Abstract
Methods for profiling genes at the single-cell level have revolutionized our ability to study several biological processes and systems including development, differentiation, response programmes and disease progression. In many of these studies, cells are profiled over time in order to infer dynamic changes in cell states and types, sets of expressed genes, active pathways and key regulators. However, time-series single-cell RNA sequencing (scRNA-seq) also raises several new analysis and modelling issues. These issues range from determining when and how deep to profile cells, linking cells within and between time points, learning continuous trajectories, and integrating bulk and single-cell data for reconstructing models of dynamic networks. In this Review, we discuss several approaches for the analysis and modelling of time-series scRNA-seq, highlighting their steps, key assumptions, and the types of data and biological questions they are most appropriate for.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Code availability
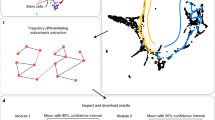
The code to reproduce the plots shown in Fig. 3 is available at http://dinglab.rimuhc.ca:8080/nrg/.
Data availability
The primary data for the analyses shown in Fig. 3 are from Treutlein et al.69, and the processed data presented in the figure are available at http://dinglab.rimuhc.ca:8080/nrg/.
References
Gasch, A. P. et al. Single-cell RNA sequencing reveals intrinsic and extrinsic regulatory heterogeneity in yeast responding to stress. PLoS Biol. 15, e2004050 (2017).
Zou, Z. et al. A single-cell transcriptomic atlas of human skin aging. Dev. Cell 56, 383–397.e8 (2021).
Stuart, T. & Satija, R. Integrative single-cell analysis. Nat. Rev. Genet. 20, 257–272 (2019).
Alavi, A., Ruffalo, M., Parvangada, A., Huang, Z. & Bar-Joseph, Z. A web server for comparative analysis of single-cell RNA-seq data. Nat. Commun. 9, 4768 (2018).
Dumitrascu, B., Villar, S., Mixon, D. G. & Engelhardt, B. E. Optimal marker gene selection for cell type discrimination in single cell analyses. Nat. Commun. 12, 1186 (2021).
Efremova, M., Vento-Tormo, M., Teichmann, S. A. & Vento-Tormo, R. CellPhoneDB: inferring cell–cell communication from combined expression of multi-subunit ligand–receptor complexes. Nat. Protoc. 15, 1484–1506 (2020).
Bendall, S. C. et al. Single-cell trajectory detection uncovers progression and regulatory coordination in human B cell development. Cell 157, 714–725 (2014).
Bar-Joseph, Z., Gitter, A. & Simon, I. Studying and modelling dynamic biological processes using time-series gene expression data. Nat. Rev. Genet. 13, 552–564 (2012).
La Manno, G. et al. RNA velocity of single cells. Nature 560, 494–498 (2018). This paper introduces the idea of RNA velocity and presents the first method to infer velocity from scRNA-seq data.
Herzog, V. A. et al. Thiol-linked alkylation of RNA to assess expression dynamics. Nat. Methods 14, 1198–1204 (2017).
Erhard, F. et al. scSLAM-seq reveals core features of transcription dynamics in single cells. Nature 571, 419–423 (2019).
Hendriks, G.-J. et al. NASC-seq monitors RNA synthesis in single cells. Nat. Commun. 10, 3138 (2019).
Cao, J., Zhou, W., Steemers, F., Trapnell, C. & Shendure, J. Sci-fate characterizes the dynamics of gene expression in single cells. Nat. Biotechnol. 38, 980–988 (2020).
Qiu, Q. et al. Massively parallel and time-resolved RNA sequencing in single cells with scNT-seq. Nat. Methods 17, 991–1001 (2020).
Schofield, J. A., Duffy, E. E., Kiefer, L., Sullivan, M. C. & Simon, M. D. TimeLapse-seq: adding a temporal dimension to RNA sequencing through nucleoside recoding. Nat. Methods 15, 221 (2018).
Kiefer, L., Schofield, J. A. & Simon, M. D. Expanding the nucleoside recoding toolkit: revealing RNA population dynamics with 6-thioguanosine. J. Am. Chem. Soc. 140, 14567–14570 (2018).
Matsushima, W. et al. SLAM-ITseq: sequencing cell type-specific transcriptomes without cell sorting. Development 145, dev164640 (2018).
Byrnes, L. E. et al. Lineage dynamics of murine pancreatic development at single-cell resolution. Nat. Commun. 9, 3922 (2018).
Gehart, H. et al. Identification of enteroendocrine regulators by real-time single-cell differentiation mapping. Cell 176, 1158–1173.e16 (2019). This study, by determining the change in the ratios between two fluorescent proteins with different half-lives over time, establishes a real-time scale bar for virtual pseudo-time maps.
Reizel, Y. et al. Colon stem cell and crypt dynamics exposed by cell lineage reconstruction. PLoS Genet. 7, e1002192 (2011).
McKenna, A. et al. Whole-organism lineage tracing by combinatorial and cumulative genome editing. Science 353, aaf7907 (2016). This study uses CRISPR–Cas9 to introduce mutations at specific loci on the genome, which remain as ‘scars’ that register the cell lineage.
Alemany, A., Florescu, M., Baron, C. S., Peterson-Maduro, J. & Van Oudenaarden, A. Whole-organism clone tracing using single-cell sequencing. Nature 556, 108–112 (2018).
Spanjaard, B. et al. Simultaneous lineage tracing and cell-type identification using CRISPR–Cas9-induced genetic scars. Nat. Biotechnol. 36, 469–473 (2018).
Raj, B. et al. Simultaneous single-cell profiling of lineages and cell types in the vertebrate brain. Nat. Biotechnol. 36, 442–450 (2018).
Bowling, S. et al. An engineered CRISPR–Cas9 mouse line for simultaneous readout of lineage histories and gene expression profiles in single cells. Cell 181, 1410–1422.e27 (2020).
Quinn, J. J. et al. Single-cell lineages reveal the rates, routes, and drivers of metastasis in cancer xenografts. Science 371, eabc1944 (2021).
Chan, M. M. et al. Molecular recording of mammalian embryogenesis. Nature 570, 77–82 (2019).
Pearson, K. L. III. On lines and planes of closest fit to systems of points in space. Lond. Edinb. Dublin Philos. Mag. J. Sci. 2, 559–572 (1901).
Maaten, Lvd & Hinton, G. Visualizing data using t-SNE. J. Mach. Learn. Res. 9, 2579–2605 (2008).
Becht, E. et al. Dimensionality reduction for visualizing single-cell data using UMAP. Nat. Biotechnol. 37, 38–44 (2019).
Lin, C., Jain, S., Kim, H. & Bar-Joseph, Z. Using neural networks for reducing the dimensions of single-cell RNA-seq data. Nucleic Acids Res. 45, e156 (2017).
Trapnell, C. et al. The dynamics and regulators of cell fate decisions are revealed by pseudotemporal ordering of single cells. Nat. Biotechnol. 32, 381 (2014).
Qiu, X. et al. Reversed graph embedding resolves complex single-cell trajectories. Nat. Methods 14, 979 (2017).
Ji, Z. & Ji, H. TSCAN: pseudo-time reconstruction and evaluation in single-cell RNA-seq analysis. Nucleic Acids Res. 44, e117 (2016).
Street, K. et al. Slingshot: cell lineage and pseudotime inference for single-cell transcriptomics. BMC Genomics 19, 477 (2018).
Wolf, F. A. et al. PAGA: graph abstraction reconciles clustering with trajectory inference through a topology preserving map of single cells. Genome Biol. 20, 59 (2019).
Cao, J. et al. The single-cell transcriptional landscape of mammalian organogenesis. Nature 566, 496–502 (2019). This study introduces and uses Monocle 3, one of the most successful and popular methods for pseudo-time inference.
Ding, J. et al. Reconstructing differentiation networks and their regulation from time series single-cell expression data. Genome Res. 28, 383–395 (2018).
Guo, M., Bao, E. L., Wagner, M., Whitsett, J. A. & Xu, Y. SLICE: determining cell differentiation and lineage based on single cell entropy. Nucleic Acids Res. 45, e54 (2017).
Halbritter, F. et al. Epigenomics and single-cell sequencing define a developmental hierarchy in Langerhans cell histiocytosis. Cancer Discov. 9, 1406–1421 (2019).
Schiebinger, G. et al. Optimal-transport analysis of single-cell gene expression identifies developmental trajectories in reprogramming. Cell 176, 928–943.e22 (2019).
Yang, K. D. et al. Predicting cell lineages using autoencoders and optimal transport. PLoS Comput. Biol. 16, e1007828 (2020).
Forrow, A. & Schiebinger, G. LineageOT is a unified framework for lineage tracing and trajectory inference. Nat. Commun. 12, 4940 (2021).
Saelens, W., Cannoodt, R., Todorov, H. & Saeys, Y. A comparison of single-cell trajectory inference methods. Nat. Biotechnol. 37, 547–554 (2019). This comprehensive trajectory inference comparison study systematically benchmarks several different pseudo-time and trajectory inference methods.
Chen, H. et al. Single-cell trajectories reconstruction, exploration and mapping of omics data with STREAM. Nat. Commun. 10, 1903 (2019).
Megill, C. et al. cellxgene: a performant, scalable exploration platform for high dimensional sparse matrices. Preprint at bioRxiv https://doi.org/10.1101/2021.04.05.438318 (2021).
Rashid, S., Kotton, D. N. & Bar-Joseph, Z. TASIC: determining branching models from time series single cell data. Bioinformatics 33, 2504–2512 (2017).
Marco, E. et al. Bifurcation analysis of single-cell gene expression data reveals epigenetic landscape. Proc. Natl Acad. Sci. USA 111, E5643–E5650 (2014).
Lin, C. & Bar-Joseph, Z. Continuous-state HMMs for modeling time-series single-cell RNA-seq data. Bioinformatics 35, 4707–4715 (2019).
Consortium, T. M. Single-cell transcriptomics of 20 mouse organs creates a Tabula Muris. Nature 562, 367–372 (2018).
Bergen, V., Lange, M., Peidli, S., Wolf, F. A. & Theis, F. J. Generalizing RNA velocity to transient cell states through dynamical modeling. Nat. Biotechnol. 38, 1408–1414 (2020). This study generalizes RNA velocity-based trajectory inference to transient cell states.
Lange, M. et al. CellRank for directed single-cell fate mapping. Preprint at bioRxiv https://doi.org/10.1101/2020.10.19.345983 (2020).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902.e21 (2019). This paper presents Seurat, one of the most comprehensive and popular packages for the analysis and visualization of scRNA-seq data.
Hurley, K. et al. Reconstructed single-cell fate trajectories define lineage plasticity windows during differentiation of human PSC-derived distal lung progenitors. Cell Stem Cell 26, 593–608.e8 (2020). This study applies and validates methods to reconstruct dynamic regulatory networks by integrating time-series scRNA-seq data with interaction data.
Weinreb, C., Rodriguez-Fraticelli, A., Camargo, F. D. & Klein, A. M. Lineage tracing on transcriptional landscapes links state to fate during differentiation. Science 367, eaaw3381 (2020).
Hwang, B. et al. Lineage tracing using a Cas9-deaminase barcoding system targeting endogenous L1 elements. Nat. Commun. 10, 1234 (2019).
Zafar, H., Lin, C. & Bar-Joseph, Z. Single-cell lineage tracing by integrating CRISPR–Cas9 mutations with transcriptomic data. Nat. Commun. 11, 3055 (2020).
Ding, J., Hagood, J. S., Ambalavanan, N., Kaminski, N. & Bar-Joseph, Z. iDREM: interactive visualization of dynamic regulatory networks. PLoS Comput. Biol. 14, e1006019 (2018).
Ding, J. et al. Integrating multiomics longitudinal data to reconstruct networks underlying lung development. Am. J. Physiol. Lung Cell. Mol. Physiol. 317, L556–L568 (2019).
Campbell, K. R. & Yau, C. Uncovering pseudotemporal trajectories with covariates from single cell and bulk expression data. Nat. Commun. 9, 2442 (2018).
Duren, Z., Chen, X., Xin, J., Wang, Y. & Wong, W. H. Time course regulatory analysis based on paired expression and chromatin accessibility data. Genome Res. 30, 622–634 (2020).
Sanchez-Castillo, M., Blanco, D., Tienda-Luna, I. M., Carrion, M. & Huang, Y. A Bayesian framework for the inference of gene regulatory networks from time and pseudo-time series data. Bioinformatics 34, 964–970 (2018).
Hamey, F. K. et al. Reconstructing blood stem cell regulatory network models from single-cell molecular profiles. Proc. Natl Acad. Sci. USA 114, 5822–5829 (2017).
Todorov, H., Cannoodt, R., Saelens, W. & Saeys, Y. in Gene Regulatory Networks Vol. 1883 (eds Sanguinetti, G. & Huynh-Thu, V.) 235–249 (Humana, 2019).
Nguyen, H., Tran, D., Tran, B., Pehlivan, B. & Nguyen, T. A comprehensive survey of regulatory network inference methods using single cell RNA sequencing data. Brief. Bioinform. 22, bbaa190 (2021).
Peng, J. et al. SimiC: a single cell gene regulatory network inference method with similarity constraints. Preprint at bioRxiv https://doi.org/10.1101/2020.04.03.023002 (2020).
Friedman, C. E. et al. Single-cell transcriptomic analysis of cardiac differentiation from human PSCs reveals HOPX-dependent cardiomyocyte maturation. Cell Stem Cell 23, 586–598.e8 (2018).
Lin, C., Ding, J. & Bar-Joseph, Z. Inferring TF activation order in time series scRNA-seq studies. PLoS Comput. Biol. 16, e1007644 (2020).
Treutlein, B. et al. Dissecting direct reprogramming from fibroblast to neuron using single-cell RNA-seq. Nature 534, 391–395 (2016). This paper is one of the first to use time-series scRNA-seq data to study and model development.
Zhang, M. J., Ntranos, V. & Tse, D. Determining sequencing depth in a single-cell RNA-seq experiment. Nat. Commun. 11, 774 (2020).
Kleyman, M. et al. Selecting the most appropriate time points to profile in high-throughput studies. eLife 6, e18541 (2017). This paper introduces one of the first methods for designing time-series RNA-seq experiments.
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Tritschler, S. et al. Concepts and limitations for learning developmental trajectories from single cell genomics. Development 146, dev170506 (2019).
Davis, A., Gao, R. & Navin, N. E. SCOPIT: sample size calculations for single-cell sequencing experiments. BMC Bioinformatics 20, 566 (2019).
Haque, A., Engel, J., Teichmann, S. A. & Lönnberg, T. A practical guide to single-cell RNA-sequencing for biomedical research and clinical applications. Genome Med. 9, 75 (2017).
Schwabe, D., Formichetti, S., Junker, J. P., Falcke, M. & Rajewsky, N. The transcriptome dynamics of single cells during the cell cycle. Mol. Syst. Biol. 16, e9946 (2020).
Kang, H. M. et al. Multiplexed droplet single-cell RNA-sequencing using natural genetic variation. Nat. Biotechnol. 36, 89–94 (2018).
de Kok, J. B. et al. Normalization of gene expression measurements in tumor tissues: comparison of 13 endogenous control genes. Lab. Invest. 85, 154–159 (2005).
Lun, A. T., Calero-Nieto, F. J., Haim-Vilmovsky, L., Göttgens, B. & Marioni, J. C. Assessing the reliability of spike-in normalization for analyses of single-cell RNA sequencing data. Genome Res. 27, 1795–1806 (2017).
Johnson, W. E., Li, C. & Rabinovic, A. Adjusting batch effects in microarray expression data using empirical Bayes methods. Biostatistics 8, 118–127 (2007).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Tran, H. T. N. et al. A benchmark of batch-effect correction methods for single-cell RNA sequencing data. Genome Biol. 21, 12 (2020).
Moter, A. & Göbel, U. B. Fluorescence in situ hybridization (FISH) for direct visualization of microorganisms. J. Microbiol. Methods 41, 85–112 (2000).
Chen, K. H., Boettiger, A. N., Moffitt, J. R., Wang, S. & Zhuang, X. Spatially resolved, highly multiplexed RNA profiling in single cells. Science 348, aaa6090 (2015).
Moffitt, J. R. et al. High-throughput single-cell gene-expression profiling with multiplexed error-robust fluorescence in situ hybridization. Proc. Natl Acad. Sci. USA 113, 11046–11051 (2016).
Shah, S., Lubeck, E., Zhou, W. & Cai, L. In situ transcription profiling of single cells reveals spatial organization of cells in the mouse hippocampus. Neuron 92, 342–357 (2016).
Eng, C.-H. L. et al. Transcriptome-scale super-resolved imaging in tissues by RNA seqFISH+. Nature 568, 235–239 (2019).
Codeluppi, S. et al. Spatial organization of the somatosensory cortex revealed by osmFISH. Nat. Methods 15, 932–935 (2018).
Wang, X. et al. Three-dimensional intact-tissue sequencing of single-cell transcriptional states. Science 361, eaat5691 (2018).
Bergenstråhle, J., Larsson, L. & Lundeberg, J. Seamless integration of image and molecular analysis for spatial transcriptomics workflows. BMC Genomics 21, 482 (2020).
Rodriques, S. G. et al. Slide-seq: a scalable technology for measuring genome-wide expression at high spatial resolution. Science 363, 1463–1467 (2019).
Schiller, H. B. et al. The Human Lung Cell Atlas: a high-resolution reference map of the human lung in health and disease. Am. J. Respir. Cell Mol. Biol. 61, 31–41 (2019).
Park, J., Liu, C. L., Kim, J. & Susztak, K. Understanding the kidney one cell at a time. Kidney Int. 96, 862–870 (2019).
Gitter, A., Carmi, M., Barkai, N. & Bar-Joseph, Z. Linking the signaling cascades and dynamic regulatory networks controlling stress responses. Genome Res. 23, 365–376 (2013).
Wagner, D. E. et al. Single-cell mapping of gene expression landscapes and lineage in the zebrafish embryo. Science 360, 981–987 (2018).
Su, Y. et al. Single-cell analysis resolves the cell state transition and signaling dynamics associated with melanoma drug-induced resistance. Proc. Natl Acad. Sci. USA 114, 13679–13684 (2017).
Russell, A. B., Elshina, E., Kowalsky, J. R., Te Velthuis, A. J. & Bloom, J. D. Single-cell virus sequencing of influenza infections that trigger innate immunity. J. Virol. 93, e00500-19 (2019).
Rozenblatt-Rosen, O. et al. The Human Tumor Atlas Network: charting tumor transitions across space and time at single-cell resolution. Cell 181, 236–249 (2020).
Delile, J. et al. Single cell transcriptomics reveals spatial and temporal dynamics of gene expression in the developing mouse spinal cord. Development 146, dev173807 (2019).
Huisman, S. M. et al. BrainScope: interactive visual exploration of the spatial and temporal human brain transcriptome. Nucleic Acids Res. 45, e83 (2017).
Zhu, Y. et al. Spatiotemporal transcriptomic divergence across human and macaque brain development. Science 362, eaat8077 (2018).
Maniatis, S. et al. Spatiotemporal dynamics of molecular pathology in amyotrophic lateral sclerosis. Science 364, 89–93 (2019). This paper is one of the first to use time-series spatial transcriptomics to study a dynamic biological process.
Acknowledgements
The work in this article is partially supported by National Institutes of Health (NIH) grants 1R01GM122096, OT2OD026682, 1U54AG075931 and 1U24CA268108 to Z.B.-J.
Author information
Authors and Affiliations
Contributions
The authors contributed to all aspects of the article.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information
Nature Reviews Genetics thanks Hongkai Ji, Yvan Saeys and Xiuwei Zhang for their contribution to the peer review of this work.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
BioTuring: https://bioturing.com/
How Many Cells: https://satijalab.org/howmanycells/
Glossary
- Trajectory
-
A graph (often tree) structure that represents the states and their order during the biological process being studied. Cells are assigned to points on this graph.
- Pseudo-time
-
Partial ordering of cells in single-cell RNA sequencing (scRNA-seq) data that represents predecessor and descendent cell state information.
- Unique molecular identifiers
-
Sequence indices (often randomly generated) that are added to sequencing libraries before PCR amplification and enable the identification of PCR duplicates.
- Dimensionality
-
In single-cell analyses, typically refers to the high versus low number of dimensions of the data. When working with large samples where each is composed of tens of thousands of features (for example, cells and their gene expression levels), the high dimension corresponds to the original values whereas the low dimension is a compact, although lossy, way to represent the data with many fewer values. Several low-dimension representation methods have been developed and they differ in the function they attempt to optimize (such as minimizing reconstruction loss, or minimizing differences in distance between the high-dimensional and low-dimensional spaces).
- Auto-encoders
-
Neural networks whose goal is to reconstruct the input values. These networks are used for dimensionality reduction as they compress all input values through a small intermediate layer and then reconstruct them from the information in that layer.
- Graphical models
-
Computational methods that are used to represent joint probability distributions in a compact manner. These include Bayesian networks, hidden Markov models (HMMs) and more.
- Expectation–maximization
-
A widely used computational method that can be used to fill in missing data while simultaneously learning model parameters. The method iterates between the expectation step which determines expected values for missing data and the maximization step which infers parameters using the values obtained by the expectation step.
Rights and permissions
About this article
Cite this article
Ding, J., Sharon, N. & Bar-Joseph, Z. Temporal modelling using single-cell transcriptomics. Nat Rev Genet 23, 355–368 (2022). https://doi.org/10.1038/s41576-021-00444-7
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41576-021-00444-7
This article is cited by
-
CDSKNNXMBD: a novel clustering framework for large-scale single-cell data based on a stable graph structure
Journal of Translational Medicine (2024)
-
Powerful and accurate detection of temporal gene expression patterns from multi-sample multi-stage single-cell transcriptomics data with TDEseq
Genome Biology (2024)
-
scGPT: toward building a foundation model for single-cell multi-omics using generative AI
Nature Methods (2024)
-
Deciphering tumor-infiltrating dendritic cells in the single-cell era
Experimental Hematology & Oncology (2023)
-
Therapeutic uses of oxytocin in stress-related neuropsychiatric disorders
Cell & Bioscience (2023)