Abstract
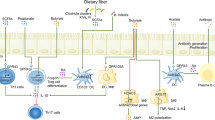
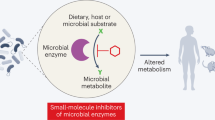
Trillions of microorganisms transit through and reside in the mammalian gastrointestinal tract each day, collectively producing thousands of small molecules and metabolites with local and systemic effects on host physiology. Identifying effector microorganisms that causally affect host phenotype and deciphering the underlying mechanisms have become foci of microbiome research and have begun to enable the development of microbiota-based therapeutics. Two complementary, reductionist approaches have commonly been used: the first starts with an immune phenotype and narrows down the microbiota to identify responsible effector bacteria, while the second starts with bacteria-derived molecules and metabolites and seeks to understand their effects on the host immune system. Together, these strategies provide the basis for the rational design of microbial and metabolite-based therapeutics that target and ameliorate immune deficits in patients.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Wampach, L. et al. Colonization and succession within the human gut microbiome by archaea, bacteria, and microeukaryotes during the first year of life. Front. Microbiol. 8, 738 (2017).
Human Microbiome Project Consortium. Structure, function and diversity of the healthy human microbiome. Nature 486, 207–214 (2012). Shows the enormous heterogeneity in phylogenetic composition of healthy human microbiota and relative stability of metabolic pathways.
Round, J. L. & Palm, N. W. Causal effects of the microbiota on immune-mediated diseases. Sci. Immunol. 3, eaao1603 (2018).
Qin, J. et al. A human gut microbial gene catalogue established by metagenomic sequencing. Nature 464, 59–65 (2010).
Lynch, S. V. & Pedersen, O. The human intestinal microbiome in health and disease. N. Engl. J. Med. 375, 2369–2379 (2016).
Levy, M., Kolodziejczyk, A. A., Thaiss, C. A. & Elinav, E. Dysbiosis and the immune system. Nat. Rev. Immunol. 17, 219–232 (2017).
Olesen, S. W. & Alm, E. J. Dysbiosis is not an answer. Nat. Microbiol. 1, 16228 (2016).
Frank, D. N. et al. Molecular-phylogenetic characterization of microbial community imbalances in human inflammatory bowel diseases. Proc. Natl Acad. Sci. USA 104, 13780–13785 (2007).
Kostic, A. D. et al. Genomic analysis identifies association of Fusobacterium with colorectal carcinoma. Genome Res. 22, 292–298 (2012).
Hsiao, E. Y. et al. Microbiota modulate behavioral and physiological abnormalities associated with neurodevelopmental disorders. Cell 155, 1451–1463 (2013).
Wen, L. et al. Innate immunity and intestinal microbiota in the development of type 1 diabetes. Nature 455, 1109–1113 (2008).
Karlsson, F. H. et al. Symptomatic atherosclerosis is associated with an altered gut metagenome. Nat. Commun. 3, 1245 (2012).
Ley, R. E., Turnbaugh, P. J., Klein, S. & Gordon, J. I. Microbial ecology: human gut microbes associated with obesity. Nature 444, 1022–1023 (2006).
Turnbaugh, P. J. et al. An obesity-associated gut microbiome with increased capacity for energy harvest. Nature 444, 1027–1031 (2006).
Turnbaugh, P. J. & Gordon, J. I. An Invitation to the marriage of metagenomics and metabolomics. Cell 134, 708–713 (2008).
Gilbert, J. A. et al. Microbiome-wide association studies link dynamic microbial consortia to disease. Nature 535, 94–103 (2016).
Blumberg, R. & Powrie, F. Microbiota, disease, and back to health: a metastable journey. Sci. Transl Med. 4, 137rv7 (2012).
Weinstock, G. M. Genomic approaches to studying the human microbiota. Nature 489, 250–256 (2012).
Lozupone, C. A., Stombaugh, J. I., Gordon, J. I., Jansson, J. K. & Knight, R. Diversity, stability and resilience of the human gut microbiota. Nature 489, 220–230 (2012).
Surana, N. K. & Kasper, D. L. Moving beyond microbiome-wide associations to causal microbe identification. Nature 552, 244 (2017).
Fischbach, M. A. Microbiome: focus on causation and mechanism. Cell 174, 785–790 (2018).
Postler, T. S. & Ghosh, S. Understanding the holobiont: how microbial metabolites affect human health and shape the immune system. Cell Metab. 26, 110–130 (2017).
Browne, H. P. et al. Culturing of ‘unculturable’ human microbiota reveals novel taxa and extensive sporulation. Nature 533, 543–546 (2016).
Lagier, J.-C. et al. Culture of previously uncultured members of the human gut microbiota by culturomics. Nat. Microbiol. 1, 16203 (2016).
Lagier, J.-C. et al. Culturing the human microbiota and culturomics. Nat. Rev. Microbiol. 16, 540–550 (2018).
Wikoff, W. R. et al. Metabolomics analysis reveals large effects of gut microflora on mammalian blood metabolites. Proc. Natl Acad. Sci. USA 106, 3698–3703 (2009).
David, L. A. et al. Diet rapidly and reproducibly alters the human gut microbiome. Nature 505, 559–563 (2014).
Cammarota, G., Ianiro, G. & Gasbarrini, A. Fecal microbiota transplantation for the treatment of clostridium difficile infection: a systematic review. J. Clin. Gastroenterol. 48, 693–702 (2014).
van Nood, E. et al. Duodenal infusion of donor feces for recurrent Clostridium difficile. N. Engl. J. Med. 368, 407–415 (2013). This study highlights the striking efficacy of FMT for treating CDI.
Smillie, C. S. et al. Strain tracking reveals the determinants of bacterial engraftment in the human gut following fecal microbiota transplantation. Cell Host Microbe 23, 229–240 (2018).
Li, S. S. et al. Durable coexistence of donor and recipient strains after fecal microbiota transplantation. Science 352, 586–589 (2016).
Bojanova, D. P. & Bordenstein, S. R. Fecal transplants: what is being transferred? PLOS Biol. 14, e1002503 (2016).
Ott, S. J. et al. Efficacy of sterile fecal filtrate transfer for treating patients with Clostridium difficile infection. Gastroenterology 152, 799–811 (2017).
Gerding, D. N. et al. Administration of spores of nontoxigenic Clostridium difficile strain M3 for prevention of recurrent C difficile infection: a randomized clinical trial. JAMA 313, 1719–1727 (2015).
Ng, S. C. et al. Systematic review: the efficacy of herbal therapy in inflammatory bowel disease. Aliment. Pharmacol. Ther. 38, 854–863 (2013).
Rossen, N. G. et al. Findings from a randomized controlled trial of fecal transplantation for patients with ulcerative colitis. Gastroenterology 149, 110–118 (2015).
Moayyedi, P. et al. Fecal microbiota transplantation induces remission in patients with active ulcerative colitis in a randomized controlled trial. Gastroenterology 149, 102–109 (2015).
Paramsothy, S. et al. Multidonor intensive faecal microbiota transplantation for active ulcerative colitis: a randomised placebo-controlled trial. Lancet 389, 1218–1228 (2017).
Kakihana, K. et al. Fecal microbiota transplantation for patients with steroid-resistant acute graft-versus-host disease of the gut. Blood 128, 2083–2088 (2016).
Van Lier, Y. F. et al. Fecal microbiota transplantation as safe and successful therapy for intestinal graft-versus-host disease. Blood 130, 1986 (2017).
Doki, N. et al. Clinical impact of pre-transplant gut microbial diversity on outcomes of allogeneic hematopoietic stem cell transplantation. Ann. Hematol. 96, 1517–1523 (2017).
Vrieze, A. et al. Transfer of intestinal microbiota from lean donors increases insulin sensitivity in individuals with metabolic syndrome. Gastroenterology 143, 913–916 (2012).
Kang, D. W. et al. Microbiota transfer therapy alters gut ecosystem and improves gastrointestinal and autism symptoms: an open-label study. Microbiome 5, 10 (2017).
Khanna, S. et al. A novel microbiome therapeutic increases gut microbial diversity and prevents recurrent Clostridium difficile infection. J. Infect. Dis. 214, 173–181 (2016).
Sherman, R. E., Li, J., Shapley, S., Robb, M. & Woodcock, J. Expediting drug development — the FDA’s new “breakthrough therapy” designation. N. Engl. J. Med. 369, 1877–1880 (2013).
Mullard, A. Leading microbiome-based therapeutic falters in phase II trial. Nat. Rev. Drug Discov. 15, 595 (2016).
Mills, J. P., Rao, K. & Young, V. B. Probiotics for prevention of Clostridium difficile infection. Curr. Opin. Gastroenterol. 34, 3–10 (2018).
Lagier, J.-C., Cadoret, F. & Raoult, D. Critical microbiological view of SER-109. J. Infect. Dis. 215, 161–162 (2017).
Ratner, M. Seres’s pioneering microbiome drug fails mid-stage trial. Nat. Biotechnol. 34, 1004–1005 (2016).
Buffie, C. G. et al. Precision microbiome reconstitution restores bile acid mediated resistance to Clostridium difficile. Nature 517, 205–208 (2015). This study shows that C. scindens protects against CDI via production of secondary bile acids.
Sorg, J. A. & Sonenshein, A. L. Bile salts and glycine as cogerminants for Clostridium difficile spores. J. Bacteriol. 190, 2505–2512 (2008).
Sorg, J. A. & Sonenshein, A. L. Inhibiting the initiation of Clostridium difficile spore germination using analogs of chenodeoxycholic acid, a bile acid. J. Bacteriol. 192, 4983–4990 (2010).
Thanissery, R., Winston, J. A. & Theriot, C. M. Inhibition of spore germination, growth, and toxin activity of clinically relevant C. difficile strains by gut microbiota derived secondary bile acids. Anaerobe 45, 86–100 (2017).
Baktash, A. et al. Mechanistic insights in the success of fecal microbiota transplants for the treatment of Clostridium difficile infections. Front. Microbiol. 9, 1242 (2018).
Theriot, C. M. & Young, V. B. Interactions between the gastrointestinal microbiome and Clostridium difficile. Annu. Rev. Microbiol. 69, 445–461 (2015).
Weingarden, A. R. et al. Ursodeoxycholic acid inhibits clostridium difficile spore germination and vegetative growth, and prevents the recurrence of ileal pouchitis associated with the infection. J. Clin. Gastroenterol. 50, 624–630 (2016).
Battaglioli, E. J. et al. Clostridioides difficile uses amino acids associated with gut microbial dysbiosis in a subset of patients with diarrhea. Sci. Transl Med. 10, eaam7019 (2018).
Ng, K. M. et al. Microbiota-liberated host sugars facilitate post-antibiotic expansion of enteric pathogens. Nature 502, 96–99 (2013).
Caballero, S. et al. Distinct but spatially overlapping intestinal niches for vancomycin-resistant Enterococcus faecium and carbapenem-resistant Klebsiella pneumoniae. PLOS Pathog. 11, e1005132 (2015).
Caballero, S. et al. Cooperating commensals restore colonization resistance to vancomycin-resistant Enterococcus faecium. Cell Host Microbe 21, 592–602 (2017).
Sorbara, M. T. et al. Inhibiting antibiotic-resistant Enterobacteriaceae by microbiota-mediated intracellular acidification. J. Exp. Med. 216, 84–98 (2018).
Seed, P. C. The human mycobiome. Cold Spring Harb. Perspect. Med. 5, a019810 (2014).
Nash, A. K. et al. The gut mycobiome of the Human Microbiome Project healthy cohort. Microbiome 5, 153 (2017).
Kernbauer, E., Ding, Y. & Cadwell, K. An enteric virus can replace the beneficial function of commensal bacteria. Nature 516, 94–98 (2014).
Cadwell, K. The virome in host health and disease. Immunity 42, 805–813 (2015).
Manrique, P. et al. Healthy human gut phageome. Proc. Natl Acad. Sci. USA 113, 10400–10405 (2016).
Chudnovskiy, A. et al. Host-protozoan interactions protect from mucosal infections through activation of the inflammasome. Cell 167, 444–456 (2016).
Lukeš, J., Stensvold, C. R., Jirků-Pomajbíková, K. & Wegener Parfrey, L. Are human intestinal eukaryotes beneficial or commensals? PLOS Pathog. 11, e1005039 (2015).
Ramanan, D. et al. Helminth infection promotes colonization resistance via type 2 immunity. Science 352, 608–612 (2016).
Gause, W. C. & Maizels, R. M. Macrobiota - helminths as active participants and partners of the microbiota in host intestinal homeostasis. Curr. Opin. Microbiol. 32, 14–18 (2016).
Sczesnak, A. et al. The genome of Th17 cell-inducing segmented filamentous bacteria reveals extensive auxotrophy and adaptations to the intestinal environment. Cell Host Microbe 10, 260–272 (2011).
Prakash, T. et al. Complete genome sequences of rat and mouse segmented filamentous bacteria, a potent inducer of Th17 cell differentiation. Cell Host Microbe 10, 273–284 (2011).
Schnupf, P. et al. Growth and host interaction of mouse segmented filamentous bacteria in vitro. Nature 520, 99–103 (2015).
Tannock, G. W., Miller, J. R. & Savage, D. C. Host specificity of filamentous, segmental microorganisms adherent to the small bowel epithelium in mice and rats. Appl. Environ. Microbiol. 47, 441–442 (1984).
Klaasen, H. L. B. M. et al. Intestinal, segmented, filamentous bacteria in a wide range of vertebrate species. Lab. Anim. 27, 141–150 (1993).
Ivanov, I. I. et al. Induction of intestinal Th17 cells by segmented filamentous bacteria. Cell 139, 485–498 (2009). This study identifies the murine commensal SFB as a potent inducer of T H 17 cells.
Gaboriau-Routhiau, V. et al. The key role of segmented filamentous bacteria in the coordinated maturation of gut helper T cell responses. Immunity 31, 677–689 (2009).
Talham, G. L., Jiang, H. Q., Bos, N. A. & Cebra, J. J. Segmented filamentous bacteria are potent stimuli of a physiologically normal state of the murine gut mucosal immune system. Infect. Immun. 67, 1992–2000 (1999).
Lécuyer, E. et al. Segmented filamentous bacterium uses secondary and tertiary lymphoid tissues to induce gut IgA and specific T helper 17 cell responses. Immunity 40, 608–620 (2014).
Umesaki, Y., Okada, Y., Matsumoto, S., Imaoka, A. & Setoyama, H. Segmented filamentous bacteria are indigenous intestinal bacteria that activate intraepithelial lymphocytes and induce MHC class II molecules and fucosyl asialo GM1 glycolipids on the small intestinal epithelial cells in the ex-germ-free mouse. Microbiol. Immunol. 39, 555–562 (1995).
Atarashi, K. et al. Treg induction by a rationally selected mixture of Clostridia strains from the human microbiota. Nature 500, 232–236 (2013). Used a gnotobiotic pipeline to identify a consortium of 17 human commensals that induce T reg cells.
Tanoue, T. et al. A defined commensal consortium elicits CD8 T cells and anti-cancer immunity. Nature 565, 600–605 (2019). Identified a consortium of 11 human commensals that induce IFNγ + CD8 + T cells.
Palm, N. W. et al. Immunoglobulin A coating identifies colitogenic bacteria in inflammatory bowel disease. Cell 158, 1000–1010 (2014).
Kau, A. L. et al. Functional characterization of IgA-targeted bacterial taxa from undernourished Malawian children that produce diet-dependent enteropathy. Sci. Transl Med. 7, 276ra24 (2015). References 83 and 84 described IgA-seq and its use in identifying microorganisms associated with health and disease.
D’Auria, G. et al. Active and secreted IgA-coated bacterial fractions from the human gut reveal an under-represented microbiota core. Sci. Rep. 3, 3515 (2013).
Wilmore, J. R. et al. Commensal microbes induce serum IgA responses that protect against polymicrobial sepsis. Cell Host Microbe 23, 302–311 (2018).
Dodd, D. et al. A gut bacterial pathway metabolizes aromatic amino acids into nine circulating metabolites. Nature 551, 648–652 (2017). This study elucidated the effects of a C. sporogenes metabolic pathway on host physiology.
Guo, C. J. et al. Depletion of microbiome-derived molecules in the host using Clostridium genetics. Preprint at bioRxiv. https://doi.org/10.1101/401489 (2018).
Lim, B., Zimmermann, M., Barry, N. A. & Goodman, A. L. Engineered regulatory systems modulate gene expression of human commensals in the gut. Cell 169, 547–558 (2017).
Plovier, H. et al. A purified membrane protein from Akkermansia muciniphila or the pasteurized bacterium improves metabolism in obese and diabetic mice. Nat. Med. 23, 107–113 (2017).
Atarashi, K. et al. Th17 cell induction by adhesion of microbes to intestinal epithelial cells. Cell 163, 367–380 (2015). Showed that epithelial adhesion by SFB promotes T H 17 cell induction and identified 20 microorganisms from humans that induce T H 17 cells.
Vedanta Biosciences. Pipeline. vadantabio https://www.vedantabio.com/pipeline (2019).
Turnbaugh, P. J. et al. A core gut microbiome in obese and lean twins. Nature 457, 480–484 (2009). Highlights the stability of metabolic pathways across phylogenetically diverse microbiotas.
Lloyd-Price, J. et al. Strains, functions and dynamics in the expanded Human Microbiome Project. Nature 550, 61–66 (2017).
Abubucker, S. et al. Metabolic reconstruction for metagenomic data and its application to the human microbiome. PLOS Comput. Biol. 8, 1002358 (2012).
Shimizu, H., Yisireyili, M., Higashiyama, Y., Nishijima, F. & Niwa, T. Indoxyl sulfate upregulates renal expression of ICAM-1 via production of ROS and activation of NF-κB and p53 in proximal tubular cells. Life Sci. 92, 143–148 (2013).
Shimizu, H. et al. Indoxyl sulfate upregulates renal expression of MCP-1 via production of ROS and activation of NF-κB, p53, ERK, and JNK in proximal tubular cells. Life Sci. 90, 525–530 (2012).
Devlin, A. S. et al. Modulation of a circulating uremic solute via rational genetic manipulation of the gut microbiota. Cell Host Microbe 20, 709–715 (2016).
Williams, B. B. et al. Discovery and characterization of gut microbiota decarboxylases that can produce the neurotransmitter tryptamine. Cell Host Microbe 16, 495–503 (2014).
Bhattarai, Y. et al. Gut microbiota-produced tryptamine activates an epithelial G-protein-coupled receptor to increase colonic secretion. Cell Host Microbe 23, 775–785 (2018).
Bäumler, A. J. & Sperandio, V. Interactions between the microbiota and pathogenic bacteria in the gut. Nature 535, 85–93 (2016).
Spiga, L. et al. An oxidative central metabolism enables Salmonella to utilize microbiota-derived succinate. Cell Host Microbe 22, 291–301 (2017).
Ferreyra, J. A. et al. Gut microbiota-produced succinate promotes C. Difficile infection after antibiotic treatment or motility disturbance. Cell Host Microbe 16, 770–777 (2014).
Curtis, M. M. et al. The gut commensal bacteroides thetaiotaomicron exacerbates enteric infection through modification of the metabolic landscape. Cell Host Microbe 16, 759–769 (2014).
Fukuda, S. et al. Bifidobacteria can protect from enteropathogenic infection through production of acetate. Nature 469, 543–549 (2011).
Kamada, N. et al. Regulated virulence controls the ability of a pathogen to compete with the gut microbiota. Science 336, 1325–1329 (2012). References 58, 105 and 106 show that the microbiota can affect susceptibility to enteropathogenic infections by producing or consuming metabolites.
Knoop, K. A., Miller, M. J. & Newberry, R. D. Transepithelial antigen delivery in the small intestine: different paths, different outcomes. Curr. Opin. Gastroenterol. 29, 112–118 (2013).
Geva-Zatorsky, N. et al. In vivo imaging and tracking of host–microbiota interactions via metabolic labeling of gut anaerobic bacteria. Nat. Med. 21, 1091–1100 (2015).
Hudak, J. E., Alvarez, D., Skelly, A., von Andrian, U. H. & Kasper, D. L. Illuminating vital surface molecules of symbionts in health and disease. Nat. Microbiol. 2, 17099 (2017). References 108 and 109 used a metabolic-labelling platform to tag and track surface molecules on commensal bacteria in vivo.
Mazmanian, S. K., Cui, H. L., Tzianabos, A. O. & Kasper, D. L. An immunomodulatory molecule of symbiotic bacteria directs maturation of the host immune system. Cell 122, 107–118 (2005).
Mazmanian, S. K., Round, J. L. & Kasper, D. L. A microbial symbiosis factor prevents intestinal inflammatory disease. Nature 453, 620–625 (2008). References 110 and 111 showed that the commensal-derived carbohydrate PSA induces T reg cells and IL-10.
Round, J. L. & Mazmanian, S. K. Inducible Foxp3+regulatory T cell development by a commensal bacterium of the intestinal microbiota. Proc. Natl Acad. Sci. USA 11, 79–80 (2010).
Dasgupta, S., Erturk-Hasdemir, D., Ochoa-Reparaz, J., Reinecker, H. C. & Kasper, D. L. Plasmacytoid dendritic cells mediate anti-inflammatory responses to a gut commensal molecule via both innate and adaptive mechanisms. Cell Host Microbe 15, 413–423 (2014).
Round, J. L. et al. The toll-like receptor 2 pathway establishes colonization by a commensal of the human microbiota. Science 332, 974–977 (2011).
Shen, Y. et al. Outer membrane vesicles of a human commensal mediate immune regulation and disease protection. Cell Host Microbe 12, 509–520 (2012).
Chu, H. et al. Gene-microbiota interactions contribute to the pathogenesis of inflammatory bowel disease. Science 352, 1116–1120 (2016).
Verma, R. et al. Cell surface polysaccharides of Bifidobacterium bifidum induce the generation of Foxp3+regulatory T cells. Sci. Immunol. 3, eaat6975 (2018).
Atarashi, K. et al. Induction of colonic regulatory T cells by indigenous Clostridium species. Science 331, 337–341 (2011).
Arpaia, N. et al. Metabolites produced by commensal bacteria promote peripheral regulatory T cell generation. Nature 504, 451–455 (2013).
Furusawa, Y. et al. Commensal microbe-derived butyrate induces the differentiation of colonic regulatory T cells. Nature 504, 446–450 (2013).
Smith, P. M. et al. The microbial metabolites, short-chain fatty acids, regulate colonic Treg cell homeostasis. Science 341, 569–573 (2013). References 119–121 provide mechanistic insight into how commensal-derived SCFAs affect host immunity.
Maslowski, K. M. et al. Regulation of inflammatory responses by gut microbiota and chemoattractant receptor GPR43. Nature 461, 1282–1286 (2009).
Macia, L. et al. Metabolite-sensing receptors GPR43 and GPR109A facilitate dietary fibre-induced gut homeostasis through regulation of the inflammasome. Nat. Commun. 6, 6734 (2015).
Fujiwara, H. et al. Microbial metabolite sensor GPR43 controls severity of experimental GVHD. Nat. Commun. 9, 3674 (2018).
Zhou, L. et al. TGF-Β-induced Foxp3 inhibits TH17 cell differentiation by antagonizing RORγt function. Nature 453, 236–240 (2008).
Sefik, E. et al. Individual intestinal symbionts induce a distinct population of ROR+regulatory T cells. Science 349, 993–997 (2015).
Ohnmacht, C. et al. The microbiota regulates type 2 immunity through RORγt+ T cells. Science 349, 989–993 (2015). References 126 and 127 show how the microbiota affects intestinal RORγt + T reg cells.
Xu, M. et al. C-MAF-dependent regulatory T cells mediate immunological tolerance to a gut pathobiont. Nature 554, 373–377 (2018).
Chai, J. N. et al. Helicobacter species are potent drivers of colonic T cell responses in homeostasis and inflammation. Sci. Immunol. 2, eaal5068 (2017).
Sano, T. et al. An IL-23R/IL-22 circuit regulates epithelial serum amyloid A to promote local effector Th17 responses. Cell 163, 381–393 (2015).
Tan, T. G. et al. Identifying species of symbiont bacteria from the human gut that, alone, can induce intestinal Th17 cells in mice. Proc. Natl Acad. Sci. USA 113, E8141–E8150 (2016).
Yang, Y. et al. Focused specificity of intestinal TH17 cells towards commensal bacterial antigens. Nature 510, 152–156 (2014).
Hall, A. B. et al. A novel Ruminococcus gnavus clade enriched in inflammatory bowel disease patients. Genome Med. 9, 103 (2017).
Wu, H. J. et al. Gut-residing segmented filamentous bacteria drive autoimmune arthritis via T helper 17 cells. Immunity 32, 815–827 (2010).
Scher, J. U. et al. Expansion of intestinal Prevotella copri correlates with enhanced susceptibility to arthritis. eLife 2, e01202 (2013).
Sakaguchi, S., Takahashi, T., Hata, H., Nomura, T. & Sakaguchi, N. SKG mice, a new genetic model of rheumatoid arthritis. Arthritis Res. Ther. 5, 10 (2003).
Maeda, Y. et al. Dysbiosis contributes to arthritis development via activation of autoreactive T cells in the intestine. Arthritis Rheumatol. 68, 2646–2661 (2016).
Lee, Y. K., Menezes, J. S., Umesaki, Y. & Mazmanian, S. K. Proinflammatory T cell responses to gut microbiota promote experimental autoimmune encephalomyelitis. Proc. Natl Acad. Sci. USA 108, 4615–4622 (2011).
Wilck, N. et al. Salt-responsive gut commensal modulates TH17 axis and disease. Nature 551, 585–589 (2017).
Mangan, P. R. et al. Transforming growth factor-β induces development of the TH17 lineage. Nature 441, 231–234 (2006).
Godinez, I. et al. Interleukin-23 orchestrates mucosal responses to Salmonella enterica serotype typhimurium in the intestine. Infect. Immun. 77, 387–398 (2009).
Happel, K. I. et al. Divergent roles of IL-23 and IL-12 in host defense against Klebsiella pneumoniae. J. Exp. Med. 202, 761–769 (2005).
Atarashi, K. et al. Ectopic colonization of oral bacteria in the intestine drives TH1 cell induction and inflammation. Science 358, 162–169 (2017). Shows that members of the oral microbiota can colonize the gut and cause intestinal inflammation.
Nordmann, P., Cuzon, G. & Naas, T. The real threat of Klebsiella pneumoniae carbapenemase-producing bacteria. Lancet Infect. Dis. 9, 228–236 (2009).
Bell, B. et al. Antibiotic Resistance Threats in the United States (Centers for Disease Control and Prevention, 2013).
Hampton, T. Report reveals scope of US antibiotic resistance threat. JAMA 310, 1661–1663 (2013).
Chassaing, B., Koren, O., Carvalho, F. A., Ley, R. E. & Gewirtz, A. T. AIEC pathobiont instigates chronic colitis in susceptible hosts by altering microbiota composition. Gut 63, 1069–1080 (2014).
Darfeuille-Michaud, A. et al. High prevalence of adherent-invasive Escherichia coli associated with ileal mucosa in Crohn’s disease. Gastroenterology 127, 412–421 (2004).
Martin, H. M. et al. Enhanced Escherichia coli adherence and invasion in Crohn’s disease and colon cancer. Gastroenterology 127, 80–93 (2004).
Devkota, S. et al. Dietary-fat-induced taurocholic acid promotes pathobiont expansion and colitis in Il10−/− mice. Nature 487, 104–108 (2012).
Olivares-Villagómez, D. & Van Kaer, L. Intestinal intraepithelial lymphocytes: sentinels of the mucosal barrier. Trends Immunol. 39, 264–275 (2018).
Gutiérrez-Vázquez, C. & Quintana, F. J. Regulation of the immune response by the aryl hydrocarbon receptor. Immunity 48, 19–33 (2018).
Li, Y. et al. Exogenous stimuli maintain intraepithelial lymphocytes via aryl hydrocarbon receptor activation. Cell 147, 629–640 (2011).
Cervantes-Barragan, L. et al. Lactobacillus reuteri induces gut intraepithelial CD4+CD8αα+T cells. Science 357, 806–810 (2017).
Sujino, T. et al. Tissue adaptation of regulatory and intraepithelial CD4+T cells controls gut inflammation. Science 352, 1581–1586 (2016).
Mucida, D. et al. Transcriptional reprogramming of mature CD4+helper T cells generates distinct MHC class II-restricted cytotoxic T lymphocytes. Nat. Immunol. 14, 281–289 (2013).
Reis, B. S., Rogoz, A., Costa-Pinto, F. A., Taniuchi, I. & Mucida, D. Mutual expression of the transcription factors Runx3 and ThPOK regulates intestinal CD4+ T cell immunity. Nat. Immunol. 14, 271–280 (2013).
Reis, B. S., Hoytema van Konijnenburg, D. P., Grivennikov, S. I. & Mucida, D. Transcription factor T-bet regulates intraepithelial lymphocyte functional maturation. Immunity 41, 244–256 (2014).
Steenholt, J. V. et al. The composition of T cell subtypes in duodenal biopsies are altered in coeliac disease patients. PLOS ONE 12, e0170270 (2017).
Carton, J., Byrne, B., Madrigal-Estebas, L., O’Donoghue, D. P. & O’Farrelly, C. CD4+CD8+human small intestinal T cells are decreased in coeliac patients, with CD8 expression downregulated on intra-epithelial T cells in the active disease. Eur. J. Gastroenterol. Hepatol. 16, 961–968 (2004).
Godfrey, D. I., Stankovic, S. & Baxter, A. G. Raising the NKT cell family. Nat. Immunol. 11, 197–206 (2010).
Olszak, T. et al. Microbial exposure during early life has persistent effects on natural killer T cell function. Science 336, 489–493 (2012).
An, D. et al. Sphingolipids from a symbiotic microbe regulate homeostasis of host intestinal natural killer T cells. Cell 156, 123–133 (2014).
Wieland Brown, L. C. et al. Production of α-galactosylceramide by a prominent member of the human gut microbiota. PLOS Biol. 11, e1001610 (2013).
Ma, C. et al. Gut microbiome–mediated bile acid metabolism regulates liver cancer via NKT cells. Science 360, eaan5931 (2018).
Iyer, S. S. et al. Dietary and microbial oxazoles induce intestinal inflammation by modulating aryl hydrocarbon receptor responses. Cell 173, 1123–1134 (2018).
Metelev, M. V. & Ghilarov, D. A. Structure, function, and biosynthesis of thiazole/oxazole-modified microcins. Mol. Biol. 48, 29–45 (2014).
Xiao, X. & Cai, J. Mucosal-associated invariant T cells: New insights into antigen recognition and activation. Front. Immunol. 8, 1540 (2017).
Gold, M. C. et al. Human mucosal associated invariant T cells detect bacterially infected cells. PLOS Biol. 8, e1000407 (2010).
Georgel, P., Radosavljevic, M., Macquin, C. & Bahram, S. The non-conventional MHC class I MR1 molecule controls infection by Klebsiella pneumoniae in mice. Mol. Immunol. 48, 769–775 (2011).
Treiner, E. et al. Selection of evolutionarily conserved mucosal-associated invariant T cells by MR1. Nature 422, 164–169 (2003).
Le Bourhis, L., Mburu, Y. K. & Lantz, O. MAIT cells, surveyors of a new class of antigen: development and functions. Curr. Opin. Immunol. 25, 174–180 (2013).
Kjer-Nielsen, L. et al. MR1 presents microbial vitamin B metabolites to MAIT cells. Nature 491, 717–723 (2012).
Corbett, A. J. et al. T cell activation by transitory neo-antigens derived from distinct microbial pathways. Nature 509, 361–365 (2014).
Chen, Z. et al. Mucosal-associated invariant T cell activation and accumulation after in vivo infection depends on microbial riboflavin synthesis and co-stimulatory signals. Mucosal Immunol. 10, 58–68 (2017).
Routy, B. et al. Gut microbiome influences efficacy of PD-1-based immunotherapy against epithelial tumors. Science 359, 91–97 (2018).
Matson, V. et al. The commensal microbiome is associated with anti-PD-1 efficacy in metastatic melanoma patients. Science 359, 104–108 (2018).
Gopalakrishnan, V. et al. Gut microbiome modulates response to anti-PD-1 immunotherapy in melanoma patients. Science 359, 97–103 (2018). References 176–178 show that gut commensals affect responses to immune checkpoint blockade therapy in patients with cancer.
Uchimura, Y. et al. Antibodies set boundaries limiting microbial metabolite penetration and the resultant mammalian host response. Immunity 49, 545–559 (2018).
Donaldson, G. P. et al. Gut microbiota utilize immunoglobulin A for mucosal colonization. Science 360, 795–800 (2018).
Bunker, J. J. et al. Natural polyreactive IgA antibodies coat the intestinal microbiota. Science 358, eaan6619 (2017).
Kim, M., Qie, Y., Park, J. & Kim, C. H. Gut microbial metabolites fuel host antibody responses. Cell Host Microbe 20, 202–214 (2016).
McLoughlin, K., Schluter, J., Rakoff-Nahoum, S., Smith, A. L. & Foster, K. R. Host selection of microbiota via differential adhesion. Cell Host Microbe 19, 550–559 (2016).
Coombes, J. L. & Powrie, F. Dendritic cells in intestinal immune regulation. Nat. Rev. Immunol. 8, 435–446 (2008).
Coombes, J. L. et al. A functionally specialized population of mucosal CD103+DCs induces Foxp3+regulatory T cells via a TGF-β– and retinoic acid–dependent mechanism. J. Exp. Med. 204, 1757–1764 (2007).
Sun, C.-M. et al. Small intestine lamina propria dendritic cells promote de novo generation of Foxp3 T reg cells via retinoic acid. J. Exp. Med. 204, 1775–1785 (2007).
Travis, M. A. et al. Loss of integrin αvβ8 on dendritic cells causes autoimmunity and colitis in mice. Nature 449, 361–365 (2007).
Jaensson-Gyllenbäck, E. et al. Bile retinoids imprint intestinal CD103 + dendritic cells with the ability to generate gut-tropic T cells. Mucosal Immunol. 4, 438–447 (2011).
Yokota, A. et al. GM-CSF and IL-4 synergistically trigger dendritic cells to acquire retinoic acid-producing capacity. Int. Immunol. 21, 361–377 (2009).
Iwata, M. et al. Retinoic acid imprints gut-homing specificity on T cells. Immunity 21, 527–538 (2004).
Tanoue, T., Atarashi, K. & Honda, K. Development and maintenance of intestinal regulatory T cells. Nat. Rev. Immunol. 16, 295–309 (2016).
Xu, L. et al. Positive and negative transcriptional regulation of the Foxp3 gene is mediated by access and binding of the Smad3 protein to enhancer I. Immunity 33, 313–325 (2010).
Zeng, R. et al. Retinoic acid regulates the development of a gut-homing precursor for intestinal dendritic cells. Mucosal Immunol. 6, 847–856 (2013).
Zeng, R., Bscheider, M., Lahl, K., Lee, M. & Butcher, E. C. Generation and transcriptional programming of intestinal dendritic cells: essential role of retinoic acid. Mucosal Immunol. 9, 183–193 (2016).
Konieczna, P. et al. Immunomodulation by Bifidobacterium infantis 35624 in the murine lamina propria requires retinoic acid-dependent and independent mechanisms. PLOS ONE 8, e62617 (2013).
Kovatcheva-Datchary, P. et al. Dietary fiber-induced improvement in glucose metabolism is associated with increased abundance of prevotella. Cell Metab. 22, 971–982 (2015).
Rubic, T. et al. Triggering the succinate receptor GPR91 on dendritic cells enhances immunity. Nat. Immunol. 9, 1261–1269 (2008).
Wu, W. et al. Microbiota metabolite short-chain fatty acid acetate promotes intestinal IgA response to microbiota which is mediated by GPR43. Mucosal Immunol. 10, 946–956 (2016).
Mezrich, J. D. et al. An interaction between kynurenine and the aryl hydrocarbon receptor can generate regulatory T cells. J. Immunol. 185, 3190–3198 (2010).
Mellor, A. L. & Munn, D. H. IDO expression by dendritic cells: tolerance and tryptophan catabolism. Nat. Rev. Immunol. 4, 762–774 (2004).
Puccetti, P. & Grohmann, U. IDO and regulatory T cells: a role for reverse signalling and non-canonical NF-κB activation. Nat. Rev. Immunol. 7, 817–823 (2007).
Kim, M. et al. Critical role for the microbiota in CX3CR1+intestinal mononuclear phagocyte regulation of intestinal T cell responses. Immunity 49, 151–163 (2018).
Diehl, G. E. et al. Microbiota restricts trafficking of bacteria to mesenteric lymph nodes by CX3CR1-hi cells. Nature 494, 116–120 (2013).
Panea, C. et al. Intestinal monocyte-derived macrophages control commensal-specific Th17 responses. Cell Rep. 12, 1314–1324 (2015).
Chang, P. V., Hao, L., Offermanns, S. & Medzhitov, R. The microbial metabolite butyrate regulates intestinal macrophage function via histone deacetylase inhibition. Proc. Natl Acad. Sci. USA 111, 2247–2252 (2014).
Mann, E. R. et al. Antibiotics induce sustained dysregulation of intestinal T cell immunity by perturbing macrophage homeostasis. Sci. Transl Med. 10, 4755 (2018).
Ji, J. et al. Microbial metabolite butyrate facilitates M2 macrophage polarization and function. Sci. Rep. 6, 24838 (2016).
Steed, A. L. et al. The microbial metabolite desaminotyrosine protects from influenza through type I interferon. Science 357, 498–502 (2017).
Karst, S. M. The influence of commensal bacteria on infection with enteric viruses. Nat. Rev. Microbiol. 14, 197–204 (2016).
Kuss, S. K. et al. Intestinal microbiota promote enteric virus replication and systemic pathogenesis. Science 334, 249–252 (2011).
Claesen, J. & Fischbach, M. A. Synthetic microbes as drug delivery systems. ACS Synth. Biol. 4, 358–364 (2015).
Kumar, H., Kawai, T. & Akira, S. Pathogen recognition by the innate immune system. Int. Rev. Immunol. 30, 16–34 (2011).
Krawczyk, C. M. et al. Toll-like receptor-induced changes in glycolytic metabolism regulate dendritic cell activation. Blood 115, 4742–4749 (2010).
Kelly, B. & O’Neill, L. A. J. Metabolic reprogramming in macrophages and dendritic cells in innate immunity. Cell Res. 25, 771–784 (2015).
Symbiotix Biotherapies. Pipeline. symbiotix-bio http://symbiotix-bio.com/research-and-development/pipeline/ (2019).
Sonnenburg, J. L. & Bäckhed, F. Diet-microbiota interactions as moderators of human metabolism. Nature 535, 56–64 (2016).
Suez, J. & Elinav, E. The path towards microbiome-based metabolite treatment. Nat. Microbiol. 2, 17075 (2017).
Stein, R. R. et al. Computer-guided design of optimal microbial consortia for immune system modulation. eLife 7, e30916 (2018).
Tramontano, M. et al. Nutritional preferences of human gut bacteria reveal their metabolic idiosyncrasies. Nat. Microbiol. 3, 514–522 (2018).
Maldonado-Gómez, M. X. et al. Stable engraftment of bifidobacterium longum AH1206 in the human gut depends on individualized features of the resident microbiome. Cell Host Microbe 20, 515–526 (2016).
Shepherd, E. S., Deloache, W. C., Pruss, K. M., Whitaker, W. R. & Sonnenburg, J. L. An exclusive metabolic niche enables strain engraftment in the gut microbiota. Nature 557, 434–438 (2018).
Gurry, T. et al. Predictability and persistence of prebiotic dietary supplementation in a healthy human cohort. Sci. Rep. 8, 12699 (2017).
Panigrahi, P. et al. A randomized synbiotic trial to prevent sepsis among infants in rural India. Nature 548, 407–412 (2017). This study shows that a synbiotic consisting of L. plantarum and fructo-oligosaccharide protects infants against sepsis.
Panigrahi, P. et al. Long-term colonization of a lactobacillus plantarum synbiotic preparation in the neonatal gut. J. Pediatr. Gastroenterol. Nutr. 47, 45–53 (2008).
Goodman, A. L. et al. Extensive personal human gut microbiota culture collections characterized and manipulated in gnotobiotic mice. Proc. Natl Acad. Sci. USA 108, 6252–6257 (2011).
Litvak, Y., Byndloss, M. X. & Bäumler, A. J. Colonocyte metabolism shapes the gut microbiota. Science 362, eaat9076 (2018).
Byndloss, M. X., Pernitzsch, S. R. & Bäumler, A. J. Healthy hosts rule within: ecological forces shaping the gut microbiota. Mucosal Immunol. 11, 1299–1305 (2018).
Byndloss, M. X. et al. Microbiota-activated PPAR-γ signaling inhibits dysbiotic Enterobacteriaceae expansion. Science 357, 570–575 (2017).
Litvak, Y., Byndloss, M. X., Tsolis, R. M. & Bäumler, A. J. Dysbiotic proteobacteria expansion: a microbial signature of epithelial dysfunction. Curr. Opin. Microbiol. 39, 1–6 (2017).
Tang, C. et al. Suppression of IL-17F, but not of IL-17A, provides protection against colitis by inducing Treg cells through modification of the intestinal microbiota. Nat. Immunol. 19, 755–765 (2018).
Cullen, T. W. et al. Antimicrobial peptide resistance mediates resilience of prominent gut commensals during inflammation. Science 347, 170–175 (2015).
Seregin, S. S. et al. NLRP6 protects Il10−/−mice from colitis by limiting colonization of akkermansia muciniphila. Cell Rep. 19, 733–745 (2017).
Png, C. W. et al. Mucolytic bacteria with increased prevalence in IBD mucosa augment in vitro utilization of mucin by other bacteria. Am. J. Gastroenterol. 105, 2420–2428 (2010).
Chelakkot, C. et al. Akkermansia muciniphila-derived extracellular vesicles influence gut permeability through the regulation of tight junctions. Exp. Mol. Med. 50, e450 (2018).
Schneeberger, M. et al. Akkermansia muciniphila inversely correlates with the onset of inflammation, altered adipose tissue metabolism and metabolic disorders during obesity in mice. Sci. Rep. 5, 16643 (2015).
Hryckowian, A. J. et al. Microbiota-accessible carbohydrates suppress Clostridium difficile infection in a murine model. Nat. Microbiol. 3, 662–669 (2018).
Steidler, L. et al. Treatment of murine colitis by Lactococcus lactis secreting interleukin-10. Science 289, 1352–1355 (2000).
Braat, H. et al. A phase I trial with transgenic bacteria expressing interleukin-10 in Crohn’s disease. Clin. Gastroenterol. Hepatol. 4, 754–759 (2006).
Álvarez, B. & Fernández, L. Á. Sustainable therapies by engineered bacteria. Microb. Biotechnol. 10, 1057–1061 (2017).
Duan, F. & March, J. C. Engineered bacterial communication prevents Vibrio cholerae virulence in an infant mouse model. Proc. Natl Acad. Sci. USA 107, 11260–11264 (2010).
Silva, A. J. & Benitez, J. A. Vibrio cholerae biofilms and cholera pathogenesis. PLOS Negl. Trop. Dis. 10, e0004330 (2016).
Hsiao, A. et al. Members of the human gut microbiota involved in recovery from Vibrio cholerae infection. Nature 515, 423–426 (2014).
Saeidi, N. et al. Engineering microbes to sense and eradicate Pseudomonas aeruginosa, a human pathogen. Mol. Syst. Biol. 7, 521 (2011).
Hwang, I. Y. et al. Engineered probiotic Escherichia coli can eliminate and prevent Pseudomonas aeruginosa gut infection in animal models. Nat. Commun. 8, 15028 (2017).
Wahlström, A., Sayin, S. I., Marschall, H. U. & Bäckhed, F. Intestinal crosstalk between bile acids and microbiota and its impact on host metabolism. Cell Metab. 24, 41–50 (2016).
Jones, B. V., Begley, M., Hill, C., Gahan, C. G. M. & Marchesi, J. R. Functional and comparative metagenomic analysis of bile salt hydrolase activity in the human gut microbiome. Proc. Natl Acad. Sci. USA 105, 13580–13585 (2008).
Kitahara, M., Takamine, F., Imamura, T. & Benno, Y. Assignment of eubacterium sp. VPI 12708 and related strains with high bile acid 7α-dehydroxylating activity to Clostridium scindens and proposal of Clostridium hylemonae sp. nov., isolated from human faeces. Int. J. Syst. Evol. Microbiol. 50, 971–978 (2000).
Kitahara, M., Takamine, F., Imamura, T. & Benno, Y. Clostridium hiranonis sp. nov., a human intestinal bacterium with bile acid 7α-dehydroxylating activity. Int. J. Syst. Evol. Microbiol. 51, 39–44 (2001).
Ridlon, J. M., Kang, D.-J. & Hylemon, P. B. Bile salt biotransformations by human intestinal bacteria. J. Lipid Res. 47, 241–259 (2006).
Ridlon, J., Kang, D., Hylemon, P. & Bajaj, J. Bile acids and the gut microbiome. Curr. Opin. Gastroenterol. 30, 332–338 (2014).
Lefebvre, P., Cariou, B., Lien, F., Kuipers, F. & Staels, B. Role of bile acids and bile acid receptors in metabolic regulation. Physiol. Rev. 89, 147–191 (2009).
Sinal, C. J. et al. Targeted disruption of the nuclear receptor FXR/BAR impairs bile acid and lipid homeostasis. Cell 102, 731–744 (2000).
Vavassori, P., Mencarelli, A., Renga, B., Distrutti, E. & Fiorucci, S. The bile acid receptor FXR is a modulator of intestinal innate immunity. J. Immunol. 183, 6251–6261 (2009).
Chen, X., Lou, G., Meng, Z. & Huang, W. TGR5: a novel target for weight maintenance and glucose metabolism. Exp. Diabetes Res. 2011, 853501 (2011).
Maruyama, T. et al. Identification of membrane-type receptor for bile acids (M-BAR). Biochem. Biophys. Res. Commun. 298, 714–719 (2002).
Haselow, K. et al. Bile acids PKA-dependently induce a switch of the IL-10/IL-12 ratio and reduce proinflammatory capability of human macrophages. J. Leukoc. Biol. 94, 1253–1264 (2013).
Keitel, V., Donner, M., Winandy, S., Kubitz, R. & Häussinger, D. Expression and function of the bile acid receptor TGR5 in Kupffer cells. Biochem. Biophys. Res. Commun. 372, 78–84 (2008).
Perino, A. et al. TGR5 reduces macrophage migration through mTOR-induced C/EBPβ differential translation. J. Clin. Invest. 124, 5424–5436 (2014).
Pols, T. W. H. et al. TGR5 activation inhibits atherosclerosis by reducing macrophage inflammation and lipid loading. Cell Metab. 14, 747–757 (2011).
Pols, T. W. H. et al. Lithocholic acid controls adaptive immune responses by inhibition of Th1 activation through the Vitamin D receptor. PLOS ONE 12, e0176715 (2017).
Smith, K., McCoy, K. D. & Macpherson, A. J. Use of axenic animals in studying the adaptation of mammals to their commensal intestinal microbiota. Semin. Immunol. 19, 59–69 (2007).
Round, J. L. & Mazmanian, S. K. The gut microbiota shapes intestinal immune responses during health and disease. Nat. Rev. Immunol. 9, 313–323 (2009).
Chung, H. et al. Gut immune maturation depends on colonization with a host-specific microbiota. Cell 149, 1578–1593 (2012). This study shows that a host-specific microbiota is necessary for immunomaturation.
Zhang, L. et al. Environmental spread of microbes impacts the development of metabolic phenotypes in mice transplanted with microbial communities from humans. ISME J. 11, 676–690 (2017).
Ridaura, V. K. et al. Gut microbiota from twins discordant for obesity modulate metabolism in mice. Science 341, 1241214 (2013).
Turnbaugh, P. J. et al. The effect of diet on the human gut microbiome: a metagenomic analysis in humanized gnotobiotic mice. Sci. Transl Med. 1, 6ra14 (2009).
Hugenholtz, F. & De Vos, W. M. Mouse models for human intestinal microbiota research: a critical evaluation. Cell. Mol. Life Sci. 75, 149–160 (2018).
Takahashi, S. et al. Cyp2c70 is responsible for the species difference in bile acid metabolism between mice and humans. J. Lipid Res. 57, 2130–2137 (2016).
Nguyen, T. L. A., Vieira-Silva, S., Liston, A. & Raes, J. How informative is the mouse for human gut microbiota research? Dis. Model. Mech. 8, 1–16 (2015).
Kim, H. J., Li, H., Collins, J. J. & Ingber, D. E. Contributions of microbiome and mechanical deformation to intestinal bacterial overgrowth and inflammation in a human gut-on-a-chip. Proc. Natl Acad. Sci. USA 113, E7–E15 (2016).
Shah, P. et al. A microfluidics-based in vitro model of the gastrointestinal human-microbe interface. Nat. Commun. 7, 11535 (2016).
Turnbaugh, P. J., Bäckhed, F., Fulton, L. & Gordon, J. I. Diet-induced obesity is linked to marked but reversible alterations in the mouse distal gut microbiome. Cell Host Microbe 3, 213–223 (2008).
Bloom, S. M. et al. Commensal Bacteroides species induce colitis in host-genotype-specific fashion in a mouse model of inflammatory bowel disease. Cell Host Microbe 9, 390–403 (2011).
Stefka, A. T. et al. Commensal bacteria protect against food allergen sensitization. Proc. Natl Acad. Sci. USA 111, 13145–13150 (2014).
Sivan, A. et al. Commensal Bifidobacterium promotes antitumor immunity and facilitates anti-PD-L1 efficacy. Science 350, 1084–1089 (2015).
Derrien, M., Vaughan, E. E., Plugge, C. M. & de Vos, W. M. Akkermansia municiphila gen. nov., sp. nov., a human intestinal mucin-degrading bacterium. Int. J. Syst. Evol. Microbiol. 54, 1469–1476 (2004).
Everard, A. et al. Cross-talk between Akkermansia muciniphila and intestinal epithelium controls diet-induced obesity. Proc. Natl Acad. Sci. USA 110, 9066–9071 (2013).
Ottman, N. et al. Pili-like proteins of Akkermansia muciniphila modulate host immune responses and gut barrier function. PLOS ONE 12, e0173004 (2017).
van den Abbeele, P. et al. Arabinoxylans and inulin differentially modulate the mucosal and luminal gut microbiota and mucin-degradation in humanized rats. Environ. Microbiol. 13, 2667–2680 (2011).
Chia, L. W. et al. Deciphering the trophic interaction between Akkermansia muciniphila and the butyrogenic gut commensal Anaerostipes caccae using a metatranscriptomic approach. Antonie Van Leeuwenhoek 111, 859–873 (2018).
Ijssennagger, N. et al. Gut microbiota facilitates dietary heme-induced epithelial hyperproliferation by opening the mucus barrier in colon. Proc. Natl Acad. Sci. USA 112, 10038–10043 (2015).
Ganesh, B. P., Klopfleisch, R., Loh, G. & Blaut, M. Commensal Akkermansia muciniphila exacerbates gut inflammation in Salmonella typhimurium-infected gnotobiotic mice. PLOS ONE 8, e74963 (2013).
Desai, M. S. et al. A dietary fiber-deprived gut microbiota degrades the colonic mucus barrier and enhances pathogen susceptibility. Cell 167, 1339–1353 (2016).
Pitt, J. M. et al. Resistance mechanisms to immune-checkpoint blockade in cancer: tumor-intrinsic and -extrinsic factors. Immunity 44, 1255–1269 (2016).
Acknowledgements
This work was supported by the Japan Agency for Medical Research and Development (AMED) LEAP under grant number JP18gm0010003, the Takeda Science Foundation, the Mitsukoshi Health and Welfare Foundation (K.H.) and Grant-in-Aid for Japan Society for the Promotion of Science (JSPS) Fellows (18F18104) (S.K.).
Reviewer information
Nature Reviews Immunology thanks N. Cerf-Bensussan, L. O’Neill and P. Turnbaugh for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
All authors were involved in discussing the content and writing this article. A.N.S., Y.S. and S.K. were involved in researching data for the article. K.H. and A.N.S. contributed to the review and editing of the article before submission.
Corresponding author
Ethics declarations
Competing interests
K.H. is co-founder of and scientific adviser to Vedanta Biosciences since 11 August 2011. The other authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Glossary
- Dysbiosis
-
The perturbation of the homeostatic microbiota composition; associated with a slew of diseases. The ambiguity of this term can encompass increases in pathobionts, decreases in commensals or changes that do not clearly distinguish healthy from unhealthy guts.
- Gnotobiotic
-
Referring to mice that are only colonized by a known, defined set of microorganisms.
- Colonization resistance
-
The mechanism by which certain microorganisms are prevented from colonizing an ecosystem (for example, the gut) by the indigenous microorganisms.
- Germ-free (GF) mice
-
Mice living in sterile conditions, uncolonized by any microorganism.
- Muricholic acids
-
A class of 6-hydroxy bile acids present in mice but not humans.
- Beta-diversity
-
The ecological diversity between environments (for example, gut microbial diversity between different individuals).
- M cells
-
Microfold cells found in Peyer’s patches that transcytose material (including antigen and IgA) across the intestinal epithelium.
- Metabolic oligosaccharide engineering
-
The process of co-opting endogenous bacterial biosynthetic machinery to introduce a synthetic non-natural sugar containing an inert chemical ‘handle’ into surface macromolecules.
- Bioorthogonal click chemistry
-
Any chemical reaction that can occur rapidly and specifically in the complex cellular milieu. Often used in conjunction with metabolic oligosaccharide engineering to functionalize a tagged macromolecule.
- Outer membrane vesicles
-
(OMVs). Vesicles containing periplasm; released into the extracellular space by Gram-negative bacteria.
- Aryl hydrocarbon receptor
-
(AhR). A cytosolic ligand-activated transcription factor that controls the expression of many genes.
- Auxotrophic
-
Relating to auxotrophy; the inability to produce a substance required for growth. Using an auxotrophic bacterial strain enables reversible colonization under experimental conditions.
- M2 macrophage phenotype
-
‘M1’ and ‘M2’ are classifications historically used to define macrophages activated in vitro as pro-inflammatory (when classically activated with IFNγ and lipopolysaccharide) or anti-inflammatory (when alternatively activated with IL-4 or IL-10), respectively. However, in vivo macrophages are highly specialized, transcriptomically dynamic and extremely heterogeneous with regards to their phenotypes and functions, which are continuously shaped by their tissue microenvironment. Therefore, the M1 or M2 classification is too simplistic to explain the true nature of in vivo macrophages, although these terms are still often used to indicate whether the macrophages in question are more pro-inflammatory or anti-inflammatory.
- Synbiotic
-
A therapeutic consisting of probiotics (bacteria) together with prebiotics (nutrients).
- Quorum-sensing
-
The ability to sense population density and respond accordingly at the gene expression level.
Rights and permissions
About this article
Cite this article
Skelly, A.N., Sato, Y., Kearney, S. et al. Mining the microbiota for microbial and metabolite-based immunotherapies. Nat Rev Immunol 19, 305–323 (2019). https://doi.org/10.1038/s41577-019-0144-5
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41577-019-0144-5
This article is cited by
-
The gut-liver axis in hepatobiliary diseases
Inflammation and Regeneration (2024)
-
Gut microbial metabolites SCFAs and chronic kidney disease
Journal of Translational Medicine (2024)
-
Pretreatment with an antibiotics cocktail enhances the protective effect of probiotics by regulating SCFA metabolism and Th1/Th2/Th17 cell immune responses
BMC Microbiology (2024)
-
Microbiota–gut–brain axis and its therapeutic applications in neurodegenerative diseases
Signal Transduction and Targeted Therapy (2024)
-
Dubosiella newyorkensis modulates immune tolerance in colitis via the L-lysine-activated AhR-IDO1-Kyn pathway
Nature Communications (2024)