Abstract
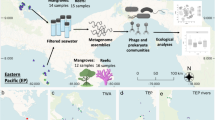
We commonly acknowledge that bacterial viruses (phages) shape the composition and evolution of bacterial communities in nature and therefore have important roles in ecosystem functioning. This view stems from studies in the 1990s to the first decade of the twenty-first century that revealed high viral abundance, high viral diversity and virus-induced microbial death in aquatic ecosystems as well as an association between collapses in bacterial density and peaks in phage abundance. The recent surge in metagenomic analyses has provided deeper insight into the abundance, genomic diversity and spatio-temporal dynamics of phages in a wide variety of ecosystems, ranging from deep oceans to soil and the mammalian digestive tract. However, the causes and consequences of variations in phage community compositions remain poorly understood. In this Review, we explore current knowledge of the composition and evolution of phage communities, as well as their roles in controlling the population and evolutionary dynamics of bacterial communities. We discuss the need for greater ecological realism in laboratory studies to capture the complexity of microbial communities that thrive in natural environments.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Hendrix, R. W., Smith, M. C. M., Burns, R. N., Ford, M. E. & Hatfull, G. F. Evolutionary relationships among diverse bacteriophages and prophages: all the world’s a phage. Proc. Natl Acad. Sci. USA 96, 2192–2197 (1999).
Simmonds, P. et al. Virus taxonomy in the age of metagenomics. Nat. Rev. Microbiol. 15, 161–168 (2017).
Suttle, C. A. Viruses: unlocking the greatest biodiversity on Earth. Genome 56, 542–544 (2013).
Whitman, W. B., Coleman, D. C. & Wiebe, W. J. Prokaryotes: the unseen majority. Proc. Natl Acad. Sci. USA 95, 6578–6583 (1998).
Wigington, C. H. et al. Re-examination of the relationship between marine virus and microbial cell abundances. Nat. Microbiol. 1, 15024 (2016).
Mushegian, A. R. Are there 10 31 virus particles on Earth, or more, or fewer? J. Bacteriol. https://doi.org/10.1128/JB.00052-20 (2020).
Bar-On, Y. M., Phillips, R. & Milo, R. The biomass distribution on Earth. Proc. Natl Acad. Sci. USA 115, 6506–6511 (2018).
Graham, E. B. et al. Untapped viral diversity in global soil metagenomes. Preprint at bioRxiv https://doi.org/10.1101/583997 (2019).
Dion, M. B., Oechslin, F. & Moineau, S. Phage diversity, genomics and phylogeny. Nat. Rev. Microbiol. 18, 125–138 (2020).
Hoyles, L. et al. Characterization of virus-like particles associated with the human faecal and caecal microbiota. Res. Microbiol. 165, 803–812 (2014).
Santiago-Rodriguez, T. M. et al. Natural mummification of the human gut preserves bacteriophage DNA. FEMS Microbiol. Lett. 363, fnv219 (2016).
Paez-Espino, D. et al. Uncovering Earth’s virome. Nature 536, 425–430 (2016).
Tara Oceans Coordinators et al. Tara Oceans: towards global ocean ecosystems biology. Nat. Rev. Microbiol. 18, 428–445 (2020).
Hatfull, G. F. Bacteriophage genomics. Curr. Opin. Microbiol. 11, 447–453 (2008).
Suttle, C. A. Marine viruses — major players in the global ecosystem. Nat. Rev. Microbiol. 5, 801–812 (2007).
Suttle, C. A. The significance of viruses to mortality in aquatic microbial communities. Microb. Ecol. 28, 237–243 (1994).
Breitbart, M., Bonnain, C., Malki, K. & Sawaya, N. A. Phage puppet masters of the marine microbial realm. Nat. Microbiol. 3, 754–766 (2018).
Roux, S. et al. Cryptic inoviruses revealed as pervasive in bacteria and archaea across Earth’s biomes. Nat. Microbiol. 4, 1895–1906 (2019).
Roux, S., Krupovic, M., Poulet, A., Debroas, D. & Enault, F. Evolution and diversity of the Microviridae viral family through a collection of 81 new complete genomes assembled from virome reads. PLoS ONE 7, e40418 (2012).
Krupovic, M. & Forterre, P. Microviridae goes temperate: microvirus-related proviruses reside in the genomes of Bacteroidetes. PLoS ONE 6, e19893 (2011).
Quaiser, A. et al. Diversity and comparative genomics of Microviridae in Sphagnum- dominated peatlands. Front. Microbiol. 6, 375 (2015).
Gregory, A. C. et al. Marine DNA viral macro- and microdiversity from pole to pole. Cell 177, 1109–1123 (2019).
Krishnamurthy, S. R. & Wang, D. Origins and challenges of viral dark matter. Virus Res. 239, 136–142 (2017).
Callanan, J. et al. Biases in viral metagenomics-based detection, cataloguing and quantification of bacteriophage genomes in human faeces, a review. Microorganisms 9, 524 (2021).
Krishnamurthy, S. R., Janowski, A. B., Zhao, G., Barouch, D. & Wang, D. Hyperexpansion of RNA bacteriophage diversity. PLoS Biol. 14, e1002409 (2016).
Callanan, J. et al. Expansion of known ssRNA phage genomes: from tens to over a thousand. Sci. Adv. 6, eaay5981 (2020).
Weitz, J. S., Li, G., Gulbudak, H., Cortez, M. H. & Whitaker, R. J. Viral invasion fitness across a continuum from lysis to latency. Virus Evol. 5, vez006 (2019).
Correa, A. M. S. et al. Revisiting the rules of life for viruses of microorganisms. Nat. Rev. Microbiol. https://doi.org/10.1038/s41579-021-00530-x (2021).
Secor, P. R. et al. Filamentous bacteriophage promote biofilm assembly and function. Cell Host Microbe 18, 549–559 (2015).
Sweere, J. M. et al. Bacteriophage trigger antiviral immunity and prevent clearance of bacterial infection. Science 363, eaat9691 (2019).
Berngruber, T. W., Froissart, R., Choisy, M. & Gandon, S. Evolution of virulence in emerging epidemics. PLoS Pathog. 9, e1003209 (2013).
Berngruber, T. W., Lion, S. & Gandon, S. Spatial structure, transmission modes and the evolution of viral exploitation strategies. PLoS Pathog. 11, e1004810 (2015).
Brum, J. R., Hurwitz, B. L., Schofield, O., Ducklow, H. W. & Sullivan, M. B. Seasonal time bombs: dominant temperate viruses affect Southern Ocean microbial dynamics. ISME J. 10, 437–449 (2016).
Liang, X. et al. Lysogenic reproductive strategies of viral communities vary with soil depth and are correlated with bacterial diversity. Soil. Biol. Biochem. 144, 107767 (2020).
Knowles, B. et al. Variability and host density independence in inductions-based estimates of environmental lysogeny. Nat. Microbiol. 2, 17064 (2017).
Kim, M.-S. & Bae, J.-W. Lysogeny is prevalent and widely distributed in the murine gut microbiota. ISME J. 12, 1127–1141 (2018).
Mathieu, A. et al. Virulent coliphages in 1-year-old children fecal samples are fewer, but more infectious than temperate coliphages. Nat. Commun. 11, 378 (2020).
Touchon, M., Bernheim, A. & Rocha, E. P. Genetic and life-history traits associated with the distribution of prophages in bacteria. ISME J. 10, 2744–2754 (2016).
Li, G., Cortez, M. H., Dushoff, J. & Weitz, J. S. When to be temperate: on the fitness benefits of lysis vs. lysogeny. Preprint at biorxiv https://doi.org/10.1101/709758 (2020).
Berngruber, T. W., Weissing, F. J. & Gandon, S. Inhibition of superinfection and the evolution of viral latency. J. Virol. 84, 10200–10208 (2010).
Kieft, K. & Anantharaman, K. Deciphering active prophages from metagenomes. Preprint at biorxiv https://www.biorxiv.org/content/10.1101/2021.01.29.428894 (2021).
Onodera, S. et al. Construction of a transducing virus from double-stranded RNA bacteriophage phi6: establishment of carrier states in host cells. J. Virol. 66, 190–196 (1992).
Pourcel, C., Midoux, C., Vergnaud, G. & Latino, L. A carrier state is established in Pseudomonas aeruginosa by phage LeviOr01, a newly isolated ssRNA levivirus. J. Gen. Virol. 98, 2181–2189 (2017).
Ripp, S. & Miller, R. V. The role of pseudolysogeny in bacteriophage-host interactions in a natural freshwater environment. Microbiology 143, 2065–2070 (1997).
Luo, E., Eppley, J. M., Romano, A. E., Mende, D. R. & DeLong, E. F. Double-stranded DNA virioplankton dynamics and reproductive strategies in the oligotrophic open ocean water column. ISME J. 14, 1304–1315 (2020).
Luo, E., Aylward, F. O., Mende, D. R. & DeLong, E. F. Bacteriophage distributions and temporal variability in the ocean’s interior. mBio 8, e01903 (2017).
Emerson, J. B. et al. Host-linked soil viral ecology along a permafrost thaw gradient. Nat. Microbiol. 3, 870–880 (2018).
Trubl, G. et al. Soil viruses are underexplored players in ecosystem carbon processing. mSystems 3, e00076–18 (2018).
Edwards, R. A. et al. Global phylogeography and ancient evolution of the widespread human gut virus crAssphage. Nat. Microbiol. 4, 1727–1736 (2019).
Shkoporov, A. N. et al. The human gut virome is highly diverse, stable, and individual specific. Cell Host Microbe 26, 527–541 (2019).
Gregory, A. C. et al. The gut virome database reveals age-dependent patterns of virome diversity in the human gut. Cell Host Microbe 28, 724–740 (2020).
Bellas, C. M., Schroeder, D. C., Edwards, A., Barker, G. & Anesio, A. M. Flexible genes establish widespread bacteriophage pan-genomes in cryoconite hole ecosystems. Nat. Commun. 11, 4403 (2020).
Aylward, F. O. et al. Diel cycling and long-term persistence of viruses in the ocean’s euphotic zone. Proc. Natl Acad. Sci. USA 114, 11446–11451 (2017).
Marston, M. F. & Martiny, J. B. H. Genomic diversification of marine cyanophages into stable ecotypes. Environ. Microbiol. 18, 4240–4253 (2016).
Ballaud, F. et al. Dynamics of viral abundance and diversity in a Sphagnum-dominated peatland: temporal fluctuations prevail over habitat. Front. Microbiol. 6, 1494 (2016).
Hevroni, G., Flores-Uribe, J., Béjà, O. & Philosof, A. Seasonal and diel patterns of abundance and activity of viruses in the Red Sea. Proc. Natl Acad. Sci. USA 117, 29738–29747 (2020).
Arkhipova, K. et al. Temporal dynamics of uncultured viruses: a new dimension in viral diversity. ISME J. 12, 199–211 (2018).
Nilsson, E. et al. Genomic and seasonal variations among aquatic phages infecting the baltic sea gammaproteobacterium Rheinheimera sp. strain BAL341. Appl. Environ. Microbiol. 85, e01003–e01019 (2019).
Lim, E. S. et al. Early life dynamics of the human gut virome and bacterial microbiome in infants. Nat. Med. 21, 1228–1234 (2015).
Dahlman, S., Avellaneda-Franco, L. & Barr, J. J. Phages to shape the gut microbiota? Curr. Opin. Biotech. 68, 89–95 (2021).
Shkoporov, A. N. et al. ΦCrAss001 represents the most abundant bacteriophage family in the human gut and infects Bacteroides intestinalis. Nat. Commun. 9, 4781 (2018).
Manrique, P. et al. Healthy human gut phageome. Proc. Natl Acad. Sci. USA 113, 10400–10405 (2016).
Roux, S. et al. Minimum information about an uncultivated virus genome (MIUViG). Nat. Biotechnol. 37, 29–37 (2019).
de Jonge, P. A., Nobrega, F. L., Brouns, S. J. J. & Dutilh, B. E. Molecular and evolutionary determinants of bacteriophage host range. Trends Microbiol. 27, 51–63 (2019).
Cazares, D. et al. A novel group of promiscuous podophages infecting diverse gammaproteobacteria from river communities exhibits dynamic intergenus host adaptation. mSystems 6, e00773–20 (2021).
Heineman, R. H., Springman, R. & Bull, J. J. Optimal foraging by bacteriophages through host avoidance. Am. Naturalist 171, 149–157 (2008).
Meyer, J. R. et al. Repeatability and contingency in the evolution of a key innovation in phage lambda. Science 335, 428–432 (2012).
Meyer, J. R. et al. Ecological speciation of bacteriophage lambda in allopatry and sympatry. Science 354, 1301–1304 (2016).
Holtzman, T. et al. A continuous evolution system for contracting the host range of bacteriophage T7. Sci. Rep. 10, 307 (2020).
Sant, D. G., Woods, L. C., Barr, J. J. & McDonald, M. J. Host diversity slows bacteriophage adaptation by selecting generalists over specialists. Nat. Ecol. Evol. 5, 350–359 (2021).
De Sordi, L., Khanna, V. & Debarbieux, L. The gut microbiota facilitates drifts in the genetic diversity and infectivity of bacterial viruses. Cell Host Microbe 22, 801–808 (2017).
Cornuault, J. K. et al. The enemy from within: a prophage of Roseburia intestinalis systematically turns lytic in the mouse gut, driving bacterial adaptation by CRISPR spacer acquisition. ISME J. 14, 771–787 (2020).
Ford, B. E. et al. Frequency and fitness consequences of bacteriophage Φ6 host range mutations. PLoS ONE 9, e113078 (2014).
Schmerer, M., Molineux, I. J. & Bull, J. J. Synergy as a rationale for phage therapy using phage cocktails. PeerJ 2, e590 (2014).
Streisinger, G. Phenotypic mixing of host range and serological specificities in bacteriophages T2 and T4. Virology 2, 388–398 (1956).
Avrani, S., Wurtzel, O., Sharon, I., Sorek, R. & Lindell, D. Genomic island variability facilitates Prochlorococcus–virus coexistence. Nature 474, 604–608 (2011).
Roux, S., Hallam, S. J., Woyke, T. & Sullivan, M. B. Viral dark matter and virus–host interactions resolved from publicly available microbial genomes. eLife 4, e08490 (2015).
Díaz-Muñoz, S. L. Viral coinfection is shaped by host ecology and virus–virus interactions across diverse microbial taxa and environments. Virus Evol. 3, vex011 (2017).
Joseph, S. B., Hanley, K. A., Chao, L. & Burch, C. L. Coinfection rates in φ6 bacteriophage are enhanced by virus-induced changes in host cells. Evol. Appl. 2, 24–31 (2009).
Weitz, J. S., Mileyko, Y., Joh, R. I. & Voit, E. O. Collective decision making in bacterial viruses. Biophys. J. 95, 2673–2680 (2008).
Erez, Z. et al. Communication between viruses guides lysis-lysogeny decisions. Nature 541, 488–493 (2017).
Trinh, J. T., Székely, T., Shao, Q., Balázsi, G. & Zeng, L. Cell fate decisions emerge as phages cooperate or compete inside their host. Nat. Commun. 8, 14341 (2017).
Refardt, D. Within-host competition determines reproductive success of temperate bacteriophages. ISME J. 5, 1451–1460 (2011).
Griffith, J. & Kornberg, A. Mini M13 bacteriophage: circular fragments of M13 DNA are replicated and packaged during normal infections. Virology 59, 139–152 (1974).
Lindqvist, B. H., Dehò, G. & Calendar, R. Mechanisms of genome propagation and helper exploitation by satellite phage P4. Microbiol. Rev. 57, 683–702 (1993).
Borges, A. L. et al. Bacteriophage cooperation suppresses CRISPR-Cas3 and Cas9 immunity. Cell 174, 917–925 (2018).
Landsberger, M. et al. Anti-CRISPR phages cooperate to overcome CRISPR-Cas Immunity. Cell 174, 908–916 (2018).
Bondy-Denomy, J. et al. Prophages mediate defense against phage infection through diverse mechanisms. ISME J. 10, 2854–2866 (2016).
Dedrick, R. M. et al. Prophage-mediated defence against viral attack and viral counter-defence. Nat. Microbiol. 2, 16251 (2017).
Turner, P. E. & Duffy, S. Evolutionary ecology of multiple phage adsorption and infection. in Bacteriophage Ecology (ed. Abedon, S. T.) 195–216 (Cambridge University Press, 2008).
Secor, P. R. & Dandekar, A. A. More than simple parasites: the sociobiology of bacteriophages and their bacterial hosts. mBio 11, e00041–20 (2020).
Díaz-Muñoz, S. L., Sanjuán, R. & West, S. Sociovirology: conflict, cooperation, and communication among viruses. Cell Host Microbe 22, 437–441 (2017).
Meier-Kolthoff, J. P., Uchiyama, J., Yahara, H., Paez-Espino, D. & Yahara, K. Investigation of recombination-intense viral groups and their genes in the Earth’s virome. Sci. Rep. 8, 11496 (2018).
Mavrich, T. N. & Hatfull, G. F. Bacteriophage evolution differs by host, lifestyle and genome. Nat. Microbiol. 2, 17112 (2017).
Kupczok, A. et al. Rates of mutation and recombination in Siphoviridae phage genome evolution over three decades. Mol. Biol. Evol. 35, 1147–1159 (2018).
de Sousa, J. A. M., Pfeifer, E., Touchon, M. & Rocha, E. P. C. Causes and consequences of bacteriophage diversification via genetic exchanges across lifestyles and bacterial taxa. Mol. Biol. Evol. 38, 2497–2512 (2021).
Eshelman, C. M. et al. Unrestricted migration favours virulent pathogens in experimental metapopulations: evolutionary genetics of a rapacious life history. Phil. Trans. R. Soc. B 365, 2503–2513 (2010).
Waller, A. S. et al. Classification and quantification of bacteriophage taxa in human gut metagenomes. ISME J. 8, 1391–1402 (2014).
Thingstad, T. F. Elements of a theory for the mechanisms controlling abundance, diversity, and biogeochemical role of lytic bacterial viruses in aquatic systems. Limnol. Oceanogr. 45, 1320–1328 (2000).
Xue, C. & Goldenfeld, N. Coevolution maintains diversity in the stochastic ‘Kill the Winner’ model. Phys. Rev. Lett. 119, 268101 (2017).
Howard-Varona, C. et al. Regulation of infection efficiency in a globally abundant marine Bacteriodetes virus. ISME J. 11, 284–295 (2017).
Manage, P. M., Kawabata, Z. & Nakano, S. Seasonal changes in densities of cyanophage infectious to Microcystis aeruginosa in a hypereutrophic pond. Hydrobiologia 411, 211–216 (1999).
Rodriguez-Brito, B. et al. Viral and microbial community dynamics in four aquatic environments. ISME J. 4, 739–751 (2010).
Faruque, S. M. et al. Seasonal epidemics of cholera inversely correlate with the prevalence of environmental cholera phages. Proc. Natl Acad. Sci. USA 102, 1702–1707 (2005).
Faruque, S. M. et al. Self-limiting nature of seasonal cholera epidemics: Role of host-mediated amplification of phage. Proc. Natl Acad. Sci. USA 102, 6119–6124 (2005).
Parsons, R. J., Breitbart, M., Lomas, M. W. & Carlson, C. A. Ocean time-series reveals recurring seasonal patterns of virioplankton dynamics in the northwestern Sargasso Sea. ISME J. 6, 273–284 (2012).
Howard-Varona, C. et al. Phage-specific metabolic reprogramming of virocells. ISME J. 14, 881–895 (2020).
Mann, N. H., Cook, A., Millard, A., Bailey, S. & Clokie, M. Bacterial photosynthesis genes in a virus. Nature 424, 741–741 (2003).
Thompson, L. R. et al. Phage auxiliary metabolic genes and the redirection of cyanobacterial host carbon metabolism. Proc. Natl Acad. Sci. USA 108, 757–764 (2011).
Lindell, D., Jaffe, J. D., Johnson, Z. I., Church, G. M. & Chisholm, S. W. Photosynthesis genes in marine viruses yield proteins during host infection. Nature 438, 86–89 (2005).
Roux, S. et al. Ecogenomics and potential biogeochemical impacts of globally abundant ocean viruses. Nature 537, 689–693 (2016).
Kieft, K. et al. Ecology of inorganic sulfur auxiliary metabolism in widespread bacteriophages. Preprint at bioRxiv https://doi.org/10.1101/2020.08.24.253096v1 (2020).
Spriewald, S. et al. Evolutionary stabilization of cooperative toxin production through a bacterium-plasmid-phage interplay. mBio 11, e00912–e00920 (2020).
De Paepe, M. et al. Carriage of λ latent virus is costly for its bacterial host due to frequent reactivation in monoxenic mouse intestine. PLoS Genet. 12, e1005861 (2016).
Frazão, N., Sousa, A., Lässig, M. & Gordo, I. Horizontal gene transfer overrides mutation in Escherichia coli colonizing the mammalian gut. Proc. Natl Acad. Sci. USA 116, 17906–17915 (2019).
Jamet, A. et al. A widespread family of polymorphic toxins encoded by temperate phages. BMC Biol. 15, 75 (2017).
van Houte, S., Buckling, A. & Westra, E. R. Evolutionary ecology of prokaryotic immune mechanisms. Microbiol. Mol. Biol. Rev. 80, 745–763 (2016).
Doron, S. et al. Systematic discovery of antiphage defense systems in the microbial pangenome. Science 359, 6379 (2018).
Gao, L. et al. Diverse enzymatic activities mediate antiviral immunity in prokaryotes. Science 369, 1077–1084 (2020).
Koskella, B. & Brockhurst, M. A. Bacteria–phage coevolution as a driver of ecological and evolutionary processes in microbial communities. FEMS Microbiol. Rev. 38, 916–931 (2014).
Meaden, S., Paszkiewicz, K. & Koskella, B. The cost of phage resistance in a plant pathogenic bacterium is context-dependent. Evolution 69, 1321–1328 (2015).
Westra, E. R. et al. Parasite exposure drives selective evolution of constitutive versus inducible defense. Curr. Biol. 25, 1043–1049 (2015).
Alseth, E. O. et al. Bacterial biodiversity drives the evolution of CRISPR-based phage resistance. Nature 574, 549–552 (2019).
Koskella, B., Lin, D. M., Buckling, A. & Thompson, J. N. The costs of evolving resistance in heterogeneous parasite environments. Proc. Biol. Sci. 279, 1896–1903 (2012).
Wright, R. C. T., Friman, V.-P., Smith, M. C. M. & Brockhurst, M. A. Resistance evolution against phage combinations depends on the timing and order of exposure. mBio 10, e01652–19 (2019).
Pal, C., Maciá, M. D., Oliver, A., Schachar, I. & Buckling, A. Coevolution with viruses drives the evolution of bacterial mutation rates. Nature 450, 1079–1081 (2007).
Betts, A., Gray, C., Zelek, M., MacLean, R. C. & King, K. C. High parasite diversity accelerates host adaptation and diversification. Science 360, 907–911 (2018).
Wielgoss, S., Bergmiller, T., Bischofberger, A. M. & Hall, A. R. Adaptation to parasites and costs of parasite resistance in mutator and nonmutator bacteria. Mol. Biol. Evol. 33, 770–782 (2016).
Chevallereau, A., Meaden, S., van Houte, S., Westra, E. R. & Rollie, C. The effect of bacterial mutation rate on the evolution of CRISPR-Cas adaptive immunity. Phil. Trans. R. Soc. B 374, 20180094 (2019).
Oliver, A., Cantón, R., Campo, P., Baquero, F. & Blázquez, J. High frequency of hypermutable pseudomonas aeruginosa in cystic fibrosis lung infection. Science 288, 1251–1253 (2000).
Gómez, P. & Buckling, A. Coevolution with phages does not influence the evolution of bacterial mutation rates in soil. ISME J. 7, 2242–2244 (2013).
Matic, I. Mutation rate heterogeneity increases odds of survival in unpredictable environments. Mol. Cell 75, 421–425 (2019).
Doulatov, S. et al. Tropism switching in Bordetella bacteriophage defines a family of diversity-generating retroelements. Nature 431, 476–481 (2004).
Roux, S. et al. Ecology and molecular targets of hypermutation in the global microbiome. Nat. Commun. 12, 3076 (2021).
O’Brien, S., Kümmerli, R., Paterson, S., Winstanley, C. & Brockhurst, M. A. Transposable temperate phages promote the evolution of divergent social strategies in Pseudomonas aeruginosa populations. P. Roy. Soc. B Biol. Sci. 286, 20191794 (2019).
Davies, E. V. et al. Temperate phages both mediate and drive adaptive evolution in pathogen biofilms. Proc. Natl Acad. Sci. USA 113, 8266–8271 (2016).
Rollie, C. et al. Targeting of temperate phages drives loss of type I CRISPR–Cas systems. Nature 578, 149–153 (2020).
Obeng, N., Pratama, A. A. & van Elsas, J. D. The significance of mutualistic phages for bacterial ecology and evolution. Trends Microbiol. 24, 440–449 (2016).
Li, D. et al. Prophage phiv142-3 enhances the colonization and resistance to environmental stresses of avian pathogenic Escherichia coli. Vet. Microbiol. 218, 70–77 (2018).
Brouwer, S. et al. Prophage exotoxins enhance colonization fitness in epidemic scarlet fever-causing Streptococcus pyogenes. Nat. Commun. 11, 5018 (2020).
Duerkop, B. A., Clements, C. V., Rollins, D., Rodrigues, J. L. M. & Hooper, L. V. A composite bacteriophage alters colonization by an intestinal commensal bacterium. Proc. Natl Acad. Sci. USA 109, 17621–17626 (2012).
Diard, M. et al. Inflammation boosts bacteriophage transfer between Salmonella spp. Science 355, 1211–1215 (2017).
Seed, K. D., Lazinski, D. W., Calderwood, S. B. & Camilli, A. A bacteriophage encodes its own CRISPR/Cas adaptive response to evade host innate immunity. Nature 494, 489–491 (2013).
Owen, S. V. et al. Prophage-encoded phage defence proteins with cognate self-immunity. Preprint at biorxiv https://doi.org/10.1101/2020.07.13.199331 (2021).
Chiang, Y. N., Penadés, J. R. & Chen, J. Genetic transduction by phages and chromosomal islands: the new and noncanonical. PLoS Pathog. 15, e1007878 (2019).
Chen, J. et al. Genome hypermobility by lateral transduction. Science 362, 207–212 (2018).
Haaber, J. et al. Bacterial viruses enable their host to acquire antibiotic resistance genes from neighbouring cells. Nat. Commun. 7, 13333 (2016).
Lang, A. S., Westbye, A. B. & Beatty, J. T. The distribution, evolution, and roles of gene transfer agents in prokaryotic genetic exchange. Annu. Rev. Virol. 4, 87–104 (2017).
Calero-Cáceres, W., Ye, M. & Balcázar, J. L. Bacteriophages as environmental reservoirs of antibiotic resistance. Trends Microbiol. 27, 570–577 (2019).
Enault, F. et al. Phages rarely encode antibiotic resistance genes: a cautionary tale for virome analyses. ISME J. 11, 237–247 (2017).
Kenzaka, T., Tani, K. & Nasu, M. High-frequency phage-mediated gene transfer in freshwater environments determined at single-cell level. ISME J. 4, 648–659 (2010).
Kleiner, M., Bushnell, B., Sanderson, K. E., Hooper, L. V. & Duerkop, B. A. Transductomics: sequencing-based detection and analysis of transduced DNA in pure cultures and microbial communities. Microbiome 8, 1–17 (2020).
Solheim, H. T., Sekse, C., Urdahl, A. M., Wasteson, Y. & Nesse, L. L. Biofilm as an environment for dissemination of stx genes by transduction. Appl. Environ. Microbiol. 79, 896–900 (2013).
Popa, O., Landan, G. & Dagan, T. Phylogenomic networks reveal limited phylogenetic range of lateral gene transfer by transduction. ISME J. 11, 543–554 (2017).
Touchon, M., Moura de Sousa, J. A. & Rocha, E. P. Embracing the enemy: the diversification of microbial gene repertoires by phage-mediated horizontal gene transfer. Curr. Opin. Microbiol. 38, 66–73 (2017).
De Sordi, L., Lourenço, M. & Debarbieux, L. “I will survive”: a tale of bacteriophage-bacteria coevolution in the gut. Gut Microbes 10, 92–99 (2018).
Gómez, P. & Buckling, A. Bacteria-phage antagonistic coevolution in soil. Science 332, 106–109 (2011).
Buckling, A. & Rainey, P. B. Antagonistic coevolution between a bacterium and a bacteriophage. Proc. Biol. Sci. 269, 931–936 (2002).
Galtier, M. et al. Bacteriophages targeting adherent invasive Escherichia coli strains as a promising new treatment for Crohn’s disease. J. Crohns Colitis 11, 840–847 (2017).
Lourenço, M. et al. The spatial heterogeneity of the gut limits predation and fosters coexistence of bacteria and bacteriophages. Cell Host Microbe 28, 390–401 (2020).
Koskella, B. Phage-mediated selection on microbiota of a long-lived host. Curr. Biol. 23, 1256–1260 (2013).
Laanto, E., Hoikkala, V., Ravantti, J. & Sundberg, L.-R. Long-term genomic coevolution of host-parasite interaction in the natural environment. Nat. Commun. 8, 111 (2017).
LeGault, K. N. et al. Temporal shifts in antibiotic resistance elements govern virus-pathogen conflicts. Preprint at biorxiv https://www.biorxiv.org/content/ 10.1101/2020.12.16.423150 (2020).
Makarova, K. S., Wolf, Y. I., Snir, S. & Koonin, E. V. Defense islands in bacterial and archaeal genomes and prediction of novel defense systems. J. Bacteriol. 193, 6039–6056 (2011).
Millman, A. et al. Bacterial retrons function in anti-phage defense. Cell 183, 1551–1561 (2020).
Bernheim, A. et al. Prokaryotic viperins produce diverse antiviral molecules. Nature 589, 120–124 (2020).
Ofir, G. et al. Antiviral activity of bacterial TIR domains via signaling molecules that trigger cell death. biorxiv https://www.biorxiv.org/content/10.1101/2021.01.06.425286v1 (2021).
Cohen, D. et al. Cyclic GMP–AMP signalling protects bacteria against viral infection. Nature 574, 691–695 (2019).
Kuzmenko, A. et al. DNA targeting and interference by a bacterial Argonaute nuclease. Nature 587, 632–637 (2020).
Hampton, H. G., Watson, B. N. J. & Fineran, P. C. The arms race between bacteria and their phage foes. Nature 577, 327–336 (2020).
Pinilla-Redondo, R. et al. Discovery of multiple anti-CRISPRs highlights anti-defense gene clustering in mobile genetic elements. Nat. Commun. 11, 5652 (2020).
Brugman, S. et al. A comparative review on microbiota manipulation: lessons from fish, plants, livestock, and human research. Front. Nutr. 5, 80 (2018).
Chan, B. K., Abedon, S. T. & Loc-Carrillo, C. Phage cocktails and the future of phage therapy. Future Microbiol. 8, 769–783 (2013).
Rasmussen, T. S. et al. Faecal virome transplantation decreases symptoms of type 2 diabetes and obesity in a murine model. Gut 69, 2122–2130 (2020).
León, M. & Bastías, R. Virulence reduction in bacteriophage resistant bacteria. Front. Microbiol. 6, 343 (2015).
Friman, V.-P. et al. High temperature and bacteriophages can indirectly select for bacterial pathogenicity in environmental reservoirs. PLoS ONE 6, e17651 (2011).
Roach, D. R. et al. Synergy between the host immune system and bacteriophage is essential for successful phage therapy against an acute respiratory pathogen. Cell Host Microbe 22, 38–47 (2017).
Meir, M. et al. Competition between social cheater viruses is driven by mechanistically different cheating strategies. Sci. Adv. 6, eabb7990 (2020).
Chevallereau, A. et al. Exploitation of the cooperative behaviors of anti-CRISPR phages. Cell Host Microbe 27, 189–198 (2020).
Medvedeva, S. et al. Virus-borne mini-CRISPR arrays are involved in interviral conflicts. Nat. Commun. 10, 5204 (2019).
Pausch, P. et al. CRISPR-CasΦ from huge phages is a hypercompact genome editor. Science 369, 333–337 (2020).
Hays, S. G. & Seed, K. D. Dominant Vibrio cholerae phage exhibits lysis inhibition sensitive to disruption by a defensive phage satellite. eLife 9, e53200 (2020).
Wang, X. et al. Phage combination therapies for bacterial wilt disease in tomato. Nat. Biotech. 37, 1513–1520 (2019).
Koskella, B. & Taylor, T. B. Multifaceted impacts of bacteriophages in the plant microbiome. Annu. Rev. Phytopathol. 56, 361–380 (2018).
Barr, J. J. et al. Bacteriophage adhering to mucus provide a non-host-derived immunity. Proc. Natl Acad. Sci. USA 110, 10771–10776 (2013).
Green, S. I. et al. Targeting of mammalian glycans enhances phage predation in the gastrointestinal tract. mBio 12, e03474–20 (2021).
Mori, K., Kubo, T., Yuki, K., Okhuma, T. & Akira, K. Anti-vaccinia virus effect of M13 bacteriophage DNA. Antivir. Res. 31, 79–86 (1996).
Górski, A. et al. Bacteriophages and transplantation tolerance. Transplant. Proc. 38, 331–333 (2006).
Górski, A. et al. Bacteriophage translocation. FEMS Immunol. Med. Mic. 46, 313–319 (2006).
Jahn, M. T. et al. A phage protein aids bacterial symbionts in eukaryote immune evasion. Cell Host Microbe 26, 542–550 (2019).
Verma, N. K., Brandt, J. M., Verma, D. J. & Lindberg, A. A. Molecular characterization of the O-acetyl transferase gene of converting bacteriophage SF6 that adds group antigen 6 to Shigella flexneri. Mol. Microbiol. 5, 71–75 (1991).
Koonin, E. V., Makarova, K. S. & Wolf, Y. I. Evolutionary genomics of defense systems in archaea and bacteria. Annu. Rev. Microbiol. 71, 233–261 (2017).
Bull, J. J. et al. Phage-bacterial dynamics with spatial structure: self organization around phage sinks can promote increased cell densities. Antibiotics 7, 8 (2018).
Eriksen, R. S., Svenningsen, S. L., Sneppen, K. & Mitarai, N. A growing microcolony can survive and support persistent propagation of virulent phages. Proc. Natl Acad. Sci. USA 115, 337–342 (2018).
Simmons, E. L. et al. Biofilm structure promotes coexistence of phage-resistant and phage-susceptible bacteria. mSystems 5, e00877–19 (2020).
Bull, J. J., Vegge, C. S., Schmerer, M., Chaudhry, W. N. & Levin, B. R. Phenotypic resistance and the dynamics of bacterial escape from phage control. PLoS ONE 9, e94690 (2014).
Bryan, D., El-Shibiny, A., Hobbs, Z., Porter, J. & Kutter, E. M. Bacteriophage T4 infection of stationary phase E. coli: life after log from a phage perspective. Front. Microbiol. 7, 1391 (2016).
Aidley, J., Sørensen, M. C. H., Bayliss, C. D. & Brøndsted, L. Phage exposure causes dynamic shifts in the expression states of specific phase-variable genes of Campylobacter jejuni. Microbiology 163, 911–919 (2017).
Høyland-Kroghsbo, N. M., Maerkedahl, R. B. & Svenningsen, S. L. A quorum-sensing-induced bacteriophage defense mechanism. mBio https://doi.org/10.1128/mBio.00362-12 (2013).
Tan, D., Svenningsen, S. L. & Middelboe, M. Quorum sensing determines the choice of antiphage defense strategy in Vibrio anguillarum. mBio 6, e00627 (2015).
Manning, A. J. & Kuehn, M. J. Contribution of bacterial outer membrane vesicles to innate bacterial defense. BMC Microbiol. 11, 258 (2011).
Reyes-Robles, T. et al. Vibrio cholerae outer membrane vesicles inhibit bacteriophage infection. J. Bacteriol. 200, e00792–17 (2018).
Bru, J.-L. et al. PQS produced by the Pseudomonas aeruginosa stress response repels swarms away from bacteriophage and antibiotics. J. Bacteriol. 201, e00383–19 (2019).
Pearl, S., Gabay, C., Kishony, R., Oppenheim, A. & Balaban, N. Q. Nongenetic individuality in the host–phage interaction. PLoS Biol. 6, e120 (2008).
Acknowledgements
A.C. has received funding from the European Union’s Horizon 2020 research and innovation programme under the Marie Skłodowska-Curie Actions (grant agreement no. 834052, anti-CRISPR). E.R.W. has received funding from the European Research Council (ERC-STG-2016-714478 — EVOIMMECH), the Leverhulme Trust (RPG-2018-380) and the UK Natural Environment Research Council (NE/M018350/1 and NE/S001921/1). S.v.H. has received funding from the UK Biotechnology and Biological Sciences Research Council (BB/S017674/1 and BB/R010781/10).
Author information
Authors and Affiliations
Contributions
A.C. and B.J.P. researched data for the article. A.C. and E.R.W. substantially contributed to discussion of the content. A.C. and B.J.P wrote the manuscript. E.R.W and S.V.H. reviewed and edited the manuscript before submission.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests
Additional information
Peer review information
Nature Reviews Microbiology thanks B. Koskella and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Glossary
- Mosaicism
-
The description of a phage genome as composed of gene blocks similar to other phage genomes and flanked by dissimilar sequences. This mosaic genome architecture is due to horizontal transfer of genetic material.
- Lysogenic conversion
-
Phenotypic changes that result from the acquisition and expression of prophage-encoded genes.
- Transduction
-
Transfer of bacterial DNA from one bacterium to another, mediated through phage infection.
- Viral dark matter
-
A viral species (or genome) that has not yet been characterized but for which the existence has been revealed by environmental metagenomic sequencing. More restrictively, viral dark matter can also refer to phage genes that have no assigned functions.
- Prophage
-
Latent form of a phage in which lytic functions are repressed and that is often integrated into the host chromosome.
- Lysogeny
-
Phenomenon where a temperate phage parasitizes a host bacterium (hence called a ‘lysogen’), often by integrating its genetic material into the host DNA, but without producing phage particles or triggering host lysis.
- Satellite phages
-
Phages that lack the ability to replicate autonomously and require the products of infection generated by a helper phage (for example, capsid proteins).
- Superinfection
-
Infection of a host that already accommodates a phage from an earlier infection, which can be engaged either in a lytic or in a lysogenic cycle and integrated as a prophage.
- Copiotroph species
-
Fast-growing organisms found in nutrient-rich environments.
- Homoimmunity
-
Mechanism by which a prophage inhibits the secondary infection of its host by a closely related phage.
- Viromes
-
Ensembles of viral genomes found in an organism or within a given environment.
- Temperate phages
-
Phages that replicate either through a lytic cycle or through a lysogenic cycle.
- Virulent phages
-
Phages that replicate exclusively through a lytic cycle, which ultimately triggers the death of the host cell, releasing new phage particles.
- Phageome
-
Phage fraction of the virome. Phages are generally the main constituents of viromes.
- Host range
-
The range of genetically distinct bacteria in which a phage can replicate. Usually, phages that infect a single or a few genetically close strains are referred to as ‘specialists’, whereas phages that infect many strains of the same species, or even multiple different species or genera, are called ‘generalists’.
- Co-infection
-
Infection where two phages are simultaneously present within the same host cell. This definition therefore widely includes phenomena such as superinfection or polylysogeny.
- Lysis inhibition
-
Delay of the lysis of an infecting phage (this extension of phage latent period results in an increased burst size) induced by secondary adsorptions of additional phages.
- Auxiliary metabolic genes
-
(AMGs). Phage-encoded genes that originate from bacterial cells and can modulate host cell metabolism, likely resulting in improved phage replication.
- Virocells
-
Refers to a concept stating that the reproductive, living form of a virus is the infected cell (called virocell) as opposed to its dissemination form, the virion.
- Phase variation
-
Switch of gene expression from an ‘on’ phase to an ‘off’ phase, generally caused by the introduction of a mutation into a hypermutable DNA region. A secondary mutation in the same region can reactivate gene expression, rendering this phenomenon reversible. Phase variation generates phenotypic diversity in bacterial populations.
- Phage-inducible chromosomal islands
-
Class of mobile genetic elements that specifically exploit (and interfere with) temperate phages for their horizontal transfer.
- Fluctuating selection dynamics
-
Type of phage–bacterium coevolutionary dynamics where phage and host genotype frequencies oscillate over time because of negative frequency-dependent selection.
- Arms race dynamics
-
Type of phage–bacterium coevolutionary dynamics where phage infectivity and host resistance generally increase over time.
Rights and permissions
About this article
Cite this article
Chevallereau, A., Pons, B.J., van Houte, S. et al. Interactions between bacterial and phage communities in natural environments. Nat Rev Microbiol 20, 49–62 (2022). https://doi.org/10.1038/s41579-021-00602-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41579-021-00602-y
This article is cited by
-
Integrative metagenomic analysis reveals distinct gut microbial signatures related to obesity
BMC Microbiology (2024)
-
Inhibitors of bacterial immune systems: discovery, mechanisms and applications
Nature Reviews Genetics (2024)
-
Bacteriophages from human skin infecting coagulase-negative Staphylococcus: diversity, novelty and host resistance
Scientific Reports (2024)
-
Phage Paride can kill dormant, antibiotic-tolerant cells of Pseudomonas aeruginosa by direct lytic replication
Nature Communications (2024)
-
Discovery and description of novel phage genomes from urban microbiomes sampled by the MetaSUB consortium
Scientific Reports (2024)