Abstract
The conserved and essential DEAD-box RNA helicase Ded1p from yeast and its mammalian orthologue DDX3 are critical for the initiation of translation1. Mutations in DDX3 are linked to tumorigenesis2,3,4 and intellectual disability5, and the enzyme is targeted by a range of viruses6. How Ded1p and its orthologues engage RNAs during the initiation of translation is unknown. Here we show, by integrating transcriptome-wide analyses of translation, RNA structure and Ded1p–RNA binding, that the effects of Ded1p on the initiation of translation are connected to near-cognate initiation codons in 5′ untranslated regions. Ded1p associates with the translation pre-initiation complex at the mRNA entry channel and repressing the activity of Ded1p leads to the accumulation of RNA structure in 5′ untranslated regions, the initiation of translation from near-cognate start codons immediately upstream of these structures and decreased protein synthesis from the corresponding main open reading frames. The data reveal a program for the regulation of translation that links Ded1p, the activation of near-cognate start codons and mRNA structure. This program has a role in meiosis, in which a marked decrease in the levels of Ded1p is accompanied by the activation of the alternative translation initiation sites that are seen when the activity of Ded1p is repressed. Our observations indicate that Ded1p affects translation initiation by controlling the use of near-cognate initiation codons that are proximal to mRNA structure in 5′ untranslated regions.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Sharma, D. & Jankowsky, E. The Ded1/DDX3 subfamily of DEAD-box RNA helicases. Crit. Rev. Biochem. Mol. Biol. 49, 343–360 (2014).
Bol, G. M., Xie, M. & Raman, V. DDX3, a potential target for cancer treatment. Mol. Cancer 14, 188 (2015).
Oh, S. et al. Medulloblastoma-associated DDX3 variant selectively alters the translational response to stress. Oncotarget 7, 28169–28182 (2016).
Pugh, T. J. et al. Medulloblastoma exome sequencing uncovers subtype-specific somatic mutations. Nature 488, 106–110 (2012).
Snijders Blok, L. et al. Mutations in DDX3X are a common cause of unexplained intellectual disability with gender-specific effects on Wnt signaling. Am. J. Hum. Genet. 97, 343–352 (2015).
Valiente-Echeverría, F., Hermoso, M. A. & Soto-Rifo, R. RNA helicase DDX3: at the crossroad of viral replication and antiviral immunity. Rev. Med. Virol. 25, 286–299 (2015).
Ingolia, N. T., Brar, G. A., Rouskin, S., McGeachy, A. M. & Weissman, J. S. The ribosome profiling strategy for monitoring translation in vivo by deep sequencing of ribosome-protected mRNA fragments. Nat. Protocols 7, 1534–1550 (2012).
Putnam, A. A. et al. Division of labor in an oligomer of the DEAD-box RNA helicase Ded1p. Mol. Cell 59, 541–552 (2015).
Burckin, T. et al. Exploring functional relationships between components of the gene expression machinery. Nat. Struct. Mol. Biol. 12, 175–182 (2005).
Chuang, R. Y., Weaver, P. L., Liu, Z. & Chang, T. H. Requirement of the DEAD-box protein Ded1p for messenger RNA translation. Science 275, 1468–1471 (1997).
Sen, N. D., Zhou, F., Ingolia, N. T. & Hinnebusch, A. G. Genome-wide analysis of translational efficiency reveals distinct but overlapping functions of yeast DEAD-box RNA helicases Ded1 and eIF4A. Genome Res. 25, 1196–1205 (2015).
Heyer, E. E. & Moore, M. J. Redefining the translational status of 80S monosomes. Cell 164, 757–769 (2016).
Hinnebusch, A. G., Ivanov, I. P. & Sonenberg, N. Translational control by 5′-untranslated regions of eukaryotic mRNAs. Science 352, 1413–1416 (2016).
Ingolia, N. T., Ghaemmaghami, S., Newman, J. R. & Weissman, J. S. Genome-wide analysis in vivo of translation with nucleotide resolution using ribosome profiling. Science 324, 218–223 (2009).
Brar, G. A. et al. High-resolution view of the yeast meiotic program revealed by ribosome profiling. Science 335, 552–557 (2012).
Kolitz, S. E., Takacs, J. E. & Lorsch, J. R. Kinetic and thermodynamic analysis of the role of start codon/anticodon base pairing during eukaryotic translation initiation. RNA 15, 138–152 (2008).
Berthelot, K., Muldoon, M., Rajkowitsch, L., Hughes, J. & McCarthy, J. E. Dynamics and processivity of 40S ribosome scanning on mRNA in yeast. Mol. Microbiol. 51, 987–1001 (2004).
Zubradt, M. et al. DMS-MaPseq for genome-wide or targeted RNA structure probing in vivo. Nat. Methods 14, 75–82 (2017).
Huppertz, I. et al. iCLIP: protein-RNA interactions at nucleotide resolution. Methods 65, 274–287 (2014).
Hinnebusch, A. G. The scanning mechanism of eukaryotic translation initiation. Annu. Rev. Biochem. 83, 779–812 (2014).
Gavin, A. C. et al. Proteome survey reveals modularity of the yeast cell machinery. Nature 440, 631–636 (2006).
Krogan, N. J. et al. High-definition macromolecular composition of yeast RNA-processing complexes. Mol. Cell 13, 225–239 (2004).
Kozak, M. Downstream secondary structure facilitates recognition of initiator codons by eukaryotic ribosomes. Proc. Natl Acad. Sci. USA 87, 8301–8305 (1990).
Gao, Z. et al. Coupling between the DEAD-box RNA helicases Ded1p and eIF4A. eLife 5, e16408 (2016).
Hilliker, A., Gao, Z., Jankowsky, E. & Parker, R. The DEAD-box protein Ded1 modulates translation by the formation and resolution of an eIF4F-mRNA complex. Mol. Cell 43, 962–972 (2011).
Putnam, A. A. & Jankowsky, E. AMP sensing by DEAD-box RNA helicases. J. Mol. Biol. 425, 3839–3845 (2013).
Jain, S. et al. ATPase-modulated stress granules contain a diverse proteome and substructure. Cell 164, 487–498 (2016).
Andrews, S. J. & Rothnagel, J. A. Emerging evidence for functional peptides encoded by short open reading frames. Nat. Rev. Genet. 15, 193–204 (2014).
Tang, H. L. et al. Translation of a yeast mitochondrial tRNA synthetase initiated at redundant non-AUG codons. J. Biol. Chem. 279, 49656–49663 (2004).
Aylett, C. H., Boehringer, D., Erzberger, J. P., Schaefer, T. & Ban, N. Structure of a yeast 40S–eIF1–eIF1A–eIF3–eIF3j initiation complex. Nat. Struct. Mol. Biol. 22, 269–271 (2015).
Hu, W., Sweet, T. J., Chamnongpol, S., Baker, K. E. & Coller, J. Co-translational mRNA decay in Saccharomyces cerevisiae. Nature 461, 225–229 (2009).
Smith, J. E. et al. Translation of small open reading frames within unannotated RNA transcripts in Saccharomyces cerevisiae. Cell Reports 7, 1858–1866 (2014).
Goecks, J., Nekrutenko, A. & Taylor, J. Galaxy: a comprehensive approach for supporting accessible, reproducible, and transparent computational research in the life sciences. Genome Biol. 11, R86 (2010).
Andreev, D. E. et al. Translation of 5′ leaders is pervasive in genes resistant to eIF2 repression. eLife 4, e03971 (2015).
Matz, M. et al. Amplification of cDNA ends based on template-switching effect and step-out PCR. Nucleic Acids Res. 27, 1558–1560 (1999).
Nagalakshmi, U. et al. The transcriptional landscape of the yeast genome defined by RNA sequencing. Science 320, 1344–1349 (2008).
König, J. et al. iCLIP reveals the function of hnRNP particles in splicing at individual nucleotide resolution. Nat. Struct. Mol. Biol. 17, 909–915 (2010).
Subtelny, A. O., Eichhorn, S. W., Chen, G. R., Sive, H. & Bartel, D. P. Poly(A)-tail profiling reveals an embryonic switch in translational control. Nature 508, 66–71 (2014).
Licatalosi, D. D. et al. HITS-CLIP yields genome-wide insights into brain alternative RNA processing. Nature 456, 464–469 (2008).
Park, D., Morris, A. R., Battenhouse, A. & Iyer, V. R. Simultaneous mapping of transcript ends at single-nucleotide resolution and identification of widespread promoter-associated non-coding RNA governed by TATA elements. Nucleic Acids Res. 42, 3736–3749 (2014).
Robinson, J. T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Eden, E., Navon, R., Steinfeld, I., Lipson, D. & Yakhini, Z. GOrilla: a tool for discovery and visualization of enriched GO terms in ranked gene lists. BMC Bioinformatics 10, 48 (2009).
Gruber, A. R., Bernhart, S. H. & Lorenz, R. The ViennaRNA web services. Methods Mol. Biol. 1269, 307–326 (2015).
The R Core Team. A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria., http://www.R-project.org (2013).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. Series B Stat. Methodol. 57, 289–300 (1995).
Miyasaka, H. The positive relationship between codon usage bias and translation initiation AUG context in Saccharomyces cerevisiae. Yeast 15, 633–637 (1999).
Nielsen, K. H. et al. Synergistic activation of eIF4A by eIF4B and eIF4G. Nucleic Acids Res. 39, 2678–2689 (2011).
Floor, S. N., Condon, K. J., Sharma, D., Jankowsky, E. & Doudna, J. A. Autoinhibitory interdomain interactions and subfamily-specific extensions redefine the catalytic core of the human DEAD-box protein DDX3. J. Biol. Chem. 291, 2412–2421 (2016).
Siepel, A. et al. Evolutionarily conserved elements in vertebrate, insect, worm, and yeast genomes. Genome Res. 15, 1034–1050 (2005).
Acknowledgements
This study was supported by the National Institutes of Health (NIH) (GM118088 to E.J., GM107331 to D.D.L.) and by a postdoctoral fellowship from the German Research Council (GU 1146/1-1 to U.-P.G.). D.P.B. and J.S.W. are investigators of the Howard Hughes Medical Institute. We thank M. Adams for help with the initial data analysis; N. Al-Huseini and J. Coller for discussion and materials; A. Tambe, M. Hannigan, and R. Backofen for advice on bioinformatic data analysis; B. Klaus for critical advice on statistical analysis and M. Hentze for discussion.
Reviewer information
Nature thanks P. Todd and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
U.-P.G.: conceptualization, material generation, ribosome profiling, XL-RAP–seq, iCLIP, molecular biology techniques, bioinformatic analysis, statistical analysis and manuscript writing; D.E.W.: conceptualization and iCLIP; M.M.Z.: conceptualization, DMS-MaPseq and bioinformatic analysis; F.A.T.: iCLIP, bioinformatic analysis, western blots; B.N.S.: material generation and northern blots; L.L.Z. and D.D.L.: iCLIP; G.A.B.: material generation and meiosis strains; D.P.B.: conceptualization and study supervision; J.S.W.: conceptualization and study supervision.; E.J.: conceptualization, study supervision and manuscript writing. All authors commented on and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 mRNA expression, and translation profiles in wild-type DED1 and ded1-95.
a, Correlation of ribosome footprint counts between two biological replicates in wild-type DED1 at 30 °C (n = 5,523). b, Correlation of mRNA expression levels between two biological replicates in wild-type DED1 at 30 °C (n = 5,372). c, Correlation of ribosome footprint counts between two biological replicates in wild-type DED1, 5 min after temperature shift to 37 °C (n = 5,523). d, Correlation of mRNA expression levels between two biological replicates in wild-type DED1, 5 min after temperature shift to 37 °C (n = 5,372). e, Correlation of ribosome footprint counts between two biological replicates in ded1-95 at 30 °C (n = 5,523). f, Correlation of mRNA expression levels between two biological replicates in ded1-95 at 30 °C (n = 5,372). g, Correlation of ribosome footprint counts between two biological replicates in ded1-95, 5 min after temperature shift to 37 °C (n = 5,523). h, Correlation of mRNA expression levels between two biological replicates in ded1-95, 5 min after temperature shift to 37 °C (n = 5,372). i, Correlation of mRNA expression levels between wild-type DED1 and ded1-95 at 30 °C (n = 2,976). Each data point represents the average of at least two replicates. j, Correlation of translational efficiencies between wild-type DED1 and ded1-95 at 30 °C (n = 2,976). Each data point represents the average of at least two replicates. k, Representative polysome profiles of wild-type DED1 and ded1-95 strains at 30 °C and 5 min after shift to 37 °C. Similar results were obtained in three independent experiments. l, Changes in translational efficiencies (∆TE) for mRNAs in ded1-95, compared to wild-type DED1, 5 min after temperature shift (mean of two biological replicates). The dotted line indicates no change. m, Fraction of 18S rRNA in polysome fractions, compared to the entire sample, at 30 °C and 5 min after temperature shift to 37 °C. Each bar represents the average of three independent experiments. Empty circles represent each replicate. n, Cumulative distribution of translational efficiencies of wild-type DED1 and ded1-95, 5 min after temperature shift to 37 °C (n = 2,976). Each data point represents the average of at least two replicates.
Extended Data Fig. 2 A subset of mRNAs is largely insensitive to Ded1p.
a, mRNA groups defined by Gene Ontology term with translation that is strongly affected (green) or largely unaffected by Ded1p (blue). Box plots (group median) of change in translational efficiencies. Box boundaries, upper and lower quartiles; error bars, 1.5 × interquartile range. The black box plot marks changes in translational efficiencies for all mRNAs (Fig. 1b). mRNAs for each Gene Ontology term were extracted from the Saccharomyces Genome Database (https://www.yeastgenome.org). The false discovery rate q value indicates the enrichment P value according to a hypergeometric model after correction for multiple testing using the Benjamini and Hochberg method46. b, Box plots (as in a) of 5′ UTR lengths and median of the shift in the normalized centre of ribosome density (Fig. 1b) for Gene Ontology term defined mRNA groups, colour-coded as in a.
Extended Data Fig. 3 Activation of ATIS in ded1-95 upon temperature shift.
a, Fraction of ribosome footprints on 5′ UTRs in wild-type DED1 and ded1-95, (5 min, 37 °C, n = 3,273). The red line indicates the mean. Statistical significance for the difference between ded1-95 and wild-type DED1: P = 1.2 × 10−119 (two tailed t-test). A similar result was obtained in an independent replicate (P = 5.4 × 10−47). b, Changes in the fraction of ribosomes on 5′ UTRs for all mRNAs (n = 2,660) in wild-type DED1 compared to ded1-95, 5 min after temperature shift to 37 °C. The values on the x-axis represent the ratio (log2) of the fraction of ribosomes on each 5′ UTR in the wild type, divided by the fraction of ribosomes on the same 5′ UTR in ded1-95. Each value represents the mean of two independent biological replicates. c, Representative northern blots of PSA1 after sucrose gradient centrifugation for wild-type DED1 and ded1-95, at 30 °C. A similar result was obtained in an independent biological replicate. d, Fraction of ribosome footprints on 5′ UTRs in wild-type DED1 and ded1-95, (5 min, 37 °C), measured only in 80S monosomes (n = 973, reads from two independent experiments combined). Statistical significance for the difference between ded1-95 and wild-type DED1: P = 1.2×10−50 (two tailed t-test). e, Mean ribosome occupancy within 10 nt 3′ and 5′ of the high-confidence ATIS in 5′ UTRs (moving average ± 1 nt, 5 min, 37 °C), measured without cycloheximide. f, Mean ribosome occupancy within 10 nt 3′ and 5′ of the high-confidence ATIS in 5′ UTRs (n = 274) in 80S monosomes (moving average ± 1 nt, 5 min, 37 °C). g, Mean ribosome occupancy within 10 nt 3′ and 5′ of all near-cognate initiation codons (n = 61,614; excluding medium-confidence ATIS) in 5′ UTRs (moving average ± 1 nt, 5 min, 37 °C). h, Ribosome occupancy of 3′ ends in small upstream open reading frames (smORFs) initiating at high-confidence ATIS in ded1-95 before (t = 0) and after (t = 5 min) temperature shift. smORFs were included in this analysis if its length exceeds three codons and if the smORF terminates at least 11 nt upstream of the main AUG codon (n = 76). i, Ribosome occupancy of 3′ ends of smORFs defined in e in wild-type DED1 before (t = 0) and after (t = 5 min) temperature shift. j, Ribosome occupancy 4 nt 5′ and 20 nt 3′ of high-confidence ATIS on 5′ UTRs (n = 274) for ded1-95, 5 min after temperature shift. The dashed lines indicate the first nucleotide of the marked in-frame codons. k, Ribosome occupancy 4 nt 5′ and 20 nt 3′ of the main AUG (mAUG) of mRNAs containing high-confidence ATIS on 5′ UTRs for ded1-95, 5 min after temperature shift. For mRNAs with multiple high-confidence ATIS in their 5′ UTR, the main AUG was counted only once. The dashed lines indicate the first nucleotide of the marked in-frame codons. l, Ribosome occupancy 4 nt 5′ and 20 nt 3′ of high-confidence ATIS-matched random positions (averaged from five randomizations) on 5′ UTRs (n = 274) for ded1-95, 5 min after temperature shift. The dashed line indicates the first nucleotide.
Extended Data Fig. 4 Characteristics of small open reading frames associated with activated ATISs.
a, Enrichment or depletion of each near-cognate codon in ATISs over the background distribution of the codon. P values determined using a two-tailed t-test. b, Mean translation initiation site score (positions −6 to +6, excluding +1 to +3), calculated according to previously published methods47, for high-stringency ATIS (n = 274, red), and TIS of main ORFs (n = 4,972, grey). A TIS score exceeding 0.01 is considered a potential translational initiation site14. c, Changes in translational efficiencies (ΔTE) for mRNAs in ded1-95, compared to wild-type DED1, 5 min after temperature shift for all mRNAs (Fig. 1b) and ATIS-containing mRNAs. d, Length of the small open reading frames (smORFs) associated with ded1-95-activated ATIS. smORFs encoding N-terminal extensions were excluded from the analysis. e, Type of smORFs associated with ded1-95-activated ATIS. The bar graphs show the fraction of smORFs that falls into each category. The distribution of changes in translation efficiency (ΔTE) for RNAs with each type of smORF did not differ significantly.
Extended Data Fig. 5 mRNA structure unwinding by Ded1p in cells using DMS MaPSeq.
a, Schematic for DMS-MaPseq approach to monitor RNA structure unwinding by Ded1p. All DMS-MapSeq experiments were performed 5 min after temperature shift. b, Representative DMS MaPSeq tracks in the PSA1 5′ UTR 5 min after temperature shift for wild-type DED1 (grey) and ded1-95 (red). Bars show normalized reverse transcription stops. A similar result was obtained in an independent replicate. The average Pearson’s correlation coefficient of DMS-MaPseq counts per 5′ UTR between two replicates (5 min after temperature shift to 37 °C) were R = 0.57 (n = 864) for the wild type and R = 0.63 (n = 692) for ded1-95. The ribosome occupancy track for ded1-95 is shown for reference. c, Unwinding of mRNA structure by Ded1p for different mRNA regions. Similar results were obtained in two independent experiments.
Extended Data Fig. 6 XL-RAP–seq and iCLIP
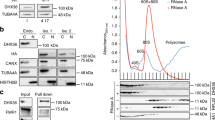
a, Correlation of sequence reads (fragments per kilobase of exon per million fragments mapped, FPKM) per mRNA for two independent biological XL-RAP–seq replicates (n = 2,992) b, Fraction of mRNA (40%) and rRNA (44%) cross-linked to wild-type Ded1p as a fraction of all sequencing reads (mean of two independent experiments). n = 4,280 mRNAs exceed a minimal read count of FPKM ≥ 10. c, Correlation of the number of reverse transcription stops (FPKM) per mRNA for the two independent iCLIP approaches. Replicate 1, FLAG-tagged Ded1p; replicate 2, HTBH-tagged Ded1p; n = 4,007.
Extended Data Fig. 7 Ded1p binding sites on 18S RNA and mRNAs.
a, Ded1p binding sites on helix 720 (exit) and helix 16 (entry) (red) are in close proximity to the binding sites of eIF3c (purple) and eIF3b (green) on the 40S ribosomal subunit42 (rRNA, grey; ribosomal proteins, cyan). b, Localization of Ded1p (apricot) on helix 16 of the PIC. Schematic model of the yeast PIC with eIF3 (http://www.bangroup.ethz.ch/research/eukaryotic_translation_initiation.html and references therein); the positioning of eIF4G572-853 and eIF4A is derived from previously published work48. The position of the eIF4G C terminus is hypothetical. The helicase core of Ded1p was modelled in analogy to the DDX3 core structure49, with the RNA binding site in contact with helix 16 at the main iCLIP cross-link sites. The position of the low-complexity N terminus of Ded1p is hypothetical. c, Sequence logo of Ded1p binding sites on mRNAs. Sets of 104 binding sites were randomly sampled from all Ded1p cross-linking sites and used as input to create a sequence logo (http://weblogo.berkeley.edu). All subsets yielded essentially the same sequence logo as shown here. Position zero denotes the reverse transcription stop.
Extended Data Fig. 8 Representative northern blots of PSA1 (Δ2°) and ATP5 (ΔATIS, Δ2°) for wild-type DED1 and ded1-95.
a, Representative RNA blots (5 min, 37 °C) for the PSA1 mRNA with altered secondary structure, 3′ of the ATIS (Δ2°). Similar results were obtained in three independent biological replicates. b, Quantification of RNA blots for accumulation of the PSA1 Δ2° mRNA in monosomes in ded1-95, compared to wild-type DED1. The line indicates the average. The P value for the difference in monosome accumulation was determined using a one-tailed t-test. c, Representative ribosome profiling tracks for the 5′ UTR of ATP5 in wild-type DED1 and ded1-95 (at 30 °C and after temperature shift). The near-cognate initiation codon is highlighted by a star. For comparison, Ded1p cross-linking (yellow track) and differential DMS-MaPseq tracks (log2(ded1-95/wild type), unwound mRNA regions are marked by red bars and have negative values) for the 5′ UTR of the ATP5 mRNA are also shown. Similar results were obtained in two independent experiments. d, DMS-MapSeq-constrained secondary-structure model of a fragment of the ATP5 mRNA 5′ UTR. The ATIS is marked by a line. Ded1p cross-linking (iCLIP) and unwinding (DMS-MapSeq) for each nucleotide are indicated. The ratio of normalized DMS-MapSeq counts of wild type/ded1-95 in two categories: yellow triangles, 0.6 – 1.0 (moderately unwound) and red triangles, >1.0 (strongly unwound). e, Representative RNA blots (5 min, 37 °C) for wild-type ATP5 mRNA and the same mRNA with mutations in the ATIS (ΔATIS) or with altered secondary structure 3′ of the ATIS (Δ2°) for wild-type DED1 and ded1-95. Similar results were obtained in three independent experiments. f, Quantification of RNA blots for accumulation of the ATP5 ΔATIS and Δ2° mRNA in monosomes in ded1-95, compared to wild-type DED1. Lines indicate averages and P values were determined using a one-tailed t-test.
Extended Data Fig. 9 Ded1p binding and mRNA remodelling can occur without decreased translation efficiency if no near-cognate initiation codon is present.
Ded1p iCLIP track, differential DMS-MaPseq track (5 min, 37 °C) and ribosome occupancy tracks (5 min, 37 °C) of wild-type DED1 and ded1-95 for ADH3 mRNA, the translation of which is largely unaffected by Ded1p (ΔTE = −0.1). 5′ UTR and ORF are marked. iCLIP and DMS-MaPseq tracks show Ded1p binding and remodelling of the 5′ UTR, ribosome profiling tracks indicate no significant accumulation of ribosomes in the 5′ UTR. Similar results were obtained in two independent experiments.
Extended Data Fig. 10 ATIS conservation across fungi and Ded1p-mediated activation of upstream ORFs starting from near-cognate initiation codons.
a, Sequence conservation in fungi around high-confidence ATIS (moving average of ±1 nt). Positive values indicate higher sequence conservation than the average of five randomly chosen positions on the same 5′ UTR for each ATIS, negative values indicate less sequence conservation (conservation scores were obtained from the sacCer3 phastCons7way dataset, on the basis of sequence homology between the following species: S. cerevisiae, S. paradoxus, S. mikatae, S. kudriavzevii, S. bayanus, Naumoovozyma castellii and Lachancea kluyveri)50. b, Ribosome occupancy tracks (30 °C and 5 min, 37 °C) of wild-type DED1 and ded1-95 for ALA1 mRNA. 5′ UTR and ORF are indicated. The ACG initiation codon (−25, marked) has been previously shown to function as an ATIS for the mitochondrial isoform of Ala1p29. Similar results were obtained in two independent biological replicates for each experiment. c, Ribosome occupancy tracks (5 min, 37 °C) of ded1-95 and wild-type DED1 (vegetative control and anaphase II) for PSA1 mRNA. ATISs are marked by dashed lines. Similar results were obtained in two (vegetative control) and four (anaphase II) independent experiments. d, Ribosome-protected fragments mapping to DED1 in vegetative cells and cells in anaphase II. Data are the mean of two (vegetative) and four (anaphase II) independent experiments, circles represent each replicate.
Supplementary information
Supplementary Figures
This file contains Supplementary Figure 1, the gel source data.
Supplementary Tables
This file contains Supplementary Tables 1-4.
Source data
Rights and permissions
About this article
Cite this article
Guenther, UP., Weinberg, D.E., Zubradt, M.M. et al. The helicase Ded1p controls use of near-cognate translation initiation codons in 5′ UTRs. Nature 559, 130–134 (2018). https://doi.org/10.1038/s41586-018-0258-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0258-0
This article is cited by
-
The molecular basis of translation initiation and its regulation in eukaryotes
Nature Reviews Molecular Cell Biology (2024)
-
Dynamic regulation of messenger RNA structure controls translation
Nature (2023)
-
Pervasive downstream RNA hairpins dynamically dictate start-codon selection
Nature (2023)
-
Secondary structures that regulate mRNA translation provide insights for ASO-mediated modulation of cardiac hypertrophy
Nature Communications (2023)
-
Kaposi’s sarcoma-associated herpesvirus induces specialised ribosomes to efficiently translate viral lytic mRNAs
Nature Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.