J. C. & Cimprich, K. A. Functional uncoupling of MCM helicase and DNA polymerase activities activates the ATR-dependent checkpoint. Genes Dev. 19, 1040–1052 (2005).
Wohlschlegel, J. A., Dhar, S. K., Prokhorova, T. A., Dutta, A. & Walter, J. C. Xenopus Mcm10 binds to origins of DNA replication after Mcm2-7 and stimulates origin binding of Cdc45. Mol. Cell 9, 233–240 (2002).
Pacek, M., Tutter, A. V., Kubota, Y., Takisawa, H. & Walter, J. C. Localization of MCM2-7, Cdc45, and GINS to the site of DNA unwinding during eukaryotic DNA replication. Mol. Cell 21, 581–587 (2006).
Kochaniak, A. B. et al. Proliferating cell nuclear antigen uses two distinct modes to move along DNA. J. Biol. Chem. 284, 17700–17710 (2009).
Rosado, I. V., Langevin, F., Crossan, G. P., Takata, M. & Patel, K. J. Formaldehyde catabolism is essential in cells deficient for the Fanconi anemia DNA-repair pathway. Nat. Struct. Mol. Biol. 18, 1432–1434 (2011).
Budzowska, M., Graham, T. G., Sobeck, A., Waga, S. & Walter, J. C. Regulation of the Rev1-pol ζ complex during bypass of a DNA interstrand cross-link. EMBO J. 34, 1971–1985 (2015).
Michel, M. A. et al. Assembly and specific recognition of K29- and K33-linked polyubiquitin. Mol. Cell 58, 95–109 (2015).
Knipscheer, P., Räschle, M., Schärer, O. D. & Walter, J. C. Replication-coupled DNA interstrand cross-link repair in Xenopus egg extracts. Methods Mol. Biol. 920, 221–243 (2012).
Graham, T. G., Walter, J. C. & Loparo, J. J. Two-stage synapsis of DNA ends during non-homologous end joining. Mol. Cell 61, 850–858 (2016).
Hemsley, A., Arnheim, N., Toney, M. D., Cortopassi, G. & Galas, D. J. A simple method for site-directed mutagenesis using the polymerase chain reaction. Nucleic Acids Res. 17, 6545–6551 (1989).
Trowitzsch, S., Bieniossek, C., Nie, Y., Garzoni, F. & Berger, I. New baculovirus expression tools for recombinant protein complex production. J. Struct. Biol. 172, 45–54 (2010).
Ilves, I., Petojevic, T., Pesavento, J. J. & Botchan, M. R. Activation of the MCM2-7 helicase by association with Cdc45 and GINS proteins. Mol. Cell 37, 247–258 (2010).
Liu, M. et al. The mouse ortholog of NEIL3 is a functional DNA glycosylase in vitro and in vivo. Proc. Natl Acad. Sci. USA 107, 4925–4930 (2010).
Franken, N. A., Rodermond, H. M., Stap, J., Haveman, J. & van Bree, C. Clonogenic assay of cells in vitro. Nat. Protoc. 1, 2315–2319 (2006).
Vos, J. M. & Hanawalt, P. C. Processing of psoralen adducts in an active human gene: repair and replication of DNA containing monoadducts and interstrand cross-links. Cell 50, 789–799 (1987).
Derheimer, F. A., Hicks, J. K., Paulsen, M. T., Canman, C. E. & Ljungman, M. Psoralen-induced DNA interstrand cross-links block transcription and induce p53 in an ataxia-telangiectasia and Rad3-related-dependent manner. Mol. Pharmacol. 75, 599–607 (2009).
Walter, J., Sun, L. & Newport, J. Regulated chromosomal DNA replication in the absence of a nucleus. Mol. Cell 1, 519–529 (1998).
Klein Douwel, D. et al. XPF-ERCC1 acts in unhooking DNA interstrand crosslinks in cooperation with FANCD2 and FANCP/SLX4. Mol. Cell 54, 460–471 (2014).
Long, D. T., Räschle, M., Joukov, V. & Walter, J. C. Mechanism of RAD51-dependent DNA interstrand cross-link repair. Science 333, 84–87 (2011).
Fu, Y. V. et al. Selective bypass of a lagging strand roadblock by the eukaryotic replicative DNA helicase. Cell 146, 931–941 (2011).
Sonneville, R. et al. CUL-2LRR-1 and UBXN-3 drive replisome disassembly during DNA replication termination and mitosis. Nat. Cell Biol. 19, 468–479 (2017).
Acknowledgements
We thank D. Pellman and members of the Walter laboratory for comments on the manuscript, and K. Arnett for help with biolayer interferometry experiments. J.C.W. is supported by NIH grant HL098316 and a gift from the family of Jonathan G. Wiseman. R.A.W. is supported by American Cancer Society postdoctoral fellowship 131415-PF-17-168-01-DMC, D.R.S. by NIH award K99GM129422, D.R.S. and G.C. by Jane Coffin Childs postdoctoral fellowships, J.L.S. by a Damon Runyon postdoctoral fellowship, M.W. by the Cancer Research UK Clinician Scientist Fellowship, and E.L. by NIH award F31GM122277. J.C.W. is a Howard Hughes Medical Institute Investigator and an American Cancer Society Research Professor.
Reviewer information
Nature thanks Daniel Durocher, Michael Seidman and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
R.A.W. identified TRAIP as the E3 ligase that ubiquitylates CMG and characterized the role of TRAIP in cisplatin-ICL repair. D.R.S. characterized the role of CMG ubiquitylation in AP-ICL repair and performed the structure–function analysis of NEIL3. A.N.K.-L., M.R.H. and M.W. generated Fig. 4d and Extended Data Fig. 9 under the supervision of K.J.P. O.V.K. generated Fig. 1c, d and Extended Data Figs. 2i and 6a, b. G.C. and E.L. prepared Xenopus laevis rCMG. R.A. generated Fig. 1e and Extended Data Fig. 3c, d. J.L.S. generated Fig. 2e and Extended Data Fig. 4c. L.D. generated Extended Data Fig. 10b, c. C.A.M. helped to generate Extended Data Fig. 8b, d, e. J.C.W., R.A.W. and D.R.S. designed experiments, analysed the data and wrote the paper with input from the other authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 DNA replication and ICL repair in Xenopus egg extracts.
a, Schematic of pICL replication in the nucleus-free Xenopus egg extract system49. Incubation of the plasmid in high-speed supernatant supports the recruitment of inactive MCM2–MCM7 double hexamers (red hexamers; ‘Licensing’). Addition of NPE activates replication initiation, including the assembly of active CMG helicases (green hexamers), and elongation of nascent strands (red lines) leads to convergence of forks at the ICL. b, Intermediates generated during replication-coupled repair of a cisplatin-ICL. Top, progression through the incision-dependent Fanconi anaemia repair pathway generates distinct intermediates resulting from fork convergence, CMG unloading, leading strand approach to the ICL, fork reversal, incisions, and repair of the double strand break by homologous recombination. Bottom, deproteinization of the DNA intermediates depicted along the top yields DNA structures that travel with characteristic mobilities during native agarose gel electrophoresis. These structures are indicated along the side of the gel and with coloured dots and/or arrows in Fig. 1b and other figures. The slow-figure-8 species arise upon fork convergence on the ICL (Fig. 1b, purple dot). Conversion of slow- to fast-figure-8 species results from CMG unloading and an accompanying change in plasmid topology9 (Fig. 1b, orange dot). Next, a species of intermediate mobility appears (Fig. 1b, green arrow), which represents reversed forks, as shown by electron microscopy9. After XPF-dependent unhooking of the reversed structure9,50, double-strand DNA break repair generates joining products that barely enter the gel51 (Fig. 1b, well product). Some of these species are resolved into monomeric, supercoiled plasmids that represent the final, fully repaired product (Fig. 1b, SC) that is sensitive to SapI digestion. c, Intermediates generated during replication-coupled repair of an AP-ICL (pICLAP). Top, progression through the NEIL3 repair pathway generates intermediates resulting from fork convergence, NEIL3-dependent N-glycosyl bond cleavage, nascent strand ligation, decatenation, and TLS. Bottom, deproteinization of the DNA intermediates depicted along the top yields DNA structures that travel with characteristic mobilities during native gel electrophoresis. These structures are indicated along the side of the gel and with coloured dots in Fig. 3a and other figures.
Extended Data Fig. 2 Recombinant TRAIP supports CMG unloading at cisplatin-ICLs.
a, NPE immunodepleted of TRAIP was loaded alongside a dilution series of mock-depleted NPE and analysed for TRAIP. A relative loading amount of 100 corresponds to 2 µl of NPE. Non-specifically detected proteins are marked with asterisks. Four independent experiments. b, Bacterially expressed wild-type rTRAIP, rTRAIP(R18C), His6-SUMO-rTRAIP(wild type), and His6-SUMO-rTRAIP(ΔPIP) (comprising residues 1–455) were partially purified, resolved by SDS–PAGE and visualized with Coomassie blue staining. Note that His6-SUMO-rTRAIP is obscured by co-migrating, contaminating proteins. Bacterially expressed rTRAIP was used for all subsequent experiments, unless otherwise indicated. Results are typical of at least two independent purifications. c, Wild-type rTRAIP and rTRAIP(R18C) were expressed in Sf9 insect cells and purified using an N-terminal 3×Flag tag. The tag was then cleaved using 3C protease. The recombinant proteins, along with a buffer sample containing 3C protease only, were resolved by SDS–PAGE and visualized with Coomassie blue staining. Results are typical of at least three independent purifications. d, Mock-depleted and TRAIP-depleted extracts supplemented with wild-type rTRAIP or rTRAIP(R18C) used in Fig. 1b were analysed by immunoblotting for TRAIP. The absence of the non-specific bands seen in a may be due to shorter incubation with the TRAIP antibody. The concentration of added recombinant TRAIP relative to endogenous TRAIP fluctuates among experiments (for example, compare d and Extended Data Fig. 3b). We ascribe this difference to variations in non-specific removal of endogenous TRAIP from extracts during the mock-depletion procedure, and possibly also in the delivery of recombinant TRAIP into the extract. Seven independent experiments. e, pICLPt was replicated in mock-depleted or TRAIP-depleted extracts supplemented with [α-32P]dATP and Sf9-expressed wild-type rTRAIP, rTRAIP(R18C) or 3C protease alone and analysed as in Fig. 1b. Two independent experiments. f, Extracts used in the replication reaction shown in e were analysed as in d. Two independent experiments. g, pICLPt was replicated in the indicated egg extracts with [α-32P]dATP and analysed as in Fig. 1b. Two independent experiments. h, Extracts used in the replication reaction shown in g were analysed as in d. Note that deleting the C-terminal PIP box disrupts the epitope for the TRAIP antibody used for immunoblotting. Therefore, to assess the activity of TRAIP(ΔPIP) in ICL repair relative to wild-type TRAIP, His6-SUMO-tagged proteins were added back to TRAIP-depleted extract and assayed in g. The relative amounts of His6-SUMO-TRAIP(wild type) and His6-SUMO-TRAIP(ΔPIP) were compared by detecting the histidine tag. By blotting the same extracts for TRAIP, a comparison of the relative concentrations of His6-SUMO-TRAIP(ΔPIP) and endogenous TRAIP was made. Two independent experiments. i, Mock-depleted and TRAIP-depleted extracts used in the replication reactions shown in Fig. 1c, d were analysed as in d. Three independent experiments.
Extended Data Fig. 3 TRAIP, but not FANCM, ATR or BRCA1, is required for CMG unloading at cisplatin-ICLs.
a, Left, schematic of nascent strands generated at ICLs. When forks converge on an ICL, nascent strands stall about 20 nucleotides from the ICL on either side of the lesion owing to the footprint of CMG (green hexamer). AflIII cuts 144 nucleotides to the left and 534 nucleotides to the right of the ICL, generating characteristic products for the leftward and rightward leading strands upon fork convergence, CMG unloading, and leading strand extension. Right, nascent strand analysis of pICLPt replication in the indicated extracts. After replication with [α-32P]dATP, nascent strands were extracted, digested with AflIII and resolved on a denaturing polyacrylamide gel alongside a sequencing ladder and visualized by autoradiography. As seen previously2,52, when replication forks converged on the ICL in mock-depleted egg extracts, leading strands initially stalled 20–40 nucleotides from the lesion (lane 1) and then advanced to the −1 position (lanes 2–6), which depends on CMG dissociation21,52. By contrast, in TRAIP-depleted egg extracts, the −20 footprint persisted for three hours (lanes 7–12). This effect was rescued with wild-type rTRAIP but not rTRAIP(R18C) (lanes 13–24). Two independent experiments. b, Extracts used in the replication reaction shown in a were analysed for TRAIP. Two independent experiments. c, Mock-depleted or FANCM-depleted extracts were analysed for FANCM. A non-specifically detected protein is marked with an asterisk. Two independent experiments. d, Nascent strand analysis of pICLPt replicating in mock-depleted or FANCM-depleted extracts was performed as in a. The CMG footprint disappeared at the ICL in FANCM-depleted egg extract, consistent with FANCM not being required for CMG unloading at ICLs. Two independent experiments. e, pICLPt was replicated in the absence or presence of ATR inhibitor ETP-46464 (ATRi), and nascent strand analysis was performed as in a. ATR inhibitor was added to the reaction 2.5 min after initiation. The CMG footprint disappeared at the ICL with or without ATR inhibitor, indicating that ATR signalling is not required for CMG unloading at ICLs. Two independent experiments. f, Extracts used in e were sampled at various time points and analysed for Xenopus CHK1 serine-344 phosphorylation to verify ATR inhibition. MCM6 was detected as a loading control. Two independent experiments. g, h, Mock-depleted and TRAIP-depleted extracts used in one of the replicate reactions quantified in Fig. 1e (g) and Fig. 1f (h) were analysed for TRAIP. Two independent experiments were performed for g and three independent experiments were performed for h. i, We previously showed that the immunodepletion of BRCA1 from egg extracts inhibits CMG unloading at ICLs, but this defect could not be rescued with recombinant BRCA1–BARD1 complex8,21. To test whether TRAIP is co-depleted with BRCA1, NPE was immunodepleted of BRCA1 with BRCA1 antiserum, loaded alongside a dilution series of mock-depleted NPE, and analysed for BRCA1 and TRAIP. A relative loading amount of 100 corresponds to 2 µl of NPE. Non-specifically detected proteins are marked with asterisks. This analysis revealed that immunodepletion of BRCA1 also removed TRAIP from NPE. We also observed TRAIP co-depletion with antibodies against other proteins (data not shown), suggesting that it interacts non-specifically with different antibodies. Two independent experiments. j, The extracts described in i were supplemented with pICLPt, [α-32P]dATP and rTRAIP, as indicated, and analysed as in Fig. 1b. Wild-type rTRAIP suppressed the stabilization of the slow-figure-8 species seen in BRCA1-depleted extract, consistent with the restoration of CMG unloading, and indicating that the unloading defect seen in BRCA1-depleted egg extracts is due primarily to the removal of TRAIP from the extract. Two independent experiments. k, The extracts used in j were analysed for TRAIP. Two independent experiments. l, To determine whether BRCA1 contributes to TRAIP-dependent CMG unloading, NPE was immunodepleted of TRAIP or TRAIP and BRCA1 using Protein A Sepharose-purified antibodies purified from antiserum. A dilution series of mock-depleted NPE was loaded alongside the depleted extracts, and extracts were analysed for BRCA1 and TRAIP. A relative loading amount of 100 corresponds to 2 µl of NPE. Two independent experiments. m, The extracts described in l were supplemented with pICLPt, [α-32P]dATP and rTRAIP, as indicated, and analysed as in Fig. 1b. Wild-type rTRAIP suppressed the accumulation of slow-figure-8 species to a similar extent in TRAIP-depleted egg extracts whether or not BRCA1 was co-depleted (lanes 19–30), indicating that BRCA1 is not needed to support TRAIP function. BRCA1 depletion reproducibly resulted in a decrease in well-product formation, suggesting a role for BRCA1 in recombination after a double-strand break is formed by ICL-unhooking incisions. Two independent experiments. n, The extracts used in m were analysed for TRAIP. Two independent experiments.
Extended Data Fig. 4 TRAIP and CRL2LRR1 promote distinct CMG unloading pathways.
a, pCtrl was replicated in NPE in the presence of Geminin or the CDC7 inhibitor PHA-767491 (CDC7i). Eight minutes after the addition of NPE, the plasmid was recovered and the indicated proteins were analysed by immunoblot. Three independent experiments. b, Cartoon depicting the effects of Geminin and CDC7i, and the step at which TRAIP is recruited to chromatin. c, Extracts used in the replication reaction shown in Fig. 2e were analysed for TRAIP. Four independent experiments. d, Top, upon termination of pCtrl replication, CMG (green) unloading depends on CRL2LRR1-mediated MCM7 ubiquitylation (purple)23,53, but it is not known whether unloading also requires TRAIP. Middle, CMGs that have converged at an ICL undergo TRAIP-dependent ubiquitylation (Fig. 1), but the involvement of CRL2LRR1 is unknown. Bottom, if two origins fire on a single plasmid, one pair of replication forks converges at the ICL and undergoes TRAIP-dependent CMG unloading, whereas a second pair undergoes CRL2LRR1-dependent unloading. Because both pairs of CMGs should undergo ubiquitylation, in some experiments we include Culi to monitor only TRAIP-dependent ubiquitylation. e, To determine whether TRAIP is required for CMG unloading during replication termination, we analysed proteins associated with plasmids 60 min after replication initiation in mock-depleted or TRAIP-depleted extracts containing p97i or Culi, as indicated. Chromatin was recovered and analysed for the indicated proteins. CMG unloading from pCtrl was unaffected by TRAIP depletion (compare lanes 4 and 7). Consistent with this, in the presence of p97i, TRAIP was not required for MCM7 ubiquitylation on pCtrl (compare lanes 2 and 5). By contrast, in the absence of TRAIP, CMG unloading from pICLPt was inhibited compared to the mock-depleted control (compare lanes 10 and 13), consistent with Fig. 1c. Similarly, TRAIP was essential for efficient MCM7 ubiquitylation on pICLPt (compare lanes 8 and 11, note the greater level of unmodified MCM7 in lane 11). The partial CMG unloading (lane 13) and residual MCM7 ubiquitylation observed on pICLPt in the absence of TRAIP (lane 11) were probably the result of termination events that occurred elsewhere on the plasmid (as described in d, bottom). Consistent with this interpretation, the combination of TRAIP depletion and Culi abolished MCM7 ubiquitylation (lane 12). Three independent experiments. f, To determine whether CRL2LRR1 contributes to CMG unloading at ICLs, pICLPt was replicated in undepleted extract containing p97i or Culi and analysed as in Fig. 1b. Culi had no effect on the accumulation of fast-figure-8 structures, consistent with CRL2LRR1 being dispensable for CMG unloading at ICLs. Two independent experiments. g, Left, to assess the effect of LRR1 depletion on CMG unloading at ICLs, NPE was immunodepleted of LRR1, loaded alongside a dilution series of mock-depleted NPE, and analysed for LRR1 and CUL2. A relative loading amount of 100 corresponds to 2 µl of NPE. Non-specifically detected protein is marked with an asterisk. Right, pICLPt was replicated in mock-depleted or LRR1-depleted egg extracts and analysed as in Fig. 1b. The absence of LRR1 had no effect on the formation of fast-figure-8 structures, supporting the idea that CRL2LRR1 is dispensable for CMG unloading at ICLs. Two independent experiments. h, Nascent strand analysis of pICLPt replicating in mock-depleted or LRR1-depleted extracts was performed as in Extended Data Fig. 3a. The CMG footprint disappeared with normal kinetics at the ICL in LRR1-depleted egg extract, consistent with CRL2LRR1 not being required for CMG unloading at ICLs. Two independent experiments.
Extended Data Fig. 5 TRAIP ubiquitylates numerous CMG subunits with heterotypically linked chains upon fork convergence at an ICL.
a, pICL-lacOPt was incubated with LacR before replication in mock- or TRAIP-depleted extracts. After 1 h of replication (to allow for termination of replication forks that do not converge on the ICL or lacO array as depicted in Figs. 2c), 0.3 mM NMS-873 was added and the extracts were incubated for 5 min to inhibit p97. IPTG (10 mM) and Sf9-expressed wild-type rTRAIP were then added as indicated and the extract was incubated for 2 h to disrupt LacR DNA binding and allow fork convergence and TRAIP-dependent ubiquitylation. The plasmid was recovered and analysed for the indicated proteins. Two independent experiments. b, The extracts described in a were analysed for TRAIP. Two independent experiments. c, pICL-lacOPt was incubated with LacR before replication in undepleted egg extracts. After 30 min of replication (to allow for termination of replication forks that do not converge on the ICL or lacO array as depicted in Figs. 2c), 0.3 mM NMS-873 was added and the extracts were incubated for 5 min to inhibit p97. IPTG (10 mM) was then added as indicated to disrupt LacR DNA binding and the extract was incubated for 1 h to allow for fork convergence. The plasmid was recovered and analysed for the indicated proteins. Two independent experiments. d, Recombinant CMG was purified, resolved by SDS–PAGE, and visualized with SYPRO Ruby staining. Results are typical of at least five independent purifications. e, rTRAIP ubiquitin ligase activity. Wild-type rTRAIP or rTRAIP(R18C) was combined with ubiquitin, E1 ligase, three E2 ligases (UbcH5a, UbcH5b, UbcH5c) and ATP as indicated. Polyubiquitin chain synthesis (top) and TRAIP autoubiquitylation (bottom) were detected by immunoblotting the reactions with ubiquitin and TRAIP antibody, respectively. rTRAIP(R18C) was much more compromised in forming free polyubiquitin chains in this assay than it was in ubiquitylating rCMG (see Fig. 2d). The data suggest that the interaction between TRAIP and CMG can suppress, to a great extent, the profound ubiquitylation defect of the R18C mutation. Three independent experiments. f, pCtrl-lacO and pICL-lacOPt were replicated in undepleted extract as in c and recovered. Samples were treated with the indicated DUBs and analysed for the indicated proteins. Three independent experiments. g, pCtrl-lacO and pICL-lacOPt pre-bound with LacR were replicated in undepleted extract as in c. At the time of IPTG addition to release the LacR array and allow for fork convergence, 100 µM recombinant ubiquitin (wild-type or various lysine-to-arginine mutants) was added to the extract (which contains around 8 µM endogenous ubiquitin) and incubated for 1 h. The plasmid was recovered and analysed for the indicated proteins. Three independent experiments. h, Extracts used in g were analysed for ubiquitin. Some ubiquitin mutants contain a di-ubiquitin species (marked with an asterisk). Whether this arises upon addition to the extract is unclear. Three independent experiments.
Extended Data Fig. 6 AP-ICL repair by NEIL3 requires TRAIP.
a, Analysis of chromatin-associated proteins during replication of pICL-lacOPt or pICL-lacOAP in undepleted extract in the presence and absence of Geminin, as indicated. At different times after replication initiation, chromatin was recovered and analysed for the indicated proteins. Two independent experiments. b, pICL-lacOPt and pICL-lacOAP were replicated in undepleted extract supplemented with Culi, Geminin and p97i, as indicated, and analysed as in Fig. 1b. Two independent experiments. c, The extracts used in the replication reaction shown in Fig. 3a were analysed for TRAIP. Five independent experiments. d, Extracts used in one of the reactions quantified in Fig. 3b were analysed as in c. Three independent experiments. e, f, pICLAP was replicated in the extracts shown in Extended Data Fig. 2h (e) and Extended Data Fig. 2f (f) with [α-32P]dATP and analysed as in Fig. 1b. Two independent experiments were performed for e and f.
Extended Data Fig. 7 ICL repair by NEIL3 requires the association of CMG with chromatin.
a, Analysis of proteins associated with pICLAP during replication with NEIL3-depleted extract supplemented with wild-type rNEIL3 or catalytically inactive rNEIL3(K60A), p97i and Culi. At the indicated times after replication initiation, chromatin was recovered and analysed for the indicated proteins. Consistent with NEIL3 dissociating rapidly after unhooking, rNEIL3(K60A) was recovered more efficiently with pICLAP than was wild-type rNEIL3. Two independent experiments. b, pICLAP was replicated with NEIL3-depleted extract supplemented with wild-type rNEIL3 or rNEIL3(K60A), p97i and Culi, as indicated, and analysed as in Fig. 1b. Two independent experiments. c, The extracts used in Fig. 3c were analysed for TRAIP. Non-specifically detected proteins are marked with asterisks. Four independent experiments. d, If NEIL3 activity is coupled to ubiquitylated CMG, NEIL3 should function only before CMG has been unloaded. To test this prediction, we inhibited all unhooking events by depleting egg extracts of NEIL3 (to block the NEIL3 pathway) and FANCD2 (to block the Fanconi anaemia pathway). At a late time point (90 min), we added back rNEIL3 to extract in which CMG had been allowed to unload (−p97i), or extract in which CMG unloading was prevented (+p97i). Our model predicts that rNEIL3 should unhook the ICL only in the latter setting. Top, schematic illustrating the late addition of rNEIL3 to NEIL3-depleted and FANCD2-depleted egg extracts in the absence (left) or presence (right) of p97i. Bottom, replication of pICLAP in mock-depleted, NEIL3-depleted, or both NEIL3- and FANCD2-depleted extracts in the presence of [α-32P]dATP. Extracts were supplemented with p97i as indicated and rNEIL3 was added at 90 min where indicated (black arrowheads). Replication intermediates were resolved and visualized as in Fig. 1b. Depletion of NEIL3 and FANCD2 blocked all unhooking of the AP-ICL, resulting in an accumulation of reversed forks (lane 9, green arrowhead). Addition of rNEIL3 at 90 min in the absence of p97i (after CMG unloading) failed to induce unhooking, based on the persistence of the reversed forks (lanes 21–24). By contrast, when CMG unloading was prevented with p97i (lanes 25–30; note the persistence of slow-figure-8 intermediates), late rNEIL3 addition led to efficient ICL unhooking, as seen from the rapid conversion of slow-figure-8 intermediates to open circular and supercoiled species (lanes 34–36). This gel image has been compressed vertically to fit the page. Two independent experiments. e, To confirm the presence or absence of CMG at the AP-ICL, DNA was recovered from the reactions described in d and subjected to nascent strand analysis as in Extended Data Fig. 3a. Top, extension products and nascent strands of the leftward fork. Bottom, nascent strands of the rightward fork. Black arrowheads, rNEIL3 addition. Depletion of NEIL3 and FANCD2 did not affect loss of the CMG footprint at −20 and caused persistence of nascent DNA strands at −1 (lanes 13–24), which is indicative of failure to unhook the ICL. Late addition of NEIL3 failed to stimulate further nascent strand extension (lanes 21–24), indicating that unhooking did not occur. Treatment with p97i led to persistence of the CMG footprint at −20 (lanes 25–30), consistent with retention of CMG at the ICL, and late addition of NEIL3 stimulated formation of full-length nascent strand extension products (lanes 34–36), indicative of efficient unhooking. Taken together, the data in d and e strongly suggest that NEIL3 activity is coupled to the presence of CMG at the site of the ICL, although we cannot rule out that NEIL3 activity is suppressed by downstream events, such as fork reversal, that depend on CMG unloading. Two independent experiments.
Extended Data Fig. 8 The zinc-finger domains of NEIL3 contribute to its recruitment to the replication fork.
a, Left, to determine whether rNEIL3(∆291) is catalytically active, a model AP-ICL substrate comprising a synthetic 5′-radiolabelled 24mer oligonucleotide crosslinked to a ~3mer was mixed with rNEIL3(∆291) or wild-type rNEIL3. Crosslinked and unhooked species were resolved by denaturing polyacrylamide gel electrophoresis and visualized by autoradiography. Asterisks indicate the 32P radiolabel. Note that the crosslinked species migrates as a doublet owing to heterogeneity in the bottom strand after RNase digestion (see Methods for details). Right, quantification of unhooking. Equivalent results were obtained in three independent experiments, which show that rNEIL3(∆291) retains full glycosylase activity. b, Interaction of the NEIL3 NPL4-type zinc-finger (residues 300–328) with ubiquitin. GST-NEIL3 NZF fusion protein (wild-type or T310L/L311V-substituted) was immobilized on a biosensor tip and monoubiquitin binding was measured by biolayer interferometry. The ubiquitin binding response was corrected for non-specific binding to GST and plotted as a function of ubiquitin concentration. Data are mean ± s.e.m. from three independent experiments. c, Extracts used in Fig. 3e were analysed for NEIL3. Black arrowheads, NEIL3-specific bands. rNEIL3∆291 is not efficiently detected by the NEIL3-specific primary antibody. Non-specifically detected proteins are marked with asterisks. Three independent experiments. d, e, To test whether the two GRF zinc-fingers in NEIL3 interact with ssDNA, we expressed each individually and performed electrophoretic mobility shift assays. rMBP-NEIL3 GRF zinc-finger fusion proteins (wild-type or substituted) were incubated with 5′-radiolabelled 25-mer ssDNA or dsDNA. Bound and unbound DNAs were resolved by native PAGE and visualized by autoradiography. This analysis reveals that both GRF domains bind specifically to ssDNA. Two independent experiments were performed for d and e. f, Analysis of proteins associated with pICLPt during replication in undepleted extract in the presence of p97i and Culi. Extracts were supplemented with wild-type or mutated rNEIL3. At different times, chromatin was recovered and analysed for the indicated proteins. The individual GRF substitutions modestly affected recovery of rNEIL3 upon pICL pull-down while combination of the substitutions or deletion of both GRF zinc-fingers strongly reduced rNEIL3 recovery, indicating that interactions mediated by the GRF zinc-fingers promote recruitment of NEIL3 to an ICL. Two independent experiments. g, pICLAP was replicated in mock-depleted or NEIL3-depleted extracts supplemented with wild-type or mutated NEIL3 as indicated and analysed as in Fig. 1b. Relative to wild-type rNEIL3, rNEIL3 with substitutions in either GRF zinc-finger that abolish ssDNA binding (K500E and K546E) exhibited modest defects in pICLAP unhooking that were exacerbated when the substitutions were combined, indicating that interactions between the GRF zinc-fingers and ssDNA contribute to ICL repair. Two independent experiments. h, Extracts used in the replication reactions shown in g were analysed for NEIL3. Non-specifically detected proteins are marked with asterisks. Two independent experiments. i, Model for recruitment of NEIL3 to chromatin by zinc-finger-mediated interactions. Upon replication-fork convergence at an ICL, TRAIP-dependent CMG ubiquitylation recruits NEIL3 through direct interactions between the NZF domain of NEIL3 and ubiquitin. Association of NEIL3 with chromatin is further enhanced by interactions between the tandem GRF zinc-fingers and ssDNA, possibly on the lagging strand template.
Extended Data Fig. 9 Knockout of the Fanconi anaemia and NEIL3 pathways have additive effects on ICL sensitivity in mammalian cells.
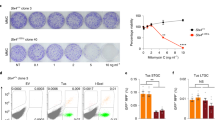
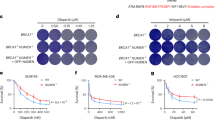
a, Immunoblot analysis of NEIL3 expression in wild-type, NEIL3-knockout and FANCL/NEIL3-knockout HAP1 cell lines. Histone H3 was detected as a loading control. Non-specifically detected proteins are marked with asterisks. Analysis performed twice. b, Schematic of FANCL targeting by CRISPR. (i) Human FANCL exon 10, single-guide (sg)RNA binding sites, and homology arm targets; (ii) FANCL-Puro targeting construct with homology arms flanking exon 10; (iii) Targeted FANCL allele with integrated puromycin-resistance cassette. c, Detection of the integrated puromycin-resistance cassette in HAP1 cells by FANCL long-range PCR. Analysis performed twice. d, Analysis of FANCD2 ubiquitylation in mitomycin-C-treated wild-type, FANCL-knockout and FANCL/NEIL3-knockout HAP1 cell lines to confirm FANCL knockout. Vinculin was detected as a loading control. FANCL is the catalytic subunit of the Fanconi anaemia core complex, which ubiquitylates FANCD2. Analysis performed twice. e, Neil3 qRT–PCR confirming gene disruption in CH12 cell lines. ND, not detected. Analysis performed once. f, Analysis of FANCD2 ubiquitylation in mitomycin-C-treated Fancb-knockout and Neil3/Fancb-knockout CH12 cell lines to confirm Fancb knockout. Vinculin was detected as a loading control. A non-specifically detected protein is marked with an asterisk. Analysis performed once. g, Cell viability of wild-type, Fancb-knockout, Neil3-knockout or Fancb/Neil3-knockout CH12 cells after exposure to trioxsalen and UVA irradiation. Two independent clones were used for the single mutants and three independent clones were used for the double mutant. Data are mean ± s.e.m. We speculate that, relative to HAP1 cells, CH12 cells may be more reliant on the Fanconi anaemia pathway to repair trioxsalen-induced damage owing to lower expression levels of NEIL3. Three independent experiments.
Extended Data Fig. 10 Model of the action of TRAIP in interphase and mitosis.
a, TRAIP (blue) travels with the replisome and associates with CMG, directly or indirectly (i–iv, top). In interphase, TRAIP is oriented such that its E2 (the identity of which is unknown) points ahead of the replisome. As a result, TRAIP cannot ubiquitylate the CMG it is associated with, but it can ubiquitylate any protein that lies in the path of the replisome. TRAIP can thus ubiquitylate CMG in trans when two replisomes meet (i), and it can ubiquitylate DPCs that block the path of the replisome (ii). We speculate that TRAIP ubiquitylates any proteinaceous structure that causes extended stalling of the replisome. In mitosis, TRAIP undergoes a conformational change so that it can ubiquitylate the CMG with which it is associated in cis. As a result, in mitosis, stalled forks undergo TRAIP-dependent CMG ubiquitylation in the absence of fork convergence (iii)4. We propose that TRAIP does not ubiquitylate terminated CMGs in interphase because they rapidly move past each other30, precluding ubiquitylation by the forward-pointing TRAIP (iv, middle). However, upon mitotic entry, terminated CMGs undergo TRAIP-dependent ubiquitylation in cis (iv, bottom)4. It is notable that TRAIP-dependent CMG unloading in interphase egg extracts is not dependent on residual CDK1 activity (b, c), which indicates that TRAIP is regulated differently in interphase and mitosis. b, In interphase egg extracts, TRAIP travels with DNA replication forks but ubiquitylates CMGs only when forks converge. In the presence of cyclin B1-CDK1, TRAIP is activated in the absence of fork convergence4 (summarized in a). We therefore wanted to know whether TRAIP-dependent CMG unloading in interphase egg extract depends on residual CDK1 activity. Top, reaction scheme to determine whether the action of TRAIP at ICLs in interphase egg extract requires residual CDK1 activity. Replication of pICL-lacOPt with a pre-assembled LacR array was initiated by addition of NPE (−55 min). Forty-five minutes after initiation (−10 min), reactions were supplemented with buffer or the CDK1 inhibitor RO-3306 (CDK1i) and allowed to incubate for an additional 10 min; CDK1i was added late to avoid inhibition of replication initiation. The LacR array was then released with the addition of IPTG to trigger fork convergence and ICL repair (0 min). In a control, buffer instead of IPTG was added to maintain the LacR array and thereby prevent fork convergence. At the indicated times after IPTG addition, samples were collected and analysed as in Fig. 1b to look for evidence of CMG unloading and ICL processing. Bottom, experiment described in top scheme. Fork stalling at the boundaries of the LacR array lead to a theta structure (lanes 1, 7, 13, 19). Upon addition of IPTG, theta was converted to slow-figure-8, fast-figure-8 (co-migrating with theta) and well products with equal efficiency in the presence and absence of CDK1i (compare lanes 7–12 and 19–24). The data suggest that CDK1 is not required for CMG unloading or ICL repair. Two independent experiments. c, Left, reaction scheme of a control experiment to ensure that the addition of CDK1i in b blocked CDK1 activity. As recently described4, when replication forks are stalled at a LacR array, the subsequent addition of cyclin B1-CDK1 promotes TRAIP-dependent CMG unloading and fork breakage, alternative end joining, and the formation of aberrant replication products that migrate in the well of an agarose gel. To verify that CDK1 was effectively inhibited in a, we used this CDK1-dependent well-product formation as an assay. To this end, replication of pCtrl-lacO with a pre-assembled LacR array was initiated (−55 min) by addition of NPE. Forty-five min after initiation (−10 min), reactions were supplemented with CDK1i or buffer and allowed to incubate for an additional 10 min. Cyclin B1-CDK1 was then added to activate fork breakage and well-product formation (0 min). Right, experiment described in left scheme. As expected, the addition of cyclin B1-CDK1 in the absence of CDK1i led to the formation of well products (lanes 25–30), but in the presence of CDK1i well products were not formed (lanes 31–36). Two independent experiments. Note that the experiments in b and c were performed in parallel; extracts were aliquoted for the various conditions described at 0 min. Therefore, the CDK1i effectively inhibited CDK1-dependent activation of TRAIP. We conclude that TRAIP activation at ICLs does not depend on residual CDK1 activity.
Supplementary information
Supplementary Table 1
This file contains the sequences of oligonucleotides mentioned in Methods
Supplementary Figure 1
This file contains uncropped source images with size marker indications
Rights and permissions
About this article
Cite this article
Wu, R.A., Semlow, D.R., Kamimae-Lanning, A.N. et al. TRAIP is a master regulator of DNA interstrand crosslink repair. Nature 567, 267–272 (2019). https://doi.org/10.1038/s41586-019-1002-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1002-0
This article is cited by
-
TRAIP suppresses bladder cancer progression by catalyzing K48-linked polyubiquitination of MYC
Oncogene (2024)
-
FLIP(C1orf112)-FIGNL1 complex regulates RAD51 chromatin association to promote viability after replication stress
Nature Communications (2024)
-
When DNA-damage responses meet innate and adaptive immunity
Cellular and Molecular Life Sciences (2024)
-
TRAIP resolves DNA replication-transcription conflicts during the S-phase of unperturbed cells
Nature Communications (2023)
-
NEIL3-mediated proteasomal degradation facilitates the repair of cisplatin-induced DNA damage in human cells
Scientific Reports (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.