Abstract
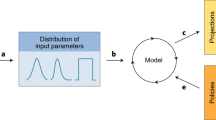
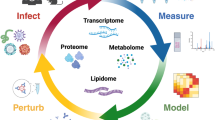
Ongoing social, political and ecological changes in the 21st century have placed more people at risk of life-threatening acute and chronic infections than ever before. The development of new diagnostic, prophylactic, therapeutic and curative strategies is critical to address this burden but is predicated on a detailed understanding of the immensely complex relationship between pathogens and their hosts. Traditional, reductionist approaches to investigate this dynamic often lack the scale and/or scope to faithfully model the dual and co-dependent nature of this relationship, limiting the success of translational efforts. With recent advances in large-scale, quantitative omics methods as well as in integrative analytical strategies, systems biology approaches for the study of infectious disease are quickly forming a new paradigm for how we understand and model host–pathogen relationships for translational applications. Here, we delineate a framework for a systems biology approach to infectious disease in three parts: discovery — the design, collection and analysis of omics data; representation — the iterative modelling, integration and visualization of complex data sets; and application — the interpretation and hypothesis-based inquiry towards translational outcomes.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
World Health Organization. WHO global health estimates 2016: disease burden by cause, age, sex, by country and by region, 2000–2016 (WHO, 2018).
Aderem, A. et al. A systems biology approach to infectious disease research: innovating the pathogen-host research paradigm. mBio 2, e00325–e00410 (2011).
Hillmer, R. A. Systems biology for biologists. PLoS Pathog. 11, e1004786 (2015). An approachable introduction to systems biology for experimentalists.
Kitano, H. Systems biology: a brief overview. Science 295, 1662–1664 (2002). A foundational introduction to the principles of systems biology.
Ideker, T., Galitski, T. & Hood, L. A new approach to decoding life: systems biology. Annu. Rev. Genomics Hum. Genet. 2, 343–372 (2001).
Casadevall, A. & Pirofski, L. A. Host-pathogen interactions: redefining the basic concepts of virulence and pathogenicity. Infect. Immun. 67, 3703–3713 (1999).
Fischbach, M. A. & Krogan, N. J. The next frontier of systems biology: higher-order and interspecies interactions. Genome Biol. 11, 208 (2010).
[No authors listed] Pathogenesis: of host and pathogen. Nat. Immunol. 7, 217 (2006).
Westerhoff, H. V. & Palsson, B. O. The evolution of molecular biology into systems biology. Nat. Biotechnol. 22, 1249–1252 (2004).
Hasin, Y., Seldin, M. & Lusis, A. Multi-omics approaches to disease. Genome Biol. 18, 83 (2017).
Vidova, V. & Spacil, Z. A review on mass spectrometry-based quantitative proteomics: Targeted and data independent acquisition. Anal. Chim. Acta 964, 7–23 (2017).
Aebersold, R. & Mann, M. Mass-spectrometric exploration of proteome structure and function. Nature 537, 347–355 (2016).
Bensimon, A., Heck, A. J. & Aebersold, R. Mass spectrometry-based proteomics and network biology. Annu. Rev. Biochem. 81, 379–405 (2012).
Klemm, S. L., Shipony, Z. & Greenleaf, W. J. Chromatin accessibility and the regulatory epigenome. Nat. Rev. Genet. 20, 207–220 (2019).
Rinschen, M. M., Ivanisevic, J., Giera, M. & Siuzdak, G. Identification of bioactive metabolites using activity metabolomics. Nat. Rev. Mol. Cell Biol. 20, 353–367 (2019).
Doench, J. G. Am I ready for CRISPR? A user’s guide to genetic screens. Nat. Rev. Genet. 19, 67–80 (2018).
Johnson, C. H., Ivanisevic, J. & Siuzdak, G. Metabolomics: beyond biomarkers and towards mechanisms. Nat. Rev. Mol. Cell Biol. 17, 451–459 (2016).
Needham, E. J., Parker, B. L., Burykin, T., James, D. E. & Humphrey, S. J. Illuminating the dark phosphoproteome. Sci. Signal. 12, eaau8645 (2019).
Saliba, A. E., Vonkova, I. & Gavin, A. C. The systematic analysis of protein-lipid interactions comes of age. Nat. Rev. Mol. Cell Biol. 16, 753–761 (2015).
Wang, D. & Bodovitz, S. Single cell analysis: the new frontier in ‘omics’. Trends Biotechnol. 28, 281–290 (2010).
Goodwin, S., McPherson, J. D. & McCombie, W. R. Coming of age: ten years of next-generation sequencing technologies. Nat. Rev. Genet. 17, 333–351 (2016).
Ideker, T. & Krogan, N. J. Differential network biology. Mol. Syst. Biol. 8, 565 (2012).
Greco, T. M. & Cristea, I. M. Proteomics tracing the footsteps of infectious disease. Mol. Cell Proteom. 16, S5–S14 (2017).
Jean Beltran, P. M., Federspiel, J. D., Sheng, X. & Cristea, I. M. Proteomics and integrative omic approaches for understanding host-pathogen interactions and infectious diseases. Mol. Syst. Biol. 13, 922 (2017).
Oxford, K. L. et al. The landscape of viral proteomics and its potential to impact human health. Expert. Rev. Proteomics 13, 579–591 (2016).
Shah, P. S., Wojcechowskyj, J. A., Eckhardt, M. & Krogan, N. J. Comparative mapping of host-pathogen protein-protein interactions. Curr. Opin. Microbiol. 27, 62–68 (2015).
Puschnik, A. S., Majzoub, K., Ooi, Y. S. & Carette, J. E. A CRISPR toolbox to study virus-host interactions. Nat. Rev. Microbiol. 15, 351–364 (2017).
Grubaugh, N. D. et al. Tracking virus outbreaks in the twenty-first century. Nat. Microbiol. 4, 10–19 (2019).
Houldcroft, C. J., Beale, M. A. & Breuer, J. Clinical and biological insights from viral genome sequencing. Nat. Rev. Microbiol. 15, 183–192 (2017).
Newsom, S. N. & McCall, L. I. Metabolomics: Eavesdropping on silent conversations between hosts and their unwelcome guests. PLoS Pathog. 14, e1006926 (2018).
Bernstein, B. E. et al. The NIH roadmap epigenomics mapping consortium. Nat. Biotechnol. 28, 1045–1048 (2010).
Legrain, P. et al. The human proteome project: current state and future direction. Mol. Cell. Proteomics 10, M111.009993 (2011).
Lander, E. S. Initial impact of the sequencing of the human genome. Nature 470, 187–197 (2011).
Mortazavi, A., Williams, B. A., McCue, K., Schaeffer, L. & Wold, B. Mapping and quantifying mammalian transcriptomes by RNA-seq. Nat. Methods 5, 621–628 (2008). Pioneering work demonstrating the use of RNA sequencing to quantify changes in the mammalian transcriptome.
Berns, K. et al. A large-scale RNAi screen in human cells identifies new components of the p53 pathway. Nature 428, 431–437 (2004).
Paddison, P. J. et al. A resource for large-scale RNA-interference-based screens in mammals. Nature 428, 427–431 (2004).
Wang, T., Wei, J. J., Sabatini, D. M. & Lander, E. S. Genetic screens in human cells using the CRISPR-Cas9 system. Science 343, 80–84 (2014).
Shalem, O. et al. Genome-scale CRISPR-Cas9 knockout screening in human cells. Science 343, 84–87 (2014).
Pan, C., Kumar, C., Bohl, S., Klingmueller, U. & Mann, M. Comparative proteomic phenotyping of cell lines and primary cells to assess preservation of cell type-specific functions. Mol. Cell. Proteomics 8, 443–450 (2009).
Sandberg, R. & Ernberg, I. The molecular portrait of in vitro growth by meta-analysis of gene-expression profiles. Genome Biol. 6, R65 (2005).
Ross, D. T. et al. Systematic variation in gene expression patterns in human cancer cell lines. Nat. Genet. 24, 227–235 (2000).
Fux, C. A., Shirtliff, M., Stoodley, P. & Costerton, J. W. Can laboratory reference strains mirror “real-world” pathogenesis? Trends Microbiol. 13, 58–63 (2005).
Jenkins, J. What is the key best practice for collaborating with a computational biologist? Cell Syst. 3, 7–11 (2016).
Lapatas, V., Stefanidakis, M., Jimenez, R. C., Via, A. & Schneider, M. V. Data integration in biological research: an overview. J. Biol. Res. 22, 9 (2015).
Elde, N. C. et al. Poxviruses deploy genomic accordions to adapt rapidly against host antiviral defenses. Cell 150, 831–841 (2012).
Rauch, B. J. et al. Inhibition of CRISPR-Cas9 with bacteriophage proteins. Cell 168, 150–158.e10 (2017).
Weekes, M. P. et al. Quantitative temporal viromics: an approach to investigate host-pathogen interaction. Cell 157, 1460–1472 (2014).
Huttenhain, R. et al. ARIH2 is a Vif-dependent regulator of CUL5-mediated APOBEC3G degradation in HIV infection. Cell Host Microbe 26, 86–99.e7 (2019).
Jean Beltran, P. M., Mathias, R. A. & Cristea, I. M. A portrait of the human organelle proteome in space and time during cytomegalovirus infection. Cell Syst. 3, 361–373.e6 (2016).
Holgate, S. A. How to collaborate. Science https://www.sciencemag.org/careers/2012/07/how-collaborate (2012).
Du, Y. et al. Genome-wide identification of interferon-sensitive mutations enables influenza vaccine design. Science 359, 290–296 (2018). A systems analysis of interferon sensitivity in influenza A viruses made possible by the design of new vaccine approaches, with proof of principle in animal models.
Elde, N. C., Child, S. J., Geballe, A. P. & Malik, H. S. Protein kinase R reveals an evolutionary model for defeating viral mimicry. Nature 457, 485–489 (2009).
Collins, J. et al. Dietary trehalose enhances virulence of epidemic Clostridium difficile. Nature 553, 291–294 (2018).
Carey, A. F. et al. TnSeq of Mycobacterium tuberculosis clinical isolates reveals strain-specific antibiotic liabilities. PLoS Pathog. 14, e1006939 (2018).
Integrative, H. M. P. R. N. C. The integrative human microbiome project. Nature 569, 641–648 (2019).
Liu, R. et al. Homozygous defect in HIV-1 coreceptor accounts for resistance of some multiply-exposed individuals to HIV-1 infection. Cell 86, 367–377 (1996). An early example of population genomics in infectious disease; this is the first report of the Δ32 mutation in human CCR5 conferring natural resistance to HIV-1 infection.
Bryant, J. M. et al. Whole-genome sequencing to identify transmission of Mycobacterium abscessus between patients with cystic fibrosis: a retrospective cohort study. Lancet 381, 1551–1560 (2013).
Bengsch, B. et al. Epigenomic-guided mass cytometry profiling reveals disease-specific features of exhausted CD8 T cells. Immunity 48, 1029–1045.e5 (2018).
Hamdane, N. et al. HCV-induced epigenetic changes associated with liver cancer risk persist after sustained virologic response. Gastroenterology 156, 2313–2329.e7 (2019).
Kennedy, E. M. et al. Posttranscriptional m(6)A editing of HIV-1 mRNAs enhances viral gene expression. Cell Host Microbe 22, 830 (2017).
Arvey, A. et al. An atlas of the Epstein-Barr virus transcriptome and epigenome reveals host-virus regulatory interactions. Cell Host Microbe 12, 233–245 (2012).
Jeng, E. E. et al. Systematic identification of host cell regulators of Legionella pneumophila pathogenesis using a genome-wide CRISPR screen. Cell Host Microbe 26, 551–563.e6 (2019).
Pillay, S. et al. An essential receptor for adeno-associated virus infection. Nature 530, 108–112 (2016).
Marceau, C. D. et al. Genetic dissection of Flaviviridae host factors through genome-scale CRISPR screens. Nature 535, 159–163 (2016).
Hultquist, J. F. et al. A Cas9 ribonucleoprotein platform for functional genetic studies of HIV-host interactions in primary human T cells. Cell Rep. 17, 1438–1452 (2016).
Park, R. J. et al. A genome-wide CRISPR screen identifies a restricted set of HIV host dependency factors. Nat. Genet. 49, 193–203 (2017).
Hoffmann, H. H. et al. Diverse viruses require the calcium transporter SPCA1 for maturation and spread. Cell Host Microbe 22, 460–470.e5 (2017).
Korbee, C. J. et al. Combined chemical genetics and data-driven bioinformatics approach identifies receptor tyrosine kinase inhibitors as host-directed antimicrobials. Nat. Commun. 9, 358 (2018).
Zhou, P. et al. Alpha-kinase 1 is a cytosolic innate immune receptor for bacterial ADP-heptose. Nature 561, 122–126 (2018). A host- and pathogen-based systems approach allows the paired identification of a new bacterial pathogen-associated molecular pattern and its receptor in human cells.
Patrick, K. L. et al. Quantitative yeast genetic interaction profiling of bacterial effector proteins uncovers a role for the human retromer in salmonella infection. Cell Syst. 7, 323–338 e326 (2018).
Ramage, H. R. et al. A combined proteomics/genomics approach links hepatitis C virus infection with nonsense-mediated mRNA decay. Mol. Cell 57, 329–340 (2015).
Hultquist, J. F. et al. CRISPR-Cas9 genome engineering of primary CD4+ T cells for the interrogation of HIV-host factor interactions. Nat. Protoc. 14, 1–27 (2019).
Brass, A. L. et al. Identification of host proteins required for HIV infection through a functional genomic screen. Science 319, 921–926 (2008). A pioneering, RNA interference-based, functional genomics screen for the identification of host factors required for HIV-1 replication in human cells.
Michlmayr, D. et al. Comprehensive innate immune profiling of chikungunya virus infection in pediatric cases. Mol. Syst. Biol. 14, e7862 (2018).
Thompson, E. G. et al. Host blood RNA signatures predict the outcome of tuberculosis treatment. Tuberculosis 107, 48–58 (2017).
Sychev, Z. E. et al. Integrated systems biology analysis of KSHV latent infection reveals viral induction and reliance on peroxisome mediated lipid metabolism. PLoS Pathog. 13, e1006256 (2017).
Lupberger, J. et al. Combined analysis of metabolomes, proteomes, and transcriptomes of hepatitis C virus-infected cells and liver to identify pathways associated with disease development. Gastroenterology 157, 537–551 e539 (2019).
Bradley, T., Ferrari, G., Haynes, B. F., Margolis, D. M. & Browne, E. P. Single-cell analysis of quiescent HIV infection reveals host transcriptional profiles that regulate proviral latency. Cell Rep. 25, 107–117.e3 (2018).
Russell, A. B., Trapnell, C. & Bloom, J. D. Extreme heterogeneity of influenza virus infection in single cells. eLife 7, e32303 (2018).
Diep, J. et al. Enterovirus pathogenesis requires the host methyltransferase SETD3. Nat. Microbiol. 4, 2523–2537 (2019). A combined functional genomics and proteomics approach allows the identification of a new enterovirus host factor, with validation in primary human cells and translationally focused extension into an animal model.
Shah, P. S. et al. Comparative flavivirus-host protein interaction mapping reveals mechanisms of dengue and zika virus pathogenesis. Cell 175, 1931–1945.e18 (2018).
Tripathi, S. et al. Meta- and orthogonal integration of influenza “OMICs” data defines a role for UBR4 in virus budding. Cell Host Microbe 18, 723–735 (2015).
Mirrashidi, K. M. et al. Global mapping of the Inc-human interactome reveals that retromer restricts chlamydia infection. Cell Host Microbe 18, 109–121 (2015).
Jager, S. et al. Global landscape of HIV-human protein complexes. Nature 481, 365–370 (2011). A pioneering study systematically identifying the physical interactions of all HIV-1 proteins and polyproteins with host proteins using affinity tagging and purification mass spectrometry.
Penn, B. H. et al. An Mtb-human protein-protein interaction map identifies a switch between host antiviral and antibacterial responses. Mol. Cell 71, 637–648.e5 (2018).
Davis, Z. H. et al. Global mapping of herpesvirus-host protein complexes reveals a transcription strategy for late genes. Mol. Cell 57, 349–360 (2015).
Kane, J. R. et al. Lineage-specific viral hijacking of non-canonical E3 ubiquitin ligase cofactors in the evolution of Vif anti-APOBEC3 activity. Cell Rep. 11, 1236–1250 (2015).
Babu, M. et al. Global landscape of cell envelope protein complexes in Escherichia coli. Nat. Biotechnol. 36, 103–112 (2018).
Batra, J. et al. Protein interaction mapping identifies RBBP6 as a negative regulator of Ebola virus replication. Cell 175, 1917–1930.e13 (2018).
Eckhardt, M. et al. Multiple routes to oncogenesis are promoted by the human papillomavirus-host protein network. Cancer Discov. 8, 1474–1489 (2018).
Zampieri, M. et al. High-throughput metabolomic analysis predicts mode of action of uncharacterized antimicrobial compounds. Sci. Transl Med. 10, eaal3973 (2018). A metabolomics approach to decipher the mechanism of action of small-molecule antimicrobial compounds with translational potential.
Rother, M. et al. Combined human genome-wide RNAi and metabolite analyses identify IMPDH as a host-directed target against chlamydia infection. Cell Host Microbe 23, 661–671.e8 (2018).
Yuan, S. et al. SREBP-dependent lipidomic reprogramming as a broad-spectrum antiviral target. Nat. Commun. 10, 120 (2019).
Fontaine, K. A., Sanchez, E. L., Camarda, R. & Lagunoff, M. Dengue virus induces and requires glycolysis for optimal replication. J. Virol. 89, 2358–2366 (2015).
Brazma, A. Minimum information about a microarray experiment (MIAME)–successes, failures, challenges. ScientificWorldJournal 9, 420–423 (2009).
Barrett, T. et al. NCBI GEO: archive for functional genomics data sets–update. Nucleic Acids Res. 41, D991–D995 (2013).
Bustin, S. A. et al. The MIQE guidelines: minimum information for publication of quantitative real-time PCR experiments. Clin. Chem. 55, 611–622 (2009).
Kahl, G. in The Dictionary of Genomics, Transcriptomics, and Proteomics (Wiley-VCH, 2015).
Sansone, S. A. et al. FAIRsharing as a community approach to standards, repositories and policies. Nat. Biotechnol. 37, 358–367 (2019). An updated call for FAIR data sharing practices as a community approach to improving scientific research integrity.
Eisenberg, D., Marcotte, E. M., Xenarios, I. & Yeates, T. O. Protein function in the post-genomic era. Nature 405, 823–826 (2000).
Ma’ayan, A., Blitzer, R. D. & Iyengar, R. Toward predictive models of mammalian cells. Annu. Rev. Biophys. Biomol. Struct. 34, 319–349 (2005).
Gosak, M. et al. Network science of biological systems at different scales: a review. Phys. Life Rev. 24, 118–135 (2018).
Ideker, T. & Nussinov, R. Network approaches and applications in biology. PLoS Comput. Biol. 13, e1005771 (2017).
Wickham, H. Tidy data. J. Stat. Softw. https://doi.org/10.18637/jss.v059.i10 (2014). A fundamental treatise on the clear organization and management of data in modelling and statistics.
Chavan, S. S., Shaughnessy, J. D. Jr. & Edmondson, R. D. Overview of biological database mapping services for interoperation between different ‘omics’ datasets. Hum. Genomics 5, 703–708 (2011).
Zhang, Y. et al. Influenza research database: an integrated bioinformatics resource for influenza virus research. Nucleic Acids Res. 45, D466–D474 (2017).
Robertson, D. L. et al. HIV-1 nomenclature proposal. Science 288, 55–56 (2000).
Parker, T. G., Tindall, B. J. & Garrity, G. M. International Code of Nomenclature of Prokaryotes. Int. J. Syst. Evol. Microbiol. 69, S1–S111 (2019).
Kim, M. & Tagkopoulos, I. Data integration and predictive modeling methods for multi-omics datasets. Mol. Omics 14, 8–25 (2018).
D’Argenio, V. The high-throughput analyses era: are we ready for the data struggle? High Throughput 7, 8 (2018).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Sarajlic, A., Malod-Dognin, N., Yaveroglu, O. N. & Przulj, N. Graphlet-based characterization of directed networks. Sci. Rep. 6, 35098 (2016).
Hagberg, A. A., Swart, P. & Schult, D. Exploring network structure, dynamics, and function using NetworkX. in Proc. 7th Python Sci. Conf. (2008).
Huang, S., Chaudhary, K. & Garmire, L. X. More is better: recent progress in multi-omics data integration methods. Front. Genet. 8, 84 (2017).
Tarca, A. L., Carey, V. J., Chen, X. W., Romero, R. & Draghici, S. Machine learning and its applications to biology. PLoS Comput. Biol. 3, e116 (2007).
Zhou, Y. et al. Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat. Commun. 10, 1523 (2019).
Szklarczyk, D. et al. STRING v11: protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 47, D607–D613 (2019).
Huang, D. W. et al. The DAVID gene functional classification tool: a novel biological module-centric algorithm to functionally analyze large gene lists. Genome Biol. 8, R183 (2007).
Mootha, V. K. et al. PGC-1alpha-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat. Genet. 34, 267–273 (2003).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005). The first peer-reviewed report of enrichment analysis as a supervised approach for the interpretation of large biological data sets.
Kanehisa, M. & Goto, S. KEGG: Kyoto Encyclopedia of Genes and Genomes. Nucleic Acids Res. 28, 27–30 (2000).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat. Genet. 25, 25–29 (2000). The first report of the widely used Gene Ontology classifications for human genes to allow standardized interpretation and supervised analysis of genetic data sets.
Foulger, R. E. et al. Representing virus-host interactions and other multi-organism processes in the gene ontology. BMC Microbiol. 15, 146 (2015).
The Gene Ontology Consortium. The gene ontology resource: 20 years and still going strong. Nucleic Acids Res. 47, D330–D338 (2019).
Kavvas, E. S. et al. Machine learning and structural analysis of Mycobacterium tuberculosis pan-genome identifies genetic signatures of antibiotic resistance. Nat. Commun. 9, 4306 (2018).
Hofree, M., Shen, J. P., Carter, H., Gross, A. & Ideker, T. Network-based stratification of tumor mutations. Nat. Methods 10, 1108–1115 (2013).
Leiserson, M. D. et al. Pan-cancer network analysis identifies combinations of rare somatic mutations across pathways and protein complexes. Nat. Genet. 47, 106–114 (2015).
Cotto, K. C. et al. DGIdb 3.0: a redesign and expansion of the drug-gene interaction database. Nucleic Acids Res. 46, D1068–D1073 (2018).
Li, Y. H. et al. Therapeutic target database update 2018: enriched resource for facilitating bench-to-clinic research of targeted therapeutics. Nucleic Acids Res. 46, D1121–D1127 (2018).
Wishart, D. S. et al. DrugBank 5.0: a major update to the DrugBank database for 2018. Nucleic Acids Res. 46, D1074–D1082 (2018).
Whirl-Carrillo, M. et al. Pharmacogenomics knowledge for personalized medicine. Clin. Pharmacol. Ther. 92, 414–417 (2012).
Gaulton, A. et al. The ChEMBL database in 2017. Nucleic Acids Res. 45, D945–D954 (2017).
Janes, J. et al. The ReFRAME library as a comprehensive drug repurposing library and its application to the treatment of cryptosporidiosis. Proc. Natl Acad. Sci. USA 115, 10750–10755 (2018).
Miller, C. H., Nisa, S., Dempsey, S., Jack, C. & O’Toole, R. Modifying culture conditions in chemical library screening identifies alternative inhibitors of mycobacteria. Antimicrob. Agents Chemother. 53, 5279–5283 (2009).
Couture, J. L., Blake, R. E., McDonald, G. & Ward, C. L. A funder-imposed data publication requirement seldom inspired data sharing. PLoS One 13, e0199789 (2018).
Alsheikh-Ali, A. A., Qureshi, W., Al-Mallah, M. H. & Ioannidis, J. P. Public availability of published research data in high-impact journals. PLoS One 6, e24357 (2011).
Vines, T. H. et al. The availability of research data declines rapidly with article age. Curr. Biol. 24, 94–97 (2014).
Savage, C. J. & Vickers, A. J. Empirical study of data sharing by authors publishing in PLoS journals. PLoS One 4, e7078 (2009).
Wilkinson, M. D. et al. The FAIR guiding principles for scientific data management and stewardship. Sci. Data 3, 160018 (2016).
Goncalves, R. S. & Musen, M. A. The variable quality of metadata about biological samples used in biomedical experiments. Sci. Data 6, 190021 (2019).
Chelliah, V. et al. BioModels: ten-year anniversary. Nucleic Acids Res. 43, D542–D548 (2015).
Juty, N. et al. BioModels: content, features, functionality, and use. CPT Pharmacomet. Syst. Pharmacol. 4, e3 (2015).
Pillich, R. T., Chen, J., Rynkov, V., Welker, D. & Pratt, D. NDEx: a community resource for sharing and publishing of biological networks. Methods Mol. Biol. 1558, 271–301 (2017).
Barrett, T. et al. BioProject and BioSample databases at NCBI: facilitating capture and organization of metadata. Nucleic Acids Res. 40, D57–D63 (2012).
Courtot, M. et al. BioSamples database: an updated sample metadata hub. Nucleic Acids Res. 47, D1172–D1178 (2019).
Acknowledgements
J.F.H. is supported by amfAR grant 109504-61-RKRL with funds raised by generationCURE, the Gilead Sciences Research Scholars Program in HIV, US National Institutes of Health (NIH) grant K22 AI136691, a supplement from the NIH-supported Third Coast Center for AIDS Research (P30 AI117943) and a supplement from the NIH-sponsored HARC Center (P50 AI150476). R.M.K. is supported by the NIH-sponsored HARC Center (P50 AI150476) and the NIH-sponsored Host-Pathogen Mapping Initiative (U19 AI135990). R.H. is supported by the US Department of Defense Advanced Research Projects Agency (HR0011-19-2-0020). N.J.K. is supported by the NIH-sponsored HARC Center (P50 AI150476), the NIH-sponsored Host-Pathogen Mapping Initiative (U19 AI135990), the NIH-sponsored FluOMICs consortium (U19 AI135972) and NIH grant P01 AI063302.
Author information
Authors and Affiliations
Contributions
M.E., J.F.H, R.M.K. and R.H. researched the literature. M.E., J.F.H, R.M.K., R.H. and N.J.K. wrote the article, provided substantial contributions to discussions of the content and reviewed and/or edited the manuscript before submission.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information
Nature Reviews Genetics thanks T. Baumert and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
FAIRsharing: https://fairsharing.org/
Scientific Datarecommended data repositories: https://www.nature.com/sdata/policies/repositories
Glossary
- Primary model systems
-
Types of host models that rely on cells taken directly from living tissue (such as from biopsy material or blood) for growth and maintenance ex vivo.
- Laboratory-adapted strain
-
A genetically distinct strain of a pathogen that has been selected for enhanced fitness ex vivo and for use in laboratory experiments even though it is not found as a major strain in the natural world.
- Clinical isolates
-
Genetic strains of pathogens isolated directly from patients or clinical samples.
- Technical replicates
-
Repeated experiments analysing the same sample with the same instrumentation to measure the variability inherent in the testing protocol.
- Biological replicates
-
Repeated experiments analysing different samples that represent the same thing (such as samples collected from different patients with the same disease outcome) to determine the variability in the sample pools.
- Confounding effects
-
The influence of one or more unmonitored variables on a system’s components or the relationships between those components that can alter experimental interpretation.
- Saturating mutagenesis
-
A genetic screening technique wherein a codon or set of codons is randomized to produce all possible amino acids at a position or positions.
- Host–pathogen co-evolution
-
Iterative rounds of adaptation and counter-adaptation between a pathogen and its host over evolutionary history as a result of the ability of pathogens to elicit selective pressure on their host populations and vice versa.
- Transposon mutagenesis
-
A method for the random disruption of gene function by the untargeted insertion of transposable retroelements into a genome.
- Metadata
-
Information that describes a set of data.
- Multiplicity of infection
-
The ratio of infectious agents (such as virions or bacteria) to infection targets (such as cells).
- Nodes
-
A connection point in a network representing a component of the system.
- Edges
-
A connection between nodes in a network representing a relationship between two components.
- Enrichment analysis
-
An approach for identifying over-represented classifications of components by comparing the frequency of a given annotation in a data set with a predefined reference list.
- k-means clustering
-
A method of data clustering that aims to partition a set of components into a total of k clusters, wherein each component belongs to the cluster with the nearest mean value.
- Principal component analysis
-
A statistical procedure often used in the development of predictive models, which describes a data set as a series of uncorrelated variables called ‘principal components’ that account for sources of variability.
- Support vector machines
-
A machine learning method related to regression analysis that seeks to identify the separation boundary between clusters of data given predefined clusters in a prelabelled set of input data.
- Neural networks
-
A machine learning method that seeks to cluster and classify data on the basis of similarities and differences extracted from a prelabelled set of input data.
- Random forests
-
A machine learning algorithm that seeks to cluster and classify data on the basis of the ensemble output of a series of decision trees formulated from a prelabelled set of input data.
- Mutual information
-
A measurement of dependency between two variables that is used in machine learning to determine how much can be assumed about one component on the basis of the observed behaviour of another.
- Phenotypic selection
-
Isolation of a given cell population based on an observed trait or characteristic (such as fluorescence or resistance to a toxic compound).
Rights and permissions
About this article
Cite this article
Eckhardt, M., Hultquist, J.F., Kaake, R.M. et al. A systems approach to infectious disease. Nat Rev Genet 21, 339–354 (2020). https://doi.org/10.1038/s41576-020-0212-5
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41576-020-0212-5
This article is cited by
-
Development and psychometric evaluation of nurse’s intention to care for patients with infectious disease scale: an exploratory sequential mixed method study
BMC Nursing (2024)
-
Next-generation proteomics for quantitative Jumbophage-bacteria interaction mapping
Nature Communications (2023)
-
Prioritizing non-communicable diseases in the post-pandemic era based on a comprehensive analysis of the GBD 2019 from 1990 to 2019
Scientific Reports (2023)
-
Proteomic and genetic analyses of influenza A viruses identify pan-viral host targets
Nature Communications (2023)
-
From systems to structure — using genetic data to model protein structures
Nature Reviews Genetics (2022)