Abstract
Immune recognition of pathogen-associated molecular patterns (PAMPs) by pattern recognition receptors often activates proinflammatory NF-κB signalling1. Recent studies indicate that the bacterial metabolite d-glycero-β-d-manno-heptose 1,7-bisphosphate (HBP) can activate NF-κB signalling in host cytosol2,3,4, but it is unclear whether HBP is a genuine PAMP and the cognate pattern recognition receptor has not been identified. Here we combined a transposon screen in Yersinia pseudotuberculosis with biochemical analyses and identified ADP-β-d-manno-heptose (ADP-Hep), which mediates type III secretion system-dependent NF-κB activation and cytokine expression. ADP-Hep, but not other heptose metabolites, could enter host cytosol to activate NF-κB. A CRISPR–Cas9 screen showed that activation of NF-κB by ADP-Hep involves an ALPK1 (alpha-kinase 1)–TIFA (TRAF-interacting protein with forkhead-associated domain) axis. ADP-Hep directly binds the N-terminal domain of ALPK1, stimulating its kinase domain to phosphorylate and activate TIFA. The crystal structure of the N-terminal domain of ALPK1 and ADP-Hep in complex revealed the atomic mechanism of this ligand–receptor recognition process. HBP was transformed by host adenylyltransferases into ADP-heptose 7-P, which could activate ALPK1 to a lesser extent than ADP-Hep. ADP-Hep (but not HBP) alone or during bacterial infection induced Alpk1-dependent inflammation in mice. Our findings identify ALPK1 and ADP-Hep as a pattern recognition receptor and an effective immunomodulator, respectively.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Pandey, S., Kawai, T. & Akira, S. Microbial sensing by Toll-like receptors and intracellular nucleic acid sensors. Cold Spring Harb. Perspect. Biol. 7, a016246 (2014).
Gaudet, R. G. et al. Cytosolic detection of the bacterial metabolite HBP activates TIFA-dependent innate immunity. Science 348, 1251–1255 (2015).
Milivojevic, M. et al. ALPK1 controls TIFA/TRAF6-dependent innate immunity against heptose-1,7-bisphosphate of gram-negative bacteria. PLoS Pathog. 13, e1006224 (2017).
Zimmermann, S. et al. ALPK1- and TIFA-dependent innate immune response triggered by the Helicobacter pylori type IV secretion system. Cell Rep. 20, 2384–2395 (2017).
Auerbuch, V., Golenbock, D. T. & Isberg, R. R. Innate immune recognition of Yersinia pseudotuberculosis type III secretion. PLoS Pathog. 5, e1000686 (2009).
Bruno, V. M. et al. Salmonella typhimurium type III secretion effectors stimulate innate immune responses in cultured epithelial cells. PLoS Pathog. 5, e1000538 (2009).
Hii, C. S. et al. Interleukin-8 induction by Burkholderia pseudomallei can occur without Toll-like receptor signaling but requires a functional type III secretion system. J. Infect. Dis. 197, 1537–1547 (2008).
Litvak, Y. et al. Epithelial cells detect functional type III secretion system of enteropathogenic Escherichia coli through a novel NF-κB signaling pathway. PLoS Pathog. 13, e1006472 (2017).
Lu, Q. et al. An iron-containing dodecameric heptosyltransferase family modifies bacterial autotransporters in pathogenesis. Cell Host Microbe 16, 351–363 (2014).
Yao, Q. et al. A structural mechanism for bacterial autotransporter glycosylation by a dodecameric heptosyltransferase family. eLife 3, e03714 (2014).
Kneidinger, B. et al. Biosynthesis pathway of ADP-l-glycero-β-d-manno-heptose in Escherichia coli. J. Bacteriol. 184, 363–369 (2002).
Takatsuna, H. et al. Identification of TIFA as an adapter protein that links tumor necrosis factor receptor-associated factor 6 (TRAF6) to interleukin-1 (IL-1) receptor-associated kinase-1 (IRAK-1) in IL-1 receptor signaling. J. Biol. Chem. 278, 12144–12150 (2003).
Huang, C. C. et al. Intermolecular binding between TIFA-FHA and TIFA-pT mediates tumor necrosis factor alpha stimulation and NF-κB activation. Mol. Cell. Biol. 32, 2664–2673 (2012).
Ea, C. K., Sun, L., Inoue, J. & Chen, Z. J. TIFA activates IκB kinase (IKK) by promoting oligomerization and ubiquitination of TRAF6. Proc. Natl Acad. Sci. USA 101, 15318–15323 (2004).
Heine, M. et al. α-kinase 1, a new component in apical protein transport. J. Biol. Chem. 280, 25637–25643 (2005).
Keestra, A. M. et al. Manipulation of small Rho GTPases is a pathogen-induced process detected by NOD1. Nature 496, 233–237 (2013).
Keestra, A. M. et al. A Salmonella virulence factor activates the NOD1/NOD2 signaling pathway. MBio 2, e00266-11 (2011).
Yamaguchi, H., Matsushita, M., Nairn, A. C. & Kuriyan, J. Crystal structure of the atypical protein kinase domain of a TRP channel with phosphotransferase activity. Mol. Cell 7, 1047–1057 (2001).
Zhao, Y. & Shao, F. Diverse mechanisms for inflammasome sensing of cytosolic bacteria and bacterial virulence. Curr. Opin. Microbiol. 29, 37–42 (2016).
Tang, W. et al. d-Sedoheptulose-7-phosphate is a common precursor for the heptoses of septacidin and hygromycin B. Proc. Natl Acad. Sci. USA 115, 2818–2823 (2018).
Gall, A., Gaudet, R. G., Gray-Owen, S. D. & Salama, N. R. TIFA signaling in gastric epithelial cells initiates the cag type 4 secretion system-dependent innate immune response to Helicobacter pylori infection. MBio 8, e01168-17 (2017).
Stein, S. C. et al. Helicobacter pylori modulates host cell responses by CagT4SS-dependent translocation of an intermediate metabolite of LPS inner core heptose biosynthesis. PLoS Pathog. 13, e1006514 (2017).
Ge, J. et al. A Legionella type IV effector activates the NF-κB pathway by phosphorylating the IκB family of inhibitors. Proc. Natl Acad. Sci. USA 106, 13725–13730 (2009).
Li, H. et al. The phosphothreonine lyase activity of a bacterial type III effector family. Science 315, 1000–1003 (2007).
Zhang, L. et al. Cysteine methylation disrupts ubiquitin-chain sensing in NF-κB activation. Nature 481, 204–208 (2011).
Li, S. et al. Pathogen blocks host death receptor signalling by arginine GlcNAcylation of death domains. Nature 501, 242–246 (2013).
Zamyatina, A. et al. Efficient chemical synthesis of both anomers of ADP l-glycero- and d-glycero-d-manno-heptopyranose. Carbohydr. Res. 338, 2571–2589 (2003).
Borio, A., Hofinger, A., Kosma, P. & Zamyatina, A. Chemical synthesis of the innate immune modulator-bacterial d-glycero-β-d-manno-heptose-1,7-bisphosphate (HBP). Tetrahedr. Lett. 58, 2826–2829 (2017).
Dong, N. et al. Structurally distinct bacterial TBC-like GAPs link Arf GTPase to Rab1 inactivation to counteract host defenses. Cell 150, 1029–1041 (2012).
Xu, H. et al. Innate immune sensing of bacterial modifications of Rho GTPases by the Pyrin inflammasome. Nature 513, 237–241 (2014).
Murphy, K. C. & Campellone, K. G. Lambda Red-mediated recombinogenic engineering of enterohemorrhagic and enteropathogenic E. coli. BMC Mol. Biol. 4, 11 (2003).
Sorg, J. A., Miller, N. C., Marketon, M. M. & Schneewind, O. Rejection of impassable substrates by Yersinia type III secretion machines. J. Bacteriol. 187, 7090–7102 (2005).
Rietsch, A., Wolfgang, M. C. & Mekalanos, J. J. Effect of metabolic imbalance on expression of type III secretion genes in Pseudomonas aeruginosa. Infect. Immun. 72, 1383–1390 (2004).
Shi, J. et al. Cleavage of GSDMD by inflammatory caspases determines pyroptotic cell death. Nature 526, 660–665 (2015).
Zhao, Y. et al. The NLRC4 inflammasome receptors for bacterial flagellin and type III secretion apparatus. Nature 477, 596–600 (2011).
Sanjana, N. E., Shalem, O. & Zhang, F. Improved vectors and genome-wide libraries for CRISPR screening. Nat. Methods 11, 783–784 (2014).
Hao, Z. et al. In State-of-the-Art and Emerging Technologies for Therapeutic Monoclonal Antibody Characterization Volume 3. Defining the Next Generation of Analytical and Biophysical Techniques (eds. Schiel, J. E. et al.) Ch. 10, 289–315 (American Chemical Society, 2015).
Yang, B. et al. Identification of cross-linked peptides from complex samples. Nat. Methods 9, 904–906 (2012).
Ding, Y. H. et al. Modeling protein excited-state structures from “over-length” chemical cross-links. J. Biol. Chem. 292, 1187–1196 (2017).
Sin, Y. M., Sedgwick, A. D., Chea, E. P. & Willoughby, D. A. Mast cells in newly formed lining tissue during acute inflammation: a six day air pouch model in the mouse. Ann. Rheum. Dis. 45, 873–877 (1986).
Acknowledgements
We thank R. Zhou, X. Tan, F. Wang, H. Huang, X. Tian, Y. Ma, J. Li and the Metabolomics Facility at Tsinghua University Technology Center for Protein Sciences for technical assistance. The work was supported by grants from China NSFC (81788101), MOST of China (2017YFA0505900 and 2016YFA0501500), the Chinese Academy of Sciences (XDB08020202) and the Austrian Science Fund (P-28915).
Reviewer information
Nature thanks S. Hur and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
P.Z. and F.S. conceived the study; P.Z. did most experiments; Y.S., supervised by J.D. and D.-C.W., determined the structures and performed some biochemical assays; H.H. and W.G. performed mouse assays; S.L., X.D., Y.C. and M.D. did mass spectrometry experiments; A.B. and A.Z. synthesized the sugars; N.D., Q.W., Y.X. and P.L. provided technical assistance; P.Z., J.D. and F.S. analysed the data and wrote the manuscript. All authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Analyses of the ADP-Hep biosynthesis pathway and ADP-Hep-induced NF-κB activation.
a, Low calcium-induced secretion of the T3SS substrates in the hldE transposon mutant. yscC is required for T3SS assembly. b, Schematic of the classical ADP-Hep biosynthesis pathway in Gram-negative bacteria. c, ADP-Hep-dependent autotransporter heptosylation in ΔgmhB mutant. A fragment of the AIDA-I autotransporter (GST–AIDA-I50–600–Flag) and its heptosyltransferase (AAH) were expressed in Y. pseudotuberculosis Δ6 deleted of a gene in the ADP-Hep biosynthesis pathway. AIDA-I heptosylation was assessed using the ECL glycoprotein detection kit. d, NF-κB activation in 293T cells electroporated with HBP or H1P (obtained by treating HBP with recombinant GmhB for the indicated times). e, Effects of ALPK1 knockout on other known NF-κB pathways. NOD1, NOD2 and MYD88 were transfected into the cells. Wild-type cells and two ALPK1−/− 293T clones (KO-1/2) were treated with ADP-LD-Hep (100 μM), TNF (20 ng ml–1), C12-iE-DAP (10 ng ml–1) or MDP (10 ng ml–1). f, g, Effects of NOD1/2 deficiency on ADP-Hep-induced NF-κB activation and TIFA foci formation. Wild-type HeLa cells and two NOD1/2 double-knockout clones (DKO-1/2) were assayed. eGFP–TIFA was transfected into the cells (g). Scale bar, 20 µm. d–f, NF-κB activation was assessed by luciferase reporter assay (mean ± s.d. from three technical replicates); two-tailed unpaired Student’s t-test was performed (*P < 0.05, **P < 0.01, *** P < 0.001, NS, not significant). All data are representative of three independent experiments.
Extended Data Fig. 2 FACS-based genome-wide CRISPR–Cas9 screen identifies the ALPK1–TIFA–TRAF6 axis that mediates ADP-Hep-induced NF-κB activation.
a, Flow chart of the CRISPR–Cas9 screen for genes required for activation of NF-κB induced by extracellular ADP-LD-Hep but not TNF in 293T cells. b, Immunoblots of Flag–ALPK1 (wild type or its K/M mutant) and Flag–TIFA (wild type or its T9A mutant) expressed in knockout 293T cells. c, Requirement of T9 of TIFA for ADP-Hep-induced NF-κB activation. Flag-TIFA (wild type or T9A) was transfected into two TIFA−/− 293T clones (KO-1/2). d, Requirement of ALPK1 kinase activity for ADP-LD-Hep-induced formation of TIFA foci. eGFP–TIFA was stably expressed in wild-type or ALPK1−/− 293T cells expressing Flag–ALPK1 (wild type or the kinase-dead K/M mutant). Scale bar, 20 μm. e, Fluorescence imaging of mCherry–ALPK1 and eGFP–TIFA in 293T cells treated with or without ADP-LD-Hep. Scale bar, 10 μm. f, Phos-tag gel assay of myosin I phosphorylation in ADP-LD-Hep-treated cells. Flag–myosin I was transfected into indicated 293T cells. Immunoblots of total cell lysates separated on phos-tag gel or regular SDS gels are shown. g, Effects of ALPK1 or TIFA knockout on activation of NF-κB induced by ADP-LD or DD-Hep electroporation. ALPK1−/− and TIFA−/− 293T cells were complemented as indicated. h, ADP-LD-Hep-induced co-immunoprecipitation of TIFA with ALPK1 and TRAF6 in transfected 293T cells. c, g, NF-κB activation was assessed by luciferase reporter assay (mean ± s.d. from three technical replicates); two-tailed unpaired Student’s t-test was performed (*P < 0.05, **P < 0.01, ***P < 0.001, NS, not significant). d, e, Confocal images with Hoechst-stained nuclei. All data are representative of three independent experiments.
Extended Data Fig. 3 The ALPK1–TIFA pathway mediates T3SS-dependent and -independent activation of NF-κB by various bacterial pathogens.
a, b, Requirement of ALPK1 kinase activity and TIFA for Y. pseudotuberculosis-induced activation of NF-κB. Wild-type or ALPK1−/− 293T cells expressing Flag–ALPK1 (wild type or K/M) or TIFA−/− 293T cells expressing Flag–TIFA were infected with indicated Y. pseudotuberculosis strains. NF-kB activation was assessed by luciferase reporter assay (a) or eGFP reporter expression (b). Scale bar, 50 μm (b). c–e, Requirement of ALPK1 kinase activity for Y. pseudotuberculosis (or S. flexneri 2457T)-induced phosphorylation of TIFA (c, d) or formation of eGFP–TIFA foci (e). Flag– or eGFP–TIFA was expressed in wild-type or ALPK1−/− 293T cells expressing Flag–ALPK1 (wild type or K/M). TIFA phosphorylation was assessed by anti-pT9-TIFA immunoblotting (c, d). Y. pseudotuberculosis Δ6 was labelled with mCherry (scale bar, 20 μm) (e). f, g, NF-κB luciferase reporter activation and formation of eGFP–TIFA foci induced by T3SS-negative bacteria. Wild-type, ALPK1−/− or Flag–ALPK1-complemented ALPK1−/− 293T cells were infected with DAEC 2787, ETEC H10407, B. cenocepacia J2315 or their ΔhldE mutants. Scale bar, 20 μm. a, f, Luciferase data are shown as mean ± s.d. from three technical replicates; two-tailed unpaired Student’s t-test was performed (*P < 0.05, **P < 0.01, *** P < 0.001). b, e, g, Confocal images with Hoechst-stained nuclei. All data shown are representative of three independent experiments.
Extended Data Fig. 4 Co-expression of ALPK1-NTD and ALPK1-KD can support ADP-Hep and Y. pseudotuberculosis-induced activation of NF-κB and conservation of ADP-Hep-binding residues in ALPK1-NTD.
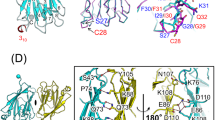
a, c, Immunoblots of ALPK1 N- and C-terminal truncation mutants expression in indicated 293T cells. NF-κB luciferase reporter activation induced by ADP-LD-Hep and Y. pseudotuberculosis Δ6 are in Fig. 2h. b, Mapping the minimal N- and C-terminal regions of ALPK1 sufficient for ADP-LD-Hep-induced activation of NF-κB. Fluorescence images of NF-κB–eGFP expression are shown. Scale bar, 50 μm. d, e, Assay of ALPK1-NTD and ALPK1-KD co-expression in mediating ADP-LD-Hep (d) or Y. pseudotuberculosis (e)-induced TIFA phosphorylation. Immunoblots of total cell lysates using indicated antibodies are shown. f, Multiple sequence alignment of the NTDs of ALPK1 from indicated organisms. Red residues are those involved in ADP-Hep binding, revealed by the crystal structure of human ALPK1-NTD–ADP-LD-Hep complex (Fig. 3g). Data shown are representative of three (a–c) or two (d, e) independent experiments.
Extended Data Fig. 5 Direct binding of ADP-Hep to ALPK1-NTD.
a, E. coli-purified ALPK1-(N+K) complex on a Coomassie blue-stained SDS–PAGE gel. b–e, HPLC–MS fractionation, NF-κB-inducing activity and mass spectrometry of small-molecule extracts from His6–ALPK1-NTD purified from wild-type E. coli BL21 (DE3). The small-molecule extracts, obtained by protein denaturing and precipitation (95 °C for 5 min), were analysed by HPLC–MS with 17 fractions obtained (b). The 17 fractions were used to treat 293T cells (c) or cells expressing Flag–TIFA (d); NF-κB luciferase activity (mean ± s.d. from three technical replicates) and anti-pT9-TIFA immunoblotting are in c and d, respectively. e, Mass spectrometry of fraction 6 identified three major ions corresponding to AMP, ADP and ADP-Hep. f, LC–MS/MS of ADP-Hep in fraction 6 (b, e) or synthetic ADP-LD-Hep standard. g, MS/MS spectra of the [M+H]+ product ions of ADP-Hep in fraction 6 in comparison with those of synthetic ADP-LD-Hep. The heptose of ADP-Hep was not shown owing to neutral loss. h, BLI assay of ADP-LD-Hep or S7P binding to ALPK1-(N+K). ALPK1-(N+K) complexes purified from E. coli BL21 (DE3) ΔhldE were biotinylated in vitro. Sensorgrams of the binding to ALPK1-(N+K) by different concentrations of the indicated sugar (colour lines) are shown. Grey lines are from model fits. Data shown are representative of three (a, h) or two (b–g) independent experiments.
Extended Data Fig. 6 HBP binding is insufficient to render ALPK1-KD competent for substrate recognition and phosphate transfer.
a, BLI assay of HBP binding to in vitro-biotinylated apo-ALPK1-(N+K). Sensorgrams of the binding in different concentrations of HBP (colour lines) are shown. Grey lines are from model fits. b, Excess HBP competitively inhibits activation of ALPK1 by ADP-Hep. Flag–ALPK1 purified from 293T cells was incubated with purified TIFA–His6 and ADP-Hep in the presence of a titrating amount of HBP. c, d, Effects of HBP binding on ALPK1 phosphorylation and recognition of a 15-residue peptide substrate derived from TIFA. The N-terminal 15-residue sequences of TIFA were fused to GST (GST–TIFA.N15). d, GST-pulldown assay of ALPK1-(N+K) binding to GST–TIFA.N15. e, Effect of HBP binding on ATP hydrolysis activity of ALPK1. Apo-ALPK1-(N+K) (wild type or K/M) was incubated with HBP or ADP-LD-Hep, and further reacted with ATP. Percentages of ATP consumption at indicated reaction conditions are shown (mean ± s.d. from three technical replicates). Two-tailed unpaired Student’s t-test was performed (*P < 0.05; **P < 0.01; ***P < 0.001). f, Graphical representation of DSS-crosslinked residues between ALPK1-NTD and ALPK1-KD identified by chemical cross-linking coupled with mass spectrometry. Apo-ALPK1-(N+K) was left untreated, or incubated with HBP or ADP-LD-Hep. Crosslinking connections are depicted by straight lines, and the corresponding raw mass spectrometry data are in Supplementary Table 3. g, Crystal structure of TRPM7 kinase domain in complex with AMPPNP (PDB code, 1IA9). TRPM7 is shown in cartoons and AMPPNP is in sticks. The loop containing K1727 and I1736 (shown in sticks, corresponding to K1140 and K1149 in human ALPK1, respectively) is in yellow. Phosphorylation of TIFA and GST-TIFA.N15 was assayed by anti-pT9-TIFA immunoblotting (b, c). Apo-ALPK1-(N+K), TIFA–His6, and GST–TIFA.N15 were purified from E. coli BL21 (DE3) ΔhldE (a–f). Data shown are representative of three (a–e) and two (f) independent experiments.
Extended Data Fig. 7 HBP can be transformed into ALPK1 activation-competent ADP-heptose 7-P by bacterial or host adenylyltransferase.
a, b, Requirement of ALPK1 kinase activity for cytosolic HBP-induced phosphorylation of TIFA (a) and activation of NF-κB (b). HBP was electroporated into wild-type or ALPK1−/− 293T cells expressing Flag–ALPK1 (wild-type or K/M). c, d, Enzymatically synthesized HBP could induce ALPK1-mediated NF-κB activation (c) and TIFA phosphorylation in vitro (d). S7P was reacted with recombinant GmhA or HldE or both. Following protein denaturing and precipitation, reaction supernatants were added to wild-type or ALPK1−/− 293T cells containing an empty vector or Flag–ALPK1 (c). Enzymatically synthesized HBP product (HBPenzymatic), synthetic ADP-LD-Hep or HBP was incubated with ALPK1-(N+K) in the presence of TIFA–His6 (d). e, Schematic of HldE adenylyltransferase synthesis of ADP-Hep and ADP-heptose 7-P from H1P and HBP, respectively. f, g, NF-κB activation by (f) and LC–MS of (g) enzymatically synthesized HBP product. Indicated reaction products were added to wild-type or ALPK1−/− 293T cells (f) or analysed by LC–MS (g). h, i, NF-κB (h) and in vitro ALPK1 activation (i) by ADP-heptose 7-P. HPLC-purified ADP-heptose 7-P was added to wild-type or ALPK1−/− 293T cells expressing Flag–ALPK1 (wild-type or K/M). i, Equal amounts of the indicated sugars were incubated with ALPK1-(N+K) in the presence of TIFA–His6. j, Ultra-performance liquid chromatography with tandem mass spectrometry of the reaction product of HBP and NMNAT1. Purified ADP-heptose 7-P, synthesized by reacting HBP with HldE, was used as the standard. k, Effect of NMNAT1 overexpression on activation of NF-κB by electroporation of HBP into 293T cells. The immunoblots show NMNAT1 expression. Apo-ALPK1-(N+K) was purified from E. coli BL21 (DE3) ΔhldE and phosphorylation of TIFA was assessed by anti-pT9-TIFA immunoblotting (d, i). b, c, f, h, k, NF-κB activation was assessed by luciferase reporter assay (mean ± s.d. from three technical replicates); two-tailed unpaired Student’s t-test was performed (*P < 0.05; **P < 0.01; ***P < 0.001). Data shown are representative of three (a–d, h, i, k) or two (f, g, j) independent experiments.
Extended Data Fig. 8 ALPK1 Q67A/Y68A mutants that can discriminate cytosolic HBP from ADP-Hep resist activation by ADP-heptose 7-P.
a–c, Effects of ALPK1 Q67A, Y68A or Q67A/Y68A double mutations on activation of NF-κB by HBP, ADP-Hep and ADP-heptose 7-P. ALPK1−/− 293T cells expressing the indicated Flag–ALPK1 mutants were electroporated with excess HBP or ADP-LD-Hep (a), or treated with excess ADP-heptose 7-P or ADP-LD-Hep (c). Anti-Flag immunoblots (b) show expression of the ALPK1 mutants. d, In vitro TIFA phosphorylation assay of ALPK1 Q67A/Y68A activation by HBP, ADP-heptose 7-P, or ADP-LD-Hep. Flag–ALPK1 Q67A/Y68A was purified from 293T cells. e, f, In vitro TIFA phosphorylation assay of ALPK1 Q67A/Y68A activation by small-molecule extracts from HBP-electroporated cells (SEHBP). 293T cells electroporated with HBP were used to prepare the small-molecule extracts, and synthetic HBP was included as the control. Apo-ALPK1-(N+K) (e) and Flag–ALPK1 (f) were purified from E. coli BL21 (DE3) ΔhldE and 293T cells, respectively. g, Effects of ALPK1 Q67A/Y68A double mutations on Y. pseudotuberculosis-induced activation of NF-κB. ALPK1−/− 293T cells expressing Flag–ALPK1 (wild-type or the Q67A/Q68A mutant) were left uninfected or infected with indicated Y. pseudotuberculosis strains. a, c, g, NF-κB activation was assessed by luciferase reporter assay (mean ± s.d. from three technical replicates); two-tailed unpaired Student’s t-test was performed (*P < 0.05; **P < 0.01; ***P < 0.001). All data are representative of at least two independent experiments.
Extended Data Fig. 9 ADP-Hep but not HBP induces Alpk1-dependent cytokine expression in mice.
HBP or ADP-LD-Hep (2 mg kg–1) or the saline control were injected into the dorsal air pouches of wild-type (a–d) or Alpk1−/− (d) mice (C57BL/6). n (biologically independent animals) = 5 for each group of treatment. Cytokine profiling was determined by the multiplex immunoassay (a–c) or ELISA (d). a, b, Heat maps of indicated cytokine concentrations in the air pouch (a) and the serum (b) of injected mice. c, d, Cytokine concentrations in the serum shown as mean ± s.e.m. (two-tailed unpaired Student’s t test, *P < 0.05, **P < 0.01, ***P < 0.001, ns, not significant). All data are representative of two independent experiments.
Supplementary information
Supplementary Information
This file contains Supplementary Text, Supplementary Tables 1-3 and the uncropped blots from the main and Extended Data Figures.
Source data
Rights and permissions
About this article
Cite this article
Zhou, P., She, Y., Dong, N. et al. Alpha-kinase 1 is a cytosolic innate immune receptor for bacterial ADP-heptose. Nature 561, 122–126 (2018). https://doi.org/10.1038/s41586-018-0433-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0433-3
Keywords
This article is cited by
-
Bacterial peptidoglycan acts as a digestive signal mediating host adaptation to diverse food resources in C. elegans
Nature Communications (2024)
-
Characterization of the ADP-β-d-manno-heptose biosynthetic enzymes from two pathogenic Vibrio strains
Applied Microbiology and Biotechnology (2024)
-
Alpha‐kinase1 promotes tubular injury and interstitial inflammation in diabetic nephropathy by canonical pyroptosis pathway
Biological Research (2023)
-
Genome-wide CRISPR screens identify ILF3 as a mediator of mTORC1-dependent amino acid sensing
Nature Cell Biology (2023)
-
In vitro kinase assay reveals ADP-heptose-dependent ALPK1 autophosphorylation and altered kinase activity of disease-associated ALPK1 mutants
Scientific Reports (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.