Abstract
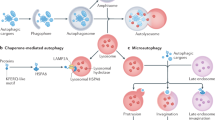
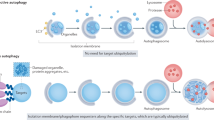
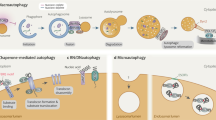
Chaperone-mediated autophagy (CMA) was the first studied process that indicated that degradation of intracellular components by the lysosome can be selective — a concept that is now well accepted for other forms of autophagy. Lysosomes can degrade cellular cytosol in a nonspecific manner but can also discriminate what to target for degradation with the involvement of a degradation tag, a chaperone and a sophisticated mechanism to make the selected proteins cross the lysosomal membrane through a dedicated translocation complex. Recent studies modulating CMA activity in vivo using transgenic mouse models have demonstrated that selectivity confers on CMA the ability to participate in the regulation of multiple cellular functions. Timely degradation of specific cellular proteins by CMA modulates, for example, glucose and lipid metabolism, DNA repair, cellular reprograming and the cellular response to stress. These findings expand the physiological relevance of CMA beyond its originally identified role in protein quality control and reveal that CMA failure with age may aggravate diseases, such as ageing-associated neurodegeneration and cancer.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Green, D. R. & Levine, B. To be or not to be? How selective autophagy and cell death govern cell fate. Cell 157, 65–75 (2014).
Galluzzi, L. et al. Molecular definitions of autophagy and related processes. EMBO J. 36, 1811–1836 (2017).
Stolz, A., Ernst, A. & Dikic, I. Cargo recognition and trafficking in selective autophagy. Nat. Cell Biol. 16, 495–501 (2014).
De Duve, C. & Wattiaux, R. Functions of lysosomes. Annu. Rev. Physiol. 28, 435–492 (1966).
Marzella, L., Ahlberg, J. & Glaumann, H. Autophagy, heterophagy, microautophagy and crinophagy as the means for intracellular degradation. Virchows Archiv B Cell Pathol. Incl. Mol. Pathol. 36, 219–234 (1981).
Roberts, P. et al. Piecemeal microautophagy of nucleus in Saccharomyces cerevisiae. Mol. Biol. Cell 14, 129–141 (2003).
Sakai, Y., Koller, A., Rangell, L., Keller, G. & Subramani, S. Peroxisome degradation by microautophagy in Pichia pastoris. Identification of specific steps and morphological intermediates. J. Cell Biol. 141, 625–636 (1998).
Sahu, R. et al. Microautophagy of cytosolic proteins by late endosomes. Dev. Cell 20, 131–139 (2011). First report of endosomal microautophagy.
Dice, J. F. Peptide sequences that target cytosolic proteins for lysosomal proteolysis. Trends Biochem. Sci. 15, 305–309 (1990). First characterization of the properties of the KFERQ-like signal.
Terlecky, S. R., Chiang, H.-L., Olson, T. S. & Dice, J. F. Protein and peptide binding and stimulation of in vitro lysosomal proteolysis by the 73-kDa heat shock cognate protein. J. Biol. Chem. 267, 9202–9209 (1992).
Cuervo, A. M., Terlecky, S. R., Dice, J. F. & Knecht, E. Selective binding and uptake of ribonuclease A and glyceraldehyde-3-phosphate dehydrogenase by isolated rat liver lysosomes. J. Biol. Chem. 269, 26374–26380 (1994).
Eskelinen, E. L. et al. Unifying nomenclature for the isoforms of the lysosomal membrane protein LAMP-2. Traffic 6, 1058–1061 (2005).
Massey, A. C., Kaushik, S., Sovak, G., Kiffin, R. & Cuervo, A. M. Consequences of the selective blockage of chaperone-mediated autophagy. Proc. Natl Acad. Sci. USA 103, 5805–5810 (2006).
Bandyopadhyay, U., Sridhar, S., Kaushik, S., Kiffin, R. & Cuervo, A. M. Identification of regulators of chaperone-mediated autophagy. Mol. Cell 39, 535–547 (2010).
Bandyopadhyay, U., Kaushik, S., Varticovski, L. & Cuervo, A. M. The chaperone-mediated autophagy receptor organizes in dynamic protein complexes at the lysosomal membrane. Mol. Cell. Biol. 28, 5747–5763 (2008). Identification of the CMA translocation complex at the lysosomal membrane.
Arias, E. et al. Lysosomal mTORC2/PHLPP1/Akt regulate chaperone-mediated autophagy. Mol. Cell 59, 270–284 (2015).
Cuervo, A. M., Stefanis, L., Fredenburg, R., Lansbury, P. T. & Sulzer, D. Impaired degradation of mutant alpha-synuclein by chaperone-mediated autophagy. Science 305, 1292–1295 (2004). First connection of CMA malfunction with a human disorder (neurodegeneration).
Kon, M. et al. Chaperone-mediated autophagy is required for tumor growth. Sci. Transl Med. 3, 109ra117 (2011).
Schneider, J. L., Suh, Y. & Cuervo, A. M. Deficient chaperone-mediated autophagy in liver leads to metabolic dysregulation. Cell Metab. 20, 417–432 (2014). First mouse model with tissue-specific CMA blockage in vivo.
Schneider, J. L. et al. Loss of hepatic chaperone-mediated autophagy accelerates proteostasis failure in aging. Aging Cell 14, 249–264 (2015).
Valdor, R. et al. Chaperone-mediated autophagy regulates T cell responses through targeted degradation of negative regulators of T cell activation. Nat. Immunol. 15, 1046–1054 (2014).
Dice, J. F. Altered degradation of proteins microinjected into senescent human fibroblasts. J. Biol. Chem. 257, 14624–14627 (1982).
Kaushik, S. & Cuervo, A. M. AMPK-dependent phosphorylation of lipid droplet protein PLIN2 triggers its degradation by CMA. Autophagy 12, 432–438 (2016).
Park, C., Suh, Y. & Cuervo, A. M. Regulated degradation of Chk1 by chaperone-mediated autophagy in response to DNA damage. Nat. Commun. 6, 6823 (2015).
Quintavalle, C. et al. Phosphorylation-regulated degradation of the tumor-suppressor form of PED by chaperone-mediated autophagy in lung cancer cells. J. Cell. Physiol. 229, 1359–1368 (2014).
Zhou, J. et al. Chaperone-mediated autophagy regulates proliferation by targeting RND3 in gastric cancer. Autophagy 12, 515–528 (2016).
Lv, L. et al. Acetylation targets the M2 isoform of pyruvate kinase for degradation through chaperone-mediated autophagy and promotes tumor growth. Mol. Cell 42, 719–730 (2011).
Bonhoure, A. et al. Acetylation of translationally controlled tumor protein promotes its degradation through chaperone-mediated autophagy. Eur. J. Cell Biol. 96, 83–98 (2017).
Ferreira, J. V., Soares, A. R., Ramalho, J. S., Pereira, P. & Girao, H. K63 linked ubiquitin chain formation is a signal for HIF1A degradation by chaperone-mediated autophagy. Sci. Rep. 5, 10210 (2015).
Li, L. et al. Deacetylation of tumor-suppressor MST1 in Hippo pathway induces its degradation through HBXIP-elevated HDAC6 in promotion of breast cancer growth. Oncogene 35, 4048–4057 (2016).
Chiang, H., Terlecky, S., Plant, C. & Dice, J. F. A role for a 70-kilodalton heat shock protein in lysosomal degradation of intracellular proteins. Science 246, 382–385 (1989). Identification of HSC70 as the chaperone for CMA.
Ferreira, J. V. et al. STUB1/CHIP is required for HIF1A degradation by chaperone-mediated autophagy. Autophagy 9, 1349–1366 (2013).
Agarraberes, F. & Dice, J. F. A molecular chaperone complex at the lysosomal membrane is required for protein translocation. J. Cell Sci. 114, 2491–2499 (2001).
Arndt, V. et al. Chaperone-assisted selective autophagy is essential for muscle maintenance. Curr. Biol. 20, 143–148 (2010).
Cuervo, A. M., Dice, J. F. & Knecht, E. A population of rat liver lysosomes responsible for the selective uptake and degradation of cytosolic proteins. J. Biol. Chem. 272, 5606–5615 (1997).
Salvador, N., Aguado, C., Horst, M. & Knecht, E. Import of a cytosolic protein into lysosomes by chaperone-mediated autophagy depends on its folding state. J. Biol. Chem. 275, 27447–27456 (2000).
Agarraberes, F., Terlecky, S. & Dice, J. An intralysosomal hsp70 is required for a selective pathway of lysosomal protein degradation. J. Cell Biol. 137, 825–834 (1997).
Kaushik, S., Massey, A., Mizushima, N. & Cuervo, A. M. Constitutive activation of chaperone-mediated autophagy in cells with impaired macroautophagy. Mol. Biol. Cell 19, 2179–2192 (2008).
Cuervo, A. M. & Dice, J. F. A receptor for the selective uptake and degradation of proteins by lysosomes. Science 273, 501–503 (1996). Identification of the lysosomal membrane receptor for CMA.
Cuervo, A. M., Knecht, E., Terlecky, S. R. & Dice, J. F. Activation of a selective pathway of lysosomal proteolysis in rat liver by prolonged starvation. Am. J. Physiol. 269, C1200–C1208 (1995).
Cuervo, A. M. & Dice, J. F. Unique properties of lamp2a compared to other lamp2 isoforms. J. Cell Sci. 113, 4441–4450 (2000).
Li, J. & Pfeffer, S. R. Lysosomal membrane glycoproteins bind cholesterol and contribute to lysosomal cholesterol export. eLife 5, e21635 (2016).
Cuervo, A. M. & Dice, J. F. Regulation of lamp2a levels in the lysosomal membrane. Traffic 1, 570–583 (2000).
Rout, A. K., Strub, M. P., Piszczek, G. & Tjandra, N. Structure of transmembrane domain of lysosome-associated membrane protein type 2a (LAMP-2A) reveals key features for substrate specificity in chaperone-mediated autophagy. J. Biol. Chem. 289, 35111–35123 (2014).
Kiffin, R., Christian, C., Knecht, E. & Cuervo, A. M. Activation of chaperone-mediated autophagy during oxidative stress. Mol. Biol. Cell 15, 4829–4840 (2004).
Hubbi, M. E. et al. Cyclin-dependent kinases regulate lysosomal degradation of hypoxia-inducible factor 1alpha to promote cell-cycle progression. Proc. Natl Acad. Sci. USA 111, E3325–E3334 (2014).
Anguiano, J. et al. Chemical modulation of chaperone-mediated autophagy by retinoic acid derivatives. Nat. Chem. Biol. 9, 374–382 (2013).
Sardiello, M. et al. A gene network regulating lysosomal biogenesis and function. Science 325, 473–477 (2009).
Kaushik, S., Massey, A. C. & Cuervo, A. M. Lysosome membrane lipid microdomains: novel regulators of chaperone-mediated autophagy. EMBO J. 25, 3921–3933 (2006). Identification of the mechanisms that determine LAMP2A stability in lysosomes.
Wilke, S., Krausze, J. & Bussow, K. Crystal structure of the conserved domain of the DC lysosomal associated membrane protein: implications for the lysosomal glycocalyx. BMC Biol. 10, 62 (2012).
Cuervo, A. M., Mann, L., Bonten, E. J., d’Azzo, A. & Dice, J. F. Cathepsin A regulates chaperone-mediated autophagy through cleavage of the lysosomal receptor. EMBO J. 22, 12–19 (2003).
Rodriguez-Navarro, J. A. et al. Inhibitory effect of dietary lipids on chaperone-mediated autophagy. Proc. Natl Acad. Sci. USA 109, E705–E714 (2012).
Kiffin, R. et al. Altered dynamics of the lysosomal receptor for chaperone-mediated autophagy with age. J. Cell Sci. 120, 782–791 (2007).
Napolitano, G. et al. Impairment of chaperone-mediated autophagy leads to selective lysosomal degradation defects in the lysosomal storage disease cystinosis. EMBO Mol. Med. 7, 158–174 (2015).
Zhang, J. et al. Cystinosin, the small GTPase Rab11, and the Rab7 effector RILP regulate intracellular trafficking of the chaperone-mediated autophagy receptor LAMP2A. J. Biol. Chem. 292, 10328–10346 (2017).
Tang, F. L. et al. VPS35 in dopamine neurons is required for endosome-to-Golgi retrieval of Lamp2a, a receptor of chaperone-mediated autophagy that is critical for alpha-synuclein degradation and prevention of pathogenesis of Parkinson’s disease. J. Neurosci. 35, 10613–10628 (2015).
Koga, H., Martinez-Vicente, M., Macian, F., Verkhusha, V. V. & Cuervo, A. M. A photoconvertible fluorescent reporter to track chaperone-mediated autophagy. Nat. Commun. 2, 386 (2011).
Dohi, E. et al. Hypoxic stress activates chaperone-mediated autophagy and modulates neuronal cell survival. Neurochem. Int. 60, 431–442 (2012).
Finn, P. F. & Dice, J. F. Ketone bodies stimulate chaperone-mediated autophagy. J. Biol. Chem. 280, 25864–25870 (2005).
Cuervo, A. M., Hildebrand, H., Bomhard, E. M. & Dice, J. F. Direct lysosomal uptake of alpha 2-microglobulin contributes to chemically induced nephropathy. Kidney Int. 55, 529–545 (1999).
Martinez-Vicente, M. et al. Dopamine-modified alpha-synuclein blocks chaperone-mediated autophagy. J. Clin. Invest. 118, 777–788 (2008).
Zhang, C. & Cuervo, A. M. Restoration of chaperone-mediated autophagy in aging liver improves cellular maintenance and hepatic function. Nat. Med. 14, 959–965 (2008).
Mizushima, N., Yamamoto, A., Matsui, M., Yoshimori, T. & Ohsumi, Y. In vivo analysis of autophagy in response to nutrient starvation using transgenic mice expressing a fluorescent autophagosome marker. Mol. Biol. Cell 15, 1101–1111 (2004).
Singh, R. et al. Autophagy regulates lipid metabolism. Nature 458, 1131–1135 (2009).
Singh, V. et al. Salmonella co-opts host cell chaperone-mediated autophagy for intracellular growth. J. Biol. Chem. 292, 1847–1864 (2017).
Wang, B. et al. Essential control of mitochondrial morphology and function by chaperone-mediated autophagy through degradation of PARK7. Autophagy 12, 1215–1228 (2016).
Kaushik, S. & Cuervo, A. M. Degradation of lipid droplet-associated proteins by chaperone-mediated autophagy facilitates lipolysis. Nat. Cell Biol. 17, 759–770 (2015). Identification of the regulatory role of CMA in lipid metabolism.
Lu, W. et al. Dual proteolytic pathways govern glycolysis and immune competence. Cell 159, 1578–1590 (2014).
Cuervo, A. M., Hu, W., Lim, B. & Dice, J. F. IkappaB is a substrate for a selective pathway of lysosomal proteolysis. Mol. Biol. Cell 9, 1995–2010 (1998).
Franch, H. A., Sooparb, S., Du, J. & Brown, N. S. A mechanism regulating proteolysis of specific proteins during renal tubular cell growth. J. Biol. Chem. 276, 19126–19131 (2001).
Yang, Q. et al. Regulation of neuronal survival factor MEF2D by chaperone-mediated autophagy. Science 323, 124–127 (2009).
Zhang, L. et al. Disruption of chaperone-mediated autophagy-dependent degradation of MEF2A by oxidative stress-induced lysosome destabilization. Autophagy 10, 1015–1035 (2014).
Hu, M. M. et al. Sumoylation promotes the stability of the DNA sensor cGAS and the adaptor STING to regulate the kinetics of response to DNA virus. Immunity 45, 555–569 (2016).
Cuervo, A. M. & Dice, J. F. Age-related decline in chaperone-mediated autophagy. J. Biol. Chem. 275, 31505–31513 (2000).
Rodriguez-Muela, N. et al. Balance between autophagic pathways preserves retinal homeostasis. Aging Cell 12, 478–488 (2013).
Cai, Z. et al. Chaperone-mediated autophagy: roles in neuroprotection. Neurosci. Bull. 31, 452–458 (2015).
Cuervo, A. M. & Wong, E. Chaperone-mediated autophagy: roles in disease and aging. Cell Res. 24, 92–104 (2014).
Mak, S. K., McCormack, A. L., Manning-Bog, A. B., Cuervo, A. M. & Di Monte, D. A. Lysosomal degradation of alpha-synuclein in vivo. J. Biol. Chem. 285, 13621–13629 (2010).
Malkus, K. A. & Ischiropoulos, H. Regional deficiencies in chaperone-mediated autophagy underlie alpha-synuclein aggregation and neurodegeneration. Neurobiol. Dis. 46, 732–744 (2012).
Vogiatzi, T., Xilouri, M., Vekrellis, K. & Stefanis, L. Wild type alpha-synuclein is degraded by chaperone-mediated autophagy and macroautophagy in neuronal cells. J. Biol. Chem. 283, 23542–23556 (2008).
Orenstein, S. J. et al. Interplay of LRRK2 with chaperone-mediated autophagy. Nat. Neurosci. 16, 394–406 (2013).
Kabuta, T., Furuta, A., Aoki, S., Furuta, K. & Wada, K. Aberrant interaction between Parkinson disease-associated mutant UCH-L1 and the lysosomal receptor for chaperone-mediated autophagy. J. Biol. Chem. 283, 23731–23738 (2008).
Andersson, F. I. et al. The effect of Parkinson’s-disease-associated mutations on the deubiquitinating enzyme UCH-L1. J. Mol. Biol. 407, 261–272 (2011).
Wang, Y. et al. Tau fragmentation, aggregation and clearance: the dual role of lysosomal processing. Hum. Mol. Genet. 18, 4153–4170 (2009).
Huang, C. C. et al. Metabolism and mis-metabolism of the neuropathological signature protein TDP-43. J. Cell Sci. 127, 3024–3038 (2014).
Bauer, P. O. et al. Harnessing chaperone-mediated autophagy for the selective degradation of mutant huntingtin protein. Nat. Biotechnol. 28, 256–263 (2010).
Koga, H. et al. Constitutive upregulation of chaperone-mediated autophagy in Huntington’s disease. J. Neurosci. 31, 18492–18505 (2011).
Qi, L. et al. The role of chaperone-mediated autophagy in huntingtin degradation. PLoS ONE 7, e46834 (2012).
Xilouri, M., Vogiatzi, T., Vekrellis, K., Park, D. & Stefanis, L. Abberant alpha-synuclein confers toxicity to neurons in part through inhibition of chaperone-mediated autophagy. PLoS ONE 4, e5515 (2009).
Xilouri, M. et al. Impairment of chaperone-mediated autophagy induces dopaminergic neurodegeneration in rats. Autophagy 12, 2230–2247 (2016).
Xilouri, M. et al. Boosting chaperone-mediated autophagy in vivo mitigates alpha-synuclein-induced neurodegeneration. Brain 136, 2130–2146 (2013). First evidence of antidegenerative effect of genetic enhancement of CMA.
Gan, L., Vargas, M. R., Johnson, D. A. & Johnson, J. A. Astrocyte-specific overexpression of Nrf2 delays motor pathology and synuclein aggregation throughout the CNS in the alpha-synuclein mutant (A53T) mouse model. J. Neurosci. 32, 17775–17787 (2012).
Magalhaes, J. et al. Autophagic lysosome reformation dysfunction in glucocerebrosidase deficient cells: relevance to Parkinson disease. Hum. Mol. Genet. 25, 3432–3445 (2016).
Alvarez-Erviti, L. et al. Chaperone-mediated autophagy markers in Parkinson disease brains. Arch. Neurol. 67, 1464–1472 (2010).
Murphy, K. E. et al. Reduced glucocerebrosidase is associated with increased alpha-synuclein in sporadic Parkinson’s disease. Brain 137, 834–848 (2014).
Pang, S., Chen, D., Zhang, A., Qin, X. & Yan, B. Genetic analysis of the LAMP-2 gene promoter in patients with sporadic Parkinson’s disease. Neurosci. Lett. 526, 63–67 (2012).
Alvarez-Erviti, L. et al. Influence of microRNA deregulation on chaperone-mediated autophagy and alpha-synuclein pathology in Parkinson’s disease. Cell Death Dis. 4, e545 (2013).
Park, J. S., Kim, D. H. & Yoon, S. Y. Regulation of amyloid precursor protein processing by its KFERQ motif. BMB Rep. 49, 337–342 (2016).
Liu, H., Wang, P., Song, W. & Sun, X. Degradation of regulator of calcineurin 1 (RCAN1) is mediated by both chaperone-mediated autophagy and ubiquitin proteasome pathways. FASEB J. 23, 3383–3392 (2009).
Rothenberg, C. et al. Ubiquilin functions in autophagy and is degraded by chaperone-mediated autophagy. Hum. Mol. Genet. 19, 3219–3232 (2010).
Saha, T. LAMP2A overexpression in breast tumors promotes cancer cell survival via chaperone-mediated autophagy. Autophagy 8, 1643–1656 (2012).
Zhang, Y. et al. Acetylation targets HSD17B4 for degradation via the CMA pathway in response to estrone. Autophagy 13, 538–553 (2017).
Du, C. et al. 5-Fluorouracil targets histone acetyltransferases p300/CBP in the treatment of colorectal cancer. Cancer Lett. 400, 183–193 (2017).
Ding, Z. B. et al. Lamp2a is required for tumor growth and promotes tumor recurrence of hepatocellular carcinoma. Int. J. Oncol. 49, 2367–2376 (2016).
Wu, J. H. et al. CMA down-regulates p53 expression through degradation of HMGB1 protein to inhibit irradiation-triggered apoptosis in hepatocellular carcinoma. World J. Gastroenterol. 23, 2308–2317 (2017).
Chava, S. et al. Chaperone-mediated autophagy compensates for impaired macroautophagy in the cirrhotic liver to promote hepatocellular carcinoma. Oncotarget 8, 40019–40036 (2017).
Guo, B. et al. M2 tumor-associated macrophages produce interleukin-17 to suppress oxaliplatin-induced apoptosis in hepatocellular carcinoma. Oncotarget 8, 44465–44476 (2017).
Ali, A. B., Nin, D. S., Tam, J. & Khan, M. Role of chaperone mediated autophagy (CMA) in the degradation of misfolded N-CoR protein in non-small cell lung cancer (NSCLC) cells. PLoS ONE 6, e25268 (2011).
Vakifahmetoglu-Norberg, H. et al. Chaperone-mediated autophagy degrades mutant p53. Genes Dev. 27, 1718–1730 (2013).
Xie, W. et al. Chaperone-mediated autophagy prevents apoptosis by degrading BBC3/PUMA. Autophagy 11, 1623–1635 (2015).
Suzuki, J., Nakajima, W., Suzuki, H., Asano, Y. & Tanaka, N. Chaperone-mediated autophagy promotes lung cancer cell survival through selective stabilization of the pro-survival protein, MCL1. Biochem. Biophys. Res. Commun. 482, 1334–1340 (2017).
Xia, H. G. et al. Degradation of HK2 by chaperone-mediated autophagy promotes metabolic catastrophe and cell death. J. Cell Biol. 210, 705–716 (2015).
Galan-Acosta, L., Xia, H., Yuan, J. & Vakifahmetoglu-Norberg, H. Activation of chaperone-mediated autophagy as a potential anticancer therapy. Autophagy 11, 2370–2371 (2015).
Gomes, L. R., Menck, C. F. M. & Cuervo, A. M. Chaperone-mediated autophagy prevents cellular transformation by regulating MYC proteasomal degradation. Autophagy 13, 928–940 (2017).
Lu, T. L. et al. Hispolon promotes MDM2 downregulation through chaperone-mediated autophagy. Biochem. Biophys. Res. Commun. 398, 26–31 (2010).
Garg, A. D., Dudek, A. M. & Agostinis, P. Calreticulin surface exposure is abrogated in cells lacking, chaperone-mediated autophagy-essential gene, LAMP2A. Cell Death Dis. 4, e826 (2013).
Kaushik, S. & Cuervo, A. M. Methods to monitor chaperone-mediated autophagy. Methods Enzymol. 452, 297–324 (2009).
Li, P. et al. Degradation of AF1Q by chaperone-mediated autophagy. Exp. Cell Res. 327, 48–56 (2014).
Gao, L. et al. Oxidation of survival factor MEF2D in neuronal death and Parkinson’s disease. Antioxid. Redox Signal 20, 2936–2948 (2014).
Li, G. et al. Targeted suppression of chaperone-mediated autophagy by miR-320a promotes alpha-synuclein aggregation. Int. J. Mol. Sci. 15, 15845–15857 (2014).
Wu, G. et al. Altered expression of autophagic genes in the peripheral leukocytes of patients with sporadic Parkinson’s disease. Brain Res. 1394, 105–111 (2011).
Sala, G. et al. Reduced expression of the chaperone-mediated autophagy carrier hsc70 protein in lymphomonocytes of patients with Parkinson’s disease. Brain Res. 1546, 46–52 (2014).
Wang, H. et al. Overexpression of PLK3 mediates the degradation of abnormal prion proteins dependent on chaperone-mediated autophagy. Mol. Neurobiol. 54, 4401–4413 (2017).
Pedrozo, Z. et al. Cardiomyocyte ryanodine receptor degradation by chaperone-mediated autophagy. Cardiovasc. Res. 98, 277–285 (2013).
Fidzianska, A., Walczak, E. & Walski, M. Abnormal chaperone-mediated autophagy (CMA) in cardiomyocytes of a boy with Danon disease. Folia Neuropathol. 45, 133–139 (2007).
Metrailler, S., Schorderet, D. F. & Cottet, S. Early apoptosis of rod photoreceptors in Rpe65(−/−) mice is associated with the upregulated expression of lysosomal-mediated autophagic genes. Exp. Eye Res. 96, 70–81 (2012).
Li, Y., Lu, L., Luo, N., Wang, Y. Q. & Gao, H. M. Inhibition of PI3K/AKt/mTOR signaling pathway protects against d-galactosamine/lipopolysaccharide-induced acute liver failure by chaperone-mediated autophagy in rats. Biomed. Pharmacother. 92, 544–553 (2017).
Das, S. et al. Purinergic receptor X7 is a key modulator of metabolic oxidative stress-mediated autophagy and inflammation in experimental nonalcoholic steatohepatitis. Am. J. Physiol. Gastrointest. Liver Physiol. 305, G950–G963 (2013).
Sharma, S., Mells, J. E., Fu, P. P., Saxena, N. K. & Anania, F. A. GLP-1 analogs reduce hepatocyte steatosis and improve survival by enhancing the unfolded protein response and promoting macroautophagy. PLoS ONE 6, e25269 (2011).
Cai, Y. et al. The detrimental role played by lipocalin-2 in alcoholic fatty liver in mice. Am. J. Pathol. 186, 2417–2428 (2016).
Lee, C. H., Lee, K. H., Jang, A. H. & Yoo, C. G. The impact of autophagy on the cigarette smoke extract-induced apoptosis of bronchial epithelial cells. Tuberc. Respir. Dis. 80, 83–89 (2017).
Sooparb, S., Price, S. R., Shaoguang, J. & Franch, H. A. Suppression of chaperone-mediated autophagy in the renal cortex during acute diabetes mellitus. Kidney Int. 65, 2135–2144 (2004).
Cacciottolo, M., Nogalska, A., D’Agostino, C., Engel, W. K. & Askanas, V. Chaperone-mediated autophagy components are upregulated in sporadic inclusion-body myositis muscle fibres. Neuropathol. Appl. Neurobiol. 39, 750–761 (2013).
Su, M. et al. HDAC6 regulates the chaperone-mediated autophagy to prevent oxidative damage in injured neurons after experimental spinal cord injury. Oxid. Med. Cell Longev. 2016, 7263736 (2016).
Park, Y. et al. Chaperone-mediated autophagy after traumatic brain injury. J. Neurotrauma 32, 1449–1457 (2015).
Macri, C. et al. Modulation of deregulated chaperone-mediated autophagy by a phosphopeptide. Autophagy 11, 472–486 (2015).
Venugopal, B. et al. Chaperone-mediated autophagy is defective in mucolipidosis type IV. J. Cell. Physiol. 219, 344–353 (2009).
Author information
Authors and Affiliations
Contributions
S.K. and A.M.C. researched data for the article, contributed to discussion of the content, wrote the article and reviewed and/or edited the manuscript before submission.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Glossary
- Chaperone
-
A protein that assists during folding of other proteins until they reach their functional conformations. Chaperone substrates include both de novo synthesized native proteins and previously folded proteins that undergo partial unfolding.
- Proteostasis networks
-
Intracellular components, mainly chaperones and proteolytic systems, that control each of the processes that occur from protein synthesis to degradation to prevent protein aggregation and ensure maintenance of a stable proteome.
- HIF1α
-
The hypoxia-inducible factor 1α is a transcription factor that modulates the cellular response to hypoxia. HIF1α orchestrates the transcription of a large set of genes involved in cell proliferation, cell survival, and glucose and iron metabolism.
- E3 ubiquitin-protein ligase
-
Intracellular enzyme that participates in the covalent attachment of ubiquitin moieties to cargo proteins. This tagging is commonly used for the targeting of the protein to the proteasome system for degradation.
- MST1
-
The mammalian STE20-like protein kinase 1 is a protein component of the Hippo signalling pathway that modulates cell proliferation and differentiation.
- Cochaperones
-
Proteins that assist chaperones in their functions by modulating their ATP hydrolytic capability.
- HSP90
-
Heat shock protein 90 is a chaperone that stabilizes proteins in transient conformations and facilitates their final folding. HSP90 often acts in conjunction with other intracellular chaperones such as HSC70.
- Lysosome-associated membrane protein
-
(LAMP). Single-span membrane proteins at the lysosomal membrane with a short C-terminus (12 amino acids) exposed to the cytosol while the rest of the protein is in the lysosomal lumen.
- Isoelectric point
-
The pH at which the net charge on a protein is zero.
- Nuclear retinoic acid receptor-α
-
Nuclear receptor activated by retinoic acid that activates or represses gene expression. Its wide range of targets include genes involved in development, apoptosis, differentiation, autophagy and circadian regulation.
- Lipid microdomains
-
Cholesterol and/or glycosphingolipid-rich regions in membranes that present higher order and density than the surrounding membrane. They can be transient and highly dynamic in terms of resident proteins and lipids and in their size.
- Cathepsin
-
Protease located in the lysosomal lumen that is maximally active at acidic pH.
- Cystinosis
-
Lysosomal storage disorder characterized by the abnormal accumulation of cysteine inside lysosomes owing to a defect in its normal export from this organelle.
- Retromer
-
Multiprotein complex made up of membrane-associated sorting nexin and vacuolar-protein-sorting proteins that recycle transmembrane proteins from endosomes to the Golgi complex and the plasma membrane.
- AKT1
-
Intracellular kinase that participates in signalling pathways that regulate a wide array of intracellular processes, including proliferation, cell survival, metabolism, growth and angiogenesis.
- TOR complex 2
-
(TORC2). One of the two functional multiprotein complexes containing the nutrient sensing kinase TOR. TORC2 plays regulatory roles in actin cytoskeleton dynamics, proliferation, growth and metabolism.
- Perilipins
-
Proteins that cover the surface of lipid droplets and shield the hydrophobic lipid core from the aqueous cytosol. Perilipins also regulate the rate of consumption of the lipids (lipolysis) in the lipid droplet.
- Warburg effect
-
Unique metabolic characteristic of many cancer cells whereby they sustain very high levels of glycolysis followed by lactic acid fermentation rather than the common usage of oxidation of pyruvate in mitochondria that follows glycolysis in most cells.
- Immunogenic cell death
-
Functionally peculiar variant of regulated cell death that — in immunocompetent syngeneic hosts — is sufficient to activate an adaptive immune response against dead cell-associated antigens.
Rights and permissions
About this article
Cite this article
Kaushik, S., Cuervo, A.M. The coming of age of chaperone-mediated autophagy. Nat Rev Mol Cell Biol 19, 365–381 (2018). https://doi.org/10.1038/s41580-018-0001-6
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41580-018-0001-6
This article is cited by
-
Prominent tauopathy and intracellular β-amyloid accumulation triggered by genetic deletion of cathepsin D: implications for Alzheimer disease pathogenesis
Alzheimer's Research & Therapy (2024)
-
Ferroptosis in cancer: From molecular mechanisms to therapeutic strategies
Signal Transduction and Targeted Therapy (2024)
-
Role of protein degradation systems in colorectal cancer
Cell Death Discovery (2024)
-
LUBAC promotes angiogenesis and lung tumorigenesis by ubiquitinating and antagonizing autophagic degradation of HIF1α
Oncogenesis (2024)
-
The Role of Pyroptosis and Autophagy in the Nervous System
Molecular Neurobiology (2024)