Abstract
Recent discoveries of contemporary genotypes of hepatitis B virus and parvovirus B19 in ancient human remains demonstrate that little genetic change has occurred in these viruses over 4,500–6,000 years. Endogenous viral elements in host genomes provide separate evidence that viruses similar to many major contemporary groups circulated 100 million years ago or earlier. In this Opinion article, we argue that the extraordinary conservation of virus genome sequences is best explained by a niche-filling model in which fitness optimization is rapidly achieved in their specific hosts. Whereas short-term substitution rates reflect the accumulation of tolerated sequence changes within adapted genomes, longer-term rates increasingly resemble those of their hosts as the evolving niche moulds and effectively imprisons the virus in co-adapted virus–host relationships. Contrastingly, viruses that jump hosts undergo strong and stringent adaptive selection as they maximize their fit to their new niche. This adaptive capability may paradoxically create evolutionary stasis in long-term host relationships. While viruses can evolve and adapt rapidly, their hosts may ultimately shape their longer-term evolution.
Similar content being viewed by others
Introduction
Viruses, with their often small genomes and error-prone replication mechanisms, possess extraordinary adaptive abilities and can display rates of sequence change that are orders of magnitude greater than those of the hosts they infect. They display evolution in real time as they acquire antiviral drug resistance, mediate persistent infection through escape from T and B cell immune system responses to infection or, at the experimental level, rapidly adapt to different cell culture conditions, new receptors and new hosts. Although biologists since the time of Darwin have convincingly inferred the existence of natural selection from the current species distributions of animals and plants, and their genetic relationships, this evidence is almost always indirect and observational. By contrast, virologists have access to the remarkable field of experimental evolution, such that adaptive processes that may occur over centuries or millennia in larger organisms can be observed in viruses over days or weeks.
The paradox that this Opinion article aims to address is the increasing evidence for extreme genetic conservation of viruses over longer periods of evolution. Newly developed methods to characterize viruses from ancient DNA (aDNA) samples have revealed that viruses that circulated in ancient times do not substantially differ genetically from those that currently circulate in humans. Furthermore, the discovery of endogenous viral elements (EVEs) in the genomes of mammals, birds and other eukaryotes shows that viruses similar to contemporary virus species existed tens of millions of years ago.
In this Opinion article, we describe a niche-filling model of virus evolution that aims to reconcile these conflicting aspects of virus evolutionary histories over different evolutionary timescales — a framework in which the host represents the primary driver of the longer-term evolution of viruses.
Rapid virus sequence change
A large body of literature documents remarkably high nucleotide substitution rates in virus genomes, up to 0.1–1.0% per year in HIV-1 and a wide range of mammalian RNA viruses1,2,3 and small DNA viruses such as parvoviruses4,5,6,7. Virus sequence change occurs so quickly that phylogenetic trees of their genes are often temporally structured — viruses from older samples show systematically less divergence from the most recent common ancestor (MRCA) than those collected more recently. As an early example of this phenomenon, the distance from the tree root of sequences of enterovirus 70 isolates collected through the 1970s and 1980s showed a linear relationship with collection date; the calculated nucleotide substitution rate of 5 × 10−3 substitutions per site per year (SSY) allowed the start of the outbreak to be dated to 1967 (ref.8). The availability of virus samples collected over relatively wide date ranges, often stretching back to the 1950s or 1960s, has enabled more sophisticated Bayesian methods (for example, Bayesian evolutionary analysis by sampling trees (BEAST)9,10) to estimate dates of origins and substitution rates for a wide range of viruses associated with recent outbreaks. Furthermore, these evolutionary timescales have often been linked to historical events. Among many examples, a nucleotide substitution rate of 5 × 10−4 SSY in the hepatitis C virus (HCV) genome was used to calculate dates of emergence of various genotype 2 subtypes to 1470, a finding that might explain the association of genotype 2 infections in areas where the slave trade operated several hundred years ago11. On the basis of its substitution rate and geography of currently circulating genotypes, hepatitis B virus (HBV) was proposed to have originated in native South American populations and spread into Europe and elsewhere after contact with the Europeans in the 1500s12. Likewise, the origin of the four genotypes of hepatitis E virus (HEV) infecting humans was estimated to be between 536 and 1,344 years ago13, and this was suggested to be associated with the spread of pig farming; HEV strains of genotypes 3 and 4 in Japan apparently originated from the 1900s, when pigs were first imported from Yorkshire in England14.
Virus sequence change is often dominated by synonymous substitutions in coding regions that leave sequences of the encoded proteins unaltered. Fixation of these changes may be facilitated by repeated transmission bottlenecks that reduce effective population sizes that occur as the virus transmits between hosts15. Sequence change may be augmented by adaptive changes. For example, influenza A virus shows rapid, antibody-driven antigenic drift of the haemagglutinin gene that enables it to escape from neutralizing antibodies16. Both HIV-1 and HCV fix several amino acid changes in immunodominant T cell epitopes during primary infection that prevent antigen presentation to cytotoxic T cells, contributing to their ability to replicate and transmit17,18.
These observations contribute to a general perception of the ephemeral nature of RNA viruses and a broader idea that viruses are rapidly evolving entities with perhaps frequent recent origins5,19,20. This appears particularly applicable to those emerging viruses responsible for the numerous recent and often severe disease outbreaks that have afflicted humans, animals and plants. This impression is reinforced by what we know about the origins of particular viruses; the emergence of HIV-1 is indeed documented to be recent, originating from multiple cross-species transmissions of a chimpanzee lentivirus into humans in the late 19th century in Gabon and the Congo21. This was followed by various genomic changes associated with human adaptation and increases in human-to-human transmissibility in the subsequent decades that enabled its spread out of Africa in the 1970s to become a global pandemic22,23. Recent outbreaks of influenza A virus, Nipah virus, Hendra virus, Middle East respiratory syndrome coronavirus and severe acute respiratory syndrome-related coronavirus similarly have zoonotic origins with the associated public health concern of host adaptation and the permanent establishment of these viruses in human populations24.
A darkening cloud of uncertainty
Methods that predict the temporal dynamics and phylogeography of recent virus emergence have been remarkably effective in reconstructing recent virus evolutionary histories. Although extrapolation of these substitution rates to longer periods seemingly provides the means to reconstruct much deeper evolutionary histories of viruses, a series of recent developments challenges the applicability of such methods to viruses and, more disturbingly, the widely accepted concepts of the evolutionary timescales of viruses.
An early and convincing example of potential problems with extrapolating substitution rates was found in estimates of the dates of divergence of simian immunodeficiency virus (SIV) strains that were the source of HIV-1 and HIV-2 infections in humans and of SIV variants infecting various monkey species21,25. Relatively rapid substitution rates, such as the 1.38 × 10−3 SSY (range 1.03–1.73 × 10−3) calculated for SIV strains infecting African green monkeys25, predicted time spans of hundreds of years for these divergence events and strengthened concepts of their relatively recent origins. However, a subsequent study of SIV strains infecting isolated populations of Old World monkeys on the island of Bioko, Equatorial Guinea, 32 km off the coast of Africa, was entirely incompatible with this recent origin hypothesis26. Although post-glacial sea level rises separated the island from the African landmass over 10,000 years ago, SIV strains were found to be minimally divergent from those infecting the same species in mainland Africa monkey populations. These observations lowered the minimum substitution rates of each of the SIV strains by over two orders of magnitude and, extrapolated back, predicted an MRCA for SIVs infecting different host species to around 80,000 years before the present (bp).
This isolated (literally) geological separation event provided a single opportunity to look at longer timescales for virus evolution. However, further systematic investigation has been hampered by the general unavailability of suitably stored (that is, frozen) samples dating back to much before the 1960s or 1970s from which viruses can be reliably recovered. Without the opportunity to investigate long-term substitution rates, the paradigm of RNA viruses being highly mutable emerged and has dominated much of the thinking about their evolution over many decades. Many have noted the depiction of what looks like poliomyelitis in a man on an Ancient Egyptian stele that dates to the 18th dynasty (reviewed with other possible depictions in Ancient Egypt in ref.27), but could poliovirus have existed in the 14th century bc? By conventional extrapolation, the emergence of the Enterovirus C species (to which poliovirus belongs) would be dated to only a few hundred years ago28 and not >3,000 years ago.
Two recent developments have provided the means to look further back into virus evolutionary histories. These challenge current thoughts about virus nucleotide substitution rates and the time depths for their evolution.
Findings from ancient DNA and archaeovirology
DNA degrades after the death of the host, but it can be effectively sequenced by next-generation sequencing methods. These newly developed methods have allowed the genomes of ancient human populations to be sequenced and have enabled direct analyses of genetic relationships between contemporary humans, Neanderthals and Denisovans and other archaic human population groups over the past hundred thousand years29,30,31. aDNA-based studies have also contributed to investigations of the longer-term evolution of viruses over historical timescales, including the analysis of parvovirus B19 (B19V) in human remains dating from the Second World War in Russia32, the pandemic 1918 influenza A virus H1N1 strain from Alaskan permafrost33 and HBV and smallpox in mummified material from the 1600s34,35. The timescales over which aDNA sequences can be recovered have now been extended by three recent reports of the detection of viruses in human samples dating back to the early Neolithic (5000 bc)36,37,38.
Two recent studies report the detection of HBV in several individuals in European and Central Asian populations as early as the Bronze Age and Neolithic (2500–3000 bc36,37). Viruses circulating in these prehistoric times in many cases matched currently circulating HBV genotypes (genotypes A, B and D) and were only 1.3–3.0% divergent from modern strains. This indicates a long-term substitution rate ranging from 8.04 × 10−6 SSY to 1.51 × 10−5 SSY, which is around 100-fold lower than that measured in contemporary samples (7.72 × 10−4 SSY39). Similar samples also provided evidence for the circulation of B19V in humans from Central Asia 5000 bc and in Vikings from Sweden ad 100038. These strains closely matched contemporary genotypes (type 1 and type 2), and a similarly revised lower substitution rate estimate was observed. Whereas an early study of sequence change in B19V (ref.7) predicted a time of origin of current genotype 1 strains to the 1960s or 1970s, the aDNA study indicated that this genotype was actually alive and kicking in Eurasia in the early Neolithic era, nearly 7,000 years ago. Further analyses of progressively older aDNA sequence libraries will undoubtedly reveal more insights into the pace of virus evolution for ever-widening collections of human, animal and plant viruses.
Findings from endogenous viral elements and paleovirology
A second and, again, entirely unanticipated opportunity to study virus evolution over even longer periods was provided by the discovery that copies of DNA and RNA viruses can become integrated in the genomes of animals and plants40,41,42,43,44 (Box 1 and Supplementary Fig. 1). Once endogenized, EVEs are genetically stable and preserve information about the circulation of ancient viruses that is impossible to infer from examination of contemporary virus populations. For example, lentiviruses were originally considered as a recently emerged group of viruses on the basis of the very recent origins of HIV-1 itself and measured substitution rates that place the origins of lentiviruses to a few thousand years ago21. However, endogenous lentiviruses in rabbits45, ferrets, Madagascan lemurs and colugos demonstrate the circulation of lentiviruses over almost the entire time span of mammalian evolution46,47,48. In addition to retroviruses, other RNA and DNA viruses have also adventitiously integrated into host germ lines and created records of ancient infections. On the basis of their distribution in descendant species, filoviruses44,49, parvoviruses, circoviruses and bornaviruses must have all circulated over long periods during mammalian evolution44. In addition, the detection of reptilian hepadnaviruses provides evidence for the circulation of these viruses in the early Mesozoic, >200 million years (Myr) ago, long before the radiation of mammals50.
The presence of EVEs in contemporary host genomes provides irrefutable evidence that viruses recognizably similar to contemporary strains have been continuously infecting their hosts over timescales spanning tens of millions of years.
Virus–host co-evolution
Predictions on the longevity of virus lineages from the EVE fossil record are further supported by observations of the apparent co-speciation of viruses and hosts51; these observations can inform predictions about the even earlier origin of specific viral groups. For example, the phylogeny of spumaviruses closely follows that of their mammalian, amphibian and piscine hosts, consistent with virus–host co-speciation over 450 Myr52,53. The proposed co-evolution of papillomaviruses with their hosts suggests their similarly ancient origins of 400–600 Myr54. Increasingly divergent homologues of HBV have been observed as EVEs in birds and reptiles50, and exogenous hepadna-like viruses have recently been found in fish genomic libraries55. The authors of the latter study propose a co-evolutionary scenario in which the ancestor of currently extant HBV-like viruses may have existed >400 Myr. In a similar but even more extreme example, homologues of polyomaviruses have been detected in DNA libraries of vertebrates and scorpions and spiders56, implying a Precambrian origin before the common ancestor of deuterostomes and protostomes ~650 Myr.
In the following sections, we aim to clarify how the remarkable similarity of ancient viruses discovered through archaeovirology and paleovirology to contemporary sequences can be explained given the extraordinary rates of evolutionary change that viruses can undergo.
Rates of viral evolution
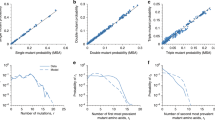
When viral evolution is measured over short timescales, rapid rates of sequence change are typically observed. However, over longer timescales, viral evolutionary rates are several orders of magnitude slower, approaching those of their hosts. Rather than a simple dichotomy between short and long timescales, viral evolutionary rates appear to decrease continuously with the timescale of measurement57, with a decay rate that is strikingly consistent with a power law relationship between substitution rate and observational period53 (Fig. 1; data sources are listed in Supplementary information). Over the longest timescales (100 million to 1 billion years), substitution rates for DNA and RNA viruses of any configuration were remarkably similar: rates of 1–5 × 10−9 SSY; these in turn closely match the 2.2 × 10−9 SSY mean substitution rate calculated for mammalian genes58. At the other end of the scale, short-term substitution rates varied by virus group, with slower rates for double-stranded DNA (dsDNA) viruses (4 × 10−4 SSY) than RNA viruses (8 × 10−3 SSY for those with positive-strand RNA genomes), with a degree of virus lineage-specific variability in short-term rates within each Baltimore group (discussed in ref.57). However, for each Baltimore group, rate decay over time was comparable. Remarkably, the recently obtained substitution rates from aDNA studies superimpose directly upon the regression line inferred from other methods (Fig. 1; blue dots).
Plots of substitution rates of DNA and RNA viruses calculated over different time periods using different methods are shown. These include Bayesian evolutionary reconstructions and rates inferred from instances of virus–host co-evolution (see the figure key). Data used in the figure are based on a previous analysis of published virus substitution rates with different genomic configurations57 and expanded with more recent published data (listed in full in Supplementary information). Three groups are depicted: double-stranded DNA (dsDNA) viruses in Baltimore group I (part a), single-stranded DNA (ssDNA) viruses in Baltimore group II (part b) and reverse transcribing (RT) viruses in Baltimore groups VI and VII57 (part c). These groups showed a remarkably similar relationship between substitution rate (y axis) and observation times over which substitution rates were calculated (plotted on a log-transformed scale on the x axis) despite their intrinsic differences in replication error rates and evolutionary histories. The regression line is based on substitution rates calculated from co-evolution and phylogeny methods. Rates inferred from very ancient co-evolutionary scenarios among RT viruses show a potential flattening of substitution rates as they approach those of host genes (mean value 2.2 × 10−9 substitutions per site per year (SSY)58). Evolutionary rates estimated from ancient DNA (aDNA) sequences of variola virus34, hepatitis B virus (HBV)36 and parvovirus B19 (ref.38) (blue circles) superimpose directly onto rates calculated by other methods. Maximum substitution rates (aDNA – maximum rate) for other HBV sequences35,37 were calculated from their divergence to the most closely related contemporary HBV strains (blue diamonds). TBK and LBK are the pottery-derived terms Trichterbecher (funnel beaker) and Linearbandkeramik (linear band ware), respectively, used to describe European Neolithic populations. bp, before the present; SIV, simian immunodeficiency virus.
Several hypotheses have been proposed to account for the time-dependent rate phenomenon (TDRP)41,59, many of which have been developed to account for substitution rate variability in other organisms (reviewed in ref.59). Using inappropriate substitution models frequently leads to underestimations of age through, for example, the effects of saturation60. However, it is unlikely that even the most complex currently available models can accurately capture nuances of viral genome evolution (for example, the effects of gene overlap, epistasis and nucleotide biases) and reconcile these disparities in age estimations. Sequencing errors, now rare in next-generation sequencing data, could also elevate recent rate estimates, but this effect cannot scale over the longer timescales, over which rate variation is observed. Explanations positing changes in biology over time have also been put forward, such as variance in the fidelity of viral polymerases61, but it is difficult to see how such features could explain the wide-ranging observation of the phenomenon across taxa and over time. Perhaps the most widely accepted explanation is that short-term rate measurements capture population-level processes including transient deleterious mutations and transient beneficial but short-sighted adaptations for their current host62,63 that do not survive in the longer term, whereas long-term rates more closely represent the true fixation rate of mutations over macroevolutionary timescales57,64. Although this explanation could account for the TDRP over short timescales, it is not clear whether deleterious mutations persist for long enough to explain the effect over timescales spanning millions of years.
Although these explanations have been of considerable value in accounting for the TDRP in hosts59, none appear to provide an adequate explanatory framework for the >1 million-fold range in virus substitution rates over different observation periods (Fig. 1) and the long-term extreme conservation of virus genomes. These findings beg the question: what prevents viruses with their seemingly unlimited evolutionary potential from forever diversifying? An overarching model that reconciles both the high rates of sequence change over short timescales and what appear to be implausibly early origins for many virus groups at the other extreme is currently lacking. Although the wide-ranging existence of the TDRP across viral groups and timescales provides an observational description of how apparent viral evolutionary rates vary over time57, we lack a biologically realistic functional model that could account for the apparent ubiquity of this phenomenon.
Host-driven virus evolution
As an alternative explanatory model, we developed ideas originating from niche-filling models65,66,67,68 that emphasize the role of host interactions in shaping virus evolution. This approach contrasts with the typically virus-centric accounts of their evolution in the literature and provides the means to account for the remarkably different trajectories of their evolution at different ends of the observational timescale. Including the host in our model does, however, place unfamiliar constraints on the concept of progressive and diversifying virus evolution.
In this model, high error rates and large population sizes achieved on infection of macroscopic hosts provide viruses with extraordinary adaptive abilities that enable them to maximize fitness in whatever host environments they find themselves (Box 2; Fig. 2). As viruses can rapidly evolve to a fitness peak in a given host environment, this may have the paradoxical effect of restricting sequence change rather than accelerating it in any period other than the short term. Infection of the same host over tens or hundreds of years or perhaps even millennia may drive the evolution of each host-adapted virus to evolutionary stasis — an optimized genome that is maximized in those aspects of its fitness that maintain infections in the host population (Fig. 3). This idea is consistent with the model proposed many years ago that close cooperation between RNA virus proteins and host proteins requires their co-evolution and thus limits their divergence69. However, this stasis may extend much further, not just to the amino acid co-variation within virus proteins but also to the preservation of nucleotide sites at synonymous coding positions and non-coding regions that preserve codon choices, RNA secondary structures and replication elements. Once fully adapted to their niche, the intensity of peer competition may create virus genomes with few genuinely phenotypically neutral sites.
The host niche, depicted as a simplified, spatial representation of the host environment that a virus occupies (see Box 2 for an outline of the typical host elements defining a niche), is shown. The range of host factors exploited by the virus and those associated with host response are depicted as pressure points (filled circles) on the virus that restrict divergence in virus regions involved in these cellular interactions. The blue area represents variable extents of sequence space in which sequence change may occur without phenotypic cost (neutral space).
Viruses remain associated and highly adapted to their host, even as the hosts themselves evolve and speciate over long periods (tens of millions or potentially hundreds of millions of years). Viruses continue to infect cells in each host lineage, but they themselves must evolve in concert with their host to retain fitness and host adaptation as the niche they occupy gradually changes. After a prolonged period of co-evolution, viruses acquire very different virus ‘shapes’ and a phylogeny that resembles in part that of their host. Viruses involved in this co-evolutionary process display long-term substitution rates that approach those of their hosts.
Host adaptation
The process of host adaptation generates viruses that are primarily shaped by the constraints of the niche and less by the ancestry of the virus. If we take parvovirus B19V and HBV as examples of viruses showing evidence for long-term presence in their host populations, their genotypes typically show diversity in the 10−15% nucleotide sequence divergence range, which is represented figuratively as the blue area of potential sequence ‘wobble’ in the virus niche (Fig. 2). This pattern of within-species diversity typifies a wide range of other human, veterinary and plant viruses; examples of the former include individual serotypes of alphaviruses, flaviviruses, measles virus, mumps virus, most of the paramyxoviruses and coronaviruses, and so on. This pattern is also the norm for the vast range of virus species infecting arthropods and fungi, and represents the fraction of genome sites not under selection for fitness optimization. Variation at this level represents the majority of what is captured in temporal sampling and may underlie the generally rapid substitution rates reported for RNA and small DNA viruses over short observation periods. However, the sequence space is small and restrictive — changes at those few neutral sites may saturate at much lower divergence levels than evolutionary models typically expect. We might describe this constraint as a cage — not in the sense of the limited genome size of RNA viruses70 but reflecting those host-imposed constraints on virus sequence change that create the appearance of much less sequence divergence and hence temporal depth than is actually present.
Over much longer periods, virus genome sequence change driven by host change resembles niche-filling models developed for phenotypic trait evolution in cellular organisms67,68; traits evolve adaptively to fit the niche in which a viral species finds itself rather than, for example, via a random-walk model in which traits evolve continuously and progressively over time and lead to clock-like sequence change. The niche is defined by the host organism that the virus infects, the viral sequence defines the phenotype, and changes are primarily adaptive. Short-term substitution rates simply reflect a virus exploring the limits of its cage at rates linked to their error rates and demography; longer-term diversification of RNA and DNA viruses calculated from aDNA and EVE data (Fig. 1) reflects how viruses adapt as the niche shape changes (Fig. 3). These changes ultimately drive the long-term evolution of viruses and explain why their nucleotide substitution rates ultimately approach those of their hosts.
Host jumps
The model equates virus jumps with the occupancy of a new niche and hence a rapid adaptation of trait values to fit this niche (Fig. 4). Host jumps are associated with periods of accelerated sequence change as the virus remodels and regains fitness in an altered environment, very much as conceptualized in bacterial evolution71. Host adaptation after cross-species transmission is associated with rapid amino acid sequence changes of viral genes, typically those associated with receptor interactions and the evasion of innate immunity72,73,74,75,76 but often pervasive throughout the entire virus genome77. Larger-scale gene modifications, such as the repurposing of the HIV-1 accessory protein Vpu to antagonize the cellular antiviral protein tetherin was a key adaptive change that enhanced the replication ability of HIV-1 in humans following its zoonotic transfer from chimpanzees22. The diversification of HIV-1 populations in the 100 or more years since its zoonotic introduction might indeed be interpreted as an ongoing process of fitness optimization. The gradual attenuation of disease severity in HIV-1 infections78 perhaps anticipates a time when HIV-1 diversity is substantially lessened following niche adaptation and the evolution of fitness-optimized, less pathogenic and fully host-adapted HIV-1 strains. HIV-1 population structures and diversity may ultimately match the endemic and tolerated SIV strains that have infected and adapted to many Old World monkey species over much longer periods.
A virus adapted to host A may be able to infect an alternative host (host B), but it may be initially poorly adapted to any available niches. Rapid fixation of adaptive changes improves virus fitness associated with sequence diversification. Fitness competition over a relatively short period of adaptive evolution leads to the emergence of a highly adapted virus strain that is genetically distinct from the founder virus. The red crosses label lineages that have become extinct over the period of virus–host adaptation.
In vertebrates, further adaptive change is driven by their highly polymorphic adaptive immune system. The heterogeneity of the major histocompatibility complex (MHC) molecules between individual hosts defines virus epitope recognition and hence the adaptive changes required to avoid antibody or T cell recognition17,18. Immediately after infection, immune escape of viruses in different individuals may drive rapid antigenic diversification. However, the sequential transit of a virus through dozens or many hundreds of individuals may lead to a static cycle of adaptation on infection and reversion on transmission through different MHC repertoires. At the population level, there may be no net sequence change, an interesting variant of the Red Queen hypothesis79,80. This larger adaptive space (but still a cage) feeds into a complex dynamic of population susceptibility, transmission rates, neutralization escape and changes in receptor use that perpetuates infections in hosts with adaptive immunity. The elaborate serotype and antigenic shift and/or drift population structures of mammalian viruses in particular may be its direct consequence.
Conclusions
In this Opinion article, we present a model of virus sequence change that links substitution rates to those of their long-term hosts, providing an alternative paradigm for understanding virus evolution and adaptation and the associated TDRP. Although it is known that viruses evolve under constraints and adapt to hosts on transmission, the perspective we offer casts viruses and their genetic relationships to each other as being primarily conditioned by hosts they infect. Their own genetic history that is emphasized so much in virus-centric accounts of their evolution over short periods is quite subservient to the shaping forces of host-driven evolution. Similarly, although existing accounts of virus sequence change are so much focused on their seemingly unlimited evolutionary potential and adaptability, the range of viruses that are able to successfully infect and maintain transmission in their hosts appears limited and is more a function of the host niches a virus can exploit65. For example, the wide range of viruses that infect humans possess specific tissue tropisms, pathologies and transmission routes. However, homologues of these viruses in other mammalian species typically reproduce very closely, and appear restricted by, these same virus–host interactions. As further evidence of host-induced constraints, virus replication ability, transmissibility and successful establishment of zoonoses are predicated, at least in part, on the degree of relatedness of the hosts involved in the host jump81,82,83,84. Host relatedness indeed underpins the distribution and pathogenicity of lentiviruses infecting primates and humans85,86. If viruses were genuinely able to adapt and innovate in any host environment, these regularities and apparent niche restrictions across viruses infecting different hosts should not occur.
Although this moulding process equates ultimate virus evolutionary rates to those of their hosts, the niche perspective is also fully consistent with the hypothesis of neutral evolution of viruses over the much shorter periods of virus evolution observed in contemporary virus samples (as discussed in ref.87). Indeed, more than any other factor, the idea that host-adapted viruses are exploring space around a small cage of tolerated substitutions accounts best for the absurdly different short-term and long-term substitution rates they display over differing evolutionary timescales. That small cage and the consequent isolation of virus populations from each other may frequently underpin what are classified as virus species in virus taxonomy88,89, which we may now regard as constrained, separate virus populations with often highly demarcated host ranges. The model of host-driven virus evolution thus places viruses as long-term residents of the hosts they infect, perhaps over millions of years or longer, a concept that accords with the general host specificity that virus species display. The majority of their differences from each other are driven by their host adaptation; niche-filling models accord with the growing evidence of the role of selection and adaptation as the driving forces behind longer-term evolution and speciation elsewhere in biology90.
There seems to be a beautiful paradox in virus evolution — the same remarkable ability of viruses to rapidly adapt to new hosts and escape from innate and adaptive immune responses may also help to create the evolutionary stasis of viruses in long-term host relationships. It is the viruses in their niches that are conservative, and it is their hosts that force them to change.
References
Jenkins, G. M., Rambaut, A., Pybus, O. G. & Holmes, E. C. Rates of molecular evolution in RNA viruses: a quantitative phylogenetic analysis. J. Mol. Evol. 54, 156–165 (2002).
Lemey, P., Rambaut, A. & Pybus, O. G. HIV evolutionary dynamics within and among hosts. AIDS Rev. 8, 125–140 (2006).
Sanjuan, R. From molecular genetics to phylodynamics: evolutionary relevance of mutation rates across viruses. PLOS Pathog. 8, e1002685 (2012).
Hanada, K., Suzuki, Y. & Gojobori, T. A large variation in the rates of synonymous substitution for RNA viruses and its relationship to a diversity of viral infection and transmission modes. Mol. Biol. Evol. 21, 1074–1080 (2004).
Duffy, S., Shackelton, L. A. & Holmes, E. C. Rates of evolutionary change in viruses: patterns and determinants. Nat. Rev. Genet. 9, 267–276 (2008).
Shackelton, L. A., Parrish, C. R., Truyen, U. & Holmes, E. C. High rate of viral evolution associated with the emergence of carnivore parvovirus. Proc. Natl Acad. Sci. USA 102, 379–384 (2005).
Shackelton, L. A. & Holmes, E. C. Phylogenetic evidence for the rapid evolution of human B19 erythrovirus. J. Virol. 80, 3666–3669 (2006).
Takeda, N., Tanimura, M. & Miyamura, K. Molecular evolution of the major capsid protein VP1 of enterovirus 70. J. Virol. 68, 854–862 (1994).
Drummond, A. J., Suchard, M. A., Xie, D. & Rambaut, A. Bayesian phylogenetics with BEAUti and the BEAST 1.7. Mol. Biol. Evol. 29, 1969–1973 (2012).
Drummond, A. J. & Rambaut, A. BEAST: Bayesian evolutionary analysis by sampling trees. BMC. Evol. Biol. 7, 214 (2007).
Markov, P. V. et al. Phylogeography and molecular epidemiology of hepatitis C virus genotype 2 in Africa. J. Gen. Virol. 90, 2086–2096 (2009).
Bollyky, P. L., Rambaut, A., Grassly, N., Carman, W. F. & Holmes, E. C. Hepatitis B virus has a new world evolutionary origin. Hepatology 26, 765–765 (1997).
Purdy, M. A. & Khudyakov, Y. E. Evolutionary history and population dynamics of hepatitis E virus. PLOS ONE 5, e14376 (2010).
Tanaka, Y. et al. Molecular tracing of Japan-indigenous hepatitis E viruses. J. Gen. Virol. 87, 949–954 (2006).
McCrone, J. T. & Lauring, A. S. Genetic bottlenecks in intraspecies virus transmission. Curr. Opin. Virol. 28, 20–25 (2018).
Fitch, W. M., Leiter, J. M., Li, X. Q. & Palese, P. Positive Darwinian evolution in human influenza A viruses. Proc. Natl Acad. Sci. USA 88, 4270–4274 (1991).
Leslie, A. J. et al. HIV evolution: CTL escape mutation and reversion after transmission. Nat. Med. 10, 282–289 (2004).
Timm, J. et al. CD8 epitope escape and reversion in acute HCV infection. J. Exp. Med. 200, 1593–1604 (2004).
Holland, J. J. Transitions in understanding of RNA viruses: a historical perspective. Curr. Top. Microbiol. Immunol. 299, 371–401 (2006).
Pybus, O. G. & Rambaut, A. Evolutionary analysis of the dynamics of viral infectious disease. Nat. Rev. Genet. 10, 540–550 (2009).
Sharp, P. M. et al. Origins and evolution of AIDS viruses: estimating the time-scale. Biochem. Soc. Trans. 28, 275–282 (2000).
Sauter, D. et al. Tetherin-driven adaptation of Vpu and Nef function and the evolution of pandemic and nonpandemic HIV-1 strains. Cell Host Microbe 6, 409–421 (2009).
Wain, L. V. et al. Adaptation of HIV-1 to its human host. Mol. Biol. Evol. 24, 1853–1860 (2007).
Woolhouse, M. & Gaunt, E. Ecological origins of novel human pathogens. Crit. Rev. Microbiol. 33, 231–242 (2007).
Wertheim, J. O. & Worobey, M. Dating the age of the SIV lineages that gave rise to HIV-1 and HIV-2. PLOS Comput. Biol. 5, e1000377 (2009).
Worobey, M. et al. Island biogeography reveals the deep history of SIV. Science 329, 1487 (2010).
Galassi, F. M., Habicht, M. E. & Ruhli, F. J. Poliomyelitis in Ancient Egypt? Neurol. Sci. 38, 375 (2017).
Lukashev, A. N. & Vakulenko, Y. A. Molecular evolution of types in non-polio enteroviruses. J. Gen. Virol. 98, 2968–2981 (2017).
Olalde, I. et al. The Beaker phenomenon and the genomic transformation of northwest Europe. Nature 555, 190–196 (2018).
Meyer, M. et al. A high-coverage genome sequence from an archaic Denisovan individual. Science 338, 222–226 (2012).
Noonan, J. P. et al. Sequencing and analysis of Neanderthal genomic DNA. Science 314, 1113–1118 (2006).
Toppinen, M. et al. Bones hold the key to DNA virus history and epidemiology. Sci. Rep. 5, 17226 (2015).
Reid, A. H., Fanning, T. G., Hultin, J. V. & Taubenberger, J. K. Origin and evolution of the 1918 “Spanish” influenza virus hemagglutinin gene. Proc. Natl Acad. Sci. USA 96, 1651–1656 (1999).
Duggan, A. T. et al. 17(th) century variola virus reveals the recent history of smallpox. Curr. Biol. 26, 3407–3412 (2016).
Patterson Ross, Z. et al. The paradox of HBV evolution as revealed from a 16th century mummy. PLOS Pathog. 14, e1006750 (2018).
Muhlemann, B. et al. Ancient hepatitis B viruses from the Bronze Age to the Medieval period. Nature 557, 418–423 (2018).
Krause-Kyora, B. et al. Neolithic and Medieval virus genomes reveal complex evolution of Hepatitis B. eLife 7, e36666 (2018).
Muhlemann, B. et al. Ancient human parvovirus B19 in Eurasia reveals its long-term association with humans. Proc. Natl Acad. Sci. USA 115, 7557–7562 (2018).
Zhou, Y. & Holmes, E. C. Bayesian estimates of the evolutionary rate and age of hepatitis B virus. J. Mol. Evol. 65, 197–205 (2007).
Kryukov, K., Ueda, M. T., Imanishi, T. & Nakagawa, S. Systematic survey of non-retroviral virus-like elements in eukaryotic genomes. Virus Res. https://doi.org/10.1016/j.virusres.2018.02.002 (2018).
Aiewsakun, P. & Katzourakis, A. Endogenous viruses: connecting recent and ancient viral evolution. Virology 479–480, 26–37 (2015).
Belyi, V. A., Levine, A. J. & Skalka, A. M. Sequences from ancestral single-stranded DNA viruses in vertebrate genomes: the parvoviridae and circoviridae are more than 40 to 50 million years old. J. Virol. 84, 12458–12462 (2010).
Horie, M. et al. Endogenous non-retroviral RNA virus elements in mammalian genomes. Nature 463, 84–87 (2010).
Katzourakis, A. & Gifford, R. J. Endogenous viral elements in animal genomes. PLOS Genet. 6, e1001191 (2010).
Katzourakis, A., Tristem, M., Pybus, O. G. & Gifford, R. J. Discovery and analysis of the first endogenous lentivirus. Proc. Natl Acad. Sci. USA 104, 6261–6265 (2007).
Han, G. Z. & Worobey, M. Endogenous lentiviral elements in the weasel family (Mustelidae). Mol. Biol. Evol. 29, 2905–2908 (2012).
Gifford, R. J. et al. A transitional endogenous lentivirus from the genome of a basal primate and implications for lentivirus evolution. Proc. Natl Acad. Sci. USA 105, 20362–20367 (2008).
Hron, T., Farkasova, H., Padhi, A., Paces, J. & Elleder, D. Life history of the oldest lentivirus: characterization of ELVgv integrations in the dermopteran genome. Mol. Biol. Evol. 33, 2659–2669 (2016).
Taylor, D. J., Leach, R. W. & Bruenn, J. Filoviruses are ancient and integrated into mammalian genomes. BMC Evol. Biol. 10, 193 (2010).
Suh, A. et al. Early mesozoic coexistence of amniotes and hepadnaviridae. PLOS Genet. 10, e1004559 (2014).
Sharp, P. M. & Simmonds, P. Evaluating the evidence for virus/host co-evolution. Curr. Opin. Virol. 1, 436–441 (2011).
Katzourakis, A., Gifford, R. J., Tristem, M., Gilbert, M. T. & Pybus, O. G. Macroevolution of complex retroviruses. Science 325, 1512 (2009).
Aiewsakun, P. & Katzourakis, A. Marine origin of retroviruses in the early Palaeozoic era. Nat. Commun. 8, 13954 (2017).
Van Doorslaer, K. et al. Unique genome organization of non-mammalian papillomaviruses provides insights into the evolution of viral early proteins. Virus Evol. 3, vex027 (2017).
Lauber, C. et al. Deciphering the origin and evolution of hepatitis B viruses by means of a family of non-enveloped fish viruses. Cell Host Microbe 22, 387–399 (2017).
Buck, C. B. et al. The ancient evolutionary history of polyomaviruses. PLOS Pathog. 12, e1005574 (2016).
Aiewsakun, P. & Katzourakis, A. Time-dependent rate phenomenon in viruses. J. Virol. 90, 7184–7195 (2016).
Kumar, S. & Subramanian, S. Mutation rates in mammalian genomes. Proc. Natl Acad. Sci. USA 99, 803–808 (2002).
Ho, S. Y. et al. Time-dependent rates of molecular evolution. Mol. Ecol. 20, 3087–3101 (2011).
Wertheim, J. O. & Kosakovsky Pond, S. L. Purifying selection can obscure the ancient age of viral lineages. Mol. Biol. Evol. 28, 3355–3365 (2011).
Gilbert, C. & Feschotte, C. Genomic fossils calibrate the long-term evolution of hepadnaviruses. PLOS Biol. 8, e1000495 (2010).
Lythgoe, K. A., Gardner, A., Pybus, O. G. & Grove, J. Short-sighted virus evolution and a germline hypothesis for chronic viral infections. Trends Microbiol. 25, 336–348 (2017).
Lythgoe, K. A., Pellis, L. & Fraser, C. Is HIV short-sighted? Insights from a multistrain nested model. Evolution 67, 2769–2782 (2013).
Duchene, S., Holmes, E. C. & Ho, S. Y. Analyses of evolutionary dynamics in viruses are hindered by a time-dependent bias in rate estimates. Proc. Biol. Sci. 281, 20140732 (2014).
Cooper, N., Jetz, W. & Freckleton, R. P. Phylogenetic comparative approaches for studying niche conservatism. J. Evol. Biol. 23, 2529–2539 (2010).
Holt, R. D. Bringing the Hutchinsonian niche into the 21st century: ecological and evolutionary perspectives. Proc. Natl Acad. Sci. USA 106 (Suppl. 2), 19659–19665 (2009).
Price, T. Correlated evolution and independent contrasts. Philos. Trans. R. Soc. Lond. B. Biol. Sci. 352, 519–529 (1997).
Harvey, P. H. & Rambaut, A. Comparative analyses for adaptive radiations. Philos. Trans. R. Soc. Lond. B. Biol. Sci. 355, 1599–1605 (2000).
Koonin, E. V. & Gorbalenya, A. E. Evolution of RNA genomes: does the high mutation rate necessitate high rate of evolution of viral proteins? J. Mol. Evol. 28, 524–527 (1989).
Belshaw, R., Gardner, A., Rambaut, A. & Pybus, O. G. Pacing a small cage: mutation and RNA viruses. Trends Ecol. Evol. 23, 188–193 (2008).
Sheppard, S. K., Guttmann, D. S. & Fitzgerald, J. R. Population genomics of bacterial host adaptation. Nat. Rev. Genet. 19, 549–565 (2018).
Sawyer, S. L., Emerman, M. & Malik, H. S. Ancient adaptive evolution of the primate antiviral DNA-editing enzyme APOBEC3G. PLOS Biol. 2, E275 (2004).
Taubenberger, J. K. & Kash, J. C. Influenza virus evolution, host adaptation, and pandemic formation. Cell Host Microbe 7, 440–451 (2010).
Li, W. et al. Receptor and viral determinants of SARS-coronavirus adaptation to human ACE2. EMBO J. 24, 1634–1643 (2005).
Urbanowicz, R. A. et al. Human adaptation of ebola virus during the West African outbreak. Cell 167, 1079–1087 (2016).
Allison, A. B. et al. Host-specific parvovirus evolution in nature is recapitulated by in vitro adaptation to different carnivore species. PLOS Pathog. 10, e1004475 (2014).
Bhatt, S. et al. The evolutionary dynamics of influenza A virus adaptation to mammalian hosts. Philos. Trans. R. Soc. Lond. B. Biol. Sci. 368, 20120382 (2013).
Blanquart, F. et al. A transmission-virulence evolutionary trade-off explains attenuation of HIV-1 in Uganda. eLife 5, e20492 (2016).
Clarke, D. K. et al. The red queen reigns in the kingdom of RNA viruses. Proc. Natl Acad. Sci. USA 91, 4821–4824 (1994).
Tedder, R. S., Bissett, S. L., Myers, R. & Ijaz, S. The ‘Red Queen’ dilemma—running to stay in the same place: reflections on the evolutionary vector of HBV in humans. Antivir. Ther. 18, 489–496 (2013).
Faria, N. R., Suchard, M. A., Rambaut, A., Streicker, D. G. & Lemey, P. Simultaneously reconstructing viral cross-species transmission history and identifying the underlying constraints. Philos. Trans. R. Soc. Lond. B. Biol. Sci. 368, 20120196 (2013).
Streicker, D. G. et al. Host phylogeny constrains cross-species emergence and establishment of rabies virus in bats. Science 329, 676–679 (2010).
Longdon, B., Hadfield, J. D., Webster, C. L., Obbard, D. J. & Jiggins, F. M. Host phylogeny determines viral persistence and replication in novel hosts. PLOS Pathog. 7, e1002260 (2011).
Davies, T. J. & Pedersen, A. B. Phylogeny and geography predict pathogen community similarity in wild primates and humans. Proc. Biol. Sci. 275, 1695–1701 (2008).
Nyamweya, S. et al. Comparing HIV-1 and HIV-2 infection: lessons for viral immunopathogenesis. Rev. Med. Virol. 23, 221–240 (2013).
Charleston, M. A. & Robertson, D. L. Preferential host switching by primate lentiviruses can account for phylogenetic similarity with the primate phylogeny. Syst. Biol. 51, 528–535 (2002).
Frost, S. D. W., Magalis, B. R. & Kosakovsky Pond, S. L. Neutral theory and rapidly evolving viral pathogens. Mol. Biol. Evol. 35, 1348–1354 (2018).
Van Regenmortel, M. H. Virus species and virus identification: past and current controversies. Infect. Genet. Evol. 7, 133–144 (2007).
Simmonds, P. A clash of ideas - the varying uses of the ‘species’ term in virology and their utility for classifying viruses in metagenomic datasets. J. Gen. Virol. 99, 277–287 (2018).
Kern, A. D. & Hahn, M. W. The neutral theory in light of natural selection. Mol. Biol. Evol. 35, 1366–1371 (2018).
Keckesova, Z., Ylinen, L. M., Towers, G. J., Gifford, R. J. & Katzourakis, A. Identification of a RELIK orthologue in the European hare (Lepus europaeus) reveals a minimum age of 12 million years for the lagomorph lentiviruses. Virology 384, 7–11 (2009).
Daugherty, M. D., Young, J. M., Kerns, J. A. & Malik, H. S. Rapid evolution of PARP genes suggests a broad role for ADP-ribosylation in host-virus conflicts. PLOS Genet. 10, e1004403 (2014).
van der Lee, R., Wiel, L., van Dam, T. J. P. & Huynen, M. A. Genome-scale detection of positive selection in nine primates predicts human-virus evolutionary conflicts. Nucleic Acids Res. 45, 10634–10648 (2017).
Sawyer, S. L., Emerman, M. & Malik, H. S. Discordant evolution of the adjacent antiretroviral genes TRIM22 and TRIM5 in mammals. PLOS Pathog. 3, e197 (2007).
Munk, C., Willemsen, A. & Bravo, I. G. An ancient history of gene duplications, fusions and losses in the evolution of APOBEC3 mutators in mammals. BMC Evol. Biol. 12, 71 (2012).
Acknowledgements
The authors thank J. Metcalf, Princeton University, for reviewing and providing helpful comments on the manuscript before submission.
Reviewer information
Nature Reviews Microbiology thanks P. Lemey, A. Vandamme and other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
P.S., P.A. and A.K. researched the data for the article. P.S., P.A. and A.K. substantially contributed to discussion of content. P.S. and A.K. wrote the article. P.S., P.A. and A.K. reviewed and edited the manuscript before submission.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Simmonds, P., Aiewsakun, P. & Katzourakis, A. Prisoners of war — host adaptation and its constraints on virus evolution. Nat Rev Microbiol 17, 321–328 (2019). https://doi.org/10.1038/s41579-018-0120-2
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41579-018-0120-2
This article is cited by
-
Surveillance and characterization of avian-origin H3N2 canine influenza viruses in 2021 in China
One Health Advances (2024)
-
Ultraconserved bacteriophage genome sequence identified in 1300-year-old human palaeofaeces
Nature Communications (2024)
-
Untangling an insect’s virome from its endogenous viral elements
BMC Genomics (2023)
-
Molecular Epidemiology of Begomoviruses Infecting Mungbean from Yellow Mosaic Disease Hotspot Regions of India
Applied Biochemistry and Biotechnology (2023)
-
Metagenomic and Molecular Detection of Novel Fecal Viruses in Free-Ranging Agile Wallabies
EcoHealth (2023)