Abstract
Many organisms capture or sense sunlight using rhodopsin pigments1,2, which are integral membrane proteins that bind retinal chromophores. Rhodopsins comprise two distinct protein families1, type-1 (microbial rhodopsins) and type-2 (animal rhodopsins). The two families share similar topologies and contain seven transmembrane helices that form a pocket in which retinal is linked covalently as a protonated Schiff base to a lysine at the seventh transmembrane helix2,3. Type-1 and type-2 rhodopsins show little or no sequence similarity to each other, as a consequence of extensive divergence from a common ancestor or convergent evolution of similar structures1. Here we report a previously unknown and diverse family of rhodopsins—which we term the heliorhodopsins—that we identified using functional metagenomics and that are distantly related to type-1 rhodopsins. Heliorhodopsins are embedded in the membrane with their N termini facing the cell cytoplasm, an orientation that is opposite to that of type-1 or type-2 rhodopsins. Heliorhodopsins show photocycles that are longer than one second, which is suggestive of light-sensory activity. Heliorhodopsin photocycles accompany retinal isomerization and proton transfer, as in type-1 and type-2 rhodopsins, but protons are never released from the protein, even transiently. Heliorhodopsins are abundant and distributed globally; we detected them in Archaea, Bacteria, Eukarya and their viruses. Our findings reveal a previously unknown family of light-sensing rhodopsins that are widespread in the microbial world.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Spudich, J. L., Yang, C. S., Jung, K. H. & Spudich, E. N. Retinylidene proteins: structures and functions from archaea to humans. Annu. Rev. Cell Dev. Biol. 16, 365–392 (2000).
Ernst, O. P. et al. Microbial and animal rhodopsins: structures, functions, and molecular mechanisms. Chem. Rev. 114, 126–163 (2014).
Govorunova, E. G., Sineshchekov, O. A., Li, H. & Spudich, J. L. Microbial rhodopsins: diversity, mechanisms, and optogenetic applications. Annu. Rev. Biochem. 86, 845–872 (2017).
Yutin, N. & Koonin, E. V. Proteorhodopsin genes in giant viruses. Biol. Direct 7, 34 (2012).
Philosof, A. & Béjà, O. Bacterial, archaeal and viral-like rhodopsins from the Red Sea. Environ. Microbiol. Rep. 5, 475–482 (2013).
Béjà, O. et al. Bacterial rhodopsin: evidence for a new type of phototrophy in the sea. Science 289, 1902–1906 (2000).
Béjà, O., Spudich, E. N., Spudich, J. L., Leclerc, M. & DeLong, E. F. Proteorhodopsin phototrophy in the ocean. Nature 411, 786–789 (2001).
Finkel, O. M., Béjà, O. & Belkin, S. Global abundance of microbial rhodopsins. ISME J. 7, 448–451 (2013).
Venter, J. C. et al. Environmental genome shotgun sequencing of the Sargasso Sea. Science 304, 66–74 (2004).
Sabehi, G. et al. New insights into metabolic properties of marine bacteria encoding proteorhodopsins. PLoS Biol. 3, e273 (2005).
Rusch, D. B. et al. The Sorcerer II Global Ocean Sampling expedition: northwest Atlantic through eastern tropical Pacific. PLoS Biol. 5, e77 (2007).
Atamna-Ismaeel, N. et al. Widespread distribution of proteorhodopsins in freshwater and brackish ecosystems. ISME J. 2, 656–662 (2008).
Sharma, A. K. et al. Actinorhodopsin genes discovered in diverse freshwater habitats and among cultivated freshwater Actinobacteria. ISME J. 3, 726–737 (2009).
Koh, E. Y. et al. Proteorhodopsin-bearing bacteria in Antarctic sea ice. Appl. Environ. Microbiol. 76, 5918–5925 (2010).
Martínez, A., Bradley, A. S., Waldbauer, J. R., Summons, R. E. & DeLong, E. F. Proteorhodopsin photosystem gene expression enables photophosphorylation in a heterologous host. Proc. Natl Acad. Sci. USA 104, 5590–5595 (2007).
Pushkarev, A. & Béjà, O. Functional metagenomic screen reveals new and diverse microbial rhodopsins. ISME J. 10, 2331–2335 (2016).
Altschul, S. F. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25, 3389–3402 (1997).
Broome-Smith, J. K. & Spratt, B. G. A vector for the construction of translational fusions to TEM β-lactamase and the analysis of protein export signals and membrane protein topology. Gene 49, 341–349 (1986).
Broome-Smith, J. K., Tadayyon, M. & Zhang, Y. β-lactamase as a probe of membrane protein assembly and protein export. Mol. Microbiol. 4, 1637–1644 (1990).
Brum, J. R. et al. Patterns and ecological drivers of ocean viral communities. Science 348, 1261498 (2015).
Sunagawa, S. et al. Structure and function of the global ocean microbiome. Science 348, 1261359 (2015).
Braiman, M. S. et al. Vibrational spectroscopy of bacteriorhodopsin mutants: light-driven proton transport involves protonation changes of aspartic acid residues 85, 96, and 212. Biochemistry 27, 8516–8520 (1988).
Palczewski, K. et al. Crystal structure of rhodopsin: a G protein-coupled receptor. Science 289, 739–745 (2000).
Toyama, D. et al. Metagenomics analysis of microorganisms in freshwater lakes of the Amazon basin. Genome Announc. 4, e01440-16 (2016).
Yan, Q. et al. Impacts of the Three Gorges Dam on microbial structure and potential function. Sci. Rep. 5, 8605 (2015).
Wright, J. J., Lee, S., Zaikova, E., Walsh, D. A. & Hallam, S. J. DNA extraction from 0.22 μM Sterivex filters and cesium chloride density gradient centrifugation. J. Vis. Exp. 31, 1352 (2009).
Gibson, D. G. et al. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat. Methods 6, 343–345 (2009).
Gibson, D. G., Smith, H. O., Hutchison, C. A., III, Venter, J. C. & Merryman, C. Chemical synthesis of the mouse mitochondrial genome. Nat. Methods 7, 901–903 (2010).
Bankevich, A. et al. SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 19, 455–477 (2012).
Marchler-Bauer, A. et al. CDD/SPARCLE: functional classification of proteins via subfamily domain architectures. Nucleic Acids Res. 45, D200–D203 (2017).
Söding, J., Biegert, A. & Lupas, A. N. The HHpred interactive server for protein homology detection and structure prediction. Nucleic Acids Res. 33, W244–W248 (2005).
Alva, V., Nam, S. Z., Söding, J. & Lupas, A. N. The MPI bioinformatics Toolkit as an integrative platform for advanced protein sequence and structure analysis. Nucleic Acids Res. 44, W410–W415 (2016).
Kelley, L. A., Mezulis, S., Yates, C. M., Wass, M. N. & Sternberg, M. J. The Phyre2 web portal for protein modeling, prediction and analysis. Nat. Protocols 10, 845–858 (2015).
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Krogh, A., Larsson, B., von Heijne, G. & Sonnhammer, E. L. Predicting transmembrane protein topology with a hidden Markov model: application to complete genomes. J. Mol. Biol. 305, 567–580 (2001).
Käll, L., Krogh, A. & Sonnhammer, E. L. A combined transmembrane topology and signal peptide prediction method. J. Mol. Biol. 338, 1027–1036 (2004).
Reynolds, S. M., Käll, L., Riffle, M. E., Bilmes, J. A. & Noble, W. S. Transmembrane topology and signal peptide prediction using dynamic Bayesian networks. PLOS Comput. Biol. 4, e1000213 (2008).
Viklund, H., Bernsel, A., Skwark, M. & Elofsson, A. SPOCTOPUS: a combined predictor of signal peptides and membrane protein topology. Bioinformatics 24, 2928–2929 (2008).
Zaremba-Niedzwiedzka, K. et al. Asgard archaea illuminate the origin of eukaryotic cellular complexity. Nature 541, 353–358 (2017).
Brown, C. T. et al. Unusual biology across a group comprising more than 15% of domain Bacteria. Nature 523, 208–211 (2015).
Anantharaman, K. et al. Thousands of microbial genomes shed light on interconnected biogeochemical processes in an aquifer system. Nat. Commun. 7, 13219 (2016).
Dereeper, A. et al. Phylogeny.fr: robust phylogenetic analysis for the non-specialist. Nucleic Acids Res. 36, W465–W469 (2008).
Guindon, S. et al. New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst. Biol. 59, 307–321 (2010).
Buchfink, B., Xie, C. & Huson, D. H. Fast and sensitive protein alignment using DIAMOND. Nat. Methods 12, 59–60 (2015).
Suzek, B. E., Wang, Y., Huang, H., McGarvey, P. B. & Wu, C. H. UniRef clusters: a comprehensive and scalable alternative for improving sequence similarity searches. Bioinformatics 31, 926–932 (2015).
McKinney, W. Data structures for statistical computing in Python. In Proc. 9th Python in Science Conference. (eds van der Walt, S. & Millman, J.) 51–56 (SciPy, Austin, 2010).
Kluyver, T. et al. Juypter notebooks – a publishing format for reproducible computational workflows. In Positioning and Power in Academic Publishing: Players, Agents and Agendas. Proc. 20th International Conference on Electronic Publishing. (eds Loizides, F. & Schmidt, B.) 87–90 (IOS, Amsterdam, 2016).
Inoue, K. et al. A light-driven sodium ion pump in marine bacteria. Nat. Commun. 4, 1678 (2013).
Inoue, K. et al. A natural light-driven inward proton pump. Nat. Commun. 7, 13415 (2016).
Yamauchi, Y. et al. Molecular properties of a DTD channelrhodopsin from Guillardia theta. Biophys. Physicobiol. 14, 57–66 (2017).
Kato, H. E. et al. Structural basis for Na+ transport mechanism by a light-driven Na+ pump. Nature 521, 48–53 (2015).
Kawanabe, A., Furutani, Y., Jung, K. H. & Kandori, H. FTIR study of the photoisomerization processes in the 13-cis and all-trans forms of Anabaena sensory rhodopsin at 77 K. Biochemistry 45, 4362–4370 (2006).
Inoue, K., Koua, F. H., Kato, Y., Abe-Yoshizumi, R. & Kandori, H. Spectroscopic study of a light-driven chloride ion pump from marine bacteria. J. Phys. Chem. B 118, 11190–11199 (2014).
Krebs, R. A., Alexiev, U., Partha, R., DeVita, A. M. & Braiman, M. S. Detection of fast light-activated H+ release and M intermediate formation from proteorhodopsin. BMC Physiol. 2, 5 (2002).
Tanimoto, T., Furutani, Y. & Kandori, H. Structural changes of water in the Schiff base region of bacteriorhodopsin: proposal of a hydration switch model. Biochemistry 42, 2300–2306 (2003).
Furutani, Y. et al. FTIR spectroscopy of the O photointermediate in pharaonis phoborhodopsin. Biochemistry 43, 5204–5212 (2004).
Camacho, C. et al. BLAST+: architecture and applications. BMC Bioinformatics 10, 421 (2009).
Acknowledgements
We thank G. Hevroni and M. Diamant for technical help with sampling, D. Cohen for help with the HPC ATLAS cluster, and M. Habib for help in constructing the protein fusions. This work was supported by Israel Science Foundation – F.I.R.S.T. (Bikura) Individual grant (545/17), the I-CORE Program of the Planning and Budgeting Committee and the Grand Technion Energy Program (GTEP), and is part of the Leona M. and Harry B. Helmsley Charitable Trust reports on Alternative Energy series of the Technion Israel Institute of Technology and the Weizmann Institute of Science, the Lorry I. Lokey Interdisciplinary Center for Life Sciences and Engineering, the Russell Berrie Nanotechnology Institute, the Louis and Lyra Richmond Memorial Chair in Life Sciences (to O.B.), the Japanese Ministry of Education, Culture, Sports, Science and Technology (26708001, 26115706, 26620005 to K.I. and 25104009, 15H02391 to H.K.), JST PRESTO grants (JPMJPR15P2 to K.I. and JPMJPR1688 to S.P.T.) and JST CREST grant (JPMJCR1753 to H.K.).
Reviewer information
Nature thanks S. Balashov, D. Kirchman, D. Oprian and J. Spudich for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
A.Pu. and O.B. devised the initial idea for the project, and together with K.I. and H.K., conceived the experiments. A.Pu. collected the DNA from Lake Kinneret, and detected and initiated preliminary characterization of heliorhodopsin; S.L. performed the topology experiments; J.F.-U., A.Ph., I.S., N.Y., E.V.K. and O.B. performed the bioinformatic analyses; and K.I., M.S., M.K., S.T., S.I., R.N., S.P.T. and H.K. performed the biophysical measurements. O.B., K.I. and H.K. prepared the manuscript with contributions from all of the authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Full alignment of heliorhodopsin and type-1 rhodopsin.
a, Multiple amino acid alignment of different heliorhodopsins. Positions 23, 80, 104 and 107 (heliorhodopsin 48C12 numbering) are marked with black arrows, as well as position 237 (grey arrow) and lysine in position 241 (red arrow). b, Multiple amino acid alignment of heliorhodopsin (48C12) with green absorbing proteorhodopsin (GPR), bacteriorhodopsin (BR), sensory rhodopsin I from Halobacterium salinarum (HsSRI) and Salinibacter ruber (SrSRI), sensory rhodopsin II from H. salinarum (HsSRII) and Natronomonas pharaonis (NpSRII), Anabaena sensory rhodopsin (ASR), xenorhodopsin from Parvularcula oceani (PoXeR), halorhodopsin from H. salinarum (HsHR), eubacterial chloride-pump rhodopsin from Nonlabens marinus (NmClR), sodium-pump rhodopsin from Krokinobacter eikastus (KR2) (K. eikastus is also known as Dokdonia eikasta), anion channelrhodopsin from Guillardia theta (GtACR1) and cation channelrhodopsin2 from Chlamydomonas reinhardtii (C-terminal side omitted, ChR2 ΔC-term). Positions 85, 89 and 96 (bacteriorhodopsin numbering) are marked with black arrows, as well as position 212 (grey arrow) and lysine in position 216 (red arrow).
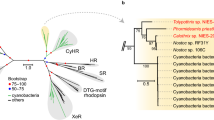
Extended Data Fig. 2 Relationship and depth distribution of heliorhodopsins and type-1 rhodopsins.
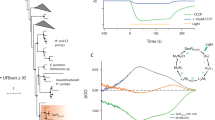
a, Clustering analysis separates heliorhodopsins (purple leaves in dendrogram tree) from type-1 rhodopsins (grey leaves in dendrogram tree) based on per cent identity obtained using protein–protein blast v.2.7.157. The hierarchical clustering was performed using the clustermap function from the Python package seaborn, with default parameters for metric and linkage. The code used to generate the figure is available as a Jupyter notebook at https://github.com/BejaLab/heliorhodopsin. b, Detailed phylogenetic relationships within the heliorhodopsin family. White circles represent bootstrap values of >80%. The scale bar indicates the average number of amino acid substitutions per site. Coloured circles indicate heliorhodopsins expressed in this study. c, Depth profiles of the relative abundance of type-1 rhodopsins and heliorhodopsins from marine metagenomes collected during the Tara Oceans expedition.
Extended Data Fig. 3 Heliorhodopsin membrane topology.
a, Predictions of membrane topology for type-1 rhodopsins, type-2 rhodopsins and heliorhodopsin. Upper panel, suggested schematic topologies for type-1 rhodopsins, type-2 rhodopsins and heliorhodopsins. Positively charged amino acids are labelled. N, amino-terminal tail, C, carboxy-terminal tail. Middle panel, membrane topology predictions by TMHMM35, Phobius36, Philius37 and SPOCTOPUS38. Lower panel, the rhodopsin sequences used for membrane topology predictions. b, Arrangement of heliorhodopsin 48C12 protein across the E. coli membrane. The heliorhodopsin–β-lactamase fusion sites resulting in ampicillin resistance (ampr) and ampicillin sensitivity (amps) are indicated. This experiment was repeated twice.
Extended Data Fig. 4 Ion-transport activity assay of heliorhodopsins.
a, Ion-transport activity assay of six kinds of heliorhodopsin (48C12, and contigs 172728, 2376895, 381806, 1205911 and 161490), and a comparison with a type-1 rhodopsin (a light-driven proton pump; green-absorbing proteorhodopsin, GPR) in the absence and presence of the protonophore CCCP. Light was present for the time region indicated by the orange bars. b, Patch-clamp assay of heliorhodopsin 48C12. The photocurrent was measured at 60, 0 or −60 mV by whole-cell mode. Left, 48C12-expressing cells were illuminated with 550-nm light (2.6 mW/mm2), indicated by a yellow bar (n = 10 cells). Right, cells expressing GtCCR4 were illuminated with 520-nm light (2.4 mW/mm2), indicated by a green bar.
Extended Data Fig. 5 UV-visible absorption of heliorhodopsin 48C12 at different pH values.
a, Deprotonation of the retinal Schiff base of heliorhodopsin 48C12 at alkaline pH. Difference absorption spectra (left) and absorption change at λ = 553 nm (right, orange solid circles) of heliorhodopsin 48C12 upon pH change from 8.5 to higher values. The deprotonated form of retinal Schiff base showed the absorption at λ = 373 nm. b, Red-shift of UV-visible absorption spectrum of heliorhodopsin 48C12, and protonation of the counterion. UV-visible absorption spectra (left) and the λmax (right, orange solid circles) of heliorhodopsin 48C12 at pH 2.8–8.4. When pH is lowered a red-shift of the absorption is observed, which is commonly reported for many type-1 rhodopsins and reflects the protonation of counterions. Thus, the red-shift of heliorhodopsin 48C12 originates from protonation of E107, which is fitted with the Henderson–Hasselbalch equation (blue dashed line), and the pKa of counterion (E107) is estimated to be 3.7. At pH values of less than 2.8, a large blue-shift to 443 nm is observed, presumably owing to the acid denaturation of the protein. The pKa values in right panels indicate mean ± s.d.
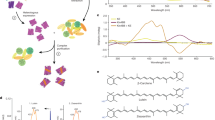
Extended Data Fig. 6 HPLC pattern of retinal extracted from heliorhodopsin 48C12.
HPLC pattern of retinal extracted from heliorhodopsin 48C12 in the dark (blue) and under illumination at λ = 540 ± 10 nm (green). Most of the retinal (>97%) bound to heliorhodopsin 48C12 adopts an all-trans configuration in the dark (n = 4). When the retinal is extracted after illumination (λ > 500 nm), the proportion of the 13-cis form increased to 59 ± 5% (mean ± s.d., n = 4).
Extended Data Fig. 7 Photocycle of heliorhodopsins.
a, The photocycle of heliorhodopsin 48C12 determined by the multi-exponential fitting for the time evolution of transient absorption change shown in Fig. 4a, b. The lifetimes of the intermediates are indicated as mean ± s.d. b, c, Time evolution of transient absorption change of photo-excited contigs 381806 (b) and 172728 (c).
Extended Data Fig. 8 Light-induced FTIR difference spectra of heliorhodopsin 48C12.
a, Light-induced FTIR spectra of heliorhodopsin 48C12 at 77 K (upper spectrum), 240 K (middle spectrum) and 277 K (lower spectrum) in the 1,800–900 cm−1, in which the intermediate produced is the K, M and O intermediate, respectively. Spectra are measured in H2O (black) and D2O (red). The amide-I vibration of the peptide backbone and the C=N stretch vibration of the Schiff base appear at 1,700–1,600 cm−1. Peaks at 1,659(−) and 1,629(+) cm−1 in H2O, and at 1,637(−) and 1,613(+) cm−1 in D2O at 77 K probably originate from a C=N stretch of the Schiff base, and small changes in amide-I suggest that the primary K intermediate does not undergo a major conformational change. This is also the case for the M intermediate, for which negative peaks at 1,656 cm−1 in H2O and at 1,638 cm−1 in D2O originate from the C=N stretch of the Schiff base, and changes in amide-I vibrations are small. By contrast, the appearance of strong peaks at 1,694 and 1,675 cm−1 in the O intermediate indicates extensive conformational changes. b, Light-induced difference FTIR absorption spectra of the K intermediate of heliorhodopsin 48C12 and bacteriorhodopsin. Light-induced difference FTIR absorption spectra of heliorhodopsin 48C12 (upper spectrum) and bacteriorhodopsin (lower spectrum) at 77 K in the 1,230–1,150 cm−1 region, which were measured in H2O. The bands at 1,200(−) and 1,188(+) cm−1 were observed at 77 K, similar to the type-1 rhodopsin bacteriorhodopsin, which suggests that the primary photochemical reaction of heliorhodopsin is the all-trans to 13-cis isomerization.
Extended Data Fig. 9 Transient absorption change of heliorhodopsin 48C12 mutants.
a–c, Transient absorption spectra of photo-excited heliorhodopsin 48C12 E107Q (a), H23F (b) and H80F (c) mutants. The M intermediate is formed for all these mutants, which suggests that none of these residues is the proton acceptor of the Schiff base. In the case of the H80F mutation, the O intermediate is not formed. d–f, Time evolution of transient absorption change of photo-excited heliorhodopsin 48C12 D36N (d), E62D (e) and E107D (f) mutants. The fact that the M intermediate forms in these mutants excludes the possibility that one of these residues acts as the proton acceptor for the Schiff base.
Extended Data Fig. 10 Proton release and uptake observed with cresol red.
a, UV-visible absorption spectra of cresol red (left) and its absorbance at 429 nm (blue circles) and 573 nm (green circles) (right), at different pH values in 100 mM NaCl and 6-mix buffer (10 mM citrate, 10 mM MES, 10 mM HEPES, 10 mM MOPS, 10 mM CHES and 10 mM CAPS). Global fitting for absorption at different pH values with the Henderson–Hasselbalch equation showed the pKa of cresol red to be 8.123 ± 0.004 (mean ± s.d.). b, c, Time evolutions of transient absorption change of GPR (b) and heliorhodopsin 48C12 (c) in unbuffered 100 mM NaCl and 0.1% DDM. The pH value was adjusted to be approximately 8.1 by addition of NaOH. The transient absorption change of cresol red was calculated by the subtracting transient absorption changes obtained with cresol red from those without cresol red at 429 and 573 nm (corresponding to accumulations of protonated and deprotonated state of cresol red, respectively).
Supplementary Information
Supplementary Table 1
Accession numbers of the analyzed metagenomes used in Figure 2b and Extended Data Figure 2c
Supplementary Table 2
Rhodopsin DIAMOND database containing 89 protein sequences from rhodopsin type-1 and heliorhodopsin representatives. Used in Figure 2b and Extended Data Figure 2c
Supplementary Table 3
UniRef50 rhodopsin clusters. Used for Figure 2b and Extended Data Figure 2c
Supplementary Table 4
Primers used to clone heliorhodopsin 48C12 into pET-9d expression vector
Supplementary Table 5
Primers used in to construct heliorhodopsin 48C12 β-lactamase fusions
Supplementary Data
This zipped file contains Supplementary Data files 1-8 and a Supplementary Data guide
Rights and permissions
About this article
Cite this article
Pushkarev, A., Inoue, K., Larom, S. et al. A distinct abundant group of microbial rhodopsins discovered using functional metagenomics. Nature 558, 595–599 (2018). https://doi.org/10.1038/s41586-018-0225-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0225-9
This article is cited by
-
Microbial circadian clocks: host-microbe interplay in diel cycles
BMC Microbiology (2023)
-
Engineering artificial photosynthesis based on rhodopsin for CO2 fixation
Nature Communications (2023)
-
Light and prey influence the abundances of two rhodopsins in the dinoflagellate Oxyrrhis marina
Protoplasma (2023)
-
Phototrophy by antenna-containing rhodopsin pumps in aquatic environments
Nature (2023)
-
Ecogenomics sheds light on diverse lifestyle strategies in freshwater CPR
Microbiome (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.