Abstract
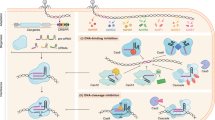
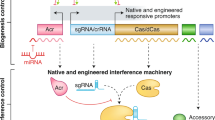
The number and diversity of known CRISPR–Cas systems have substantially increased in recent years. Here, we provide an updated evolutionary classification of CRISPR–Cas systems and cas genes, with an emphasis on the major developments that have occurred since the publication of the latest classification, in 2015. The new classification includes 2 classes, 6 types and 33 subtypes, compared with 5 types and 16 subtypes in 2015. A key development is the ongoing discovery of multiple, novel class 2 CRISPR–Cas systems, which now include 3 types and 17 subtypes. A second major novelty is the discovery of numerous derived CRISPR–Cas variants, often associated with mobile genetic elements that lack the nucleases required for interference. Some of these variants are involved in RNA-guided transposition, whereas others are predicted to perform functions distinct from adaptive immunity that remain to be characterized experimentally. The third highlight is the discovery of numerous families of ancillary CRISPR-linked genes, often implicated in signal transduction. Together, these findings substantially clarify the functional diversity and evolutionary history of CRISPR–Cas.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Komor, A. C., Badran, A. H. & Liu, D. R. CRISPR-based technologies for the manipulation of eukaryotic genomes. Cell 168, 20–36 (2017).
Pickar-Oliver, A. & Gersbach, C. A. The next generation of CRISPR–Cas technologies and applications. Nat. Rev. Mol. Cell Biol. 20, 490–507 (2019).
Mohanraju, P. et al. Diverse evolutionary roots and mechanistic variations of the CRISPR–Cas systems. Science 353, aad5147 (2016).
Jackson, S. A. et al. CRISPR–Cas: adapting to change. Science 356, eaal5056 (2017).
Barrangou, R. & Horvath, P. A decade of discovery: CRISPR functions and applications. Nat. Microbiol. 2, 17092 (2017).
Jiang, F. & Doudna, J. A. CRISPR–Cas9 structures and mechanisms. Annu. Rev. Biophys. 46, 505–529 (2017).
Koonin, E. V. & Makarova, K. S. Origins and evolution of CRISPR–Cas systems. Philos. Trans. R. Soc. Lond. B Biol. Sci. 374, 20180087 (2019).
Faure, G., Makarova, K. S. & Koonin, E. V. CRISPR–Cas: complex functional networks and multiple roles beyond adaptive immunity. J. Mol. Biol. 431, 3–20 (2019).
McGinn, J. & Marraffini, L. A. Molecular mechanisms of CRISPR–Cas spacer acquisition. Nat. Rev. Microbiol. 17, 7–12 (2019).
Koonin, E. V., Makarova, K. S. & Wolf, Y. I. Evolutionary genomics of defense systems in archaea and bacteria. Annu. Rev. Microbiol. 71, 233–261 (2017).
Koonin, E. V., Makarova, K. S. & Zhang, F. Diversity, classification and evolution of CRISPR–Cas systems. Curr. Opin. Microbiol. 37, 67–78 (2017).
Ishino, Y., Krupovic, M. & Forterre, P. History of CRISPR–Cas from encounter with a mysterious repeated sequence to genome editing technology. J. Bacteriol. 200, e00580-17 (2018).
Hille, F. & Charpentier, E. CRISPR–Cas: biology, mechanisms and relevance. Philos. Trans. R. Soc. Lond. B Biol. Sci. 371, 20150496 (2016).
Wright, A. V., Nunez, J. K. & Doudna, J. A. Biology and applications of CRISPR systems: harnessing nature’s toolbox for genome engineering. Cell 164, 29–44 (2016).
Klompe, S. E. & Sternberg, S. H. Harnessing ‘a billion years of experimentation’: the ongoing exploration and exploitation of CRISPR–Cas immune systems. CRISPR J. 1, 141–158 (2018).
Makarova, K. S., Wolf, Y. I. & Koonin, E. V. Classification and nomenclature of CRISPR–Cas systems: where from here? CRISPR J. 1, 325–336 (2018).
Makarova, K. S. et al. Evolution and classification of the CRISPR–Cas systems. Nat. Rev. Microbiol. 9, 467–477 (2011).
Makarova, K. S. et al. An updated evolutionary classification of CRISPR–Cas systems. Nat. Rev. Microbiol. 13, 722–736 (2015).
Shmakov, S. et al. Discovery and functional characterization of diverse class 2 CRISPR–Cas systems. Mol. Cell 60, 385–397 (2015).
Shmakov, S. et al. Diversity and evolution of class 2 CRISPR–Cas systems. Nat. Rev. Microbiol. 15, 169–182 (2017). This work demonstrates the relationships between the effectors of different types and subtypes of class 2 CRISPR–Cas systems and nucleases encoded by mobile genetic elements. On the basis of sequence comparison and phylogenetic analysis of Cas12 (type V effectors) and TnpB nucleases encoded by transposons, a scenario of independent recruitment of distinct TnpB variants, giving rise to different type V subtypes, is proposed.
Burstein, D. et al. New CRISPR–Cas systems from uncultivated microbes. Nature 542, 237–241 (2017). This work describes the metagenomic discovery of two new subtypes of type V CRISPR–Cas systems and experimental validation of their activity.
Harrington, L. B. et al. Programmed DNA destruction by miniature CRISPR–Cas14 enzymes. Science 362, 839–842 (2018). This work experimentally validates the enzymatic activity of small predicted effectors that have been assigned to subtype V-U by Shmakov et al. (2017) and are here reclassified as subtype V-F. It shows that these enzymes differ substantially from the previously characterized large type II and type V effectors and catalyse both crRNA-specific and non-specific cleavage of single-stranded DNA.
Yan, W. X. et al. Functionally diverse type V CRISPR–Cas systems. Science 363, 88–91 (2019). This article reports the experimental characterization of CRISPR–Cas subtypes V-C, V-G, V-H and V-I. Whereas Cas12c, Cas12h and Cas12i proteins all demonstrate RNA-guided double-stranded DNA interference similar to that in previously described CRISPR–Cas effectors, Cas12g is shown to function as an RNase with collateral RNase and single-strand DNase activities.
Abudayyeh, O. O. et al. C2c2 is a single-component programmable RNA-guided RNA-targeting CRISPR effector. Science 353, aaf5573 (2016).
Smargon, A. A. et al. Cas13b is a type VI-B CRISPR-associated RNA-guided RNase differentially regulated by accessory proteins Csx27 and Csx28. Mol. Cell 65, 618–630.e7 (2017).
Yan, W. X. et al. Cas13d is a compact RNA-targeting type VI CRISPR effector positively modulated by a WYL-domain-containing accessory protein. Mol. Cell 70, 327–339.e5 (2018). This study demonstrates RNA targeting by the smallest known type VI effector, Cas13d, and shows that the accessory WYL domain-containing protein stimulates this activity.
Murugan, K., Babu, K., Sundaresan, R., Rajan, R. & Sashital, D. G. The revolution continues: newly discovered systems expand the CRISPR–Cas toolkit. Mol. Cell 68, 15–25 (2017).
Stella, S., Alcon, P. & Montoya, G. Class 2 CRISPR–Cas RNA-guided endonucleases: Swiss army knives of genome editing. Nat. Struct. Mol. Biol. 24, 882–892 (2017).
Koonin, E. V. & Makarova, K. S. Mobile genetic elements and evolution of CRISPR–Cas systems: all the way there and back. Genome Biol. Evol. 9, 2812–2825 (2017).
Faure, G. et al. CRISPR–Cas in mobile genetic elements: counter-defense and beyond. Nat. Rev. Microbiol. 17, 513–525 (2019).
Shah, S. A. et al. Comprehensive search for accessory proteins encoded with archaeal and bacterial type III CRISPR-cas gene cassettes reveals 39 new cas gene families. RNA Biol. 16, 530–542 (2019). Along with Shmakov et al. (2018), this study describes a computational approach to predict proteins that are functionally linked to CRISPR–Cas systems and applies this approach to type III systems.
Shmakov, S. A., Makarova, K. S., Wolf, Y. I., Severinov, K. V. & Koonin, E. V. Systematic prediction of genes functionally linked to CRISPR–Cas systems by gene neighborhood analysis. Proc. Natl Acad. Sci. USA 115, E5307–E5316 (2018). Along with Shah et al. (2019), this article describes a computational approach for the systematic prediction of proteins that are functionally linked to CRISPR–Cas systems (‘CRISPRicity’ protocol) and applies that approach to all CRISPR–Cas types and subtypes.
Peters, J. E., Makarova, K. S., Shmakov, S. & Koonin, E. V. Recruitment of CRISPR–Cas systems by Tn7-like transposons. Proc. Natl Acad. Sci. USA 114, E7358–E7366 (2017). This study describes, for the first time, defective CRISPR–Cas systems encoded in Tn7-like transposons and predicts their function in RNA-guided transposition.
Strecker, J. et al. RNA-guided DNA insertion with CRISPR-associated transposases. Science 365, 48–53 (2019). This work validates the prediction made in Shmakov et al. (2017), by showing that V-U5 variant effector proteins, which are inactivated TnpB homologues encoded in Tn7-like transposons, form a complex with the transposase subunit and enable crRNA-dependent transposition.
Klompe, S. E., Vo, P. L. H., Halpin-Healy, T. S. & Sternberg, S. H. Transposon-encoded CRISPR–Cas systems direct RNA-guided DNA integration. Nature 571, 219–225 (2019). This work complements Strecker et al. (2019) by experimentally validating the prediction made in Peters et al. (2017) that interference-deficient subtype I-F CRISPR–Cas systems encoded in Tn7-like transposons enable crRNA-dependent transposition.
Kazlauskiene, M., Kostiuk, G., Venclovas, C., Tamulaitis, G. & Siksnys, V. A cyclic oligonucleotide signaling pathway in type III CRISPR-Cas systems. Science 357, 605–609 (2017). Along with Niewoehner et al. (2017), this article describes the signalling pathway involved in the function of type III CRISPR–Cas systems, which involves the synthesis of cyclic oligoA molecules by Cas10, binding of these signalling molecules to the CARF domain of Csm6 and activation of the second domain of Casm6, the HEPN nuclease that catalyses promiscuous RNA cleavage.
Niewoehner, O. et al. Type III CRISPR–Cas systems produce cyclic oligoadenylate second messengers. Nature 548, 543–548 (2017).
Makarova, K. S. & Koonin, E. V. Annotation and classification of CRISPR–Cas systems. Methods Mol. Biol. 1311, 47–75 (2015).
Altschul, S. F. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25, 3389–3402 (1997).
Iranzo, J., Krupovic, M. & Koonin, E. V. The double-stranded DNA virosphere as a modular hierarchical network of gene sharing. MBio 7, e00978-16 (2016).
Iranzo, J., Martincorena, I. & Koonin, E. V. Cancer-mutation network and the number and specificity of driver mutations. Proc. Natl Acad. Sci. USA 115, E6010–E6019 (2018).
Makarova, K. S., Wolf, Y. I. & Koonin, E. V. The basic building blocks and evolution of CRISPR–Cas systems. Biochem. Soc. Trans. 41, 1392–1400 (2013).
Koonin, E. V. & Makarova, K. S. Discovery of oligonucleotide signaling mediated by CRISPR-associated polymerases solves two puzzles but leaves an enigma. ACS Chem. Biol. 13, 309–312 (2018).
Silas, S. et al. On the origin of reverse transcriptase-using CRISPR–Cas systems and their hyperdiverse, enigmatic spacer repertoires. MBio 8, e00897-17 (2017).
Puigbo, P., Makarova, K. S., Kristensen, D. M., Wolf, Y. I. & Koonin, E. V. Reconstruction of the evolution of microbial defense systems. BMC Evol. Biol. 17, 94 (2017).
Garrett, R. A., Vestergaard, G. & Shah, S. A. Archaeal CRISPR-based immune systems: exchangeable functional modules. Trends Microbiol. 19, 549–556 (2011).
Reeks, J., Naismith, J. H. & White, M. F. CRISPR interference: a structural perspective. Biochem. J. 453, 155–166 (2013).
Özcan, A. et al. Type IV CRISPR RNA processing and effector complex formation in Aromatoleum aromaticum. Nat. Microbiol. 19, 89–96 (2019).
Makarova, K. S., Aravind, L., Wolf, Y. I. & Koonin, E. V. Unification of Cas protein families and a simple scenario for the origin and evolution of CRISPR–Cas systems. Biol. Direct 6, 38 (2011).
Venclovas, C. Structure of Csm2 elucidates the relationship between small subunits of CRISPR–Cas effector complexes. FEBS Lett. 590, 1521–1529 (2016).
Chylinski, K., Makarova, K. S., Charpentier, E. & Koonin, E. V. Classification and evolution of type II CRISPR–Cas systems. Nucleic Acids Res. 42, 6091–6105 (2014).
Briner, A. E. & Barrangou, R. Guide RNAs: a glimpse at the sequences that drive CRISPR–Cas systems. Cold Spring Harb. Protoc. 2016, pdb.top090902 (2016).
Faure, G. et al. Comparative genomics and evolution of trans-activating RNAs in Class 2 CRISPR–Cas systems. RNA Biol. 16, 435–448 (2019).
Chyou, T. Y. & Brown, C. M. Prediction and diversity of tracrRNAs from type II CRISPR–Cas systems. RNA Biol. 16, 423–434 (2019).
Fonfara, I., Richter, H., Bratovic, M., Le Rhun, A. & Charpentier, E. The CRISPR-associated DNA-cleaving enzyme Cpf1 also processes precursor CRISPR RNA. Nature 532, 517–521 (2016).
East-Seletsky, A. et al. Two distinct RNase activities of CRISPR–C2c2 enable guide-RNA processing and RNA detection. Nature 538, 270–273 (2016).
Liu, L. et al. Two distant catalytic sites are responsible for C2c2 RNase activities. Cell 168, 121–134 (2017).
Kunin, V., Sorek, R. & Hugenholtz, P. Evolutionary conservation of sequence and secondary structures in CRISPR repeats. Genome Biol. 8, R61 (2007).
Lange, S. J., Alkhnbashi, O. S., Rose, D., Will, S. & Backofen, R. CRISPRmap: an automated classification of repeat conservation in prokaryotic adaptive immune systems. Nucleic Acids Res. 41, 8034–8044 (2013).
Almendros, C., Nobrega, F. L., McKenzie, R. E. & Brouns, S. J. J. Cas4–Cas1 fusions drive efficient PAM selection and control CRISPR adaptation. Nucleic Acids Res. 47, 5223–5230 (2019).
Koonin, E. V. & Aravind, L. Origin and evolution of eukaryotic apoptosis: the bacterial connection. Cell Death Differ. 9, 394–404 (2002).
Vestergaard, G., Garrett, R. A. & Shah, S. A. CRISPR adaptive immune systems of Archaea. RNA Biol. 11, 156–167 (2014).
Sinkunas, T. et al. Cas3 is a single-stranded DNA nuclease and ATP-dependent helicase in the CRISPR/Cas immune system. EMBO J. 30, 1335–1342 (2011).
Al-Shayeb, B. et al. Clades of huge phage from across Earth’s ecosystems. Preprint at https://doi.org/10.1101/572362 (2019).
Leipe, D. D., Koonin, E. V. & Aravind, L. STAND, a class of P-loop NTPases including animal and plant regulators of programmed cell death: multiple, complex domain architectures, unusual phyletic patterns, and evolution by horizontal gene transfer. J. Mol. Biol. 343, 1–28 (2004).
Newire, E., Aydin, A., Juma, S., Enne, V. & Roberts, A. P. Identification of a Type IV CRISPR–Cas system located exclusively on IncHI1B/IncFIB plasmids in Enterobacteriaceae. Preprint at https://doi.org/10.1101/536375 (2019).
Makarova, K. S. et al. Predicted highly derived class 1 CRISPR–Cas system in Haloarchaea containing diverged Cas5 and Cas7 homologs but no CRISPR array. FEMS Microbiol. Lett. 366, fnz079 (2019).
Zaremba-Niedzwiedzka, K. et al. Asgard archaea illuminate the origin of eukaryotic cellular complexity. Nature 541, 353–358 (2017).
Strecker, J. et al. Engineering of CRISPR–Cas12b for human genome editing. Nat. Commun. 10, 212 (2019).
Swarts, D. C. & Jinek, M. Mechanistic insights into the cis- and trans-acting DNase activities of Cas12a. Mol. Cell 73, 589–600.e584 (2019).
Meeske, A. J., Nakandakari-Higa, S. & Marraffini, L. A. Cas13-induced cellular dormancy prevents the rise of CRISPR-resistant bacteriophage. Nature 570, 241–245 (2019).
Karvelis, T. et al. PAM recognition by miniature CRISPR–Cas14 triggers programmable double-stranded DNA cleavage. Preprint at https://doi.org/10.1101/654897 (2019).
Anantharaman, V., Makarova, K. S., Burroughs, A. M., Koonin, E. V. & Aravind, L. Comprehensive analysis of the HEPN superfamily: identification of novel roles in intra-genomic conflicts, defense, pathogenesis and RNA processing. Biol. Direct 8, 15 (2013).
Hudaiberdiev, S. et al. Phylogenomics of Cas4 family nucleases. BMC Evol. Biol. 17, 232 (2017).
Charpentier, E., Richter, H., van der Oost, J. & White, M. F. Biogenesis pathways of RNA guides in archaeal and bacterial CRISPR-Cas adaptive immunity. FEMS Microbiol. Rev. 39, 428–441 (2015).
Dudek, N. K. et al. Novel microbial diversity and functional potential in the marine mammal oral microbiome. Curr. Biol. 27, 3752–3762 e3756 (2017).
Castelle, C. J. et al. Biosynthetic capacity, metabolic variety and unusual biology in the CPR and DPANN radiations. Nat. Rev. Microbiol. 16, 629–645 (2018).
Burstein, D. et al. Major bacterial lineages are essentially devoid of CRISPR–Cas viral defence systems. Nat. Commun. 7, 10613 (2016).
Levin, B. R. Nasty viruses, costly plasmids, population dynamics, and the conditions for establishing and maintaining CRISPR-mediated adaptive immunity in bacteria. PLOS Genet. 6, e1001171 (2010).
Iranzo, J., Lobkovsky, A. E., Wolf, Y. I. & Koonin, E. V. Evolutionary dynamics of the prokaryotic adaptive immunity system CRISPR–Cas in an explicit ecological context. J. Bacteriol. 195, 3834–3844 (2013).
Iranzo, J., Lobkovsky, A. E., Wolf, Y. I. & Koonin, E. V. Immunity, suicide or both? Ecological determinants for the combined evolution of anti-pathogen defense systems. BMC Evol. Biol. 15, 43 (2015).
Gurney, J., Pleska, M. & Levin, B. R. Why put up with immunity when there is resistance: an excursion into the population and evolutionary dynamics of restriction-modification and CRISPR–Cas. Philos. Trans. R. Soc. Lond. B Biol. Sci. 374, 20180096 (2019).
Garcia-Martinez, J., Maldonado, R. D., Guzman, N. M. & Mojica, F. J. M. The CRISPR conundrum: evolve and maybe die, or survive and risk stagnation. Microb. Cell 5, 262–268 (2018).
van Houte, S. et al. The diversity-generating benefits of a prokaryotic adaptive immune system. Nature 532, 385–388 (2016).
Weinberger, A. D., Wolf, Y. I., Lobkovsky, A. E., Gilmore, M. S. & Koonin, E. V. Viral diversity threshold for adaptive immunity in prokaryotes. MBio 3, e00456-12 (2012).
Westra, E. R. et al. Parasite exposure drives selective evolution of constitutive versus inducible defense. Curr. Biol. 25, 1043–1049 (2015).
Bernheim, A., Bikard, D., Touchon, M. & Rocha, E. P. C. A matter of background: DNA repair pathways as a possible cause for the sparse distribution of CRISPR–Cas systems in bacteria. Philos. Trans. R. Soc. Lond. B Biol. Sci. 374, 20180088 (2019).
Koonin, E. V. & Krupovic, M. Evolution of adaptive immunity from transposable elements combined with innate immune systems. Nat. Rev. Genet. 16, 184–192 (2015).
Krupovic, M., Beguin, P. & Koonin, E. V. Casposons: mobile genetic elements that gave rise to the CRISPR–Cas adaptation machinery. Curr. Opin. Microbiol. 38, 36–43 (2017).
Krupovic, M., Makarova, K. S., Forterre, P., Prangishvili, D. & Koonin, E. V. Casposons: a new superfamily of self-synthesizing DNA transposons at the origin of prokaryotic CRISPR–Cas immunity. BMC Biol. 12, 36 (2014).
Kieper, S. N. et al. Cas4 facilitates PAM-compatible spacer selection during CRISPR adaptation. Cell Rep. 22, 3377–3384 (2018).
Lee, H., Zhou, Y., Taylor, D. W. & Sashital, D. G. Cas4-dependent prespacer processing ensures high-fidelity programming of CRISPR arrays. Mol. Cell 70, 48–59.e45 (2018).
Shiimori, M., Garrett, S. C., Graveley, B. R. & Terns, M. P. Cas4 nucleases define the pam, length, and orientation of DNA fragments integrated at CRISPR loci. Mol. Cell 70, 814–824.e816 (2018). This work reveals the molecular details of the involvement of Cas4, an ancillary protein that cooperates with Cas1 and Cas2 in several CRISPR–Cas subtypes, in the process of adaptation.
Burroughs, A. M., Zhang, D., Schaffer, D. E., Iyer, L. M. & Aravind, L. Comparative genomic analyses reveal a vast, novel network of nucleotide-centric systems in biological conflicts, immunity and signaling. Nucleic Acids Res. 43, 10633–10654 (2015).
Rostol, J. T. & Marraffini, L. A. Non-specific degradation of transcripts promotes plasmid clearance during type III-A CRISPR-Cas immunity. Nat. Microbiol. 4, 656–662 (2019). This work demonstrates that indiscriminate RNA cleavage by the HEPN RNase domain of the Csm6 protein of type III CRISPR–Cas systems induces growth arrest in the host bacteria, providing a backup defence mechanism.
Makarova, K. S., Wolf, Y. I., Snir, S. & Koonin, E. V. Defense islands in bacterial and archaeal genomes and prediction of novel defense systems. J. Bacteriol. 193, 6039–6056 (2011).
Kapitonov, V. V., Makarova, K. S. & Koonin, E. V. ISC, a novel group of bacterial and archaeal DNA transposons that encode Cas9 homologs. J. Bacteriol. 198, 797–807 (2015).
Athukoralage, J. S., Rouillon, C., Graham, S., Gruschow, S. & White, M. F. Ring nucleases deactivate type III CRISPR ribonucleases by degrading cyclic oligoadenylate. Nature 562, 277–280 (2018). This works expands the characterization of the signalling pathway in type III CRISPR–Cas sequences by showing that a distinct variety of CARF domain cleaves the cyclic oligoA molecules produced by Cas10 and thus regulates the pathway.
Shmakov, S. A. et al. Systematic prediction of functionally linked genes in bacterial and archaeal genomes. Nat. Protoc. 14, 3013–3031 (2019).
Acknowledgements
K.S.M., Y.I.W., J.I., S.A.S. and E.V.K. are supported through the Intramural Research Program of the US National Institutes of Health; F.J.M.M. was supported by grants BIO2014-53029-P (Ministerio de Ciencia, Innovación y Universidades, Spain), and 291815 Era-Net ANIHWA (7th Framework Programme, European Commission) and PROMETEO/2017/129 (Conselleria d'Educació, Investigació, Cultura i Esport, Generalitat Valenciana, Spain); S.A.S. was supported by RFBR (research project 18-34-00012) and a Systems Biology Fellowship from Philip Morris Sales and Marketing; S.M. was funded by funding from the Natural Sciences and Engineering Research Council of Canada (Discovery program) and holds a Tier 1 Canada Research Chair in Bacteriophages.
Author information
Authors and Affiliations
Contributions
K.S.M., Y.I.W., J.I., S.A.S., D.C., Č.V. and E.V.K. researched data for the article. K.S.M., Y.I.W., S.J.J.B., O.S.A., E.C., D.C., D.H.H., P.H., S.M., F.J.M.M., D.S., S.A.A., V.S., M.P.T., Č.V., M.F.W., A.F.Y., W.Y., F.Z., R.A.G., R.B., J.v.d.O., R.B. and E.V.K. substantially contributed to discussion of the content. K.S.M., J.I., Y.I.W. and E.V.K. wrote the article. K.S.M., Y.I.W, S.M., R.A.G, J.v.d.O., R.B. and E.V.K. reviewed and edited the manuscript before submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Glossary
- CRISPR
-
Clustered regularly interspaced short palindromic repeats, present in most archaeal and many bacterial genomes.
- Cas
-
CRISPR-associated (proteins).
- Adaptation
-
First stage of the CRISPR–Cas response that involves spacer acquisition.
- Interference
-
Final stage of the CRISPR–Cas response, which involves recognition and cleavage of the target DNA or RNA.
- Protospacer-adjacent motif
-
(PAM). A short nucleotide sequence next to the protospacer that is required for target recognition by the crRNA effector.
- Protospacer
-
Segment of DNA (typically, from a virus or plasmid) that is acquired by CRISPR–Cas systems via the activity of the adaptation complex.
- CRISPR array
-
Genomic locus containing multiple, tandem CRISPR.
- Spacer
-
Unique segment of DNA inserted between CRISPR units.
- CRISPR–cas
-
Archaeal and bacterial system of adaptive immunity that consists of a CRISPR array and cas genes.
- pre-crRNA
-
Long transcript of a CRISPR locus that is processed to yield the crRNA CRISPR–Cas system, where it is incorporated as a spacer.
- crRNAs
-
Short RNA molecules containing the spacer sequence and parts of the CRISPR, used as the guide to target and cleave cognate foreign DNA or RNA.
- Transposon
-
A mobile genetic element, typically flanked by inverted terminal repeats, that changes its location in the host genome by inserting into new sites with the help of a transposon-encoded enzyme known as transposase, integrase or recombinase.
- Casposon
-
A member of a distinct class of transposons that employ a Cas1 homologue as the transposases and are thought to be the ancestors of CRISPR–Cas adaptation modules.
Rights and permissions
About this article
Cite this article
Makarova, K.S., Wolf, Y.I., Iranzo, J. et al. Evolutionary classification of CRISPR–Cas systems: a burst of class 2 and derived variants. Nat Rev Microbiol 18, 67–83 (2020). https://doi.org/10.1038/s41579-019-0299-x
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41579-019-0299-x
This article is cited by
-
Eukaryotic-driven directed evolution of Cas9 nucleases
Genome Biology (2024)
-
GMP-manufactured CRISPR/Cas9 technology as an advantageous tool to support cancer immunotherapy
Journal of Experimental & Clinical Cancer Research (2024)
-
Molecular basis and engineering of miniature Cas12f with C-rich PAM specificity
Nature Chemical Biology (2024)
-
PtuA and PtuB assemble into an inflammasome-like oligomer for anti-phage defense
Nature Structural & Molecular Biology (2024)
-
Tail-tape-fused virion and non-virion RNA polymerases of a thermophilic virus with an extremely long tail
Nature Communications (2024)