Abstract
For more than a century, certain bacterial infections that can breach the skin and mucosal barriers have been implicated as common triggers of autoimmune syndromes, especially post-infection autoimmune diseases that include rheumatic fever and post-streptococcal glomerulonephritis. However, only in the past few years has the importance of imbalances within our own commensal microbiota communities, and within the gut, in the absence of infection, in promoting autoimmune pathogenesis become fully appreciated. A diversity of species and mechanisms have been implicated, including disruption of the gut barrier. Emerging data suggest that expansions (or blooms) of pathobiont species are involved in autoimmune pathogenesis and stimulate clonal expansion of T cells and B cells that recognize microbial antigens. This Review discusses the relationship between the gut microbiome and the immune system, and the potential consequence of disrupting the community balance in terms of autoimmune development, focusing on systemic lupus erythematosus. Notably, inter-relationships between expansions of certain members within gut microbiota communities and concurrent autoimmune responses bear features reminiscent of classical post-infection autoimmune disease. From such insights, new therapeutic opportunities are being considered to restore the balance within microbiota communities or re-establishing the gut-barrier integrity to reinforce immune homeostasis in the host.
Key points
-
Systemic lupus erythematosus (SLE) development involves many genetic and environmental factors, and emerging data implicate various gut pathobionts as inciting and exacerbating factors.
-
Emerging data highlight parallels between autoimmune diseases such as SLE and post-streptococcal illnesses including rheumatic fever.
-
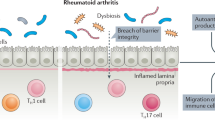
Increased permeability of the gut barrier allows microbes and microbial metabolites to interact with host immune cells, subsequently driving pathological progression.
-
Additional longitudinal studies will facilitate more in-depth elucidation of the inter-relationships between SLE and gut dysbiosis in individual patients.
-
The impact of dietary factors, antibiotics and other exposures on microbial resilience and dysbiosis, and subsequent development or flares of autoimmune disease, are also important topics for future research.
-
Modulating the gut microbiome of patients with SLE holds promise as a therapeutic intervention that could provide a more targeted and effective approach to treating the disease than broad immunosuppression.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Barber, M. R. W. et al. Global epidemiology of systemic lupus erythematosus. Nat. Rev. Rheumatol. 17, 515–532 (2021).
Dinse, G. E. et al. Increasing prevalence of antinuclear antibodies in the United States. Arthritis Rheumatol. 72, 1026–1035 (2020).
Aringer, M. et al. 2019 European League Against Rheumatism/American College of Rheumatology classification criteria for systemic lupus erythematosus. Arthritis Rheumatol. 71, 1400–1412 (2019).
Wallace, D. & Hahn, B. H. Dubois’ lupus erythematosus and related syndromes e-book: expert consult-online. (Elsevier Health Sciences, 2012).
Miyauchi, E., Shimokawa, C., Steimle, A., Desai, M. S. & Ohno, H. The impact of the gut microbiome on extra-intestinal autoimmune diseases. Nat. Rev. Immunol. 23, 9–23 (2023).
Anders, H.-J. et al. Lupus nephritis. Nat. Rev. Dis. Prim. 6, 7 (2020).
Reichlin, M., Harley, J. B. & Lockshin, M. D. Serologic studies of monozygotic twins with systemic lupus erythematosus. Arthritis Rheum. 35, 457–464 (1992).
Kuo, C. F. et al. Familial aggregation of systemic lupus erythematosus and coaggregation of autoimmune diseases in affected families. JAMA Intern. Med. 175, 1518–1526 (2015).
Ulff-Moller, C. J., Svendsen, A. J., Viemose, L. N. & Jacobsen, S. Concordance of autoimmune disease in a nationwide Danish systemic lupus erythematosus twin cohort. Semin. Arthritis Rheum. 47, 538–544 (2018).
Chen, L. et al. Genome-wide assessment of genetic risk for systemic lupus erythematosus and disease severity. Hum. Mol. Genet. 29, 1745–1756 (2020).
Langefeld, C. D. et al. Transancestral mapping and genetic load in systemic lupus erythematosus. Nat. Commun. 8, 16021 (2017).
Demkova, K., Morris, D. L. & Vyse, T. J. Genetics of SLE: does this explain susceptibility and severity across racial groups? Rheumatology 62, i15–i21 (2023).
Parks, C. G., de Souza Espindola Santos, A., Barbhaiya, M. & Costenbader, K. H. Understanding the role of environmental factors in the development of systemic lupus erythematosus. Best. Pract. Res. Clin. Rheumatol. 31, 306–320 (2017).
Azzouz, D. et al. Longitudinal gut microbiome analyses and blooms of pathogenic strains during lupus disease flares. Ann. Rheum. Dis. 82, 1315–1327 (2023).
Dogra, S. K., Doré, J. & Damak, S. Gut microbiota resilience: definition, link to health and strategies for intervention. Front. Microbiol. 11, 572921 (2020).
Cunningham MW. in: Ferretti JJ, Stevens DL, Fischetti VA, (eds.) Streptococcus Pyogenes: Basic Biology to Clinical Manifestations. (University of Oklahoma Health Sciences Center, 2016)
Muhamed, B., Parks, T. & Sliwa, K. Genetics of rheumatic fever and rheumatic heart disease’. Nat. Rev. Cardiol. 17, 145–154 (2019).
Karthikeyan, G. & Guilherme, L. Acute rheumatic fever. Lancet 392, 161–174 (2018).
Gewitz, M. H. et al. Revision of the Jones Criteria for the diagnosis of acute rheumatic fever in the era of Doppler echocardiography: a scientific statement from the American Heart Association. Circulation 131, 1806–1818 (2015).
Wells, W. C. Observations on the dropsy which succeeds scarlet fever. Trans. Soc. Imp. Med. Chir. Knowl. 3, 167–186 (1812).
Rodriguez-Iturbe, B. Autoimmunity in acute poststreptococcal GN: a neglected aspect of the disease. J. Am. Soc. Nephrol. 32, 534–542 (2021).
Cunningham, M. W. Molecular mimicry, autoimmunity, and infection: the cross-reactive antigens of group A streptococci and their sequelae. Microbiol. Spectr. 7, 7.4. 20 (2019).
Breed, E. R. & Binstadt, B. A. Autoimmune valvular carditis. Curr. Allergy Asthma Rep. 15, 491 (2015).
Teixeira, A. L., Vasconcelos, L. P., Nunes, M. & Singer, H. Sydenham’s chorea: from pathophysiology to therapeutics. Expert. Rev. Neurother. 21, 913–922 (2021).
Hanly, J. G., Kozora, E., Beyea, S. D. & Birnbaum, J. Review: nervous system disease in systemic lupus erythematosus: current status and future directions. Arthritis Rheumatol. 71, 33–42 (2019).
Psychos, D. N., Voulgari, P. V., Skopouli, F. N., Drosos, A. A. & Moutsopoulos, H. M. Erythema nodosum: the underlying conditions. Clin. Rheumatol. 19, 212–216 (2000).
Stull, C., Sprow, G. & Werth, V. P. Cutaneous involvement in systemic lupus erythematosus: a review for the rheumatologist. J. Rheumatol. 50, 27–35 (2023).
Gibofsky, A. & Zabriskie, J. B. Rheumatic fever and poststreptococcal reactive arthritis. Curr. Opin. Rheumatol. 7, 299–305 (1995).
Ceccarelli, F. et al. Joint involvement in systemic lupus erythematosus: from pathogenesis to clinical assessment. Semin. Arthritis Rheum. 47, 53–64 (2017).
Benedek, T. G. Subcutaneous nodules and the differentiation of rheumatoid arthritis from rheumatic fever. Semin. Arthritis Rheum. 13, 305–321 (1984).
Hahn, B. H., Yardley, J. H. & Stevens, M. B. “Rheumatoid” nodules in systemic lupus erythematosus. Ann. Intern. Med. 72, 49–58 (1970).
Yoshinoya, S. & Pope, R. M. Detection of immune complexes in acute rheumatic fever and their relationship to HLA-B5. J. Clin. Invest. 65, 136–145 (1980).
Kavai, M. & Szegedi, G. Immune complex clearance by monocytes and macrophages in systemic lupus erythematosus. Autoimmun. Rev. 6, 497–502 (2007).
Malik, A., Brudvig, J. M., Gadsden, B. J., Ethridge, A. D. & Mansfield, L. S. Campylobacter jejuni induces autoimmune peripheral neuropathy via Sialoadhesin and Interleukin-4 axes. Gut microbes 14, 2064706 (2022).
D’Elios, M. M., Appelmelk, B. J., Amedei, A., Bergman, M. P. & Del Prete, G. Gastric autoimmunity: the role of Helicobacter pylori and molecular mimicry. Trends Mol. Med. 10, 316–323 (2004).
Jarrot, P.-A. & Kaplanski, G. Pathogenesis of ANCA-associated vasculitis: an update. Autoimmun. Rev. 15, 704–713 (2016).
Pereira, M. S. & Kriegel, M. A. Evolving concepts of host–pathobiont interactions in autoimmunity. Curr. Opin. Immunol. 80, 102265 (2023).
Ruff, W. E., Greiling, T. M. & Kriegel, M. A. Host–microbiota interactions in immune-mediated diseases. Nat. Rev. Microbiol. 18, 521–538 (2020).
Wesemann, D. R. et al. Microbial colonization influences early B-lineage development in the gut lamina propria. Nature 501, 112–115 (2013).
Levy, M., Kolodziejczyk, A. A., Thaiss, C. A. & Elinav, E. Dysbiosis and the immune system. Nat. Rev. Immunol. 17, 219–232 (2017).
Zheng, D., Liwinski, T. & Elinav, E. Interaction between microbiota and immunity in health and disease. Cell Res. 30, 492–506 (2020).
Gillbert, J. A. et al. Current understanding of the human microbiome. Nat. Med. 24, 392–400 (2018).
Atarashi, K. et al. Induction of colonic regulatory T cells by indigenous Clostridium species. Science 331, 337–341 (2011).
Geuking, M. B. et al. Intestinal bacterial colonization induces mutualistic regulatory T cell responses. Immunity 34, 794–806 (2011).
Ohno, H. The impact of metabolites derived from the gut microbiota on immune regulation and diseases. Int. Immunol. 32, 629–636 (2020).
Arpaia, N. et al. Metabolites produced by commensal bacteria promote peripheral regulatory T-cell generation. Nature 504, 451–455 (2013).
Furusawa, Y., Obata, Y. & Hase, K. Commensal microbiota regulates T cell fate decision in the gut. Semin. Immunopathol. 37, 17–25 (2015).
Krautkramer, K. A., Fan, J. & Bäckhed, F. Gut microbial metabolites as multi-kingdom intermediates. Nat. Rev. Microbiol. 19, 77–94 (2021).
Zelante, T. et al. Tryptophan catabolites from microbiota engage aryl hydrocarbon receptor and balance mucosal reactivity via interleukin-22. Immunity 39, 372–385 (2013).
Choi, S. C. et al. Gut microbiota dysbiosis and altered tryptophan catabolism contribute to autoimmunity in lupus-susceptible mice. Sci. Transl. Med. 12, eaax2220 (2020).
Lind, N. A., Rael, V. E., Pestal, K., Liu, B. & Barton, G. M. Regulation of the nucleic acid-sensing Toll-like receptors. Nat. Rev. Immunol. 22, 224–235 (2022).
El-Zayat, S. R., Sibaii, H. & Mannaa, F. A. Toll-like receptors activation, signaling, and targeting: an overview. Bull. Natl Res. Cent. 43, 1–12 (2019).
Elloumi, N. et al. Role of innate immune receptors TLR4 and TLR2 polymorphisms in systemic lupus erythematosus susceptibility. Ann. Hum. Genet. 86, 137–144 (2022).
Wesemann, D. R. Microbes and B cell development. Adv. Immunol. 125, 155–178 (2015).
Greiling, T. M. et al. Commensal orthologs of the human autoantigen Ro60 as triggers of autoimmunity in lupus. Sci. Transl. Med. 10, eaan2306 (2018).
Maynard, C. L., Elson, C. O., Hatton, R. D. & Weaver, C. T. Reciprocal interactions of the intestinal microbiota and immune system. Nature 489, 231–241 (2012).
Kennedy, E., King, K. Y. & Baldrige, M. T. Mouse microbiota models: comparing germ-free mice and antibiotics treatment as tools for modifying gut bacteria. Front. Physiol. 9, 1534 (2018).
Chung, H. et al. Gut immune maturation depends on colonization with a host-specific microbiota. Cell 149, 1578–1593 (2012).
Ma, Y. et al. Gut microbiota promote the inflammatory response in the pathogenesis of systemic lupus erythematosus. Mol. Med. 25, 35 (2019).
Ma, Y. et al. Lupus gut microbiota transplants cause autoimmunity and inflammation. Clin. Immunol. 233, 108892 (2021).
Moeller, A. H. et al. SIV-induced instability of the chimpanzee gut microbiome. Cell Host Microbe 14, 340–345 (2013).
Lederberg, J. & McCray, A. T. Ome SweetOmics — A genealogical treasury of words. Scientist 15, 8–8 (2001).
Wells, J. M. Homeostasis of the gut barrier and potential biomarkers. Am. J. Physiol. Gastrointest. Liver Physiol. 312, G171–G193 (2017).
Sterlin, D. et al. Human IgA binds a diverse array of commensal bacteria. J. Exp. Med. 217, e20181635 (2020).
Sonnenberg, G. F. et al. Innate lymphoid cells promote anatomical containment of lymphoid-resident commensal bacteria. Science 336, 1321–1325 (2012).
Fung, T. C. et al. Lymphoid-tissue-resident commensal bacteria promote members of the IL-10 cytokine family to establish mutualism. Immunity 44, 634–646 (2016).
Manfredo Vieira, S. et al. Translocation of a gut pathobiont drives autoimmunity in mice and humans. Science 359, 1156–1161 (2018).
Azzouz, D. et al. Lupus nephritis is linked to disease-activity associated expansions and immunity to a gut commensal. Ann. Rheum. Dis. 78, 947–956 (2019).
Mishra, S. P. et al. Microbiota induces aging-related leaky gut and inflammation by dampening mucin barriers and butyrate-FFAR2/3 signaling. Preprint at bioRxiv https://doi.org/10.1101/2021.08.18.456856 (2021).
Ma, L. & Morel, L. Loss of gut barrier integrity in lupus. Front. Immunol. 13, 919792 (2022).
Natividad, J. M. & Verdu, E. F. Modulation of intestinal barrier by intestinal microbiota: pathological and therapeutic implications. Pharmacol. Res. 69, 42–51 (2013).
Flannigan, K. L. & Denning, T. L. Segmented filamentous bacteria‐induced immune responses: a balancing act between host protection and autoimmunity. Immunol 154, 537–546 (2018).
Gaboriau-Routhiau, V. et al. The key role of segmented filamentous bacteria in the coordinated maturation of gut helper T cell responses. Immunity 31, 677–689 (2009).
Ivanov, I. I. et al. Induction of intestinal Th17 cells by segmented filamentous bacteria. Cell 139, 485–498 (2009).
Valiente, G. R. et al. Gut dysbiosis is associated with acceleration of lupus nephritis. Sci. Rep. 12, 152 (2022).
Mu, Q. et al. Control of lupus nephritis by changes of gut microbiota. Microbiome 5, 73 (2017).
Zegarra-Ruiz, D. F. et al. A diet-sensitive commensal lactobacillus strain mediates TLR7-dependent systemic autoimmunity. Cell Host Microbe 25, 113–127.e6 (2019).
Thim-Uam, A. et al. Leaky-gut enhanced lupus progression in the Fc gamma receptor-IIb deficient and pristane-induced mouse models of lupus. Sci. Rep. 10, 1–18 (2020).
Brown, J. Microbiota-mediated skewing of tryptophan catabolism modulates CD4+ T cells in lupus-prone mice. iScience 25, 104241 (2022).
Yang, Y. et al. Within-host evolution of a gut pathobiont facilitates liver translocation. Nature 607, 563–570 (2022).
Nakamoto, N. et al. Gut pathobionts underlie intestinal barrier dysfunction and liver T helper 17 cell immune response in primary sclerosing cholangitis. Nat. Microbiol. 4, 492–503 (2019).
Bagavant, H. et al. Immune response to enterococcus gallinarum in lupus patients is associated with a subset of lupus-associated autoantibodies. Front. Immunol. 12, 635072 (2021).
Tajik, N. et al. Targeting zonulin and intestinal epithelial barrier function to prevent onset of arthritis. Nat. Commun. 11, 1–14 (2020).
Audo, R. et al. Rheumatoid arthritis is associated with increased gut permeability and bacterial translocation that are reversed by inflammation control. Rheumatology 62, 1264–1271 (2023).
Romero-Figueroa, M. D. S., Ramírez-Durán, N., Montiel-Jarquín, A. J. & Horta-Baas, G. Gut-joint axis: gut dysbiosis can contribute to the onset of rheumatoid arthritis via multiple pathways. Front. Cell. Infect. Microbiol. 13, 56 (2023).
Ogunrinde, E. et al. A link between plasma microbial translocation, microbiome, and autoantibody development in first-degree relatives of systemic lupus erythematosus patients. Arthritis Rheumatol. 71, 1858–1868 (2019).
Troisi, J. et al. The therapeutic use of the zonulin inhibitor AT-1001 (Larazotide) for a variety of acute and chronic inflammatory diseases. Curr. Med. Chem. 28, 5788–5807 (2021).
Fasano, A. All disease begins in the (leaky) gut: role of zonulin-mediated gut permeability in the pathogenesis of some chronic inflammatory diseases [version 1; peer review: 3 approved]. F1000Research https://doi.org/10.12688/f1000research.20510.1 (2020).
Silverman, G. J., Deng, J. & Azzouz, D. F. Sex-dependent Lupus Ruminococcus blautia gnavus strain induction of zonulin-mediated intestinal permeability and autoimmunity. Front. Immunol. 13, 897971 (2022).
Clancy, R. M. et al. Gut dysbiosis and the clinical spectrum in anti-Ro positive mothers of children with neonatal lupus. Gut Microbes 14, 2081474 (2022).
Chen, Y. et al. Gut microbiota in systemic lupus erythematosus: a fuse and a solution. J. Autoimmun. 132, 102867 (2022).
Guo, M. et al. Alteration in gut microbiota is associated with dysregulation of cytokines and glucocorticoid therapy in systemic lupus erythematosus. Gut Microbes 11, 1758–1773 (2020).
Hevia, A. et al. Intestinal dysbiosis associated with systemic lupus erythematosus. mBio 5, e01548–01514 (2014).
He, Z., Shao, T., Li, H., Xie, Z. & Wen, C. Alterations of the gut microbiome in Chinese patients with systemic lupus erythematosus. Gut Pathog. 8, 1–7 (2016).
Vujkovic-Cvijin, I. et al. Host variables confound gut microbiota studies of human disease. Nature 587, 448–454 (2020).
Luo, X. M. et al. Gut microbiota in human systemic lupus erythematosus and a mouse model of lupus. Appl. Environ. Microbiol. 84, e02288–e02317 (2018).
Bellocchi, C. et al. Identification of a shared microbiomic and metabolomic profile in systemic autoimmune diseases. J. Clin. Med. 8, 1291 (2019).
van der Meulen, T. A. et al. Shared gut, but distinct oral microbiota composition in primary Sjögren’s syndrome and systemic lupus erythematosus. J. Autoimmun. 97, 77–87 (2019).
Li, Y. et al. Disordered intestinal microbes are associated with the activity of systemic lupus erythematosus. Clin. Sci. 133, 821–838 (2019).
Zhang, H., Liao, X., Sparks, J. B. & Luo, X. M. Dynamics of gut microbiota in autoimmune lupus. Appl. Environ. Microbiol. 80, 7551–7560 (2014).
Sokol, H. et al. Faecalibacterium prausnitzii is an anti-inflammatory commensal bacterium identified by gut microbiota analysis of Crohn disease patients. Proc. Natl Acad. Sci. USA 105, 16731–16736 (2008).
Chen, B. D. et al. An autoimmunogenic and proinflammatory profile defined by the gut microbiota of patients with untreated systemic lupus erythematosus. Arthritis Rheumatol. 73, 232–243 (2021).
Silverman, G.J., Azzouz, D., Grönwall, C., Gunnarsson, I. & Svenungsson, E. Validation of a serologic antibody biomarker against a candidate gut pathobiont for the diagnosis of lupus nephritis [abstract]. Arthritis Rheumatol. 71 (2019).
Toumi, E. et al. Gut microbiota in systemic lupus erythematosus patients and lupus mouse model: a cross species comparative analysis for biomarker discovery. Front. Immunol. 13, 943241 (2022).
Boccitto, M. & Wolin, S. L. Ro60 and Y RNAs: structure, functions, and roles in autoimmunity. Crit. Rev. Biochem. Mol. Biol. 54, 133–152 (2019).
Kim, J. W., Kwok, S. K., Choe, J. Y. & Park, S. H. Recent advances in our understanding of the link between the intestinal microbiota and systemic lupus erythematosus. Int. J. Mol. Sci. 20, 4871 (2019).
Moore, W. C., Johnson, J. & Holdeman, L. Emendation of Bacteroidaceae and Butyrivibrio and descriptions of Desulfomonas gen. nov. and ten new species in the genera Desulfomonas, Butyrivibrio, Eubacterium, Clostridium, and Ruminococcus. Int. J. Syst. Evolut. Microbiol. 26, 238–252 (1976).
Togo, A. H. et al. Description of Mediterraneibacter massiliensis, gen. nov., sp. nov., a new genus isolated from the gut microbiota of an obese patient and reclassification of Ruminococcus faecis, Ruminococcus lactaris, Ruminococcus torques, Ruminococcus gnavus and Clostridium glycyrrhizinilyticum as Mediterraneibacter faecis comb. nov., Mediterraneibacter lactaris comb. nov., Mediterraneibacter torques comb. nov., Mediterraneibacter gnavus comb. nov. and Mediterraneibacter glycyrrhizinilyticus comb. nov. Antonie Van. Leeuwenhoek 111, 2107–2128 (2018).
Crost, E. H., Coletto, E., Bell, A. & Juge, N. Ruminococcus gnavus: friend or foe for human health. FEMS Microbiol. Rev. 47, fuad014 (2023).
Liu, C., Finegold, S. M., Song, Y. & Lawson, P. A. Reclassification of Clostridium coccoides, Ruminococcus hansenii, Ruminococcus hydrogenotrophicus, Ruminococcus luti, Ruminococcus productus and Ruminococcus schinkii as Blautia coccoides gen. nov., comb. nov., Blautia hansenii comb. nov., Blautia hydrogenotrophica comb. nov., Blautia luti comb. nov., Blautia producta comb. nov., Blautia schinkii comb. nov. and description of Blautia wexlerae sp. nov., isolated from human faeces. Int. J. Syst. Evol. Microbiol. 58, 1896–1902 (2008).
Qin, J. et al. A human gut microbial gene catalogue established by metagenomic sequencing. Nature 464, 59–65 (2010).
Hall, A. B. et al. A novel Ruminococcus gnavus clade enriched in inflammatory bowel disease patients. Genome Med. 9, 1–12 (2017).
Joossens, M. et al. Dysbiosis of the faecal microbiota in patients with Crohn’s disease and their unaffected relatives. Gut 60, 631–637 (2011).
Nishino, K. et al. Analysis of endoscopic brush samples identified mucosa-associated dysbiosis in inflammatory bowel disease. J. Gastroenterol. 53, 95–106 (2018).
Franzosa, E. A. et al. Gut microbiome structure and metabolic activity in inflammatory bowel disease. Nat. Microbiol. 4, 293–305 (2019).
Lloyd-Price, J. et al. Multi-omics of the gut microbial ecosystem in inflammatory bowel diseases. Nature 569, 655–662 (2019).
Breban, M. et al. Faecal microbiota study reveals specific dysbiosis in spondyloarthritis. Ann. Rheum. Dis. 76, 1614–1622 (2017).
Henke, M. T. et al. Ruminococcus gnavus, a member of the human gut microbiome associated with Crohn’s disease, produces an inflammatory polysaccharide. Proc. Natl Acad. Sci. USA 116, 12672–12677 (2019).
Henke, M. T. et al. Capsular polysaccharide correlates with immune response to the human gut microbe Ruminococcus gnavus. Proc. Natl Acad. Sci. USA 118, e2007595118 (2021).
Bell, A. et al. Elucidation of a sialic acid metabolism pathway in mucus-foraging Ruminococcus gnavus unravels mechanisms of bacterial adaptation to the gut. Nat. Microbiol. 4, 2393–2404 (2019).
Tailford, L. E. et al. Discovery of intramolecular trans-sialidases in human gut microbiota suggests novel mechanisms of mucosal adaptation. Nat. Commun. 6, 7624 (2015).
Owen, C. D. et al. Unravelling the specificity and mechanism of sialic acid recognition by the gut symbiont Ruminococcus gnavus. Nat. Commun. 8, 2196 (2017).
Thompson, K. N. et al. Alterations in the gut microbiome implicate key taxa and metabolic pathways across inflammatory arthritis phenotypes. Sci. Transl. Med. 15, eabn4722 (2023).
Vatanen, T. et al. The human gut microbiome in early-onset type 1 diabetes from the TEDDY study. Nature 562, 589–594 (2018).
McCarthy, S. et al. Altered skin and gut microbiome in hidradenitis suppurativa. J. Invest. Dermatol. 142, 459–468 e415 (2022).
Liu, Q. et al. Gut microbiota dynamics in a prospective cohort of patients with post-acute COVID-19 syndrome. Gut 71, 544–552 (2022).
Tiniakou, E., Costenbader, K. H. & Kriegel, M. A. Sex-specific environmental influences on the development of autoimmune diseases. Clin. Immunol. 149, 182–191 (2013).
Sutcliffe, I. C. & Russell, R. Lipoproteins of Gram-positive bacteria. J. Bacteriol. 177, 1123–1128 (1995).
Gisch, N. et al. Structural reevaluation of Streptococcus pneumoniae lipoteichoic acid and new insights into its immunostimulatory potency. J. Biol. Chem. 288, 15654–15667 (2013).
Gisch, N. et al. Structural analysis and immunostimulatory potency of lipoteichoic acids isolated from three Streptococcus suis serotype 2 strains. J. Biol. Chem. 293, 12011–12025 (2018).
Vujkovic-Cvijin, I. et al. The systemic anti-microbiota IgG repertoire can identify gut bacteria that translocate across gut barrier surfaces. Sci. Transl. Med. 14, eabl3927 (2022).
Chriswell, M. E. et al. Clonal IgA and IgG autoantibodies from individuals at risk for rheumatoid arthritis identify an arthritogenic strain of Subdoligranulum. Sci. Transl. Med. 14, eabn5166 (2022).
Silverman, G. J., Deng, J. & Azzouz, D. F. Sex-dependent Lupus Blautia (Ruminococcus) gnavus strain induction of zonulin-mediated intestinal permeability and autoimmunity. Front. Immunol. 13, 897971 (2022).
Amarnani, A. et al. Lupus nephritis flares linked to platelet activation during gut pathobiont blooms [abstract]. Arthritis Rheumatol. 75 (2023).
Sasso, E. H., Silverman, G. J. & Mannik, M. Human IgM molecules that bind staphylococcal protein A contain VHIII H chains. J. Immunol. 142, 2778–2783 (1989).
Graille, M. et al. Crystal structure of a Staphylococcus aureus protein A domain complexed with the Fab fragment of a human IgM antibody: structural basis for recognition of B-cell receptors and superantigen activity. Proc. Natl Acad. Sci. USA 97, 5399–5404 (2000).
Silverman, G. J. & Goodyear, C. S. Confounding B-cell defences: lessons from a staphylococcal superantigen. Nat. Rev. Immunol. 6, 465–475 (2006).
Pauli, N. T. et al. Staphylococcus aureus infection induces protein A-mediated immune evasion in humans. J. Exp. Med. 211, 2331–2339 (2014).
Ulloa-Morales, A., Goodyear, C. & Silverman, G. Essential domain-dependent roles within soluble IgG for in vivo superantigen properties of staphylococcal protein A: resolving the B-cell superantigen paradox. Front. Immunol. 9, 2011 (2018).
Bunker, J. J. et al. B cell superantigens in the human intestinal microbiota. Sci. Transl. Med. 11, eaau9356 (2019).
Borowska, M. T. The molecular characterization of antibody binding to a superantigen-like protein from a commensal microbe. Proc. Natl Acad. Sci. USA 118, e2023898118 (2021).
Piróg, A. et al. Two bacterial small heat shock proteins, IbpA and IbpB, form a functional heterodimer. J. Mol. Biol. 433, 167054 (2021).
Wen, M. et al. Correlation analysis between gut microbiota and metabolites in children with systemic lupus erythematosus. J. Immunol. Res. 2021, 5579608 (2021).
Tsigalou, C., Konstantinidis, T., Aloizou, A.-M., Bezirtzoglou, E. & Tsakris, A. in: Role of Microorganisms in Pathogenesis and Management of Autoimmune Diseases: Volume II: Kidney, Central Nervous System, Eye, Blood, Blood Vessels & Bowel. 489–520 (Springer, 2023).
Mishima, Y. & Sartor, R. B. Manipulating resident microbiota to enhance regulatory immune function to treat inflammatory bowel diseases. J. Gastroenterol. 55, 4–14 (2020).
Federici, S. et al. Targeted suppression of human IBD-associated gut microbiota commensals by phage consortia for treatment of intestinal inflammation. Cell 185, 2879–2898. e2824 (2022).
Rohlke, F. & Stollman, N. Fecal microbiota transplantation in relapsing Clostridium difficile infection. Ther. Adv. Gastroenterol. 5, 403–420 (2012).
Huang, C. et al. Safety and efficacy of fecal microbiota transplantation for treatment of systemic lupus erythematosus: an EXPLORER trial. J. Autoimmun. 130, 102844 (2022).
Sokol, H. et al. Fecal microbiota transplantation to maintain remission in Crohn’s disease: a pilot randomized controlled study. Microbiome 8, 1–14 (2020).
Zeng, J. et al. Fecal microbiota transplantation for rheumatoid arthritis: a case report. Clin. Case Rep. 9, 906–909 (2021).
Dailey, F. E., Turse, E. P., Daglilar, E. & Tahan, V. The dirty aspects of fecal microbiota transplantation: a review of its adverse effects and complications. Curr. Opin. Pharmacol. 49, 29–33 (2019).
Clark, R. L. et al. Design of synthetic human gut microbiome assembly and butyrate production. Nat. Commun. 12, 1–16 (2021).
Coras, R. et al. Baseline microbiome and metabolome are associated with response to ITIS diet in an exploratory trial in patients with rheumatoid arthritis. Clin. Transl. Med. 12, e959 (2022).
Fasano, A. Zonulin and its regulation of intestinal barrier function: the biological door to inflammation, autoimmunity, and cancer. Physiol. Rev. 91, 151–175 (2011).
Hoilat, G. J. et al. Larazotide acetate for treatment of celiac disease: a systematic review and meta-analysis of randomized controlled trials. Clin. Res. Hepatol. Gastroenterol. 46, 101782 (2022).
Hagan, T. et al. Antibiotics-driven gut microbiome perturbation alters immunity to vaccines in humans. Cell 178, 1313–1328. e1313 (2019).
Yassour, M. et al. Natural history of the infant gut microbiome and impact of antibiotic treatment on bacterial strain diversity and stability. Sci. Transl. Med. 8, 343ra381–343ra381 (2016).
Jackson, M. A. et al. Gut microbiota associations with common diseases and prescription medications in a population-based cohort. Nat. Commun. 9, 2655 (2018).
Johnson, B. M., Gaudreau, M. C., Al-Gadban, M. M., Gudi, R. & Vasu, C. Impact of dietary deviation on disease progression and gut microbiome composition in lupus-prone SNF1 mice. Clin. Exp. Immunol. 181, 323–337 (2015).
Um, C. Y. et al. Association of emulsifier and highly processed food intake with circulating markers of intestinal permeability and inflammation in the cancer prevention study-3 diet assessment sub-study. Nutr. Cancer 74, 1701–1711 (2022).
Silverman, G. J., Azzouz, D. F. & Alekseyenko, A. V. Systemic lupus erythematosus and dysbiosis in the microbiome: cause or effect or both? Curr. Opin. Immunol. 61, 80–85 (2019).
Wei, F. et al. Changes of intestinal flora in patients with systemic lupus erythematosus in northeast China. PLoS One 14, e0213063 (2019).
Garabatos, N. & Santamaria, P. Gut microbial antigenic mimicry in autoimmunity. Front. Immunol. 13, 873607 (2022).
Van Praet, J. T. et al. Commensal microbiota influence systemic autoimmune responses. EMBO J. 34, 466–474 (2015).
Lécuyer, E. et al. Segmented filamentous bacterium uses secondary and tertiary lymphoid tissues to induce gut IgA and specific T helper 17 cell responses. Immunity 40, 608–620 (2014).
Acknowledgements
The central themes presented in this Review were originally presented in a special session on “Molecular mimicry, autoimmunity and the streptococcal connection: a common cause of autoimmune conditions” at the 2022 ACR Convergence meeting. The authors would like to thank J. Weiser, M. Henke and D. Littman for their advice, as well as R. Xavier and H. Vlamakis, E. Pamer, and E. Allen-Vercoe for providing R. gnavus strains for their research studies. Finally, the authors would like to thank J. Colton and S. Colton for their generous support. The work of the authors is supported by funds from National Institutes of Health Grants R01-AR42455 (G.J.S.), National Institutes of Health Grants P50-AR070591 (G.J.S.), National Institutes of Health Contract NIAID HHSN272201400019C (G.J.S.), National Institutes of Health 5T32AR069515-07 (A.A.), Lupus Research Alliance (G.J.S.), Judith and Stewart Colton Autoimmunity Center (G.J.S.), and P. Robert Majumder Charitable Trust (G.J.S.).
Author information
Authors and Affiliations
Contributions
All authors researched data for the article, contributed substantially to discussion of the content and reviewed and/or edited the manuscript before submission. G.J.S., A.A. and N.G. wrote the article.
Corresponding author
Ethics declarations
Competing interests
G.J.S. declares that NYU has been awarded US patent 11241488B2, and additional patent applications (No. 63/533,004 and 63/354,159). N.G., D.F.A and A.A. declare no competing interests.
Peer review
Peer review information
Nature Reviews Rheumatology thanks Xin Luo, Wei Jiang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Glossary
- B cell superantigen
-
Molecules that directly bind to and stimulate B cells via their antigen receptor, often binding to sites outside the paratopic sites that bind conventional antigens.
- Commensal microbes
-
Resident microorganisms that coexist with a host without causing harm or necessarily receiving any mutual benefit.
- Microbiome
-
The collective microorganisms, such as bacteria, fungi, viruses and protozoa, that live within and on the human body.
- Pathobiont
-
A commensal microorganism that can cause disease under certain conditions, such as following a change in the host’s immune status or following an imbalance in the gut microbiota, but does not drive overt infectious disease in the host.
- Pathogen
-
A microorganism, such as a bacterium, virus, fungus or parasite, that causes disease in its host, often through tissue destruction.
- Resilience
-
The capacity of the gut microbiome to recover and regain balance in composition and functionality after being exposed to various stressors.
- Symbiont
-
An organism that lives in a mutually beneficial relationship with another organism.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Silverman, G.J., Azzouz, D.F., Gisch, N. et al. The gut microbiome in systemic lupus erythematosus: lessons from rheumatic fever. Nat Rev Rheumatol 20, 143–157 (2024). https://doi.org/10.1038/s41584-023-01071-8
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41584-023-01071-8