Abstract
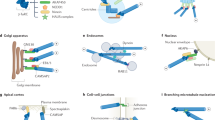
The cytoskeleton — comprising actin filaments, microtubules and intermediate filaments — serves instructive roles in regulating cell function and behaviour during development. However, a key challenge in cell and developmental biology is to dissect how these different structures function and interact in vivo to build complex tissues, with the ultimate aim to understand these processes in a mammalian organism. The preimplantation mouse embryo has emerged as a primary model system for tackling this challenge. Not only does the mouse embryo share many morphological similarities with the human embryo during its initial stages of life, it also permits the combination of genetic manipulations with live-imaging approaches to study cytoskeletal dynamics directly within an intact embryonic system. These advantages have led to the discovery of novel cytoskeletal structures and mechanisms controlling lineage specification, cell–cell communication and the establishment of the first forms of tissue architecture during development. Here we highlight the diverse organization and functions of each of the three cytoskeletal filaments during the key events that shape the early mammalian embryo, and discuss how they work together to perform key developmental tasks, including cell fate specification and morphogenesis of the blastocyst. Collectively, these findings are unveiling a new picture of how cells in the early embryo dynamically remodel their cytoskeleton with unique spatial and temporal precision to drive developmental processes in the rapidly changing in vivo environment.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Collinet, C. & Lecuit, T. Programmed and self-organized flow of information during morphogenesis. Nat. Rev. Mol. Cell. Biol. https://doi.org/10.1038/s41580-020-00318-6 (2021).
Goodwin, K. & Nelson, C. M. Mechanics of development. Dev. Cell https://doi.org/10.1016/j.devcel.2020.11.025 (2020).
Hohmann, T. & Dehghani, F. The cytoskeleton — a complex interacting meshwork. Cells 8, 362 (2019).
Simon, D. N. & Wilson, K. L. The nucleoskeleton as a genome- associated dynamic ‘network of networks’. Nat. Rev. Mol. Cell Biol. 12, 695–708 (2011).
Kunda, P. & Baum, B. The actin cytoskeleton in spindle assembly and positioning. Trends Cell Biol. 19, 174–179 (2009).
Seetharaman, S. & Etienne-Manneville, S. Cytoskeletal crosstalk in cell migration. Trends Cell Biol. https://doi.org/10.1016/j.tcb.2020.06.004 (2020).
Li, R. & Gundersen, G. G. Beyond polymer polarity: how the cytoskeleton builds a polarized cell. Nat. Rev. Mol. Cell Biol. 9, 860–873 (2008).
Parsons, J. T., Horwitz, A. R. & Schwartz, M. A. Cell adhesion: integrating cytoskeletal dynamics and cellular tension. Nat. Rev. Mol. Cell Biol. 11, 633–643 (2010).
Huber, F., Boire, A., López, M. P. & Koenderink, G. H. Cytoskeletal crosstalk: when three different personalities team up. Curr. Opin. Cell Biol. 32, 39–47 (2015).
Dogterom, M. & Koenderink, G. H. Actin–microtubule crosstalk in cell biology. Nat. Rev. Mol. Cell Biol. https://doi.org/10.1038/s41580-018-0067-1 (2018).
White, M. D., Zenker, J., Bissiere, S. & Plachta, N. Instructions for assembling the early mammalian embryo. Dev. Cell 45, 667–679 (2018).
Yamanaka, Y., Ralston, A., Stephenson, R. O. & Rossant, J. Cell and molecular regulation of the mouse blastocyst. Dev. Dyn. 235, 2301–2314 (2006).
White, M. D. & Plachta, N. Specification of the first mammalian cell lineages in vivo and in vitro. Cold Spring Harb. Perspect. Biol. 12, a035634 (2020).
Rossant, J. & Tam, P. P. L. Blastocyst lineage formation, early embryonic asymmetries and axis patterning in the mouse. Development 136, 701–713 (2009).
Chazaud, C. & Yamanaka, Y. Lineage specification in the mouse preimplantation embryo. Development 143, 1063–1074 (2016).
Ducibella, T. & Anderson, E. Cell shape and membrane changes in the eight-cell mouse embryo: prerequisites for morphogenesis of the blastocyst. Dev. Biol. 47, 45–58 (1975).
Johnson, M. H. & Ziomek, C. A. The foundation of two distinct cell lineages mouse morula. Cell 24, 71–80 (1981). A landmark article proposing a model for asymmetric inheritance of polarized regulators that specify outer and inner cells to their distinct lineages.
Vinot, S. et al. Asymmetric distribution of PAR proteins in the mouse embryo begins at the 8-cell stage during compaction. Dev. Biol. 282, 307–319 (2005).
Plusa, B. et al. Downregulation of Par3 and aPKC function directs cells towards the ICM in the pre-implantation mouse embryo. J. Cell Sci. 118, 505–515 (2005).
Thomas, F. C. et al. Contribution of JAM-1 to epithelial differentiation and tight-junction biogenesis in the mouse preimplantation embryo. J. Cell Sci. 117, 5599–5608 (2004).
Vestweber, D., Gossler, A., Boller, K. & Kemler, R. Expression and distribution of cell adhesion molecule uvomorulin in mouse preimplantation embryos. Dev. Biol. 124, 451–456 (1987).
Winkel, G. K., Ferguson, J. E., Takeichi, M. & Nuccitelli, R. Activation of protein kinase C triggers premature compaction in the four-cell stage mouse embryo. Dev. Biol. 138, 1–15 (1990).
Ohsugi, M., Ohsawa, T. & Semba, R. Similar responses to pharmacological agents of 1,2-OAG-induced compaction-like adhesion of two-cell mouse embryo to physiological compaction. J. Exp. Zool. 265, 604–608 (1993).
Shirayoshi, Y., Okada, T. S. & Takeichi, M. The calcium-dependent cell-cell adhesion system regulates inner cell mass formation and cell surface polarization in early mouse development. Cell 35, 631–638 (1983). Seminal work demonstrating the role of E-cadherin in embryo compaction and polarization.
Kemler, R., Babinet, C., Eisen, H. & Jacob, F. Surface antigen in early differentiation. Proc. Natl Acad. Sci. USA 74, 4449–4452 (1977).
Stephenson, R. O., Yamanaka, Y. & Rossant, J. Disorganized epithelial polarity and excess trophectoderm cell fate in preimplantation embryos lacking E-cadherin. Development 137, 3383–3391 (2010). An article showing that loss of both maternal and zygotic E-cadherin disrupts embryo morphology and generates overlapping apical and basal cell domains.
Anani, S., Bhat, S., Honma-Yamanaka, N., Krawchuk, D. & Yamanaka, Y. Initiation of Hippo signaling is linked to polarity rather than to cell position in the pre-implantation mouse embryo. Development 141, 2813–2824 (2014).
Korotkevich, E. et al. The apical domain is required and sufficient for the first lineage segregation in the mouse embryo. Dev. Cell 40, 1–21 (2017).
Salbreux, G., Charras, G. & Paluch, E. Actin cortex mechanics and cellular morphogenesis. Trends Cell Biol. 22, 536–545 (2012).
Thompson, D. W. On Growth and Form (Cambridge University Press, 1992).
Diz-Muñoz, A., Fletcher, D. A. & Weiner, O. D. Use the force: membrane tension as an organizer of cell shape and motility. Trends Cell Biol. 23, 47–53 (2013).
Gallicano, G. I. Composition, regulation, and function of the cytoskeleton in mammalian eggs and embryos. Front. Biosci. 6, 1089–1108 (2001).
Houliston, E., Pickering, S. J. & Maro, B. Redistribution of microtubules and pericentriolar material during the development of polarity in mouse blastomeres. J. Cell Biol. 104, 10 (1987).
Houliston, E. & Maro, B. Posttranslational modification of distinct microtubule subpopulations during cell polarization and differentiation in the mouse preimplantation embryo. J. Cell Biol. 108, 543–551 (1989).
Lehtonen, E. & Badley, R. A. Localization of cytoskeletal proteins in preimplantation mouse embryos. J. Embryol. Exp. Morphol. 55, 211–225 (1980).
Ducibella, T., Ukena, T., Karnovsky, M. & Anderson, E. Changes in cell surface and cortical cytoplasmic organization during early embryogenesis in the preimplantation mouse embryo. J. Cell Biol. 74, 153–167 (1977).
Sauvanet, C., Wayt, J., Pelaseyed, T. & Bretscher, A. Structure, regulation, and functional diversity of microvilli on the apical domain of epithelial cells. Annu. Rev. Cell Dev. Biol. 31, 593–621 (2015).
Lim, H. Y. G. et al. Keratins are asymmetrically inherited fate determinants in the mammalian embryo. Nature 585, 404–409 (2020). The first characterization of keratin filament functions within the early mouse embryo, leading to the identification of keratins as a new form of asymmetrically inherited fate regulator.
Maître, J.-L., Niwayama, R., Turlier, H., Nédélec, F. & Hiiragi, T. Pulsatile cell-autonomous contractility drives compaction in the mouse embryo. Nat. Cell Biol. 17, 849–855 (2015).
Pratt, H. P. M., Ziomek, C. A., Reeve, W. J. D. & Johnson, M. H. Compaction of the mouse embryo: an analysis of its components. J. Embryol. Exp. Morphol. 70, 113–132 (1982).
Ma, M. et al. Decreased cofilin1 expression is important for compaction during early mouse embryo development. Biochim. Biophys. Acta 1793, 1804–1810 (2009).
Fierro-González, J. C., White, M. D., Silva, J. C. & Plachta, N. Cadherin-dependent filopodia control preimplantation embryo compaction. Nat. Cell Biol. 15, 1424–1433 (2013). Discovery of actin-based cellular protrusions in the embryo important for driving compaction.
Lancaster, O. M. & Baum, B. Shaping up to divide: coordinating actin and microtubule cytoskeletal remodelling during mitosis. Semin. Cell Dev. Biol. 34, 109–115 (2014).
Stanganello, E. et al. Filopodia-based Wnt transport during vertebrate tissue patterning. Nat. Commun. 6, 1–14 (2015).
Sanders, T. A., Llagostera, E. & Barna, M. Specialized filopodia direct long-range transport of SHH during vertebrate tissue patterning. Nature 497, 628–632 (2013). Discovery of cellular protrusions directing ligand transport and cell–cell communication during chick embryo development.
Ogawa, H., Mori, T. & Shimizu, H. Effect of brefeldin-a on compaction of preimplantation mouse embryos. J. Reprod. Dev. 43, 303–310 (1997).
Sousa, P. A. D., Valdimarsson, G., Nicholson, B. J. & Kidder, G. M. Connexin trafficking and the control of gap junction assembly in mouse preimplantation embryos. Development 117, 1355–1367 (1993).
Zenker, J. et al. A microtubule-organizing center directing intracellular transport in the early mouse embryo. Science 357, 925–928 (2017).
Maro, B. & Pickering, S. J. Microtubules influence compaction in preimplantation mouse embryos. J. Embryol. Exp. Morphol. 84, 217–232 (1984).
Muroyama, A. & Lechler, T. A transgenic toolkit for visualizing and perturbing microtubules reveals unexpected functions in the epidermis. eLife 6, e29834 (2017).
Zhu, M., Leung, C. Y., Shahbazi, M. N. & Zernicka-Goetz, M. Actomyosin polarisation through PLC-PKC triggers symmetry breaking of the mouse embryo. Nat. Commun. 8, 1–16 (2017). Identification of early regulators of apical polarization in the embryo, with a proposed two-step model dissecting the interactions between actin filaments and polarity proteins.
Zhu, M. & Zernicka-Goetz, M. Building an apical domain in the early mouse embryo: Lessons, challenges and perspectives. Curr. Opin. Cell Biol. 62, 144–149 (2020).
Gerri, C. et al. Initiation of a conserved trophectoderm program in human, cow and mouse embryos. Nature https://doi.org/10.1038/s41586-020-2759-x (2020). The first demonstration of a conserved lineage specification mechanism shared by early mammalian embryos, offering novel insights into our evolutionary history.
Zhu, M. et al. Mechanism of cell polarisation and first lineage segregation in the human embryo. Preprint at bioRxiv https://doi.org/10.1101/2020.09.23.310680 (2020).
Zhu, M. et al. Developmental clock and mechanism of de novo polarization of the mouse embryo. Science 370, eabd2703 (2020).
Samarage, C. R. et al. Cortical tension allocates the first inner cells of the mammalian embryo. Dev. Cell 34, 435–447 (2015).
Ajduk, A. & Zernicka-Goetz, M. Polarity and cell division orientation in the cleavage embryo: from worm to human. Mol. Hum. Reprod. 22, 691–703 (2016).
Sutherland, A. E., Speed, T. P. & Calarco, P. G. Inner cell allocation in the mouse morula: the role of oriented division during fourth cleavage. Dev. Biol. 137, 13–25 (1990).
McDole, K., Xiong, Y., Iglesias, P. A. & Zheng, Y. Lineage mapping the pre-implantation mouse embryo by two-photon microscopy, new insights into the segregation of cell fates. Dev. Biol. 355, 239–249 (2011).
Gillies, T. E. & Cabernard, C. Cell division orientation in animals. Curr. Biol. 21, R599–R609 (2011).
Niwayama, R. et al. A tug-of-war between cell shape and polarity controls division orientation to ensure robust patterning in the mouse blastocyst. Dev. Cell 51, 564–574.e6 (2019).
Zenker, J. et al. Expanding actin rings zipper the mouse embryo for blastocyst formation. Cell 173, 776–791.e17 (2018).
McNally, F. J. Mechanisms of spindle positioning. J. Cell Biol. 200, 131–140 (2013).
Grill, S. W. & Hyman, A. A. Spindle positioning by cortical pulling forces. Dev. Cell 8, 461–465 (2005).
Hiraoka, L., Golden, W. & Magnuson, T. Spindle-pole organization during early mouse development. Dev. Biol. 133, 24–36 (1989).
Gueth-Hallonet, C. et al. γ-Tubulin is present in acentriolar MTOCs during early mouse development. J. Cell Sci. 105, 157–166 (1993).
Clift, D. & Schuh, M. A three-step MTOC fragmentation mechanism facilitates bipolar spindle assembly in mouse oocytes. Nat. Commun. 6, 7217 (2015).
Schatten, G., Simerly, C. & Schatten, H. Microtubule configurations during fertilization, mitosis, and early development in the mouse and the requirement for egg microtubule-mediated motility during mammalian fertilization. Proc. Natl Acad. Sci. USA 82, 4152–4156 (1985).
Howe, K. & Fitzharris, G. A non-canonical mode of microtubule organization operates throughout pre-implantation development in mouse. Cell Cycle 12, 1616–1624 (2013).
Manandhar, G., Schatten, H. & Sutovsky, P. Centrosome reduction during gametogenesis and its significance. Biol. Reprod. 72, 2–13 (2005).
Chaigne, A., Verlhac, M.-H. & Terret, M.-E. Spindle positioning in mammalian oocytes. Exp. Cell Res. 318, 1442–1447 (2012).
Almonacid, M., Terret, M. E. & Verlhac, M. H. Actin-based spindle positioning: new insights from female gametes. J. Cell Sci. 127, 477–483 (2014).
Schuh, M. & Ellenberg, J. A new model for asymmetric spindle positioning in mouse oocytes. Curr. Biol. 18, 1986–1992 (2008).
Dumont, J. et al. Formin-2 is required for spindle migration and for the late steps of cytokinesis in mouse oocytes. Dev. Biol. 301, 254–265 (2007).
Izumi, Y., Ohta, N., Hisata, K., Raabe, T. & Matsuzaki, F. Drosophila Pins-binding protein Mud regulates spindle-polarity coupling and centrosome organization. Nat. Cell Biol. 8, 586–593 (2006).
Siller, K. H., Cabernard, C. & Doe, C. Q. The NuMA-related Mud protein binds Pins and regulates spindle orientation in Drosophila neuroblasts. Nat. Cell Biol. 8, 594–600 (2006).
Nguyen-Ngoc, T., Afshar, K. & Gönczy, P. Coupling of cortical dynein and Gα proteins mediates spindle positioning in Caenorhabditis elegans. Nat. Cell Biol. 9, 1294–1302 (2007).
Kita, A. M. et al. Spindle–F-actin interactions in mitotic spindles in an intact vertebrate epithelium. Mol. Biol. Cell 30, 1645–1654 (2019).
Ajduk, A., Shivhare, S. B. & Zernicka-Goetz, M. The basal position of nuclei is one pre-requisite for asymmetric cell divisions in the early mouse embryo. Dev. Biol. 392, 133–140 (2014).
Watanabe, T., Biggins, J. S., Tannan, N. B. & Srinivas, S. Limited predictive value of blastomere angle of division in trophectoderm and inner cell mass specification. Development 141, 2279–2288 (2014).
Yamanaka, Y., Lanner, F. & Rossant, J. FGF signal-dependent segregation of primitive endoderm and epiblast in the mouse blastocyst. Development 137, 715–724 (2010).
Martin, A. C. & Goldstein, B. Apical constriction: themes and variations on a cellular mechanism driving morphogenesis. Development 141, 1987–1998 (2014).
Odell, G. M., Oster, G., Alberch, P. & Burnside, B. The mechanical basis of morphogenesis. Dev. Biol. 85, 446–462 (1981).
Mason, F. M., Tworoger, M. & Martin, A. C. Apical domain polarization localizes actin–myosin activity to drive ratchet-like apical constriction. Nat. Cell Biol. 15, 926–936 (2013).
Toyama, Y., Peralta, X. G., Wells, A. R., Kiehart, D. P. & Edwards, G. S. Apoptotic force and tissue dynamics during drosophila embryogenesis. Science 321, 1683–1686 (2008).
Dawes-Hoang, R. E. et al. folded gastrulation, cell shape change and the control of myosin localization. Development 134, 4507–4507 (2007).
Nishimura, T. & Takeichi, M. Shroom3-mediated recruitment of Rho kinases to the apical cell junctions regulates epithelial and neuroepithelial planar remodeling. Development 135, 1493–1502 (2008).
Maître, J.-L. et al. Asymmetric division of contractile domains couples cell positioning and fate specification. Nature 536, 344–348 (2016).
Seltmann, K., Fritsch, A. W., Käs, J. A. & Magin, T. M. Keratins significantly contribute to cell stiffness and impact invasive behavior. Proc. Natl Acad. Sci. USA 110, 18507–18512 (2013).
Ramms, L. et al. Keratins as the main component for the mechanical integrity of keratinocytes. Proc. Natl Acad. Sci. USA 110, 18513–18518 (2013).
Loranger, A. et al. Simple epithelium keratins are required for maintenance of hepatocyte integrity. Am. J. Pathol. 151, 11 (1997).
Coulombe, P. A. & Lee, C.-H. Defining keratin protein function in skin epithelia: epidermolysis bullosa simplex and its aftermath. J. Investig. Dermatol. 132, 763–775 (2012).
Weber, K. L. & Bement, W. M. F-actin serves as a template for cytokeratin organization in cell free extracts. J. Cell Sci. 115, 1373–1382 (2002).
Green, K. J. The relationship between intermediate filaments and microfilaments before and during the formation of desmosomes and adherens-type junctions in mouse epidermal keratinocytes. J. Cell Biol. 104, 1389–1402 (1987).
Kölsch, A., Windoffer, R. & Leube, R. E. Actin-dependent dynamics of keratin filament precursors. Cell Motil. Cytoskeleton 66, 976–985 (2009).
Mintz, B. Experimental genetic mosaicism in the mouse. in Preimplantation Stages of Pregnancy (eds Wolstenholme, G. E. W. & O’Connor, M.) 194–216 (John Wiley & Sons, Ltd, 1965).
Tarkowski, A. K. & Wroblewska, J. Development of blastomeres of mouse eggs isolated at the 4- and 8-cell stage. J. Embryol. Exp. Morphol. 18, 155–180 (1967).
Knoblich, J. A. Asymmetric cell division: recent developments and their implications for tumour biology. Nat. Rev. Mol. Cell Biol. 11, 849–860 (2010).
Kim-Ha, J., Smith, J. L. & Macdonald, P. M. oskar mRNA is localized to the posterior pole of the Drosophila oocyte. Cell 66, 23–35 (1991).
Betschinger, J., Mechtler, K. & Knoblich, J. A. Asymmetric segregation of the tumor suppressor brat regulates self-renewal in drosophila neural stem. Cells. Cell 124, 1241–1253 (2006).
Lu, B., Rothenberg, M., Jan, L. Y. & Jan, Y. N. Partner of numb colocalizes with numb during mitosis and directs numb asymmetric localization in drosophila neural and muscle progenitors. Cell 95, 225–235 (1998).
Skamagki, M., Wicher, K. B., Jedrusik, A., Ganguly, S. & Zernicka-Goetz, M. Asymmetric localization of Cdx2 mRNA during the first cell-fate decision in early mouse development. Cell Rep. 3, 442–457 (2013).
Paulin, D., Babinet, C., Weber, K. & Osborn, M. Antibodies as probes of cellular differentiation and cytoskeletal organization in the mouse blastocyst. Exp. Cell Res. 130, 297–304 (1980).
Jackson, B. W. et al. Formation of cytoskeletal elements during mouse embryogenesis: intermediate filaments of the cytokeratin type and desmosomes in preimplantation embryos. Differentiation 17, 161–179 (1980).
Oshima, R. G., Howe, W. E., Klier, G., Adamson, E. D. & Shevinsky, L. H. Intermediate filament protein synthesis in preimplantation murine embryos. Dev. Biol. 99, 447–455 (1983).
Chisholm, J. C. & Houliston, E. Cytokeratin filament assembly in the preimplantation mouse embryo. Development 101, 565–582 (1987).
Fleming, T. P., Garrod, D. R. & Elsmore, A. J. Desmosome biogenesis in the mouse preimplantation embryo. Development 112, 527–539 (1991).
Den, Z., Cheng, X., Merched-Sauvage, M. & Koch, P. J. Desmocollin 3 is required for pre-implantation development of the mouse embryo. J. Cell Sci. 119, 482–489 (2006).
Doi, M. & Edwards, S. F. The Theory of Polymer Dynamics. Vol. 73 (Oxford University Press, 1988).
Kas, J., Strey, H. & Sackmann, E. Direct imaging of reptation for semiflexible actin filaments. Nature 368, 226–229 (1994).
Deek, J., Maan, R., Loiseau, E. & Bausch, A. R. Reconstitution of composite actin and keratin networks in vesicles. Soft Matter 14, 1897–1902 (2018).
Moch, M. et al. Effects of plectin depletion on keratin network dynamics and organization. PLoS ONE 11, e0149106–e0149120 (2016).
Yang, Y. et al. An essential cytoskeletal linker protein connecting actin microfilaments to intermediate filaments. Cell 86, 655–665 (1996).
Kwan, R. et al. PKC412 normalizes mutation-related keratin filament disruption and hepatic injury in mice by promoting keratin-myosin binding. Hepatology 62, 1858–1869 (2015).
Wang, F., Chen, S., Liu, H. B., Parent, C. A. & Coulombe, P. A. Keratin 6 regulates collective keratinocyte migration by altering cell–cell and cell–matrix adhesion. J. Cell Biol. 217, 4314–4330 (2018).
Sasaki, H. Roles and regulations of Hippo signaling during preimplantation mouse development. Dev. Growth Differ. 59, 12–20 (2017).
Nishioka, N. et al. The Hippo signaling pathway components Lats and Yap pattern tead4 activity to distinguish mouse trophectoderm from inner cell mass. Dev. Cell 16, 398–410 (2009). Establishment of the Hippo signalling pathway as a key regulator of lineage specification differentiating the outer and inner cells of the embryo.
Nishioka, N. et al. Tead4 is required for specification of trophectoderm in pre-implantation mouse embryos. Mech. Dev. 125, 270–283 (2008).
Cockburn, K., Biechele, S., Garner, J. & Rossant, J. The Hippo pathway member Nf2 is required for inner cell mass specification. Curr. Biol. 23, 1195–1201 (2013).
Leung, C. Y. & Zernicka-Goetz, M. Angiomotin prevents pluripotent lineage differentiation in mouse embryos via Hippo pathway-dependent and -independent mechanisms. Nat. Commun. 4, 1–11 (2013).
Hirate, Y. et al. Polarity-dependent distribution of angiomotin localizes Hippo signaling in preimplantation embryos. Curr. Biol. 23, 1181–1194 (2013).
Posfai, E. et al. Position- and Hippo signaling-dependent plasticity during lineage segregation in the early mouse embryo. eLife 6, e22906 (2017). Detailed single-cell RNA sequencing analysis of Cdx2 levels in staged blastomeres, revealing their plasticity during the time of lineage specification in the early embryo.
Dupont, S. et al. Role of YAP/TAZ in mechanotransduction. Nature 474, 179–183 (2011).
Aragona, M. et al. A mechanical checkpoint controls multicellular growth through YAP/TAZ regulation by actin-processing factors. Cell 154, 1047–1059 (2013).
Elosegui-Artola, A. et al. Force triggers YAP nuclear entry by regulating transport across nuclear pores. Cell https://doi.org/10.1016/j.cell.2017.10.008 (2017). Proposal of a force-mediated change in nuclear pore morphology as a novel mechanism of YAP nuclear import.
Gundersen, G. G. & Worman, H. J. Nuclear positioning. Cell 152, 1376–1389 (2013).
Baye, L. M. & Link, B. A. Interkinetic nuclear migration and the selection of neurogenic cell divisions during vertebrate retinogenesis. J. Neurosci. 27, 10143–10152 (2007).
Alam, S. G. et al. The nucleus is an intracellular propagator of tensile forces in NIH 3T3 fibroblasts. J. Cell Sci. 128, 1901–1911 (2015).
Chancellor, T. J., Lee, J., Thodeti, C. K. & Lele, T. Actomyosin tension exerted on the nucleus through nesprin-1 connections influences endothelial cell adhesion, migration, and cyclic strain-induced reorientation. Biophys. J. 99, 115–123 (2010).
Goldman, R. D. Nuclear lamins: building blocks of nuclear architecture. Genes Dev. 16, 533–547 (2002).
Crisp, M. et al. Coupling of the nucleus and cytoplasm: role of the LINC complex. J. Cell Biol. 172, 41–53 (2006).
Kirby, T. J. & Lammerding, J. Emerging views of the nucleus as a cellular mechanosensor. Nat. Cell Biol. 20, 373–381 (2018).
Lomakin, A. J. et al. The nucleus acts as a ruler tailoring cell responses to spatial constraints. Science 370, eaba2894 (2020).
Venturini, V. et al. The nucleus measures shape changes for cellular proprioception to control dynamic cell behavior. Science 370, eaba2644 (2020).
Borsos, M. et al. Genome–lamina interactions are established de novo in the early mouse embryo. Nature 569, 729–733 (2019).
Cockburn, K. & Rossant, J. Making the blastocyst: lessons from the mouse. J. Clin. Invest. 120, 995–1003 (2010).
Eckert, J. J. & Fleming, T. P. Tight junction biogenesis during early development. Biochim. Biophys. Acta 1778, 717–728 (2008).
Watson, A. J. & Barcroft, L. C. Regulation of blastocyst formation. Obstet. Gynaecol. Publ. 58, 24 (2001).
White, J. G. & Borisy, G. G. On the mechanisms of cytokinesis in animal cells. J. Theor. Biol. 101, 289–316 (1983).
Fededa, J. P. & Gerlich, D. W. Molecular control of animal cell cytokinesis. Nat. Cell Biol. 14, 440–447 (2012).
Borowiak, M. et al. Photoswitchable inhibitors of microtubule dynamics optically control mitosis and cell death. Cell 162, 403–411 (2015).
Schwayer, C., Sikora, M., Slováková, J., Kardos, R. & Heisenberg, C.-P. Actin rings of power. Dev. Cell 37, 493–506 (2016).
Wang, H. et al. Zonula occludens-1 (ZO-1) is involved in morula to blastocyst transformation in the mouse. Dev. Biol. 318, 112–125 (2008).
Kim, J., Gye, M. C. & Kim, M. K. Role of occludin, a tight junction protein, in blastocoel formation, and in the paracellular permeability and differentiation of trophectoderm in preimplantation mouse embryos. Mol. Cell 17, 248–254 (2004).
Barcroft, L. C., Offenberg, H., Thomsen, P. & Watson, A. J. Aquaporin proteins in murine trophectoderm mediate transepithelial water movements during cavitation. Dev. Biol. 256, 342–354 (2003).
Dumortier, J. G. et al. Hydraulic fracturing and active coarsening position the lumen of the mouse blastocyst. Science 365, 465–468 (2019).
Ryan, A. Q., Chan, C. J., Graner, F. & Hiiragi, T. Lumen expansion facilitates epiblast-primitive endoderm fate specification during mouse blastocyst formation. Dev. Cell 51, 1–14 (2019).
Leonavicius, K. et al. Mechanics of mouse blastocyst hatching revealed by a hydrogel-based microdeformation assay. Proc. Natl Acad. Sci. USA 115, 10375–10380 (2018).
Chan, C. J. et al. Hydraulic control of mammalian embryo size and cell fate. Nature https://doi.org/10.1038/s41586-019-1309-x (2019).
Chan, C. J. & Hiiragi, T. Integration of luminal pressure and signalling in tissue self-organization. Development 147, dev181297 (2020).
Cadart, C., Zlotek-Zlotkiewicz, E., Le Berre, M., Piel, M. & Matthews, H. K. Exploring the function of cell shape and size during mitosis. Dev. Cell 29, 159–169 (2014).
Cutrale, F., Fraser, S. E. & Trinh, L. A. Imaging, visualization, and computation in developmental biology. Annu. Rev. Biomed. Data Sci. 2, 223–251 (2019).
McDole, K. et al. In toto imaging and reconstruction of post-implantation mouse development at the single-cell level. Cell https://doi.org/10.1016/j.cell.2018.09.031 (2018). Construction of an atlas of whole-embryo developmental dynamics at single-cell resolution, incorporating advanced microscopy and image analysis techniques.
Roca-Cusachs, P., Conte, V. & Trepat, X. Quantifying forces in cell biology. Nat. Cell Biol. 19, 742–751 (2017).
Daniels, B. R., Masi, B. C. & Wirtz, D. Probing single-cell micromechanics in vivo: the microrheology of C. elegans developing embryos. Biophys. J. 90, 4712–4719 (2006).
Wirtz, D. Particle-tracking microrheology of living cells: principles and applications. Annu. Rev. Biophys. 38, 301–326 (2009).
Sanghvi-Shah, R., Paranjpe, S., Baek, J., Dobrowolski, R. & Weber, G. A novel photoactivatable tool for intermediate filament disruption indicates a role for keratin filaments in early embryogenesis. Preprint at bioRxiv https://doi.org/10.1101/484246 (2018).
Harterink, M. et al. DeActs: genetically encoded tools for perturbing the actin cytoskeleton in single cells. Nat. Methods 14, 479–482 (2017).
Niakan, K. K., Han, J., Pedersen, R. A., Simon, C. & Pera, R. A. R. Human pre-implantation embryo development. Development 139, 829–841 (2012).
Shahbazi, M. N. Mechanisms of human embryo development: from cell fate to tissue shape and back. Development 147, dev190629 (2020).
Acknowledgements
This work was supported by an A*STAR Graduate Scholarship to H.Y.G.L. and US National Institutes of Health grant R01GM1399700-1 and HD102013-01A1 to N.P.
Author information
Authors and Affiliations
Contributions
The authors contributed equally to all aspects of the article.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information
Nature Reviews Molecular Cell Biology thanks M. Zernicka-Goetz, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Glossary
- Linker of nucleoskeleton and cytoskeleton
-
(LINC). Protein complex spanning the inner and outer nuclear membrane, connecting chromatin from within the nucleus to cytoskeletal filaments and organelles within the cytoplasm.
- Trophectoderm
-
A layer of differentiated cells located on the outer surface of the blastocyst, which eventually form the embryonic part of the placenta. These are the first epithelial-like cells that form during development.
- Inner cell mass
-
(ICM). Pluripotent cells located within the inner part of the blastocyst, which subsequently form the developing fetus and other supporting extraembryonic tissues.
- Primitive endoderm
-
One of two lineages derived from the inner cell mass, along with the epiblast, which generates the extraembryonic yolk sac that supports the growth of the fetus.
- PAR–aPKC complex
-
The PDZ domain-containing scaffold proteins PAR3 and PAR6 together with the serine/threonine kinase atypical protein kinase C (aPKC), which are key regulators of epithelial cell polarity.
- Apical domain
-
In the context of the mouse embryo, a polarized group of proteins organized in the form of a rounded patch at the contact-free exposed apical surface of 8-cell-stage blastomeres.
- Acetylated microtubules
-
A post-translational modification on tubulin that stabilizes microtubule filaments.
- Keratin
-
A class of intermediate filament proteins expressed in epithelial cells best known for its mechanical strength.
- Micropipette aspiration
-
Method of measuring cortical tension in a living cell system. A known suction force is applied through a micropipette to aspirate the cell surface, and the resulting cell deformation within the micropipette is measured.
- Filopodia
-
Actin-based cellular protrusions typically used for sensing of the cell’s external environment.
- Myosin X
-
An unconventional plus-end myosin motor localized within actin-based protrusions, including lamellipodia and filopodia.
- Laser ablation
-
Method of specifically ablating or cutting subcellular structures within a living system using a high-energy laser beam.
- Morphogens
-
Molecules whose specific spatial distribution pattern within a developing tissue or organism regulates fate specification and morphogenesis.
- RHOA
-
Ras homologue family member A, a Rho-family GTPase with key roles in promoting actin polymerization and actomyosin contractility.
- TFAP2C
-
Transcription factor AP-2γ, a DNA-binding protein regulating the transcription of many genes, including the trophectoderm lineage marker Cdx2.
- TEAD4
-
TEA domain transcription factor 4, a transcription factor involved in activating the trophectoderm lineage transcriptional programme, including Cdx2 expression.
- ARP2/3 complex
-
A protein complex containing actin-related protein 2 (ARP2) and ARP3 subunits that functions as a major nucleator of actin filaments, generating branched actin networks.
- Apical constriction
-
Myosin II-dependent coordinated narrowing of the apical part of cells located within an epithelial cell sheet, often resulting in tissue-level changes in epithelium morphology.
- Hertwig’s long-axis rule
-
A prediction of division orientation occurring along the longest cell axis.
- Centrosomes
-
Primary microtubule-organizing centre in many animal cells, made up of two centrioles and a dense mass of pericentriolar material.
- Microtubule-organizing centres
-
(MTOCs). Sites of microtubule polymerization within the cell.
- Astral microtubule arrays
-
Microtubules emanating from the centrosome towards the cell surface that aid in spindle positioning during mitosis.
- Meiosis I
-
The first of two rounds of cell division ultimately giving rise to mature gametes.
- Polar body
-
A small cell produced during the asymmetric meiotic divisions that generate the larger, mature oocyte.
- Formin 2
-
An actin nucleator promoting the polymerization of existing actin filaments.
- Neuroblasts
-
Neural stem cells in the Drosophila melanogaster embryo.
- Ventral furrow
-
The large-scale invagination of cells located on the ventral surface of the Drosophila melanogaster embryo that produces the embryonic mesoderm.
- Adherens junction
-
A class of junctional complexes mediating cell–cell adhesion via cadherin interactions, with links to the actin cytoskeleton via adaptor proteins, including α-catenin and β-catenin.
- Hippo signalling pathway
-
A major signalling pathway involved in the regulation of organ size during development and homeostasis.
- Desmosome
-
Also known as macula adherens, a hyperadhesive intercellular adhesion complex composed of extracellular cadherin family proteins and intracellular scaffold and linker proteins connecting the complex to the intermediate filament cytoskeleton.
- Plakin family
-
A family of proteins, including desmoplakin, plectin and periplakin, which function as crosslinkers between cytoskeletal filaments and connect the cytoskeleton to junctional complexes.
- Mechanosensor
-
A cellular structure/component able to detect and respond to changes in mechanical stimuli, such as by altering its molecular conformation and binding partners.
- Actin ring
-
F-actin filaments organized in the form of a ring on the apical cortex of outer blastomeres in the 16-cell-stage embryo. These rings display low levels of myosin II and expand to cell–cell junctions, where they zipper and establish mature tight and adherens junctions.
- Tight junction
-
An adhesion complex typically located at the apical-most portion of cell–cell junctions that facilitates the establishment of a sealed barrier.
- ZO1
-
Zonula occludens protein 1, a scaffolding protein within tight junction complexes that connects them to the actin cytoskeleton.
- Occludin
-
Transmembrane protein component of tight junctions, functioning together with ZO1 and the claudin family of proteins.
- Aquaporins
-
Channel proteins spanning the cell membrane, through which water can be transported in and out of the cell.
- Light-sheet microscopy
-
An imaging system using a thin sheet of illumination light that is oriented perpendicular to the detector, which greatly reduces the out-of-focus excitation and photobleaching from which traditional confocal methods suffer.
- Super-resolution microscopy
-
An imaging system providing spatial resolution higher than the diffraction limit of traditional microscopes. Common super-resolution techniques include stimulated emission depletion microscopy, photoactivated localization microscopy and stochastic optical reconstruction microscopy.
- Particle-tracking microrheology
-
A method of tracking the movement of inert, fluorescent beads within the cytoplasm of living cells to measure their viscoelastic properties.
- Optogenetic-driven techniques
-
The use of light to precisely control the subcellular localization or functions of light-sensitive proteins.
Rights and permissions
About this article
Cite this article
Lim, H.Y.G., Plachta, N. Cytoskeletal control of early mammalian development. Nat Rev Mol Cell Biol 22, 548–562 (2021). https://doi.org/10.1038/s41580-021-00363-9
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41580-021-00363-9
This article is cited by
-
Low-input lipidomics reveals lipid metabolism remodelling during early mammalian embryo development
Nature Cell Biology (2024)
-
A ubiquitin-based effector-to-inhibitor switch coordinates early brain, craniofacial, and skin development
Nature Communications (2023)
-
The nuclear lamina couples mechanical forces to cell fate in the preimplantation embryo via actin organization
Nature Communications (2023)
-
A monoastral mitotic spindle determines lineage fate and position in the mouse embryo
Nature Cell Biology (2022)
-
From the flexible to the complex: carving the zygotic pie into operational pieces
Journal of Assisted Reproduction and Genetics (2021)