Abstract
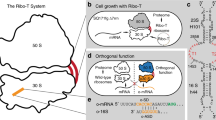
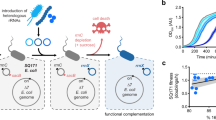
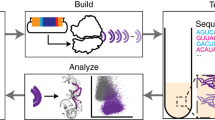
Orthogonal ribosomes are unnatural ribosomes that are directed towards orthogonal messenger RNAs in Escherichia coli, through an altered version of the 16S ribosomal RNA of the small subunit1. Directed evolution of orthogonal ribosomes has provided access to new ribosomal function, and the evolved orthogonal ribosomes have enabled the encoding of multiple non-canonical amino acids into proteins2,3,4. The original orthogonal ribosomes shared the pool of 23S ribosomal RNAs, contained in the large subunit, with endogenous ribosomes. Selectively directing a new 23S rRNA to an orthogonal mRNA, by controlling the association between the orthogonal 16S rRNAs and 23S rRNAs, would enable the evolution of new function in the large subunit. Previous work covalently linked orthogonal 16S rRNA and a circularly permuted 23S rRNA to create orthogonal ribosomes with low activity5,6; however, the linked subunits in these ribosomes do not associate specifically with each other, and mediate translation by associating with endogenous subunits. Here we discover engineered orthogonal ‘stapled’ ribosomes (with subunits linked through an optimized RNA staple) with activities comparable to that of the parent orthogonal ribosome; they minimize association with endogenous subunits and mediate translation of orthogonal mRNAs through the association of stapled subunits. We evolve cells with genomically encoded stapled ribosomes as the sole ribosomes, which support cellular growth at similar rates to natural ribosomes. Moreover, we visualize the engineered stapled ribosome structure by cryo-electron microscopy at 3.0 Å, revealing how the staple links the subunits and controls their association. We demonstrate the utility of controlling subunit association by evolving orthogonal stapled ribosomes which efficiently polymerize a sequence of monomers that the natural ribosome is intrinsically unable to translate. Our work provides a foundation for evolving the rRNA of the entire orthogonal ribosome for the encoded cellular synthesis of non-canonical biological polymers7.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The cryo-EM structure of d2d8 can be found under the PDB accession code 6HRM and the Electron Microscopy Data Bank accession number EMD-0261. Genome sequences for the strains created here are provided in Supplementary Tables 5–10. All other datasets generated and analysed here are available from the corresponding author upon reasonable request.
References
Rackham, O. & Chin, J. W. A network of orthogonal ribosome•mRNA pairs. Nat. Chem. Biol. 1, 159–166 (2005).
Wang, K., Neumann, H., Peak-Chew, S. Y. & Chin, J. W. Evolved orthogonal ribosomes enhance the efficiency of synthetic genetic code expansion. Nat. Biotechnol. 25, 770–777 (2007).
Neumann, H., Wang, K., Davis, L., Garcia-Alai, M. & Chin, J. W. Encoding multiple unnatural amino acids via evolution of a quadruplet-decoding ribosome. Nature 464, 441–444 (2010).
Wang, K. et al. Optimized orthogonal translation of unnatural amino acids enables spontaneous protein double-labelling and FRET. Nat. Chem. 6, 393–403 (2014).
Fried, S. D., Schmied, W. H., Uttamapinant, C. & Chin, J. W. Ribosome subunit stapling for orthogonal translation in E. coli. Angew. Chem. Int. Edn 54, 12791–12794 (2015).
Orelle, C. et al. Protein synthesis by ribosomes with tethered subunits. Nature 524, 119–124 (2015).
Chin, J. W. Expanding and reprogramming the genetic code. Nature 550, 53–60 (2017).
Voorhees, R. M. & Ramakrishnan, V. Structural basis of the translational elongation cycle. Annu. Rev. Biochem. 82, 203–236 (2013).
Triman, K. L., Peister, A. & Goel, R. A. Expanded versions of the 16S and 23S ribosomal RNA mutation databases (16SMDBexp and 23SMDBexp). Nucleic Acids Res. 26, 280–284 (1998).
Kitahara, K. & Suzuki, T. The ordered transcription of RNA domains is not essential for ribosome biogenesis in Escherichia coli. Mol. Cell 34, 760–766 (2009).
Szewczak, A. A. & Cech, T. R. An RNA internal loop acts as a hinge to facilitate ribozyme folding and catalysis. RNA 3, 838–849 (1997).
Nakatogawa, H. & Ito, K. The ribosomal exit tunnel functions as a discriminating gate. Cell 108, 629–636 (2002).
Vázquez-Laslop, N., Ramu, H., Klepacki, D., Kannan, K. & Mankin, A. S. The key function of a conserved and modified rRNA residue in the ribosomal response to the nascent peptide. EMBO J. 29, 3108–3117 (2010).
Barrett, O. P. & Chin, J. W. Evolved orthogonal ribosome purification for in vitro characterization. Nucleic Acids Res. 38, 2682–2691 (2010).
Youngman, E. M. & Green, R. Affinity purification of in vivo-assembled ribosomes for in vitro biochemical analysis. Methods 36, 305–312 (2005).
Vester, B. & Douthwaite, S. Macrolide resistance conferred by base substitutions in 23S rRNA. Antimicrob. Agents Chemother. 45, 1–12 (2001).
Sigmund, C. D., Ettayebi, M. & Morgan, E. A. Antibiotic resistance mutations in 16S and 23S ribosomal RNA genes of Escherichia coli. Nucleic Acids Res. 12, 4653–4664 (1984).
Wang, K. et al. Defining synonymous codon compression schemes by genome recoding. Nature 539, 59–64 (2016).
Quan, S., Skovgaard, O., McLaughlin, R. E., Buurman, E. T. & Squires, C. L. Markerless Escherichia coli rrn deletion strains for genetic determination of ribosomal binding sites. G3 (Bethesda) 5, 2555–2557 (2015).
James, N. R., Brown, A., Gordiyenko, Y. & Ramakrishnan, V. Translational termination without a stop codon. Science 354, 1437–1440 (2016).
Huter, P. et al. Structural basis for polyproline-mediated ribosome stalling and rescue by the translation elongation factor EF-P. Mol. Cell 68, 515–527 (2017).
Doerfel, L. K. et al. Entropic contribution of elongation factor P to proline positioning at the catalytic center of the ribosome. J. Am. Chem. Soc. 137, 12997–13006 (2015).
Pavlov, M. Y. et al. Slow peptide bond formation by proline and other N-alkylamino acids in translation. Proc. Natl Acad. Sci. USA 106, 50–54 (2009).
Doerfel, L. K. et al. EF-P is essential for rapid synthesis of proteins containing consecutive proline residues. Science 339, 85–88 (2013).
Ude, S. et al. Translation elongation factor EF-P alleviates ribosome stalling at polyproline stretches. Science 339, 82–85 (2013).
Maini, R. et al. Protein synthesis with ribosomes selected for the incorporation of β-amino acids. Biochemistry 54, 3694–3706 (2015).
Melo Czekster, C., Robertson, W. E., Walker, A. S., Söll, D. & Schepartz, A. In vivo biosynthesis of a β-amino acid-containing protein. J. Am. Chem. Soc. 138, 5194–5197 (2016).
Maini, R. et al. Ribosome-mediated incorporation of dipeptides and dipeptide analogues into proteins in vitro. J. Am. Chem. Soc. 137, 11206–11209 (2015).
Dedkova, L. M., Fahmi, N. E., Golovine, S. Y. & Hecht, S. M. Construction of modified ribosomes for incorporation of d-amino acids into proteins. Biochemistry 45, 15541–15551 (2006).
Terasaka, N., Hayashi, G., Katoh, T. & Suga, H. An orthogonal ribosome-tRNA pair via engineering of the peptidyl transferase center. Nat. Chem. Biol. 10, 555–557 (2014).
Engler, C., Kandzia, R. & Marillonnet, S. A one pot, one step, precision cloning method with high throughput capability. PLoS One 3, e3647 (2008).
Sachdeva, A., Wang, K., Elliott, T. & Chin, J. W. Concerted, rapid, quantitative, and site-specific dual labeling of proteins. J. Am. Chem. Soc. 136, 7785–7788 (2014).
Liu, H. & Naismith, J. H. An efficient one-step site-directed deletion, insertion, single and multiple-site plasmid mutagenesis protocol. BMC Biotechnol. 8, 91 (2008).
Peabody, D. S. & Ely, K. R. Control of translational repression by protein-protein interactions. Nucleic Acids Res. 20, 1649–1655 (1992).
LeCuyer, K. A., Behlen, L. S. & Uhlenbeck, O. C. Mutants of the bacteriophage MS2 coat protein that alter its cooperative binding to RNA. Biochemistry 34, 10600–10606 (1995).
Kwon, Y. C. & Jewett, M. C. High-throughput preparation methods of crude extract for robust cell-free protein synthesis. Sci. Rep. 5, 8663 (2015).
Yang, W. C., Patel, K. G., Wong, H. E. & Swartz, J. R. Simplifying and streamlining Escherichia coli-based cell-free protein synthesis. Biotechnol. Prog. 28, 413–420 (2012).
Labun, K., Montague, T. G., Gagnon, J. A., Thyme, S. B. & Valen, E. CHOPCHOP v2: a web tool for the next generation of CRISPR genome engineering. Nucleic Acids Res. 44 (W1), W272–W276 (2016).
Montague, T. G., Cruz, J. M., Gagnon, J. A., Church, G. M. & Valen, E. CHOPCHOP: a CRISPR/Cas9 and TALEN web tool for genome editing. Nucleic Acids Res. 42, W401–W407 (2014).
Miyazaki, K. Molecular engineering of a PheS counterselection marker for improved operating efficiency in Escherichia coli. Biotechniques 58, 86–88 (2015).
Badran, A. H. & Liu, D. R. Development of potent in vivo mutagenesis plasmids with broad mutational spectra. Nat. Commun. 6, 8425 (2015).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Cock, P. J. et al. Biopython: freely available Python tools for computational molecular biology and bioinformatics. Bioinformatics 25, 1422–1423 (2009).
Fernandez-Leiro, R. & Scheres, S. H. W. A pipeline approach to single-particle processing in RELION. Acta Crystallogr. D Struct. Biol. 73, 496–502 (2017).
Li, X. et al. Electron counting and beam-induced motion correction enable near-atomic-resolution single-particle cryo-EM. Nat. Methods 10, 584–590 (2013).
Zhang, K. Gctf: real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Scheres, S. H. Semi-automated selection of cryo-EM particles in RELION-1.3. J. Struct. Biol. 189, 114–122 (2015).
Bai, X. C., Rajendra, E., Yang, G., Shi, Y. & Scheres, S. H. Sampling the conformational space of the catalytic subunit of human γ-secretase. eLife 4, e11182 (2015).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Chen, V. B., et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D Biol. Crystallogr. 66, 12–21 (2010).
The PyMOL Molecular Graphics System v.8 (Schrödinger, 2015).
Datsenko, K. A. & Wanner, B. L. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc. Natl Acad. Sci. USA 97, 6640–6645 (2000).
Warren, D. J. Preparation of highly efficient electrocompetent Escherichia coli using glycerol/mannitol density step centrifugation. Anal. Biochem. 413, 206–207 (2011).
Acknowledgements
This work was supported by the UK Medical Research Council (MRC; grants MC_U105181009 and MC_UP_A024_1008), the Biotechnology and Biological Sciences Research Council (BBSRC; grant BB/M000842/1, for automation) and the European Research Council (ERC) Advanced Grant (grant SGCR), all to J.W.C. S.D.F. was supported by a fellowship from Kings College, Cambridge. C.D.R. was supported by a Gates Cambridge Scholarship, and supported in the laboratory of V. Ramakrishnan by the MRC (grant MC_U105184332), the Wellcome Trust (grant WT096570), the Louis-Jeantet Foundation and the Agouron Institute.
Author information
Authors and Affiliations
Contributions
W.H.S. developed riboREXER and automated parallel evolution, and analysed the resulting data. Z.T. prepared samples for electron microscopy, developed the orthogonal in vitro translation systems and analysed the resulting data. Z.T. also generated the O-stapled ribosome library, with the assistance of W.H.S., and performed and analysed the polyproline translation experiments. C.U. developed the MS2 pulldown experiments, performed the pulldowns, and analysed the data using samples prepared with the assistance of W.H.S. C.D.R performed cryo-EM and data analysis. S.D.F. cloned O-stapled ribosomes and performed some initial analysis. W.H.S. characterized the activities of O-stapled ribosomes, with assistance from C.U. and Z.T. W.H.S., Z.T., C.U. and J.W.C. wrote the paper, with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Partitioning of free large ribosomal subunits between wild-type and orthogonal small subunits, and in vivo activity of O-stapled ribosomes.
a, Gain-of-function mutations in free large subunits may confer gain-of-function phenotypes through statistical partitioning of free large subunits between wild-type (WT) and orthogonal small subunits. A mutant large subunit (LSU) can partition between WT and orthogonal (green) small subunits (SSUs) in cells that contain both WT large subunits (not shown) and mutant large subunits. Hence the mutant phenotype will be observed in the translation of both WT and orthogonal messages (see Extended Data Fig. 4d for an example). b, In vivo activity of linker-length variants of O-stapled ribosomes (using the same dataset as in Fig. 2b). GFP expression was analysed in E. coli cells containing the indicated O-ribosome, the Methanosarcina mazei PylRS synthetase/tRNACUA pair, and the O-sfGFP150TAG reporter, in the presence of 1 mM BocK (N-epsilon (tert-butoxylcarbonyl)-l-lysine). GFP fluorescence is shown as a percentage of that produced from an orthogonal ribosome with independent, non-linked subunits. O-d2d8 is highlighted in blue, and O-ribosomes with previously described subunit linkers are in dark grey. Statistics are detailed in the Methods.
Extended Data Fig. 2 MS2-tagged O-stapled ribosomes.
a, Tagging O-stapled ribosomes with an MS2 stem loop minimally perturbs in vivo ribosome activity. We measured in vivo ribosome activities via GFP production from an O-sfGFP150TAG reporter, in cells expressing an intact or MS2-tagged linker-length variant of O-stapled ribosome along with the M. mazei pyrrolysyl-tRNA synthetase/tRNACUA pair (encoded by PylS/pylT) in the presence of 1 mM BocK. GFP fluorescence was normalized to that produced from a non-stapled O-ribosome. For numbers of replicates (n) and statistics, see Methods and Supplementary Data 2. b, Sucrose gradient analysis of an E. coli lysate with and without an O-stapled ribosome variant; n = 3 biological replicates. c, Affinity purification of a non-stapled O-ribosome depends on the presence of GST-tdMS2CP (a fusion between glutathiose-S-transferase (GST) and a mutant of a tandem dimer of the MS2 coat protein (tdMS2CP)) on resin. Affinity purification of MS2-tagged ribosomes was performed on glutathione–sepharose resin with bound GST-tdMS2CP (lanes 2–4) and without GST-tdMS2CP (lane 1). Varying amounts of total RNA were loaded in lanes 2–4. d, Affinity purification depends on the presence of the MS2 RNA stem loop on ribosomes. Pellets of O-p2d6 ribosomes were collected through sucrose cushions. Affinity purifications were performed on glutathione–sepharose resin with bound GST-tdMS2CP. e, O-d5d8-MS2 was affinity purified, and MS2 stem-loop-containing species were probed by northern blot (NB). EtBr, ethidium bromide (a fluorescent stain for nucleic acid). The experiments in c–e were each performed once. For source data regarding gels, see Supplementary Fig. 1.
Extended Data Fig. 3 Engineered O-stapled ribosome variants minimize cross-assembly.
a, Screen of 50S cross-assembly coefficients for different O-ribosomes with linked subunits. n = 2 biological replicates; each replicate is shown by a dot, and the bars represent the means (using the same dataset as in Fig. 2d). b, Screen of 30S cross-assembly coefficients. n = 2 biological replicates; each replicate is shown by a dot, and the bars represent the means (using the same dataset as in Fig. 2e). In a and b, more than 90% of ribosomes had cross-assembly coefficients between 0 and 1, as expected. Previously reported O-ribosomes with linked subunits are shown in dark grey, while O-d2d8 is highlighted in blue. c, Correlation between the means (from n = 2 biological replicates) of 50S and 30S cross-assembly coefficients for different O-stapled ribosome variants; the variation in these data is shown in a, b.
Extended Data Fig. 4 In vitro translation activity of O-stapled ribosomes upon inhibition of native ribosomes.
a, In vitro translation of T7-O-GFP in S30 extracts with and without the non-stapled O-ribosome. The data show the means (grey bars) of, respectively, 12 and 8 independent replicates (dots); the error bars show ± s.d. b, In vitro translation activities of O-stapled ribosome variants (y-axis) were measured via the GFP fluorescence produced from T7-O-GFP. In vivo activities (measured as described in Extended Data Fig. 1b) are shown on the x-axis. Individual replicates are shown in Extended Data Fig. 1b, and statistics are described in the Methods. c, In vitro translation of T7-O-GFP (O-RBS) or T7-GFP (wt-RBS) in S30 extracts containing the non-stapled O-ribosome in the presence of spectinomycin. The O-16S rRNA of the O-ribosome contains the C1192U mutation, which confers resistance to spectinomycin. The data show the mean of n = 4 independent replicates; error bars are ± s.d. From this, we conclude that 10 μM of spectinomycin is sufficient to inhibit the translational activity of wild-type small subunits in the S30 extract, but has minimal effect on spectinomycin-resistant subunits. rfu, relative fluorescence units. d, In vitro translation of T7-O-GFP (O-RBS) or T7-GFP (wt-RBS) in S30 extracts containing the non-stapled O-ribosome in the presence of erythromycin. The 23S rRNA that is co-expressed with the O-16S rRNA contains the A2058G mutation, which confers resistance to erythromycin. In vitro translation of T7-GFP (wt-RBSWTS30) was also performed in S30 extracts without the O-ribosome. The data show the means of n = 4 independent replicates; error bars show ± s.d. We found that 50 μM erythromycin reduces translation from the wild-type ribosome-binding site to 18.5% of the level seen without erythromycin, and reduces translation from the orthogonal ribosome-binding site to 30% of the level without erythromycin. We conclude that 50 μM erythromycin is sufficient to inhibit wild-type large subunits in the S30 extract. e, Ratios of GFP produced from T7-O-GFP versus T7-GFP in S30 extracts containing the non-stapled O-ribosome are shown as a function of spectinomycin and erythromycin concentration. The data are means of n = 4 independent replicates; statistics are in the Methods.
Extended Data Fig. 5 Activities of O-stapled-ribosome linker-length variants in cell-free protein synthesis.
a, Activities of lysates that contain a mixture of host and O-stapled ribosomes on a WT-GFP reporter, showing that the translation machinery in the tested lysates is equally active. Every data point shows the activity of an independently produced S30 extract from independently grown cells. b, Activities of the same set of lysates as in a on an O-GFP reporter, normalized to their activity on the WT-GFP reporter. Every data point shows the activity of an independently produced S30 extract from independently grown cells. Detailed statistics are shown in Supplementary Data 3. c, The same dataset as in b, represented as a heatmap for ease of comparison. d, Activity of lysates containing a mixture of host and O-stapled ribosomes on the WT-GFP reporter. Every data point shows independently measured activities of O-stapled ribosome variants in independently prepared S30 extracts. e, Activity of the same set of the lysates as in d on the O-GFP reporter in the presence of 10 μM spectinomycin and 50 μM erythromycin, normalized to their activity on WT-GFP. Data points are independently measured activities of O-stapled ribosome variants in independently prepared S30 extracts. Detailed statistics are given in Supplementary Data 4. f, The same dataset as in e, represented as a heatmap for ease of comparison. In a, b, d and e, the error bars show ± s.d., the grey bar shows the mean of the number (n) of independent replicates. Values for n are given in the Methods.
Extended Data Fig. 6 In vitro translation activity of O-stapled ribosome variants upon inhibition of native ribosome subunits.
This figure uses the same dataset as in Fig. 2g, and shows the in vitro translation of T7-O-GFP in S30 extracts containing different O-stapled ribosome variants in the presence of 10 μM spectinomycin and 50 μM erythromycin (which inhibit contributions to translation from endogenous subunits). The dots indicate individual data for each tested O-stapled ribosome, and the grey bars show means, for the number of independent replicates given in the Methods. O-d2d8 is highlighted in blue, and the error bars indicate ± s.d.
Extended Data Fig. 7 Genomically encoded stapled ribosomes.
a, Colony PCR products reflect the genomic exchanges seen in E. coli strains with a genomically encoded, stapled ribosome rRNA operon integrated by ribo-REXER. Top, direct amplification of the genomic region upstream of the rrnE locus. Integration of the landing site (SacB/CAT) to create the strain SQ110CAT-SacB increases the length of the region upstream of rrnE from 1.3 kb to 3.5 kb. After ribo-REXER, this cassette is lost again, leading to a 1.3-kb band. Bottom, amplification from nucleotide 2,630 of 23S to the rrnE terminator region shows integration of the PheS*/HygR cassette after integration of the wild-type rrnB operon, indicated by ‘integration cassette wt’. Integration of a stapled-ribosome cassette increases the length of the PCR product further, as it includes the 16S 3′ and internal transcribed spacer (ITS) regions, leading to the bands indicated by ‘integration cassette stapled’. The experiment was performed once. b, Denaturing RNA gel electrophoresis reveals the expression of rRNA (around 4,500 nucleotides) from intact stapled ribosomes as the predominant RNA species in all strains. The experiment was performed once. c, Growth rates of strains after successful ribo-REXER. WT describes the integration of a non-stapled rrnB operon. For statistics, see Methods. d, Denaturing RNA gel electrophoresis shows the expression of rRNA from intact stapled ribosomes (around 4,500 nucleotides) as the predominant RNA species in all evolved Δ6 d2d8 strains. The experiment was performed once. e, Sucrose gradient analyses of stapled ribosomes isolated from cells under ribosome-associating conditions (10 mM MgCl2); the experiments were repeated three times with similar results. For source data regarding gels, see Supplementary Fig. 1.
Extended Data Fig. 8 Purification of the d2d8 stapled ribosome and structural determination by cryo-EM.
a, Isolation of the d2d8 stapled ribosome on a sucrose gradient for cryo-EM. Fractions corresponding to the middle of the 70S peak (grey shading) were collected and used in structural studies. The inset shows RNA agarose gel analysis of the stapled ribosome at the different purification stages (30S extract preparation and salt wash) as well as the combined 70S fraction sample. The arrow indicates the position of the stapled rRNA. For source data regarding gels, see Supplementary Fig. 1. The data represent n = 2 independent preparations. b, Fourier shell correlation (FSC) curve, calculated between independent half-maps. The resolution is estimated from the map-to-map correlation at FSC = 0.143 (for a detailed description, see Extended Data Table 1). The electron-microscopy map is coloured according to local resolution. c, Workflow showing the three-dimensional classification and refinement of cryo-EM particles.
Extended Data Fig. 9 Selection of O-ribosome variants competent in translation of polyproline sequences.
a, PCR characterization of an E. coli TOP10 Δefp strain. PCR was carried out, using the indicated primer pairs (128/144 or 128/129), on genomic DNA from a TOP10 strain with intact efp, or a strain in which the efp locus was disrupted by a hygromycin-B-resistance gene (hph). For source data regarding gels, see Supplementary Fig. 1. The experiment was performed twice. b, Cell-growth assay, showing the translation activity of O-d2d8 on O-(P)n-CAT reporters in wild-type (+) or Δefp (−) E. coli TOP10 cells and with varying concentrations of chloramphenicol. The experiment was performed once. c, Activity of the PTC-library hits and the parental O-d2d8 ribosome on p15A-O-(P)4-GFP and p15A-O-(P)7-GFP reporters in E. coli TOP10 Δefp cells. n = 3 biological replicates; error bars indicate ± s.d. d, Translation activity of exit-tunnel library hits in the absence of proline-rich sequences. p15A-O-GFP was used as a reporter. n = 7 biological replicates; error bars indicate ± s.d. e, GFP fluorescence resulting from translation of O-(P)4-GFP in E. coli TOP10 Δefp cells. n = 3 biological replicates; error bars indicate ± s.d. f, As for e, except that the activity of the evolved mutants was tested on an O-(P)7-GFP reporter in E. coli with and without efp. n = 3 biological replicates; error bars represent ± s.d. g, Mutations in both the PTC and the exit tunnel are required to confer on O-d2d8 the ability to translate O-(P)4-GFP and O-(P)7-GFP in TOP10 Δefp cells. O-d2d8PTC-36 and O-d2d8ET-5 contain selected mutations in the PTC and the exit tunnel respectively; O-d2d8-5 contains mutations in both. See Supplementary Data 12 for details on mutations. n = 6 biological replicates; error bars represent ± s.d. h, Electrospray ionization spectra of O-(P)7-GFP synthesized by O-d2d8-5 in Δefp E. coli. This experiment was performed once.
Supplementary information
Supplementary Information
This file contains a guide to Supplementary Data files 1-12, Supplementary Figure 1, a Supplementary Note and Supplementary References.
Supplementary Data
This zipped file contains Supplementary Data files 1-12 – see Supplementary Information document for full legends.
Rights and permissions
About this article
Cite this article
Schmied, W.H., Tnimov, Z., Uttamapinant, C. et al. Controlling orthogonal ribosome subunit interactions enables evolution of new function. Nature 564, 444–448 (2018). https://doi.org/10.1038/s41586-018-0773-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0773-z
Keywords
This article is cited by
-
Adding α,α-disubstituted and β-linked monomers to the genetic code of an organism
Nature (2024)
-
Bioorthogonal Reactions in Bioimaging
Topics in Current Chemistry (2024)
-
Genetically programmed cell-based synthesis of non-natural peptide and depsipeptide macrocycles
Nature Chemistry (2023)
-
Community science designed ribosomes with beneficial phenotypes
Nature Communications (2023)
-
Cell-free Biosynthesis of Peptidomimetics
Biotechnology and Bioprocess Engineering (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.