Abstract
Mammalian cells are surrounded by neighbouring cells and extracellular matrix (ECM), which provide cells with structural support and mechanical cues that influence diverse biological processes1. The Hippo pathway effectors YAP (also known as YAP1) and TAZ (also known as WWTR1) are regulated by mechanical cues and mediate cellular responses to ECM stiffness2,3. Here we identified the Ras-related GTPase RAP2 as a key intracellular signal transducer that relays ECM rigidity signals to control mechanosensitive cellular activities through YAP and TAZ. RAP2 is activated by low ECM stiffness, and deletion of RAP2 blocks the regulation of YAP and TAZ by stiffness signals and promotes aberrant cell growth. Mechanistically, matrix stiffness acts through phospholipase Cγ1 (PLCγ1) to influence levels of phosphatidylinositol 4,5-bisphosphate and phosphatidic acid, which activates RAP2 through PDZGEF1 and PDZGEF2 (also known as RAPGEF2 and RAPGEF6). At low stiffness, active RAP2 binds to and stimulates MAP4K4, MAP4K6, MAP4K7 and ARHGAP29, resulting in activation of LATS1 and LATS2 and inhibition of YAP and TAZ. RAP2, YAP and TAZ have pivotal roles in mechanoregulated transcription, as deletion of YAP and TAZ abolishes the ECM stiffness-responsive transcriptome. Our findings show that RAP2 is a molecular switch in mechanotransduction, thereby defining a mechanosignalling pathway from ECM stiffness to the nucleus.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Humphrey, J. D., Dufresne, E. R. & Schwartz, M. A. Mechanotransduction and extracellular matrix homeostasis. Nat. Rev. Mol. Cell Biol. 15, 802–812 (2014).
Meng, Z., Moroishi, T. & Guan, K. L. Mechanisms of Hippo pathway regulation. Genes Dev. 30, 1–17 (2016).
Halder, G., Dupont, S. & Piccolo, S. Transduction of mechanical and cytoskeletal cues by YAP and TAZ. Nat. Rev. Mol. Cell Biol. 13, 591–600 (2012).
Aragona, M. et al. A mechanical checkpoint controls multicellular growth through YAP/TAZ regulation by actin-processing factors. Cell 154, 1047–1059 (2013).
Codelia, V. A., Sun, G. & Irvine, K. D. Regulation of YAP by mechanical strain through Jnk and Hippo signaling. Curr. Biol. 24, 2012–2017 (2014).
Wada, K., Itoga, K., Okano, T., Yonemura, S. & Sasaki, H. Hippo pathway regulation by cell morphology and stress fibers. Development 138, 3907–3914 (2011).
Dupont, S. et al. Role of YAP/TAZ in mechanotransduction. Nature 474, 179–183 (2011).
Cherfils, J. & Zeghouf, M. Regulation of small GTPases by GEFs, GAPs, and GDIs. Physiol. Rev. 93, 269–309 (2013).
Wei, S. C. et al. Matrix stiffness drives epithelial–mesenchymal transition and tumour metastasis through a TWIST1–G3BP2 mechanotransduction pathway. Nat. Cell Biol. 17, 678–688 (2015).
Benham-Pyle, B. W., Pruitt, B. L. & Nelson, W. J. Mechanical strain induces E-cadherin-dependent Yap1 and β-catenin activation to drive cell cycle entry. Science 348, 1024–1027 (2015).
de Rooij, J. et al. PDZ-GEF1, a guanine nucleotide exchange factor specific for Rap1 and Rap2. J. Biol. Chem. 274, 38125–38130 (1999).
Monteiro, A. C. et al. Trans-dimerization of JAM-A regulates Rap2 and is mediated by a domain that is distinct from the cis-dimerization interface. Mol. Biol. Cell 25, 1574–1585 (2014).
Gloerich, M. et al. Rap2A links intestinal cell polarity to brush border formation. Nat. Cell Biol. 14, 793–801 (2012).
Carloni, V., Romanelli, R. G., Pinzani, M., Laffi, G. & Gentilini, P. Focal adhesion kinase and phospholipase C gamma involvement in adhesion and migration of human hepatic stellate cells. Gastroenterology 112, 522–531 (1997).
Zhang, X. et al. Focal adhesion kinase promotes phospholipase C-γ1 activity. Proc. Natl Acad. Sci. USA 96, 9021–9026 (1999).
Plouffe, S. W. et al. Characterization of Hippo pathway components by gene inactivation. Mol. Cell 64, 993–1008 (2016).
Meng, Z. et al. MAP4K family kinases act in parallel to MST1/2 to activate LATS1/2 in the Hippo pathway. Nat. Commun. 6, 8357 (2015).
Myagmar, B. E. et al. PARG1, a protein-tyrosine phosphatase-associated RhoGAP, as a putative Rap2 effector. Biochem. Biophys. Res. Commun. 329, 1046–1052 (2005).
Machida, N. et al. Mitogen-activated protein kinase kinase kinase kinase 4 as a putative effector of Rap2 to activate the c-Jun N-terminal kinase. J. Biol. Chem. 279, 15711–15714 (2004).
Taira, K. et al. The Traf2- and Nck-interacting kinase as a putative effector of Rap2 to regulate actin cytoskeleton. J. Biol. Chem. 279, 49488–49496 (2004).
Qiao, Y. et al. YAP regulates actin dynamics through ARHGAP29 and promotes metastasis. Cell Reports 19, 1495–1502 (2017).
Porazinski, S. et al. YAP is essential for tissue tension to ensure vertebrate 3D body shape. Nature 521, 217–221 (2015).
Li, Q. et al. Ingestion of food particles regulates the mechanosensing misshapen–yorkie pathway in Drosophila intestinal growth. Dev. Cell 45, 433–449 (2018).
Debnath, J., Muthuswamy, S. K. & Brugge, J. S. Morphogenesis and oncogenesis of MCF-10A mammary epithelial acini grown in three-dimensional basement membrane cultures. Methods 30, 256–268 (2003).
Yoh, K. E. et al. Repression of p63 and induction of EMT by mutant Ras in mammary epithelial cells. Proc. Natl Acad. Sci. USA 113, E6107–E6116 (2016).
Dawson, P. J., Wolman, S. R., Tait, L., Heppner, G. H. & Miller, F. R. MCF10AT: a model for the evolution of cancer from proliferative breast disease. Am. J. Pathol. 148, 313–319 (1996).
Serban, M. A., Scott, A. & Prestwich, G. D. Use of hyaluronan-derived hydrogels for three-dimensional cell culture and tumor xenografts. Curr. Protoc. Cell Biol. Ch. 10, Unit 10 14 (2008).
Levental, K. R. et al. Matrix crosslinking forces tumor progression by enhancing integrin signaling. Cell 139, 891–906 (2009).
Wen, J. H. et al. Interplay of matrix stiffness and protein tethering in stem cell differentiation. Nat. Mater. 13, 979–987 (2014).
Tse, J. R. & Engler, A. J. Preparation of hydrogel substrates with tunable mechanical properties. Curr. Protoc. Cell Biol. Ch. 10, Unit 10 16 (2010).
Kaushik, G., Fuhrmann, A., Cammarato, A. & Engler, A. J. In situ mechanical analysis of myofibrillar perturbation and aging on soft, bilayered Drosophila myocardium. Biophys. J. 101, 2629–2637 (2011).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Anders, S., Pyl, P. T. & Huber, W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Dennis, G. Jr et al. DAVID: Database for annotation, visualization, and integrated discovery. Genome Biol. 4, 3 (2003).
Acknowledgements
K.C.L., A.W.H., and S.W.P. are supported by the T32 GM007752 training grant, A.K. by T32AR060712, and J.K.P. by F32HL126406. K.-L.G. is supported by grants from the NIH (CA196878, CA217642, GM51586, DEO15964) as is A.J.E. (R21CA217735, R01CA206880). A.J.E. is also supported by NSF grant 1463689, and A.K. is supported by the NSF graduate research fellowship program and an ARCS/Roche Foundation Scholar Award in Life Science. H.W.P. is supported by KHIDI grant HI17C1560.
Reviewer information
Nature thanks M. Sudol, V. Weaver and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
Z.M. and K.-L.G. conceived the project and wrote the manuscript. Z.M., K.C.L., C.F., S.L., M.P., T.M. and M.L. performed in vitro cell assays, CRISPR knockout, quantitative real-time PCR, immunofluorescence, and xenograft studies. A.K., J.K.P., K.-C.W., A.W.H., S.C. and A.J.E. assisted in manufacturing hydrogels and with immunofluorescence experiments. S.W.P. and H.W.P. provided knockout cell lines. Y.Q., Y.D., Z.Y. and B.R. performed the next-generation sequencing and bioinformatics analyses. X.W. performed the pathological analyses. F.-X.Y., C.-Y.W., B.R. and A.J.E. provided technical support.
Corresponding author
Ethics declarations
Competing interests
K.-L.G. is a co-founder of Vivace Therapeutics. The other authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 RAP2 is involved in regulation of YAP and TAZ by matrix stiffness.
a, Localization of YAP and TAZ is not significantly affected by overexpression of H-RAS, K-RAS, N-RAS, or RAP1B. HEK293A cells were cultured on high-stiffness hydrogels. Overexpression of HA-tagged H-RAS, K-RAS, N-RAS, and Flag-tagged RAP1B are indicated. Merged, combined signals of YAP and TAZ (red), transfected small GTPases (green), and DAPI (blue, staining for DNA). Scale bars, 25 μm. The images are representative of two independent experiments with similar results. b, Subcellular fractionation of wild-type and RAP2-KO HEK293A cells at low or high stiffness. The images are representative of two independent experiments with similar results. c, CRISPR-mediated deletion of RAP2A, RAP2B and RAP2C in adipocyte-derived MSCs by lentiviral transduction. sgRNAs targeting RAP2A, RAP2B, and RAP2C (sgRAP2) were individually cloned into lentiCRISPR v2 (Addgene #52961) plasmids. sgRAP2 #1 and #2 were two sets of three sgRNAs targeting RAP2A, RAP2B, and RAP2C with unique sequences for each RAP2 isoform. Adipocyte-derived MSCs were infected with the sgRAP2 lentiviruses with an MOI of 10 and selected by puromycin. After puromycin selection, the pooled cells were examined for RAP2 protein expression. The images are representative of two independent experiments with similar results. d, RAP2 is required for the low stiffness-induced cytoplasmic localization of YAP and TAZ in MSCs. MSCs with CRISPR-mediated deletion of RAP2A, RAP2B and RAP2C were seeded onto 40 kPa and 1 kPa collagen-coated hydrogels, cultured for 24 h, and then stained for YAP and TAZ. The results are representative of three biologically independent samples showing similar results. Merged, combined signals of YAP and TAZ (red), F-actin (green), and DAPI (blue). Scale bars, 25 μm. e, Distribution of YAP and TAZ localization presented as mean + s.e.m. for cells with more nuclear (N > C), more cytoplasmic (N < C), or even (N = C) distribution of YAP and TAZ. n = 3 biologically independent samples. Scale bars, 25 μm. f, RAP2 has an important role in adipocyte differentiation at low stiffness. Representative images of Oil-Red O staining of adipocyte-derived MSCs that were treated with adipocyte differentiation medium for 15 days. The MSCs were grown on 1 kPa hydrogels. Scale bars, 200 μm. g, Quantification of Oil-Red O-positive cells. The results are presented as mean ± s.e.m. Two-tailed t-test, n = 8 biological independent samples, ***P = 0.00026. h, CRISPR-mediated deletion of RAP2A, RAP2B and RAP2C in MCF7 and MDA-MB-468 by lentiviral transduction. The experiments were performed similarly to those in c. The images are representative of two independent experiments with similar results. i, RAP2 deletion promotes nuclear localization of YAP and TAZ in MCF7 cells at low stiffness. The images are from three independent experiments showing similar results. Merged, combined signals of YAP and TAZ (red), F-actin (green), and DAPI (blue). Scale bars, 25 μm. j, Quantification of YAP and TAZ localization in MCF7 cells. The distribution of YAP and TAZ localization is presented as mean + s.e.m. n = 3 biologically independent samples. k, YAP and TAZ are not significantly regulated by matrix stiffness in MDA-MB-468 cells. The images are from three independent experiments showing similar results. Merged, combined signals of YAP and TAZ (red), F-actin (green), and DAPI (blue). Scale bar, 25 μm. l, Quantification of YAP and TAZ localization in MDA-MB-468 cells. The distribution of YAP and TAZ localization is presented as mean + s.e.m. n = 3 biologically independent samples.
Extended Data Fig. 2 Activation of RAP2 at low matrix stiffness involves PDZGEF1 and PDZGEF2.
a, RAP2 has no effect on localization of TWIST1 and β-catenin at different matrix rigidities. Immunofluorescence staining of TWIST1 and β-catenin in wild-type and RAP2-KO HEK293A cells grown on soft (1 kPa) and stiff (40 kPa) fibronectin-coated hydrogels. Low stiffness induced cytoplasmic localization of TWIST1 similarly in wild-type and RAP2-KO cells. Stiffness had no significant effect on β-catenin localization in HEK293A cells. Scale bars, 25 μm. The images are representative of three independent experiments with similar results. b, GST–RalGDS-RBD specifically binds to the active RAP2A in the pull-down assay. HEK293A cells were transfected with plasmids expressing wild-type RAP2A, RAP2A-G12V (constitutively GTP-binding), or RAP2A-S17N (GTP-binding deficient) and then seeded onto soft hydrogels. The cells were lysed 24 h after seeding and the lysates were incubated with glutathione agarose beads that were pre-loaded with GST–RalGDS-RBD. The beads were washed and subjected to western blot analyses with the indicated antibodies. The images are representative of two independent experiments with similar results. c, GTP-binding and activity of RAP2A is required to induce cytoplasmic translocation of YAP and TAZ. Merged, combined signals from YAP and TAZ (red), Flag (green), and DAPI (blue). Scale bar, 25 μm. The images are representative of two independent experiments with similar results. d, ECM stiffness regulates the interaction between RAP2A and PDZGEF1. GST–RAP2A plasmids were co-transfected with Flag–Raptor (a negative control) or Flag–PDZGEF1 into HEK293A cells. The cells were thereafter seeded on stiff and soft hydrogels. Twenty-four hours after seeding, the cells were lysed and the lysates were incubated with glutathione agarose beads for 6 h. Then the beads were washed and subjected to western blot analyses. The images are representative of two independent experiments with similar results. e, Sanger DNA sequencing confirmed the homozygous deletion of a ‘T’ nucleotide in PDZGEF1 genomic DNAs in PDZGEF1/2-dKO HEK293A cells. f, Western blot showing the absence of PDZGEF2 expression in PDZGEF1/2-dKO HEK293A cells. The images are representative of two independent experiments with similar results. g, The repression of the YAP and TAZ target genes CTGF and CYR61 by low stiffness was compromised in PDZGEF1/2-dKO cells. Expression of CTGF and CYR61 in PDZGEF1/2-dKO HEK293A cells on soft and stiff matrices was measured by qPCR. ns, not significant, two-tailed t-test, n = 3 biologically independent samples. Data are represented as mean ± s.e.m. h, RAP2 is required for PDZGEF1 to induce cytoplasmic localization of YAP and TAZ. Immunofluorescence showing localization of YAP and TAZ after ectopic expression of Flag-tagged PDZGEF1 in wild-type and RAP2-KO cells at high stiffness. Merged, combined signals from Flag-PDZGEF1 (green), YAP and TAZ (red), and DAPI (blue). Scale bars, 25 μm. i, Quantification of the results in h. + denotes Flag–PDZGEF1-transfected cells, – denotes cells that were not transfected. Data are represented as mean + s.e.m. n = 3 biologically independent samples. j, Stiffness influences cellular PtdIns(4,5)P2. The PtdIns(4,5)P2 reporter GFP–PLCδ-PH domain, which binds to PtdIns(4,5)P2, was imaged with a Nikon inverted microscope in cells at low or high stiffness. Cells grown at high stiffness display diffuse PtdIns(4,5)P2 localization whereas cells at low stiffness show enrichment of PtdIns(4,5)P2 at the plasma membrane. The image is representative of two independent experiments with similar results. Scale bars, 25 μm. k, Inhibition of PLC alters cellular PtdIns(4,5)P2 distribution. Immunofluorescence of cells treated with 5 µM PLC inhibitor U73122 at high stiffness. GFP was detected with anti-GFP immunofluorescence. PtdIns(4,5)P2 enrichment, indicated by arrows, was observed. Scale bars, 25 μm. The image is representative of two independent experiments with similar results.
Extended Data Fig. 3 FAK, PLCγ1, and PLD are involved in RAP2 activation and Hippo pathway activation in response to stiffness.
a, The FAK inhibitor PF573228 and PLC inhibitor U73122 promote RAP2–GTP loading at high stiffness. The images are representative of two biologically independent experiments with similar results. b, RAP2 functions downstream of FAK and PLC to regulate localization of YAP and TAZ. YAP and TAZ were imaged in cells cultured at high stiffness and treated with 10 µM FAK inhibitor PF573228 or 5 µM PLC inhibitor U73122. Scale bars, 25 μm. The image is representative of three (wild-type treated with PF573228 or U73122; RAP2-KO control (Ctrl)) or four (wild-type control; RAP2-KO treated with PF573228 or U73122) biologically independent samples with similar results. c, Distribution of localization from b as mean + s.e.m. n = 3 (wild-type treated with PF573228 or U73122; RAP2-KO control) or four (wild-type control; RAP2-KO treated with PF573228 or U73122) biologically independent samples. d, The PLD1 and PLD2 inhibitor BML279 suppresses RAP2–GTP binding at low stiffness. The images are representative of two independent experiments with similar results. e, Inhibition of PLD increases nuclear YAP and TAZ in cells at low stiffness. Cells growing at low stiffness were treated with 5 µM PLD inhibitor BML279. Scale bars, 25 μm. Images are representative of seven (control) or five (BM279) biologically independent samples with similar results. f, Distribution of localization from e presented as mean + s.e.m. n = 7 (control) or 5 (BML279) biologically independent samples. g, Western blot showing PLCγ1 knockdown by duplex siRNAs (DsiRNAs). Two independent siRNAs were used. The image is representative of two independent experiments with similar results. h, PLCγ1 knockdown decreases nuclear YAP and TAZ at high stiffness. Quantification of YAP and TAZ localization in PLCγ1 knockdown and control cells growing on 40 kPa hydrogels. The distribution of YAP and TAZ localization is presented as mean + s.e.m. n = 3 biologically independent samples. i, Representative images of YAP and TAZ localization in cells with PLCγ1 knockdown. Scale bars, 25 μm. The images are representative of three biologically independent experiments with similar results. j, Western blot showing knockdown of PLD1 and PLD2. Two independent DsiRNAs were used. The image is representative of two independent experiments with similar results. k, Knockdown of PLD1 and PLD2 increases nuclear YAP and TAZ at low stiffness. Quantification of YAP and TAZ localization in PLD1/2 knockdown and control cells growing on 1 kPa hydrogels. The distribution of YAP and TAZ localization is presented as mean + s.e.m. The results are from three biologically independent samples. l, Representative images of YAP and TAZ localization in PLD1/2 knockdown cells. Scale bars, 25 μm. Images are representative of three biologically independent experiments with similar results. m, Wild-type, but not GTP-binding-deficient (S17N), RAP2A induces YAP phosphorylation. YAP phosphorylation was analysed by phos-tag SDS–PAGE in HEK293A cells stably expressing wild-type or S17N mutant RAP2A at high stiffness. Images are representative of two independent experiments with similar results. n, Hippo pathway components are involved in regulation of YAP and TAZ by stiffness. HEK293A cell lines in which Hippo components were deleted were cultured at low stiffness (1 kPa). Deletion of Hippo components includes LATS1/2-dKO, MST1/2-dKO, MM-5KO (MST1/2-MAP4K4/6/7-5KO), and MM-8KO (MST1/2-MAP4K1/2/3/4/6/7-8KO). Images are representative of three biologically independent experiments with similar results. Scale bars, 25 μm. o, Quantification of immunofluorescence of the samples in n. The distribution of YAP and TAZ localization is presented as mean + s.e.m. n = 3 biologically independent samples.
Extended Data Fig. 4 RAP2 activates MAP4K4 and induces phosphorylation of YAP.
a, Hippo pathway components are required for RAP2A to induce phosphorylation of YAP and TAZ. RAP2A is overexpressed in the indicated knockout HEK293A cells. Images are representative of two independent experiments with similar results. Two clones of MM-5KO cells were used for this experiment. b, RAP2A acts through the Hippo pathway to induce cytoplasmic localization of YAP and TAZ. Flag–RAP2A was transfected into HEK293A cell lines with deletion of different core Hippo pathway components as indicated. Localization of YAP and TAZ was analysed by immunofluorescence. Merged, combined signals from Flag–RAP2A (green), YAP and TAZ (red), and DAPI (blue). Scale bars, 25 μm. Images are representative of three biologically independent experiments with similar results. c, Localization of YAP and TAZ in LATS1−/−LATS2flox/flox (LATS1-KO LATS2-F/F) and LATS1−/−LATS2−/− (LATS1/2-dKO) mouse embryonic fibroblasts (MEFs). Scale bars, 25 μm. Images are representative of two biologically independent experiments with similar results. d, Localization of YAP and TAZ in NF2-KO and MOB1A/1B-dKO HEK293A cells. Scale bars, 25 μm. Images are representative of two biologically independent experiments with similar results. e, Low matrix stiffness activates MAP4K4 in a RAP2-dependent manner. Plasmids expressing HA-tagged MAP4K4 were transfected into wild-type and RAP2-KO HEK293A cells. Twenty-four hours after seeding on 40 kPa or 1 kPa hydrogels, HA–MAP4K4 proteins were immunoprecipitated and then used for an in vitro kinase assay, in which recombinant full-length GST-tagged LATS2 protein was used as a substrate. Phosphorylation of LATS2 by MAP4K4 was detected with a phosphospecific antibody recognizing the phosphorylated hydrophobic motif of LATS1 and LATS2. Results are from three biologically independent experiments. f, Quantification of the kinase assay (e) shown as mean ± s.e.m. Relative LATS2 phosphorylation was normalized to protein levels of HA–MAP4K4 and defined as MAP4K4 kinase activity. n = 3 biologically independent samples, two-tailed t-test, *P = 0.027 (wild-type, 1 kPa versus RAP2 KO, 1 kPa) or 0.037 (wild-type, 1 kPa versus wild-type, 40 kPa); ns, not significant. g, The lysates of wild-type and RAP2-KO HEK293A cells growing on stiff and soft hydrogels were analysed by western blot for MAP4K4 migration. A ‘stretched’ image was generated by vertically extending the same western image directly above it in order to better visualize the MAP4K4 mobility shift. The image is representative of two independent experiments with similar results. h, Phosphatase treatment increases MAP4K4 migration on SDS–PAGE. Lambda phosphatase was used to treat cell lysates of RAP2A-expressing HEK293A cells before western blotting. These results indicate that the altered migration of MAP4K4 was correlated with its phosphorylation. The image is representative of two independent experiments with similar results. i, RAP2A promotes MAP4K4 phosphorylation (slower band migration) dependent on its citron domain. Flag–RAP2A-G12V plasmid was co-transfected with wild-type MAP4K4 or the citron domain deletion mutant (ΔCNH) into HEK293A cells. The image is representative of two independent experiments with similar results. j, RAP2 is required for the reduction in MAP4K4 mobility. Wild-type and ΔCNH mutant MAP4K4 were transfected into wild-type and RAP2-KO HEK293A cells. The image is representative of two independent experiments with similar results.
Extended Data Fig. 5 RAP2 inhibits RhoA GTPase through ARHGAP29.
a, RAP2A expression inhibits endogenous RhoA GTP-binding. HEK293A cells with doxycycline-inducible expression of RAP2A were established and the expression of Flag–RAP2A was induced by doxycycline. RhoA activity was determined by a GST–Rhotekin-RBD pull-down assay. s.e. denotes short exposure of the RhoA western blot. The image is representative of two independent experiments with similar results. b, Western blot confirms CRISPR-mediated RhoA gene editing. HEK293A cells were transfected with CRISPR plasmids targeting RhoA and selected with puromycin for 3 days. Two sgRNAs were used to generate two RhoA knockout pools (sgRhoA #1 and #2)16. The image is representative of two independent experiments with similar results. c, CRISPR-mediated deletion of RhoA leads to increased cytoplasmic localization of YAP and TAZ in HEK293A cells at high stiffness. The localization distribution is presented as mean + s.e.m. n = 6 biologically independent samples. d, Representative immunofluorescence images from c. Scale bars, 25 μm. e, ARHGAP29 induces YAP phosphorylation in a Hippo pathway-dependent manner. MM-5KO, a HEK293A clone with deletion of MST1, MST2, MAP4K4, MAP4K6 and MAP4K7. MM-8KO, deletion of MST1, MST2, MAP4K1, MAP4K2, MAP4K3, MAP4K4, MAP4K6 and MAP4K7. YAP phosphorylation was detected by phos-tag gels. The image is representative of two independent experiments with similar results. f, Immunoblot showing deletion of ARHGAP29 in HEK293A wild-type and MAP4K4/6/7-tKO cells. The image is representative of two independent experiments with similar results. g, Deletion of ARHGAP29 compromises inhibition of RhoA by low matrix stiffness. Wild-type and ARHGAP29-KO HEK293A cells were cultured at the indicated stiffness and then assayed for RhoA activity with a GST–Rhotekin-RBD binding assay. The image is representative of two independent experiments with similar results. h, Combined deletion of ARHGAP29, MAP4K4, MAP4K6 and MAP4K7 abolishes YAP phosphorylation induced by RAP2A. HA–YAP was co-transfected with vector or Flag–RAP2A into HEK293A cells cultured at high stiffness. HA–YAP phosphorylation was detected by phos-tag SDS–PAGE. The image is representative of two independent experiments with similar results. i, Combined deletion of ARHGAP29, MAP4K4, MAP4K6 and MAP4K7 blocks low stiffness-induced cytoplasmic localization of YAP. Quantification of YAP and TAZ localization in HEK293A cells with deletion of ARHGAP29 and/or MAP4K4, MAP4K6 and MAP4K7 at low stiffness in Fig. 3e. The YAP and TAZ localization distribution is presented as mean + s.e.m. n = 3 biologically independent samples.
Extended Data Fig. 6 RAP2 contributes to cell density-induced inactivation of YAP and TAZ.
a, Cytoplasmic translocation of YAP and TAZ caused by cell contact involves RAP2 and the Hippo pathway. Scale bars, 25 μm. MM-5KO, a HEK293A clone with deletion of MST1, MST2, MAP4K4, MAP4K6 and MAP4K7. RAP2-MST-KO, deletion of RAP2A, RAP2B, RAP2C, MST1 and MST2. The images are representative of three biologically independent experiments with similar results. b, Western blot showing absence of RAP2 proteins in RAP2-KO and RAP2-MST1/2-KO cells. Note that part of these results is shown in Fig. 1b and are from the same experiment. The image is representative of two independent experiments with similar results. c, Phosphorylation of YAP and TAZ induced by cell contact requires RAP2, MST1 and MST2. The western blot shows phosphorylation of YAP in cells with low, medium, or high confluence. The image is representative of two independent experiments with similar results. d, Deletions of RAP2A, RAP2B, RAP2C, MST1 and MST2 interfere with regulation of YAP and TAZ by cell density. Subcellular fractionations were performed for wild-type, RAP2-KO, MST1/2-dKO and RAP2-MST1/2-KO HEK293A cells cultured at low (S, sparse) or high density (D, dense). GAPDH and LaminB1 are markers for the cytoplasmic and nuclear fraction, respectively. The image is representative of two independent experiments with similar results. e, RAP2A, RAP2B, RAP2C, MST1 and MST2 are required for regulation of YAP and TAZ target genes by cell density. qPCR was performed to determine the expression of the YAP and TAZ target genes CTGF and CYR61 in the above cells at low or high confluence. Data are represented as mean ± s.e.m. n = 3 biologically independent samples.
Extended Data Fig. 7 RAP2 prevents aberrant acinus growth in MCF10A cells on soft matrices.
a, RAP2 deletion selectively enhances HEK293A cell growth on soft matrices. Wild-type and RAP2-KO HEK293A cells were seeded on stiff or soft matrices, and cell numbers were recorded as mean ± s.e.m. every day. Two-way ANOVA test, ****P < 0.0001, wild-type versus RAP2-KO cells cultured at 1 kPa, n = 3 biologically independent samples. b, Model of 3D culture of MCF10A cells. c, RAP2 deletion causes abnormal acinus growth and cell polarity defects in MCF10A cells. Immunofluorescence staining of acini for cell polarity markers, Laminin V and GM130, and YAP and TAZ in wild-type and RAP2-KO MCF10A cells cultured for 6 days. Scale bars, 25 µm. Images are representative of three independent experiments with similar results. d, Western blot showing deletion of MST1 and MST2 in wild-type and RAP2-KO cells. The arrow indicates the specific band for MST1. The image is representative of two independent experiments with similar results. e, Representative images showing acinus formation by various genetically engineered MCF10A cells at 1 kPa. The red arrow indicates aberrant acini. The blue arrow indicates invasive cell morphology. Scale bars, 100 µm. The results are representative of two independent experiments with similar results. f, Quantification of aberrant, oversized, and invasive acini in e. The percentage of the cells is presented as mean ± s.e.m. Two-tailed t-tests were used for statistical analyses of aberrant and oversized clones in n = 3 biologically independent samples. Aberrant: ****P = 0.0000062; ***P = 0.0012; oversized: *P = 0.017 (wild-type versus RAP2-KO) or 0.046 (MST1/2-dKO versus RAP2-MST1/2-KO). For analysis of invasive clones, one-tailed t-test was used as no invasive clones were observed in either wild-type or MST1/2-dKO cells. *P = 0.047 (wild-type versus RAP2-KO) or 0.029 (MST1/2-dKO versus RAP2-MST1/2-KO). n = 3 biologically independent samples. g, Representative images showing acinus formation by various genetically engineered MCF10A cells at 150 Pa. Scale bars, 100 µm. Images are representative of two biologically independent experiments with similar results. h, Quantification of aberrant acini in g. The results are from three biologically independent samples. Two-tailed t-tests were used for statistical analyses of aberrant clones in n = 3 biologically independent samples. ****P = 0.000045; ***P = 0.00011.
Extended Data Fig. 8 RAP2 deletion contributes to aberrant acinus growth and tumorigenesis of MCF10A cells in a YAP- and TAZ-dependent manner.
a, Representative images showing soft-agar assays of MCF10A cells. Overexpression of the constitutively active YAP(5SA) (mutation of all five LATS1- and LATS2-phosphorylation serines to alanines in YAP) strongly promotes anchorage-independent growth. Combined deletion of RAP2, MST1 and MST2, but not either group alone, also causes anchorage-independent growth of MCF10A cells. Images are representative of three biologically independent experiments with similar results. Scale bar, 35 mm. b, RAP2 inhibits tumorigenicity of MST1/2-dKO MCF10A cells. MST1/2-dKO and RAP2-MST1/2-KO MCF10A cells were injected into NOD/SCID mice. Tumour weight on day 32 after injection is presented as mean ± s.e.m. n = 6 biologically independent xenografts, ****P = 0.000025, two-tailed t-test. c, Representative tumour sizes. Only three very small xenografts were recovered from the initial 6 subcutaneous injections for MST1/2-dKO cells. Images representative of six biologically independent xenografts for each group that were initially made in the NOD/SCID mice. d, Immunohistochemistry staining with an antibody recognizing human HLA Class I. Only the acinus structures in the xenografts were formed by MCF10A cells. Stroma cells negative for HLA class I were derived from the host mice. Images are representative of two biologically independent experiments with similar results. Scale bars, 50 µm. e, Haematoxylin and eosin staining of xenografts from MST1/2-dKO MCF10A cells shows that largely hypocellular connective tissue is observed as stroma from the host animals. By contrast, RAP2-MST1/2-KO xenografts showed MCF10A cell-derived acinar and duct structures exhibiting nuclear polymorphisms, irregular nuclear contour, hyperchromasia, prominent nucleoli (star), and pathological mitosis (arrows). Images representative of two biologically independent experiments with similar results. Scale bars, 50 µm. f, Western blot showing knockdown of YAP or TAZ by lentiviral shRNAs in RAP2-MST1/2-KO MCF10A cells. shYAP#2 and shTAZ#1 were used for the xenograft studies. The image is representative of two independent experiments with similar results. g, Knockdown of YAP and TAZ inhibits tumour growth of RAP2-MST1/2-KO MCF10A cells. Tumour weight on day 32 is presented as mean ± s.e.m. Two-tailed t-test, n = 6, ****P = 0.000010. h, Xenografts from NOD/SCID mice, in which six biologically independent xenografts were generated for each group. i, Western blot showing that MCF10A-T cells are generated by expression of the oncogenic mutant H-RAS-G12V. H-RAS-G12V expression activates ERK whereas RAP2 deletion has no effect on ERK. The comparable phosphorylation levels of ERK1 and ERK2 in wild-type and RAP2-KO MCF10A-T cells indicate that the difference in xenograft growth was not due to difference in ERK1 and ERK2 activity. Image representative of two independent experiments with similar results. j, MCF10A and MCF10A-T xenografts from nude mice, in which six biologically independent xenografts were generated for each group and yielded similar results.
Extended Data Fig. 9 RAP2 deletion selectively promotes MCF7 malignancy at low stiffness in vivo.
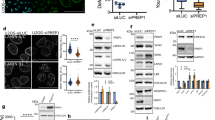
a, Western blot showing LOX overexpression in NIH3T3 fibroblasts. Ectopic LOX expression promotes cross-linking of ECM proteins and thus increases matrix stiffness28. Image representative of two independent experiments with similar results. b, RAP2 is involved in the xenograft growth enhancement caused by LOX-overexpressing fibroblasts. Xenografts were generated by co-injection of 0.4 × 106 fibroblasts (control and Flag–LOX-overexpressing) and 2.0 × 106 MCF7 cells. The xenografts were removed on day 6 and examined for collagen deposition and crosslinking and localization of YAP and TAZ. LOX overexpression led to a woven-like structure of collagen in the xenografts based on Sirius Red staining (red colour for collagen) and Masson staining (blue colour for collagen), and increased nuclear YAP and TAZ. Images representative of three biologically independent experiments with similar results. Scale bars, 50 µm. c, LOX-induced tumour growth requires RAP2. Xenografts were generated by co-injection of 0.4 × 106 NIH3T3 cells and 2.0 × 106 MCF7 cells. The growth of the xenografts with different combinations of NIH3T3 and MCF7 cells is shown as mean ± s.e.m. Deletion of RAP2A, RAP2B and RAP2C promoted MCF7 tumour growth and masked the enhancement induced by co-injected LOX-expressing fibroblasts. **two-way ANOVA test (wild-type MCF7 + wild-type NIH3T3 cells versus wild-type MCF7 +LOX-overexpressing NIH3T3 cells), n = 6 biologically independent samples, P = 0.0027. ***two-way ANOVA test (sgRAP2 MCF7 + wild-type NIH3T3 cells versus wild-type MCF7 + wild-type NIH3T3 cells), n = 6 biologically independent samples, P = 0.002. d, Tumour weights (mean ± s.e.m.). *two-tailed t-test, n = 6 biologically independent xenografts, P = 0.014 (wild-type MCF7 + wild-type NIH3T3 cells versus wild-type MCF7 + LOX-overexpressing NIH3T3 cells) or 0.029 (sgRAP2 MCF7 + wild-type NIH3T3 cells versus wild-type MCF7 + wild-type NIH3T3 cells). e, The elasticity or stiffness of the ‘soft’ and ‘stiff’ semi-synthetic hyaluronan-derived gels was measured by atomic force microscopy. Results presented as mean ± s.e.m. The measurements were made more than 55 times for each stiffness at each time. f, RAP2 inhibits expression of the YAP and TAZ target genes CTGF and CYR61 at low stiffness but not at high stiffness. Quantitative real-time PCR analyses of CTGF and CYR61 in wild-type and sgRAP2 MCF7 cells (as in Extended Data Fig. 1g) cultured in vitro for 48 h in soft or stiff hyaluronan gel. Relative mRNA levels presented as mean ± s.e.m. n = 3 biologically independent samples, two-tailed t-test. For CTGF, *sgRAP2 versus wild-type at 0.4 kPa, P = 0.020; **0.4 kPa versus 9.0 kPa for wild-type, P = 0.0032. For CYR61, *sgRAP2 versus wild-type, P = 0.025; **0.4 kPa versus 9.0 kPa for wild-type, P = 0.0033. g, Xenograft tumours. For wild-type cells grown at 0.4 kPa, eight independent xenografts were initially generated in nude mice. However, owing to animal deaths, only six xenografts were recovered. For wild-type cells grown at 9 kPa, nine independent xenografts were generated. For sgRAP2 cells grown at both 0.4 kPa and 9 kPa, eight independent xenografts were generated. h, RAP2 deletion preferentially promotes MCF7 malignancy at low stiffness. MCF7 xenografts stained with haematoxylin and eosin. Wild-type cells embedded in 0.4 kPa matrix produced mostly tubular and some cribriform structures whereas those embedded in 9 kPa matrix produced mostly solid nests, as well as more marked cellular pleomorphism and nuclear atypia. sgRAP2 cells showed more malignant architecture and morphology at 0.4 ka, while at high stiffness (9 kPa) sgRAP2 and wild-type cells exhibited similar morphology. Images representative of three biologically independent experiments with similar results. Scale bars, 50 µm. i, Immunohistochemistry for YAP and TAZ in xenografts. RAP2 deletion increased nuclear YAP and TAZ. Images representative of three biologically independent experiments with similar results. Scale bars, 50 µm.
Extended Data Fig. 10 RAP2 mediates regulation of the ECM stiffness transcriptome by the Hippo pathway.
a, MA plot of wild-type cells in low stiffness versus wild-type cells in high stiffness. Three biologically independent samples were assayed for each condition. Differentially expressed genes (adjusted P value <0.1) are coloured red. P values for deferential expression were derived using the Wald test and corrected using the Benjamini–Hochberg procedure with default functions in DESeq2. b, Dot plot showing expression of AMOTL2 and LGR5. AMOTL2 and LGR5 are YAP- and TAZ-dependent and stiffness-regulated genes. High and low denote high stiffness (40 kPa) and low stiffness (1 kPa). Data are presented as mean ± s.e.m. n = 3 biologically independent samples. c, Venn diagram comparing genes downregulated by YAP and TAZ knockout at high stiffness and genes upregulated by LATS1 and LATS2 knockout at low stiffness. d, Venn diagram comparing genes upregulated by YAP and TAZ knockout at high stiffness and genes downregulated by LATS1 and LATS2 knockout at low stiffness. e, Venn diagram comparing overlapping genes from c and genes downregulated by low stiffness. f, Venn diagram comparing overlapping genes from d and genes upregulated by low stiffness. g, Venn diagram comparing overlapping genes from e and genes upregulated by RAP2 knockout at low stiffness. h, Venn diagram comparing overlapping genes from f and genes downregulated by RAP2 knockout at low stiffness. P values for c–h: hypergeometric test. Results represent analyses of three biological replicates for each condition.
Supplementary information
Supplementary Information
This file contains Supplementary Tables 1-4 and Supplementary Figure 1 (the uncropped images of western blot data.). Supplementary Table 1 shows small GTPase screening for YAP/TAZ localization regulators (related to Fig. 1a, Extended Data Fig. 1a). HEK293A cells were transfected with plasmids expression small GTPase, and then seeded onto soft (1 kPa) and stiff (40 kPa) poly-acrylamide hydrogels. The cells were cultured on hydrogels for 24 hours, and then fixed and subjected to immunofluorescence staining with anti-YAP/TAZ antibodies. Supplementary Table 2 contains gene ontology analysis of gene regulated by ECM stiffness (related to Fig. 4i). The differentially expressed genes in HEK293A cells at 1 kPa and 40 kPa were analyzed for their biological functions. The results were derived from 3 biologically independent samples for each cell line at 1 or 40 kPa. Supplementary Table 3 contains gene ontology analysis of genes co-regulated by ECM stiffness, Hippo pathway, and RAP2 GTPase (related to Extended Data Fig. 10g,h). The genes enriched in Extended Data Fig. 10g,h were analyzed for their biological functions. The results were derived from 3 biologically independent samples for each cell line at 1 or 40 kPa. Supplementary Table 4 lists all the antibodies used in the study. The vendor information and the details of how the antibodies were used have been provided.
Rights and permissions
About this article
Cite this article
Meng, Z., Qiu, Y., Lin, K.C. et al. RAP2 mediates mechanoresponses of the Hippo pathway. Nature 560, 655–660 (2018). https://doi.org/10.1038/s41586-018-0444-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0444-0
This article is cited by
-
Hippo pathway in cell–cell communication: emerging roles in development and regeneration
Inflammation and Regeneration (2024)
-
From gene to mechanics: a comprehensive insight into the mechanobiology of LMNA mutations in cardiomyopathy
Cell Communication and Signaling (2024)
-
VPS35 promotes gastric cancer progression through integrin/FAK/SRC signalling-mediated IL-6/STAT3 pathway activation in a YAP-dependent manner
Oncogene (2024)
-
LATS2 condensates organize signalosomes for Hippo pathway signal transduction
Nature Chemical Biology (2024)
-
Liquid crystal matrix-based viscoelastic mechanical stimulation regulates nuclear localization and osteogenic differentiation of rBMSCs
Cellulose (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.