Abstract
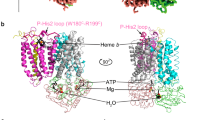
Alternative complex III (ACIII) is a key component of the respiratory and/or photosynthetic electron transport chains of many bacteria1,2,3. Like complex III (also known as the bc1 complex), ACIII catalyses the oxidation of membrane-bound quinol and the reduction of cytochrome c or an equivalent electron carrier. However, the two complexes have no structural similarity4,5,6,7. Although ACIII has eluded structural characterization, several of its subunits are known to be homologous to members of the complex iron–sulfur molybdoenzyme (CISM) superfamily8, including the proton pump polysulfide reductase9,10. We isolated the ACIII from Flavobacterium johnsoniae with native lipids using styrene maleic acid copolymer11,12,13,14, both as an independent enzyme and as a functional 1:1 supercomplex with an aa3-type cytochrome c oxidase (cyt aa3). We determined the structure of ACIII to 3.4 Å resolution by cryo-electron microscopy and constructed an atomic model for its six subunits. The structure, which contains a [3Fe–4S] cluster, a [4Fe–4S] cluster and six haem c units, shows that ACIII uses known elements from other electron transport complexes arranged in a previously unknown manner. Modelling of the cyt aa3 component of the supercomplex revealed that it is structurally modified to facilitate association with ACIII, illustrating the importance of the supercomplex in this electron transport chain. The structure also resolves two of the subunits of ACIII that are anchored to the lipid bilayer with N-terminal triacylated cysteine residues, an important post-translational modification found in numerous prokaryotic membrane proteins that has not previously been observed structurally in a lipid bilayer.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Pereira, M. M., Carita, J. N. & Teixeira, M. Membrane-bound electron transfer chain of the thermohalophilic bacterium Rhodothermus marinus: a novel multihemic cytochrome bc, a new complex III. Biochemistry 38, 1268–1275 (1999).
Yanyushin, M. F., del Rosario, M. C., Brune, D. C. & Blankenship, R. E. New class of bacterial membrane oxidoreductases. Biochemistry 44, 10037–10045 (2005).
Refojo, P. N., Sousa, F. L., Teixeira, M. & Pereira, M. M. The alternative complex III: a different architecture using known building modules. Biochim. Biophys. Acta 1797, 1869–1876 (2010).
Pereira, M. M., Refojo, P. N., Hreggvidsson, G. O., Hjorleifsdottir, S. & Teixeira, M. The alternative complex III from Rhodothermus marinus - a prototype of a new family of quinol:electron acceptor oxidoreductases. FEBS Lett. 581, 4831–4835 (2007).
Gao, X., Xin, Y., Bell, P. D., Wen, J. & Blankenship, R. E. Structural analysis of alternative complex III in the photosynthetic electron transfer chain of Chloroflexus aurantiacus. Biochemistry 49, 6670–6679 (2010).
Refojo, P. N., Ribeiro, M. A., Calisto, F., Teixeira, M. & Pereira, M. M. Structural composition of alternative complex III: variations on the same theme. Biochim. Biophys. Acta 1827, 1378–1382 (2013).
Xia, D. et al. Crystal structure of the cytochrome bc 1 complex from bovine heart mitochondria. Science 277, 60–66 (1997).
Rothery, R. A., Workun, G. J. & Weiner, J. H. The prokaryotic complex iron–sulfur molybdoenzyme family. Biochim. Biophys. Acta 1778, 1897–1929 (2008).
Dietrich, W. & Klimmek, O. The function of methyl-menaquinone-6 and polysulfide reductase membrane anchor (PsrC) in polysulfide respiration of Wolinella succinogenes. Eur. J. Biochem. 269, 1086–1095 (2002).
Hedderich, R. et al. Anaerobic respiration with elemental sulfur and with disulfides. FEMS Microbiol. Rev. 22, 353–381 (1998).
Dörr, J. M. et al. Detergent-free isolation, characterization, and functional reconstitution of a tetrameric K+ channel: the power of native nanodiscs. Proc. Natl Acad. Sci. USA 111, 18607–18612 (2014).
Postis, V. et al. The use of SMALPs as a novel membrane protein scaffold for structure study by negative stain electron microscopy. Biochim. Biophys. Acta 1848, 496–501 (2015).
Dörr, J. M. et al. The styrene–maleic acid copolymer: a versatile tool in membrane research. Eur. Biophys. J. 45, 3–21 (2016).
Parmar, M. et al. Using a SMALP platform to determine a sub-nm single particle cryo-EM membrane protein structure. Biochim. Biophys. Acta 1860, 378–383 (2018).
Bayburt, T. H. & Sligar, S. G. Membrane protein assembly into nanodiscs. FEBS Lett. 584, 1721–1727 (2010).
Jormakka, M. et al. Molecular mechanism of energy conservation in polysulfide respiration. Nat. Struct. Mol. Biol. 15, 730–737 (2008).
Majumder, E. L., King, J. D. & Blankenship, R. E. Alternative complex III from phototrophic bacteria and its electron acceptor auracyanin. Biochim. Biophys. Acta 1827, 1383–1391 (2013).
Refojo, P. N., Teixeira, M. & Pereira, M. M. The alternative complex III: properties and possible mechanisms for electron transfer and energy conservation. Biochim. Biophys. Acta 1817, 1852–1859 (2012).
Crofts, A. R. The cytochrome bc 1 complex: function in the context of structure. Annu. Rev. Physiol. 66, 689–733 (2004).
Sazanov, L. A. A giant molecular proton pump: structure and mechanism of respiratory complex I. Nat. Rev. Mol. Cell Biol. 16, 375–388 (2015).
Kovacs-Simon, A., Titball, R. W. & Michell, S. L. Lipoproteins of bacterial pathogens. Infect. Immun. 79, 548–561 (2011).
Kao, W. C. et al. The obligate respiratory supercomplex from Actinobacteria. Biochim. Biophys. Acta 1857, 1705–1714 (2016).
Graf, S. et al. Rapid electron transfer within the III–IV supercomplex in Corynebacterium glutamicum. Sci. Rep. 6, 34098 (2016).
Sousa, J. S. et al. Structural basis for energy transduction by respiratory alternative complex III. Nat. Commun. (in the press).
McBride, M. J. & Baker, S. A. Development of techniques to genetically manipulate members of the genera Cytophaga, Flavobacterium, Flexibacter, and Sporocytophaga. Appl. Environ. Microbiol. 62, 3017–3022 (1996).
Bell, A. J., Frankel, L. K. & Bricker, T. M. High yield non-detergent isolation of photosystem I-light-harvesting chlorophyll II membranes from spinach thylakoids: implications for the organization of the PS I antennae in higher plants. J. Biol. Chem. 290, 18429–18437 (2015).
Berry, E. A. & Trumpower, B. L. Simultaneous determination of hemes a, b, and c from pyridine hemochrome spectra. Anal. Biochem. 161, 1–15 (1987).
Thomas, P. E., Ryan, D. & Levin, W. An improved staining procedure for the detection of the peroxidase activity of cytochrome P-450 on sodium dodecyl sulfate polyacrylamide gels. Anal. Biochem. 75, 168–176 (1976).
Minghetti, K. C. et al. Modified, large-scale purification of the cytochrome o complex (bo-type oxidase) of Escherichia coli yields a two heme/one copper terminal oxidase with high specific activity. Biochemistry 31, 6917–6924 (1992).
Gao, X., Xin, Y. & Blankenship, R. E. Enzymatic activity of the alternative complex III as a menaquinol:auracyanin oxidoreductase in the electron transfer chain of Chloroflexus aurantiacus. FEBS Lett. 583, 3275–3279 (2009).
Ragan, C. I., Wilson, M. T., Darley-Usmar, V. M. & Lowe, P. N. Mitochondria: A Practical Approach (IRL Press, Oxford, 1987).
Carrell, C. J. et al. Generation of novel copper sites by mutation of the axial ligand of amicyanin. Atomic resolution structures and spectroscopic properties. Biochemistry 46, 1900–1912 (2007).
Ouyang, H. et al. Functional importance of a pair of conserved glutamic acid residues and of Ca2+ binding in the cbb 3-type oxygen reductases from Rhodobacter sphaeroides and Vibrio cholerae. Biochemistry 51, 7290–7296 (2012).
Bowyer, J. R., Tierney, G. V. & Crofts, A. R. Cytochrome c 2 – reaction centre coupling in chromatophores of Rhodopseudomonas sphaeroides and Rhodopseudomonas capsulata. FEBS Lett. 101, 207–212 (1979).
Dutton, P. L. Redox potentiometry: determination of midpoint potentials of oxidation-reduction components of biological electron-transfer systems. Methods Enzymol. 54, 411–435 (1978).
Fultz, M. & Durs, R. Mediator compounds for the electrochemical study of biological redox systems: a compilation. Anal. Chim. Acta 140, 1–18 (1982).
Marr, C. R., Benlekbir, S. & Rubinstein, J. L. Fabrication of carbon films with ∼500 nm holes for cryo-EM with a direct detector device. J. Struct. Biol. 185, 42–47 (2014).
Tivol, W. F., Briegel, A. & Jensen, G. J. An improved cryogen for plunge freezing. Microsc. Microanal. 14, 375–379 (2008).
Suloway, C. et al. Automated molecular microscopy: the new Leginon system. J. Struct. Biol. 151, 41–60 (2005).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Rohou, A. & Grigorieff, N. CTFFIND4: Fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Scheres, S. H. W. A Bayesian view on cryo-EM structure determination. J. Mol. Biol. 415, 406–418 (2012).
Scheres, S. H. W. Semi-automated selection of cryo-EM particles in RELION-1.3. J. Struct. Biol. 189, 114–122 (2015).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Tsirigos, K. D., Peters, C., Shu, N., Käll, L. & Elofsson, A. The TOPCONS web server for consensus prediction of membrane protein topology and signal peptides. Nucleic Acids Res. 43, W401–W407 (2015).
Drozdetskiy, A., Cole, C., Procter, J. & Barton, G. J. JPred4: a protein secondary structure prediction server. Nucleic Acids Res. 43, W389–W394 (2015).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010).
Barad, B. A. et al. EMRinger: side chain-directed model and map validation for 3D cryo-electron microscopy. Nat. Methods 12, 943–946 (2015).
Humphrey, W., Dalke, A. & Schulten, K. VMD: visual molecular dynamics. J. Mol. Graph. 14, 33–38 (1996).
Källberg, M. et al. Template-based protein structure modeling using the RaptorX web server. Nat. Protoc. 7, 1511–1522 (2012).
Madden, T. L., Tatusov, R. L. & Zhang, J. Applications of network BLAST server. Methods Enzymol. 266, 131–141 (1996).
Sievers, F. et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 7, 539 (2011).
Robert, X. & Gouet, P. Deciphering key features in protein structures with the new ENDscript server. Nucleic Acids Res. 42, W320–W324 (2014).
Jorgensen, W. L., Chandrasekhar, J., Madura, J. D., Impey, R. W. & Klein, M. L. Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 79, 926–935 (1983).
Singharoy, A. et al. Molecular dynamics-based refinement and validation for sub-5 Å cryo-electron microscopy maps. eLife 5, 61–67 (2016).
Huang, J. et al. CHARMM36m: an improved force field for folded and intrinsically disordered proteins. Nat. Methods 14, 71–73 (2017).
Best, R. B. et al. Optimization of the additive CHARMM all-atom protein force field targeting improved sampling of the backbone φ, ψ and side-chain χ(1) and χ(2) dihedral angles. J. Chem. Theory Comput. 8, 3257–3273 (2012).
Autenrieth, F., Tajkhorshid, E., Baudry, J. & Luthey-Schulten, Z. Classical force field parameters for the heme prosthetic group of cytochrome c. J. Comput. Chem. 25, 1613–1622 (2004).
Autenrieth, F., Tajkhorshid, E., Klaus Schulten, A. & Luthey-Schulten, Z. Role of water in transient cytochrome c2 docking. J. Phys. Chem. B 108, 20376–20387 (2004).
Carvalho, A. T. P. & Swart, M. Electronic structure investigation and parametrization of biologically relevant iron–sulfur clusters. J. Chem. Inf. Model. 54, 613–620 (2014). erratum 55, 1508–1508 (2015).
Allen, M. P. & Tildesley, D. J. Computer Simulation of Liquids (Oxford Univ. Press, New York, 1987).
Fletcher, R. & Reeves, C. M. Function minimization by conjugate gradients. Comput. J. 7, 149–154 (1964).
Sun, W. & Yuan, Y.-X. Optimization Theory and Methods: Nonlinear Programming (Springer, New York, 2006).
Acknowledgements
This work was supported by funds from the National Institutes of Health (R01-HL016101 to R.B.G.; P41-GM104601, U54-GM087519 and R01-GM123455 to E.T.) and Canadian Institutes of Health Research (MOP-81294 to J.L.R.); J.L.R. was supported by the Canada Research Chairs program. Some of this work was performed at the Simons Electron Microscopy Center and National Resource for Automated Molecular Microscopy, supported by grants from the Simons Foundation (349247), and the National Institute of General Medical Sciences (P41-GM103310, S10-OD019994). Molecular dynamics simulations were performed at Blue Waters (ACI-1713784 to E.T.) and XSEDE (TG-MCA06N060 to E.T.). Blue Waters is supported by the National Science Foundation (OCI-0725070 and ACI-1238993) and the State of Illinois. XSEDE is supported by the National Science Foundation (ACI-1548562). We thank M. McBride for providing us with the Flavobacterium johnsoniae UW101 strain, B. Carragher and C. Potter for facilitating access to the Titan Krios, Z. Zhang for collecting data, and M. Mazhab-Jafari for advice on building atomic models.
Reviewer information
Nature thanks Y. Cheng, M. Wikström and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
P.V. expressed, purified and characterized the protein supercomplex, S.B. prepared cryo-EM specimens and calculated cryo-EM maps, C.S. built the de novo atomic structures, Y.W. conducted the molecular dynamics simulations, S.H. performed the electrochemical characterization, J.H. performed the metal analysis, E.T. supervised molecular dynamics simulations, J.L.R. supervised the molecular structure determination, and R.B.G. conceived the project, supervised biochemical experiments, and coordinated the project. All authors participated in the manuscript preparation.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Expression and spectroscopic characterization of the ACIII-cyt aa3 supercomplex.
a, A schematic of the respiratory chain of F. johnsoniae. b, UV–visible spectrum and SDS–PAGE of the membranes from F. johnsoniae. Left, the difference spectrum of the membranes of F. johnsoniae, obtained from the spectrum of the air-oxidized membranes and the spectrum after reduction with dithionite. The wavelengths associated with the haem peaks are 605 nm, 560 nm, 552 nm and a broad peak at 630 nm for haems a, b, c and d, respectively. Right, the SDS–PAGE with the membranes followed by staining the gel for haems shows bands corresponding to the cytochrome subunits ActA (48 kDa) and ActE (20 kDa) of ACIII but no bands corresponding to the cytochrome subunit (around 35 kDa) from the cbb3 oxidase. c, The gene arrangement for the ACIII and the cytochrome oxidase aa3 genes in the F. johnsoniae genome. The genes for the subunits I and II from cyt aa3 oxidase are found immediately downstream of those for the act genes of the ACIII. Two different versions of subunit III are denoted as vI and vII. d, UV–visible spectra of the reduced and oxidized forms of the supercomplex in detergent and SMA nanodiscs. The dithionite reduced form of the samples is represented in red and shows the peaks for haem c at 524 nm and 552 nm and those for haem a at 443 nm and 605 nm. e, Pyridine haemochrome assay of the ACIII–cyt aa3 supercomplex in SMA nanodiscs. Plotted is the reduced-minus-oxidized difference spectrum of the pyridine haemochromes of the sample. Peaks at 520 nm and 550 nm are associated with haem c and the peak at 590 nm is associated with haem a. Quantification from the spectrum shows a ratio of 10.6:1 between haem c and haem a, which translates into a 3:2 ratio between ACIII and cyt aa3 assuming 7 haem c per ACIII and 2 haem a per cyt aa3. Data in b are representative of two independent experiments with similar results, and data in d and e are representative of six independent experiments with similar results.
Extended Data Fig. 2 Component and size analysis of ACIII–cyt aa3 supercomplex.
a, SDS–PAGE of the detergent-solubilized preparation followed by Coomassie staining (left) and haem staining (right). b, SDS–PAGE of the SMA nanodiscs preparation followed by Coomassie staining (left) and haem staining (right). c, Mass spectrometry results for the ACIII–cyt aa3 supercomplex preparations. d, Size-exclusion chromatography with the ACIII–cyt aa3 supercomplex from F. johnsoniae. Top left, the chromatogram of the detergent-solubilized sample, showing traces for protein at 280 nm, haem c at 412 nm and haem a at 443 nm respectively. Top right, the chromatogram of the sample isolated using the SMA copolymer, showing traces for protein at 280 nm, haem c at 410 nm and haem a at 605 nm. I and II are the two peaks corresponding to two populations of the supercomplex. Bottom left, chromatogram of the fraction containing peak I. Bottom right, chromatogram of the fraction containing peak II. e, BN–PAGE of the ACIII–cyt aa3 supercomplex. Left, the detergent-solubilized ACIII–cyt aa3 supercomplex, showing a band at around 500 kDa, a smear of possible aggregates and possibly ACIII by itself. Right, the supercomplex in SMA nanodiscs, showing two different populations. f, BN–PAGE with the two different populations of ACIII–cyt aa3 supercomplex in SMA nanodiscs purified from size-exclusion chromatography. The two chromatographic peaks correspond to the two bands observed in the BN–PAGE. Data in a, b are representative of six independent experiments and those in d–f are representative of three independent experiments with similar results.
Extended Data Fig. 3 Functional assays of the ACIII–cyt aa3 supercomplex.
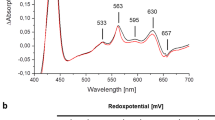
a, The EPR spectrum of the air-oxidized sample showing peaks of the [3Fe–4S]+ cluster from ACIII, the CuA from the cyt aa3 oxidase and low-spin haems with overlapping g values. Insert is a zoomed view from 3,000 G to 3,500 G to better visualize the peaks from CuA (black arrows) and the [3Fe–4S]+ cluster. The region between 4,000 G and 5,000 G is magnified ten times to show the broad gx trough of low-spin haems. The measurement condition is 10 K, 9.267 GHz, 2 mW microwave power and 20 Gauss modulation. b, The EPR spectra of the ferricyanide-oxidized sample at various temperatures. The measurement condition is 9.257 GHz, 2 mW microwave power and 5 Gauss modulation. c, The EPR spectrum of the air-oxidized sample showing peaks of iron–sulfur clusters from ACIII and low-spin haems. The measurement condition is 10 K, 9.427 GHz, 2 mW microwave power, 10 Gauss modulation. d, The EPR spectra of the air-oxidized sample at various temperatures. The measurement condition is 9.427 GHz, 2 mW microwave power, 5 Gauss modulation. e, Redox titration of the haems in the ACIII and the cyt aa3 oxidase in supercomplex in DDM. The potentiometric titration of the c haems from the ACIII (top) and the a haems from the cyt aa3 oxidase (bottom). The Em values are indicated and the solid red line represents the Nernst fitting. f, Steady-state activity of the ACIII–cyt aa3 preparations. The number of independent experiments is six for ACIII in DDM and SMA nanodiscs, and three for peak I and peak II. Data are means ± s.d. Data in a–e are representative of three independent experiments with similar results.
Extended Data Fig. 4 Single-particle cryo-EM of the ACIII–cyt aa3 supercomplex in SMA nanodiscs.
a, Sum of an aligned movie of the ACIII–cyt aa3 supercomplex in an SMA nanodisc. Scale bar, 20 nm. b, Two-dimensional class averages. Scale bar, 10 nm. c, Fourier shell coefficient curves between two independently refined half-maps for the ACIII–cyt aa3 map, ACIII map and combined map. d, Surface rendering maps coloured according to local resolution. Scale bar, 5 nm. e, Euler angle distributions of particles included in the calculation of the three final maps. Data collection and structure calculation were not repeated.
Extended Data Fig. 5 Features observed in the cryo-EM density and the de novo structure of ACIII.
a, Surface representations of ACIII, cyt aa3 and the ACIII–cyt aa3 supercomplex. The density threshold is the same for ACIII and cyt aa3. b, Different views of the ACIII density, coloured by subunit. c, Two single-span transmembrane peptides of unknown origin and sequence, denoted ActX and ActY, are present in the structure in the vicinity of ActC. These have each been modelled as a polyalanine peptide. d, α-helices 2–10 of ActC form two four-helical up-and-down bundles, coloured in two different shades of blue. α-helices 1 and 10 are coloured grey and unlabelled. e, ActB, shown in cartoon form, has contact with ActA, ActC, ActD, ActE and ActF. Surfaces are drawn from residues that are within 4 Å of ActB and coloured according to their chain. f, The transmembrane α-helices of ActC and ActF are arranged in a pseudo two-fold rotation symmetry. g, Side-by-side comparison of the polysulfide reductase (PDB 2VPZ) and the assembly of ActB and ActC. These two structures are aligned based on PsrB, the domain containing four iron–sulfur clusters.
Extended Data Fig. 6 The proposed quinone pocket in ActC.
a, Sequence alignment of the ActC from F. johnsoniae, R. marinus, and Chloroflexus aurantiacus. The transmembrane α-helices are labelled based on the structure of ACIII from F. johnsoniae. The black arrows point to conserved polar residues that are within 15 Å of the [3Fe–4S] cluster in ActB. b, Proposed quinone pocket based on the arrangement of conserved polar residues. c, Different views of the proposed quinone pocket with a docked menaquinone-1 molecule. Hydrophobic residues near the menaquinone-1 (MK1) head group are also shown. The crevice between α-helix 3 and α-helix 4 is a putative quinone entry pathway.
Extended Data Fig. 7 Fitting of the ACIII structure to cryo-EM density.
a, Fitting of cofactors into the cryo-EM density. The blue mesh is drawn with a higher density threshold to reveal metal centres. The numberings of nearby amino acid residues, which are shown along with these cofactors, are listed below each cofactor. b, Fitting of different secondary structure elements to cryo-EM density. c, Eleven identified lipids are modelled as phosphatidylethanolamine molecules. d, The triacylated cysteine at the N terminus of ActE shown along with 15 downstream amino acids. Notably, residue Tyr28 is in contact with the covalent lipid of ActE. Attachment of ActE to the membrane may also be assisted by aromatic residues Tyr30 and Phe31, which appear to be inserted into the lipid bilayer. Throughout the molecular dynamics simulation trajectory, these residues remain buried in the lipid bilayer.
Extended Data Fig. 8 Protein stability and lipid–protein interaction analysis based on molecular dynamics simulations.
a, Root-mean-square deviation (r.m.s.d.) of the protein backbone heavy atoms for the entire ACIII complex and each subunit, aligned based on ACIII backbone heavy atoms from three independent molecular dynamics simulations. b, Same as a, but aligned using the backbone heavy atoms of each subunit. c, Superposition of the initial (black) and final (coloured) conformations of each subunit after 250 ns of simulation (aligned using backbone heavy atoms). d, The lipid–protein contact number defined by the number of lipid atoms within 4 Å of the protein atoms calculated over the time course of the simulation. This contact number is either calculated for the eleven lipids resolved by cryo-EM (top) or all membrane lipids (bottom). e, The lipid–protein contact number for each of the eleven cryo-EM resolved lipids. f, Isosurfaces (50%) of the atom-occupancy map for the lipid anchors (orange), cryo-EM resolved lipids (red) and other membrane lipids (purple), calculated using the last 230 ns of the simulation trajectory. The stronger the lipid–protein interactions, the longer the local residence time, which leads to higher atom-occupancy values. ACIII subunits C, D, and F are shown in silver. For all plots, the raw data are shown as translucent thin lines and the block-averages are shown as dark lines.
Extended Data Fig. 9 Structural basis for supercomplex formation between the ACIII and the cyt aa3.
a, Two contact areas between the ACIII and the cyt aa3: the transmembrane portion of subunit III (red) and the loop from subunit III (orange). A homology model of subunit III fits the transmembrane density. The loop is modelled to the cryo-EM density. The sequence of the peptide is also shown. Trp188 and Phe189 are used to register the density to the sequence. b, Model of the ACIII–cyt aa3 supercomplex. The cyt aa3 structure from Rba. sphaeroides was positioned based on the transmembrane portion of subunit III. α-helices 1 and 2 of subunit III are omitted to avoid steric clashes with the ACIII structure. c, Sequence alignment of subunit III with a long loop (highlighted with the orange bar) between α-helix 5 and α-helix 6 (numbered according to subunit III from Rba. sphaeroides). Trp188 (black arrow) is conserved. d, Sequence alignment of ActB from organisms with a long loop in subunit III of their cyt aa3 oxidase. Arg868 (red arrow) is largely conserved with occasional substitution to lysine.
Supplementary information
Supplementary Information
This file contains Supplementary Discussions 1-14
Rights and permissions
About this article
Cite this article
Sun, C., Benlekbir, S., Venkatakrishnan, P. et al. Structure of the alternative complex III in a supercomplex with cytochrome oxidase. Nature 557, 123–126 (2018). https://doi.org/10.1038/s41586-018-0061-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0061-y
This article is cited by
-
Structures of the CcmABCD heme release complex at multiple states
Nature Communications (2022)
-
Synthesis, antifungal activity, and molecular dynamics study of novel geranyl aromatic sulfonamide compounds as potential complex III inhibitors
Medicinal Chemistry Research (2022)
-
Targeting the cytochrome bc1 complex for drug development in M. tuberculosis: review
Molecular Diversity (2022)
-
Architecture of bacterial respiratory chains
Nature Reviews Microbiology (2021)
-
Lipidomic and in-gel analysis of maleic acid co-polymer nanodiscs reveals differences in composition of solubilized membranes
Communications Biology (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.