Abstract
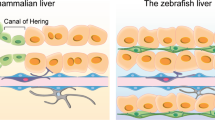
The liver is the only solid organ that uses regenerative mechanisms to ensure that the liver-to-bodyweight ratio is always at 100% of what is required for body homeostasis. Other solid organs (such as the lungs, kidneys and pancreas) adjust to tissue loss but do not return to 100% of normal. The current state of knowledge of the regenerative pathways that underlie this ‘hepatostat’ will be presented in this Review. Liver regeneration from acute injury is always beneficial and has been extensively studied. Experimental models that involve partial hepatectomy or chemical injury have revealed extracellular and intracellular signalling pathways that are used to return the liver to equivalent size and weight to those prior to injury. On the other hand, chronic loss of hepatocytes, which can occur in chronic liver disease of any aetiology, often has adverse consequences, including fibrosis, cirrhosis and liver neoplasia. The regenerative activities of hepatocytes and cholangiocytes are typically characterized by phenotypic fidelity. However, when regeneration of one of the two cell types fails, hepatocytes and cholangiocytes function as facultative stem cells and transdifferentiate into each other to restore normal liver structure. Liver recolonization models have demonstrated that hepatocytes have an unlimited regenerative capacity. However, in normal liver, cell turnover is very slow. All zones of the resting liver lobules have been equally implicated in the maintenance of hepatocyte and cholangiocyte populations in normal liver.
Key points
-
Hepatocyte proliferation during liver regeneration is controlled by multiple extracellular signals, two of which (MET and EGFR) are directly mitogenic and others only delay liver regeneration if they are bypassed.
-
Intracellular signalling pathways in hepatocytes are very rapidly (within minutes) activated after partial hepatectomy. The mechanisms triggering these pathways are not clear.
-
All hepatic cell types participate in cell proliferation during liver regeneration. No ‘stem cells’ are involved.
-
If hepatocyte or cholangiocyte proliferation is seriously impaired, then each of the two cell types can transdifferentiate into the other and function as a facultative stem cell.
-
Loss of hepatocytes occurring in chronic liver diseases triggers compensatory proliferation of the surviving hepatocytes and exposes them to potentially genotoxic injury that might lead to neoplasia.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Michalopoulos, G. K. Liver regeneration. J. Cell Physiol. 213, 286–300 (2007).
Michalopoulos, G. K. & DeFrances, M. C. Liver regeneration. Science 276, 60–66 (1997).
Michalopoulos, G. K. Principles of liver regeneration and growth homeostasis. Compr. Physiol. 3, 485–513 (2013).
Fausto, N., Campbell, J. S. & Riehle, K. J. Liver regeneration. Hepatology 43, S45–S53 (2006).
Higgins, G., Anderson, R. E., Higgins, G. M. & Anderson, R. M. Experimental pathology of the liver, 1: restoration of the liver of the white rat following partial surgical removal. Arch. Pathol. 12, 186–202 (1931).
Demetriou, A. A. et al. Transplantation of microcarrier-attached hepatocytes into 90% partially hepatectomized rats. Hepatology 8, 1006–1009 (1988).
Demetris, A. J. et al. Pathophysiologic observations and histopathologic recognition of the portal hyperperfusion or small-for-size syndrome. Am. J. Surg. Pathol. 30, 986–993 (2006).
Miyaoka, Y. & Miyajima, A. To divide or not to divide: revisiting liver regeneration. Cell Div. 8, 8 (2013).
DeLeve, L. D., Wang, X. & Wang, L. VEGF-sdf1 recruitment of CXCR7+ bone marrow progenitors of liver sinusoidal endothelial cells promotes rat liver regeneration. Am. J. Physiol. Gastrointest. Liver Physiol 310, G739–G746 (2016).
Fujii, H. et al. Contribution of bone marrow cells to liver regeneration after partial hepatectomy in mice. J. Hepatol. 36, 653–659 (2002).
Bonnardel, J. et al. Stellate cells, hepatocytes, and endothelial cells imprint the Kupffer Cell identity on monocytes colonizing the liver macrophage niche. Immunity 51, 638–654.e9 (2019).
Marubashi, S. et al. Effect of portal hemodynamics on liver regeneration studied in a novel portohepatic shunt rat model. Surgery 136, 1028–1037 (2004).
Preziosi, M., Okabe, H., Poddar, M., Singh, S. & Monga, S. P. Endothelial Wnts regulate β-catenin signaling in murine liver zonation and regeneration: a sequel to the Wnt-Wnt situation. Hepatol. Commun. 2, 845–860 (2018).
Russell, J. O. & Monga, S. P. Wnt/β-catenin signaling in liver development, homeostasis, and pathobiology. Annu. Rev. Pathol. 13, 351–378 (2018).
Rappaport, A. M. The microcirculatory hepatic unit. Microvasc. Res. 6, 212–228 (1973).
Wake, K. Hepatic stellate cells: three-dimensional structure, localization, heterogeneity and development. Proc. Jpn. Acad. Ser. B Phys. Biol. Sci. 82, 155–164 (2006).
Oda, M., Yokomori, H. & Han, J. Y. Regulatory mechanisms of hepatic microcirculatory hemodynamics: hepatic arterial system. Clin. Hemorheol. Microcirc. 34, 11–26 (2006).
Mars, W. M. et al. Immediate early detection of urokinase receptor after partial hepatectomy and its implications for initiation of liver regeneration. Hepatology 21, 1695–1701 (1995).
Kim, T. H., Mars, W. M., Stolz, D. B. & Michalopoulos, G. K. Expression and activation of pro-MMP-2 and pro-MMP-9 during rat liver regeneration. Hepatology 31, 75–82 (2000).
Kim, T. H., Mars, W. M., Stolz, D. B., Petersen, B. E. & Michalopoulos, G. K. Extracellular matrix remodeling at the early stages of liver regeneration in the rat. Hepatology 26, 896–904 (1997).
Nejak-Bowen, K., Orr, A., Bowen, W. C. Jr. & Michalopoulos, G. K. Conditional genetic elimination of hepatocyte growth factor in mice compromises liver regeneration after partial hepatectomy. PLoS ONE 8, e59836 (2013).
Mars, W. M., Kim, T. H., Stolz, D. B., Liu, M. L. & Michalopoulos, G. K. Presence of urokinase in serum-free primary rat hepatocyte cultures and its role in activating hepatocyte growth factor. Cancer Res. 56, 2837–2843 (1996).
Lindroos, P. M., Zarnegar, R. & Michalopoulos, G. K. Hepatocyte growth factor (hepatopoietin A) rapidly increases in plasma before DNA synthesis and liver regeneration stimulated by partial hepatectomy and carbon tetrachloride administration. Hepatology 13, 743–750 (1991).
Saegusa, S., Isaji, S. & Kawarada, Y. Changes in serum hyaluronic acid levels and expression of CD44 and CD44 mRNA in hepatic sinusoidal endothelial cells after major hepatectomy in cirrhotic rats. World J. Surg. 26, 694–699 (2002).
Roselli, H. T. et al. Liver regeneration is transiently impaired in urokinase-deficient mice. Am. J. Physiol. 275, G1472–G1479 (1998).
Lieber, A. et al. Adenovirus-mediated urokinase gene transfer induces liver regeneration and allows for efficient retrovirus transduction of hepatocytes in vivo. Proc. Natl Acad. Sci. USA 92, 6210–6214 (1995).
Mohammed, F. F. & Khokha, R. Thinking outside the cell: proteases regulate hepatocyte division. Trends Cell Biol. 15, 555–563 (2005).
Rudolph, K. L. et al. Differential regulation of extracellular matrix synthesis during liver regeneration after partial hepatectomy in rats. Hepatology 30, 1159–1166 (1999).
Gallai, M. et al. Proteoglycan gene expression in rat liver after partial hepatectomy. Biochem. Biophys. Res. Commun. 228, 690–694 (1996).
Weymann, A. et al. p21 is required for dextrose-mediated inhibition of mouse liver regeneration. Hepatology 50, 207–215 (2009).
Moolten, F. L. & Bucher, N. L. Regeneration of rat liver: transfer of humoral agent by cross circulation. Science 158, 272–274 (1967).
Jirtle, R. L. & Michalopoulos, G. Effects of partial hepatectomy on transplanted hepatocytes. Cancer Res. 42, 3000–3004 (1982).
Kohler, C. et al. Expression of Notch-1 and its ligand Jagged-1 in rat liver during liver regeneration. Hepatology 39, 1056–1065 (2004).
Monga, S. P., Pediaditakis, P., Mule, K., Stolz, D. B. & Michalopoulos, G. K. Changes in WNT/β-catenin pathway during regulated growth in rat liver regeneration. Hepatology 33, 1098–1109 (2001).
Stolz, D. B., Mars, W. M., Petersen, B. E., Kim, T. H. & Michalopoulos, G. K. Growth factor signal transduction immediately after two-thirds partial hepatectomy in the rat. Cancer Res. 59, 3954–3960 (1999).
Taub, R. Liver regeneration 4: transcriptional control of liver regeneration. FASEB J. 10, 413–427 (1996).
Apte, U. et al. Enhanced liver regeneration following changes induced by hepatocyte-specific genetic ablation of integrin-linked kinase. Hepatology 50, 844–851 (2009).
Desbarats, J. & Newell, M. K. Fas engagement accelerates liver regeneration after partial hepatectomy. Nat. Med. 6, 920–923 (2000).
Albrecht, J. H. et al. Involvement of p21 and p27 in the regulation of CDK activity and cell cycle progression in the regenerating liver. Oncogene 16, 2141–2150 (1998).
Bhave, V. S. et al. Genes inducing iPS phenotype play a role in hepatocyte survival and proliferation in vitro and liver regeneration in vivo. Hepatology 54, 1360–1370 (2011).
Mullany, L. K. et al. Distinct proliferative and transcriptional effects of the D-type cyclins in vivo. Cell Cycle 7, 2215–2224 (2008).
Russell, W. E., Kaufmann, W. K., Sitaric, S., Luetteke, N. C. & Lee, D. C. Liver regeneration and hepatocarcinogenesis in transforming growth factor-alpha-targeted mice. Mol. Carcinog. 15, 183–189 (1996).
Taub, R. Liver regeneration: from myth to mechanism. Nat. Rev. Mol. Cell Biol. 5, 836–847 (2004).
Paranjpe, S. et al. Combined systemic elimination of MET and epidermal growth factor receptor signaling completely abolishes liver regeneration and leads to liver decompensation. Hepatology 64, 1711–1724 (2016).
Greenbaum, L. E., Cressman, D. E., Haber, B. A. & Taub, R. Coexistence of C/EBP alpha, beta, growth-induced proteins and DNA synthesis in hepatocytes during liver regeneration. Implications for maintenance of the differentiated state during liver growth. J. Clin. Invest. 96, 1351–1365 (1995).
Wang, X. et al. Rapid hepatocyte nuclear translocation of the Forkhead Box M1B (FoxM1B) transcription factor caused a transient increase in size of regenerating transgenic hepatocytes. Gene Expr. 11, 149–162 (2003).
Klochendler, A. et al. A transgenic mouse marking live replicating cells reveals in vivo transcriptional program of proliferation. Dev. Cell 23, 681–690 (2012).
Rabes, H. M. Kinetics of hepatocellular proliferation as a function of the microvascular structure and functional state of the liver. Ciba Found. Symp. https://doi.org/10.1002/9780470720363.ch3 (1977).
Volk, A., Michalopoulos, G., Weidner, M. & Gebhardt, R. Different proliferative responses of periportal and pericentral rat hepatocytes to hepatocyte growth factor. Biochem. Biophys. Res. Commun. 207, 578–584 (1995).
Stocker, E. & Heine, W. D. Regeneration of liver parenchyma under normal and pathological conditions. Beitr. Pathol. 144, 400–408 (1971).
Biondo-Simoes Mde, L. et al. Effect of aging on liver regeneration in rats. Acta Cir. Bras. 21, 197–202 (2006).
Matsumoto, T., Wakefield, L., Tarlow, B. D. & Grompe, M. In vivo lineage tracing of polyploid hepatocytes reveals extensive proliferation during liver regeneration. Cell Stem Cell 26, 34–47.e3 (2020).
Trusolino, L., Bertotti, A. & Comoglio, P. M. MET signalling: principles and functions in development, organ regeneration and cancer. Nat. Rev. Mol. Cell Biol. 11, 834–848 (2010).
Gentile, A., Trusolino, L. & Comoglio, P. M. The Met tyrosine kinase receptor in development and cancer. Cancer Metastasis Rev. 27, 85–94 (2008).
Liu, M. L., Mars, W. M., Zarnegar, R. & Michalopoulos, G. K. Uptake and distribution of hepatocyte growth factor in normal and regenerating adult rat liver. Am. J. Pathol. 144, 129–140 (1994).
Bard-Chapeau, E. A. et al. Concerted functions of Gab1 and Shp2 in liver regeneration and hepatoprotection. Mol. Cell Biol. 26, 4664–4674 (2006).
Zarnegar, R., DeFrances, M. C., Kost, D. P., Lindroos, P. & Michalopoulos, G. K. Expression of hepatocyte growth factor mRNA in regenerating rat liver after partial hepatectomy. Biochem. Biophys. Res. Commun. 177, 559–565 (1991).
Kono, S., Nagaike, M., Matsumoto, K. & Nakamura, T. Marked induction of hepatocyte growth factor mRNA in intact kidney and spleen in response to injury of distant organs. Biochem. Biophys. Res. Commun. 186, 991–998 (1992).
Yanagita, K. et al. Lung may have an endocrine function producing hepatocyte growth factor in response to injury of distal organs. Biochem. Biophys. Res. Commun. 182, 802–809 (1992).
Broten, J., Michalopoulos, G., Petersen, B. & Cruise, J. Adrenergic stimulation of hepatocyte growth factor expression. Biochem. Biophys. Res. Commun. 262, 76–79 (1999).
Passino, M. A., Adams, R. A., Sikorski, S. L. & Akassoglou, K. Regulation of hepatic stellate cell differentiation by the neurotrophin receptor p75NTR. Science 315, 1853–1856 (2007).
Carver, R. S., Stevenson, M. C., Scheving, L. A. & Russell, W. E. Diverse expression of ErbB receptor proteins during rat liver development and regeneration. Gastroenterology 123, 2017–2027 (2002).
Paranjpe, S. et al. RNA interference against hepatic epidermal growth factor receptor has suppressive effects on liver regeneration in rats. Am. J. Pathol. 176, 2669–2681 (2010).
Odegard, J. et al. Differential effects of epidermal growth factor (EGF) receptor ligands on receptor binding, downstream signalling pathways and DNA synthesis in hepatocytes. Growth Factors 35, 239–248 (2017).
Olsen, P. S., Poulsen, S. S. & Kirkegaard, P. Adrenergic effects on secretion of epidermal growth factor from Brunner’s glands. Gut 26, 920–927 (1985).
Skov Olsen, P. et al. Influence of epidermal growth factor on liver regeneration after partial hepatectomy in rats. Hepatology 8, 992–996 (1988).
Dao, D. T., Anez-Bustillos, L., Adam, R. M., Puder, M. & Bielenberg, D. R. Heparin-binding epidermal growth factor-like growth factor as a critical mediator of tissue repair and regeneration. Am. J. Pathol. 188, 2446–2456 (2018).
Webber, E. M., FitzGerald, M. J., Brown, P. I., Bartlett, M. H. & Fausto, N. Transforming growth factor-alpha expression during liver regeneration after partial hepatectomy and toxic injury, and potential interactions between transforming growth factor-alpha and hepatocyte growth factor. Hepatology 18, 1422–1431 (1993).
Lee, D. C. et al. TACE/ADAM17 processing of EGFR ligands indicates a role as a physiological convertase. Ann. NY Acad. Sci. 995, 22–38 (2003).
Berasain, C. et al. Amphiregulin: an early trigger of liver regeneration in mice. Gastroenterology 128, 424–432 (2005).
Khai, N. C. et al. In vivo hepatic HB-EGF gene transduction inhibits Fas-induced liver injury and induces liver regeneration in mice: a comparative study to HGF. J. Hepatol. 44, 1046–1054 (2006).
Mitchell, C. et al. Heparin-binding epidermal growth factor-like growth factor links hepatocyte priming with cell cycle progression during liver regeneration. J. Biol. Chem. 280, 2562–2568 (2005).
Maretti-Mira, A. C., Wang, X., Wang, L. & DeLeve, L. D. Incomplete differentiation of engrafted bone marrow endothelial progenitor cells initiates hepatic fibrosis in the rat. Hepatology 69, 1259–1272 (2019).
Natarajan, A., Wagner, B. & Sibilia, M. The EGF receptor is required for efficient liver regeneration. Proc. Natl Acad. Sci. USA 104, 17081–17086 (2007).
Scheving, L. A., Zhang, X., Stevenson, M. C., Threadgill, D. W. & Russell, W. E. Loss of hepatocyte EGFR has no effect alone but exacerbates carbon tetrachloride-induced liver injury and impairs regeneration in hepatocyte Met-deficient mice. Am. J. Physiol. Gastrointest. Liver Physiol. 308, G364–G377 (2015).
Normanno, N. et al. Epidermal growth factor receptor (EGFR) signaling in cancer. Gene 366, 2–16 (2006).
Jo, M. et al. Cross-talk between epidermal growth factor receptor and c-Met signal pathways in transformed cells. J. Biol. Chem. 275, 8806–8811 (2000).
Tsagianni, A. et al. Combined systemic disruption of MET and epidermal growth factor receptor signaling causes liver failure in normal mice. Am. J. Pathol. 88, 2223–2235 (2018).
Limaye, P. B. et al. Mechanisms of hepatocyte growth factor-mediated and epidermal growth factor-mediated signaling in transdifferentiation of rat hepatocytes to biliary epithelium. Hepatology 47, 1702–1713 (2008).
Bhushan, B. et al. TCPOBOP-induced hepatomegaly and hepatocyte proliferation are attenuated by combined disruption of MET and EGFR signaling. Hepatology 69, 1702–1718 (2019).
Houck, K. A., Zarnegar, R., Muga, S. J. & Michalopoulos, G. K. Acidic fibroblast growth factor (HBGF-1) stimulates DNA synthesis in primary rat hepatocyte cultures. J. Cell Physiol. 143, 129–132 (1990).
Kan, M. et al. Heparin-binding growth factor type 1 (acidic fibroblast growth factor): a potential biphasic autocrine and paracrine regulator of hepatocyte regeneration. Proc. Natl Acad. Sci. USA 86, 7432–7436 (1989).
Huang, X. et al. Ectopic activity of fibroblast growth factor receptor 1 in hepatocytes accelerates hepatocarcinogenesis by driving proliferation and vascular endothelial growth factor-induced angiogenesis. Cancer Res. 66, 1481–1490 (2006).
Luo, Y. et al. Metabolic regulator βKlotho interacts with fibroblast growth factor receptor 4 (FGFR4) to induce apoptosis and inhibit tumor cell proliferation. J. Biol. Chem. 285, 30069–30078 (2010).
Padrissa-Altes, S. et al. Control of hepatocyte proliferation and survival by Fgf receptors is essential for liver regeneration in mice. Gut 64, 1444–1453 (2015).
Cicione, C., Degirolamo, C. & Moschetta, A. Emerging role of fibroblast growth factors 15/19 and 21 as metabolic integrators in the liver. Hepatology 56, 2404–2411 (2012).
Kong, B. et al. Fibroblast growth factor 15 deficiency impairs liver regeneration in mice. Am. J. Physiol. Gastrointest. Liver Physiol 306, G893–G902 (2014).
Yamada, Y., Webber, E. M., Kirillova, I., Peschon, J. J. & Fausto, N. Analysis of liver regeneration in mice lacking type 1 or type 2 tumor necrosis factor receptor: requirement for type 1 but not type 2 receptor. Hepatology 28, 959–970 (1998).
Kirillova, I., Chaisson, M. & Fausto, N. Tumor necrosis factor induces DNA replication in hepatic cells through nuclear factor κB activation. Cell Growth Differ. 10, 819–828 (1999).
Cressman, D. E. et al. Liver failure and defective hepatocyte regeneration in interleukin-6- deficient mice. Science 274, 1379–1383 (1996).
Norris, C. A. et al. Synthesis of IL-6 by hepatocytes is a normal response to common hepatic stimuli. PLoS ONE 9, e96053 (2014).
Fausto, N. Liver regeneration. J. Hepatol. 32, 19–31 (2000).
Runge, D. M., Runge, D., Foth, H., Strom, S. C. & Michalopoulos, G. K. STAT 1alpha/1beta, STAT 3 and STAT 5: expression and association with c- MET and EGF-receptor in long-term cultures of human hepatocytes. Biochem. Biophys. Res. Commun. 265, 376–381 (1999).
Cruise, J. L., Knechtle, S. J., Bollinger, R. R., Kuhn, C. & Michalopoulos, G. Alpha 1-adrenergic effects and liver regeneration. Hepatology 7, 1189–1194 (1987).
Cruise, J. L., Houck, K. A. & Michalopoulos, G. K. Induction of DNA synthesis in cultured rat hepatocytes through stimulation of alpha 1 adrenoreceptor by norepinephrine. Science 227, 749–751 (1985).
Han, C., Bowen, W. C., Michalopoulos, G. K. & Wu, T. Alpha-1 adrenergic receptor transactivates signal transducer and activator of transcription-3 (Stat3) through activation of Src and epidermal growth factor receptor (EGFR) in hepatocytes. J. Cell Physiol. 216, 486–497 (2008).
Houck, K. A., Cruise, J. L. & Michalopoulos, G. Norepinephrine modulates the growth-inhibitory effect of transforming growth factor-beta in primary rat hepatocyte cultures. J. Cell Physiol. 135, 551–555 (1988).
Huang, W. et al. Nuclear receptor-dependent bile acid signaling is required for normal liver regeneration. Science 312, 233–236 (2006).
Borude, P. et al. Hepatocyte-specific deletion of farnesoid X receptor delays but does not inhibit liver regeneration after partial hepatectomy in mice. Hepatology 56, 2344–2352 (2012).
Block, G. D. et al. Population expansion, clonal growth, and specific differentiation patterns in primary cultures of hepatocytes induced by HGF/SF, EGF and TGF alpha in a chemically defined (HGM) medium. J. Cell Biol. 132, 1133–1149 (1996).
Francavilla, A. et al. Screening for candidate hepatic growth factors by selective portal infusion after canine Eck’s fistula. Hepatology 14, 665–670 (1991).
Fafalios, A. et al. A hepatocyte growth factor receptor (Met)-insulin receptor hybrid governs hepatic glucose metabolism. Nat. Med. 17, 1577–1584 (2011).
Planas-Paz, L. et al. The RSPO-LGR4/5-ZNRF3/RNF43 module controls liver zonation and size. Nat. Cell Biol. 18, 467–479 (2016).
Capurro, M., Martin, T., Shi, W. & Filmus, J. Glypican-3 binds to Frizzled and plays a direct role in the stimulation of canonical Wnt signaling. J. Cell Sci. 127, 1565–1575 (2014).
Li, N. et al. A frizzled-like cysteine-rich domain in glypican-3 mediates wnt binding and regulates hepatocellular carcinoma tumor growth in mice. Hepatology 70, 1231–1245 (2019).
Monga, S. P. et al. Hepatocyte growth factor induces Wnt-independent nuclear translocation of beta-catenin after Met-beta-catenin dissociation in hepatocytes. Cancer Res. 62, 2064–2071 (2002).
Tetsu, O. & McCormick, F. Beta-catenin regulates expression of cyclin D1 in colon carcinoma cells. Nature 398, 422–426 (1999).
Yang, J. et al. β-catenin signaling in murine liver zonation and regeneration: a Wnt-Wnt situation! Hepatology 60, 964–976 (2014).
Ochoa, B. et al. Hedgehog signaling is critical for normal liver regeneration after partial hepatectomy in mice. Hepatology 51, 1712–1723 (2010).
Swiderska-Syn, M. et al. Hedgehog regulates yes-associated protein 1 in regenerating mouse liver. Hepatology 64, 232–244 (2016).
Liu, B. et al. Suppression of liver regeneration and hepatocyte proliferation in hepatocyte-targeted glypican 3 transgenic mice. Hepatology 52, 1060–1067 (2010).
Bhave, V. S. et al. Regulation of liver growth by glypican 3, CD81, hedgehog, and Hhex. Am. J. Pathol. 183, 153–159 (2013).
Machado, M. V. & Diehl, A. M. Hedgehog signalling in liver pathophysiology. J. Hepatol. 68, 550–562 (2018).
Li, W. et al. A homeostatic Arid1a-dependent permissive chromatin state licenses hepatocyte responsiveness to liver-injury-associated YAP signaling. Cell Stem Cell 25, 54–68 (2019).
Septer, S. et al. Yes-associated protein is involved in proliferation and differentiation during postnatal liver development. Am. J. Physiol. Gastrointest. Liver Physiol 302, G493–G503 (2012).
Halder, G. & Johnson, R. L. Hippo signaling: growth control and beyond. Development 138, 9–22 (2011).
Patel, S. H., Camargo, F. D. & Yimlamai, D. Hippo signaling in the liver regulates organ size, cell fate, and carcinogenesis. Gastroenterology 152, 533–545 (2017).
Grijalva, J. L. et al. Dynamic alterations in Hippo signaling pathway and YAP activation during liver regeneration. Am. J. Physiol. Gastrointest. Liver Physiol 307, G196–G204 (2014).
Oh, S. H., Swiderska-Syn, M., Jewell, M. L., Premont, R. T. & Diehl, A. M. Liver regeneration requires Yap1-TGFbeta-dependent epithelial-mesenchymal transition in hepatocytes. J. Hepatol. 69, 359–367 (2018).
Xue, Y. et al. Hepatitis C virus mimics effects of glypican-3 on CD81 and promotes development of hepatocellular carcinomas via activation of hippo pathway in hepatocytes. Am. J. Pathol. 188, 1469–1477 (2018).
Ferdous, Z., Wei, V. M., Iozzo, R., Hook, M. & Grande-Allen, K. J. Decorin-transforming growth factor- interaction regulates matrix organization and mechanical characteristics of three-dimensional collagen matrices. J. Biol. Chem. 282, 35887–35898 (2007).
Jirtle, R. L., Carr, B. I. & Scott, C. D. Modulation of insulin-like growth factor-II/mannose 6-phosphate receptors and transforming growth factor-beta 1 during liver regeneration. J. Biol. Chem. 266, 22444–22450 (1991).
Jakowlew, S. B. et al. Transforming growth factor-beta (TGF-beta) isoforms in rat liver regeneration: messenger RNA expression and activation of latent TGF- beta. Cell Regul. 2, 535–548 (1991).
Chari, R. S., Price, D. T., Sue, S. R., Meyers, W. C. & Jirtle, R. L. Down-regulation of transforming growth factor beta receptor type I, II, and III during liver regeneration. Am. J. Surg. 169, 126–131 (1995).
Thenappan, A. et al. Loss of transforming growth factor beta adaptor protein β-2 spectrin leads to delayed liver regeneration in mice. Hepatology 53, 1641–1650 (2011).
Pepper, M. S., Vassalli, J. D., Orci, L. & Montesano, R. Biphasic effect of transforming growth factor-beta 1 on in vitro angiogenesis. Exp. Cell Res. 204, 356–363 (1993).
Hu, J. et al. Endothelial cell-derived angiopoietin-2 controls liver regeneration as a spatiotemporal rheostat. Science 343, 416–419 (2014).
Ichikawa, T. et al. Transforming growth factor beta and activin tonically inhibit DNA synthesis in the rat liver. Hepatology 34, 918–925 (2001).
Carpentier, R. et al. Embryonic ductal plate cells give rise to cholangiocytes, periportal hepatocytes, and adult liver progenitor cells. Gastroenterology 141, 1432–1438 (2011).
Clotman, F. et al. Control of liver cell fate decision by a gradient of TGF beta signaling modulated by Onecut transcription factors. Genes. Dev. 19, 1849–1854 (2005).
Clotman, F. et al. The onecut transcription factor HNF6 is required for normal development of the biliary tract. Development 129, 1819–1828 (2002).
Grisham, J. A morphologic study of deoxyribonucleic acid synthesis and cell proliferation in regenerating liver; autoradiography with thymidine-H3. Cancer Res. 22, 842–849 (1962).
Matsumoto, K., Fujii, H., Michalopoulos, G., Fung, J. J. & Demetris, A. J. Human biliary epithelial cells secrete and respond to cytokines and hepatocyte growth factors in vitro: interleukin-6, hepatocyte growth factor and epidermal growth factor promote DNA synthesis in vitro. Hepatology 20, 376–382 (1994).
Keitel, V. & Haussinger, D. TGR5 in the biliary tree. Dig. Dis. 29, 45–47 (2011).
Pean, N. et al. The receptor TGR5 protects the liver from bile acid overload during liver regeneration in mice. Hepatology 58, 1451–1460 (2013).
Glaser, S., Han, Y., Francis, H. & Alpini, G. Melatonin regulation of biliary functions. Hepatobiliary Surg. Nutr. 3, 35–43 (2014).
Johnson, C. et al. Histamine restores biliary mass following carbon tetrachloride-induced damage in a cholestatic rat model. Dig. Liver Dis. 47, 211–217 (2015).
Michalopoulos, G. K., Barua, L. & Bowen, W. C. Transdifferentiation of rat hepatocytes into biliary cells after bile duct ligation and toxic biliary injury. Hepatology 41, 535–544 (2005).
Pepe-Mooney, B. J. et al. Single-cell analysis of the liver epithelium reveals dynamic heterogeneity and an essential role for YAP in homeostasis and regeneration. Cell Stem Cell 25, 23–38.e8 (2019).
Fouassier, L. & Fiorotto, R. Ezrin finds its groove in cholangiocytes. Hepatology 61, 1467–1470 (2015).
Ross, M. A., Sander, C. M., Kleeb, T. B., Watkins, S. C. & Stolz, D. B. Spatiotemporal expression of angiogenesis growth factor receptors during the revascularization of regenerating rat liver. Hepatology 34, 1135–1148 (2001).
LeCouter, J. et al. Angiogenesis-independent endothelial protection of liver: role of VEGFR-1. Science 299, 890–893 (2003).
Ding, B. S. et al. Inductive angiocrine signals from sinusoidal endothelium are required for liver regeneration. Nature 468, 310–315 (2010).
Rocha, A. S. et al. The angiocrine factor rspondin3 is a key determinant of liver zonation. Cell Rep. 13, 1757–1764 (2015).
Wang, L. et al. Liver sinusoidal endothelial cell progenitor cells promote liver regeneration in rats. J. Clin. Invest. 122, 1567–1573 (2012).
Ikarashi, M. et al. Distinct development and functions of resident and recruited liver Kupffer cells/macrophages. J. Leukoc. Biol. 94, 1325–1336 (2013).
Nishiyama, K. et al. Mouse CD11b+ Kupffer cells recruited from bone marrow accelerate liver regeneration after partial hepatectomy. PLoS ONE 10, e0136774 (2015).
Li, N. & Hua, J. Immune cells in liver regeneration. Oncotarget 8, 3628–3639 (2017).
Oben, J. A. et al. Hepatic fibrogenesis requires sympathetic neurotransmitters. Gut 53, 438–445 (2004).
Tsuchida, T. & Friedman, S. L. Mechanisms of hepatic stellate cell activation. Nat. Rev. Gastroenterol. Hepatol. 14, 397–411 (2017).
Gkretsi, V. et al. Liver-specific ablation of integrin-linked kinase in mice results in abnormal histology, enhanced cell proliferation, and hepatomegaly. Hepatology 48, 1932–1941 (2008).
Gkretsi, V., Bowen, W. C., Yang, Y., Wu, C. & Michalopoulos, G. K. Integrin-linked kinase is involved in matrix-induced hepatocyte differentiation. Biochem. Biophys. Res. Commun. 353, 638–643 (2007).
Donthamsetty, S. et al. Role of PINCH and its partner tumor suppressor Rsu-1 in regulating liver size and tumorigenesis. PLoS ONE 8, e74625 (2013).
Oe, S. et al. Intact signaling by transforming growth factor beta is not required for termination of liver regeneration in mice. Hepatology 40, 1098–1105 (2004).
Yang, J. et al. WNT5A inhibits hepatocyte proliferation and concludes beta-catenin signaling in liver regeneration. Am. J. Pathol. 185, 2194–2205 (2015).
Huck, I., Gunewardena, S., Espanol-Suner, R., Willenbring, H. & Apte, U. Hepatocyte nuclear factor 4 alpha activation is essential for termination of liver regeneration in mice. Hepatology 70, 666–681 (2019).
Jin, J. et al. Cooperation of C/EBP family proteins and chromatin remodeling proteins is essential for termination of liver regeneration. Hepatology 61, 315–325 (2015).
Michalopoulos, G. K. & Khan, Z. Liver stem cells: experimental findings and implications for human liver disease. Gastroenterology 149, 876–882 (2015).
Trautwein, C. et al. 2-acetaminofluorene blocks cell cycle progression after hepatectomy by p21 induction and lack of cyclin E expression. Oncogene 18, 6443–6453 (1999).
Evarts, R. P. et al. Precursor-product relationship between oval cells and hepatocytes: comparison between tritiated thymidine and bromodeoxyuridine as tracers. Carcinogenesis 17, 2143–2151 (1996).
Lu, W. Y. et al. Hepatic progenitor cells of biliary origin with liver repopulation capacity. Nat. Cell Biol. 17, 971–983 (2015).
Raven, A. et al. Cholangiocytes act as facultative liver stem cells during impaired hepatocyte regeneration. Nature 547, 350–354 (2017).
Deng, X. et al. Chronic liver injury induces conversion of biliary epithelial cells into hepatocytes. Cell Stem Cell 23, 114–122.e3 (2018).
Russell, J. O. et al. Hepatocyte-specific beta-catenin deletion during severe liver injury provokes cholangiocytes to differentiate into hepatocytes. Hepatology 69, 742–759 (2019).
Dorrell, C. et al. Prospective isolation of a bipotential clonogenic liver progenitor cell in adult mice. Genes. Dev. 25, 1193–1203 (2011).
Li, B. et al. Adult mouse liver contains two distinct populations of cholangiocytes. Stem Cell Rep. 9, 478–489 (2017).
Isse, K. et al. Preexisting epithelial diversity in normal human livers: a tissue-tethered cytometric analysis in portal/periportal epithelial cells. Hepatology 57, 1632–1643 (2013).
Aizarani, N. et al. A human liver cell atlas reveals heterogeneity and epithelial progenitors. Nature 572, 199–204 (2019).
Petersen, B. E., Zajac, V. F. & Michalopoulos, G. K. Hepatic oval cell activation in response to injury following chemically induced periportal or pericentral damage in rats. Hepatology 27, 1030–1038 (1998).
Kasprzak, A. et al. p21/Wafl/Cipl cellular expression in chronic long-lasting hepatitis C: correlation with HCV proteins (C, NS3, NS5A), other cell-cycle related proteins and selected clinical data. Folia Histochem. Cytobiol. 47, 385–394 (2009).
Sclair, S. N. et al. Increased hepatic progenitor cell response and ductular reaction in patients with severe recurrent HCV post-liver transplantation. Clin. Transpl. 30, 722–730 (2016).
Khaliq, M. et al. Stat3 regulates liver progenitor cell-driven liver regeneration in zebrafish. Gene Expr. 18, 157–170 (2018).
Limaye, P. B., Bowen, W. C., Orr, A., Apte, U. M. & Michalopoulos, G. K. Expression of hepatocytic- and biliary-specific transcription factors in regenerating bile ducts during hepatocyte-to-biliary epithelial cell transdifferentiation. Comp. Hepatol. 9, 9 (2010).
Font-Burgada, J. et al. Hybrid periportal hepatocytes regenerate the injured liver without giving rise to cancer. Cell 162, 766–779 (2015).
Michalopoulos, G. K., Bowen, W. C., Mule, K. & Luo, J. HGF-, EGF-, and dexamethasone-induced gene expression patterns during formation of tissue in hepatic organoid cultures. Gene Expr. 11, 55–75 (2003).
Michalopoulos, G. K., Bowen, W. C., Mule, K. & Stolz, D. B. Histological organization in hepatocyte organoid cultures. Am. J. Pathol. 159, 1877–1887 (2001).
Schaub, J. R. et al. De novo formation of the biliary system by TGFβ-mediated hepatocyte transdifferentiation. Nature 557, 247–251 (2018).
Yovchev, M. I., Lee, E. J., Rodriguez-Silva, W., Locker, J. & Oertel, M. Biliary obstruction promotes multilineage differentiation of hepatic stem cells. Hepatol. Commun. 3, 1137–1150 (2019).
Limaye, P. B. et al. Expression of specific hepatocyte and cholangiocyte transcription factors in human liver disease and embryonic development. Lab. Invest. 88, 865–872 (2008).
Hattoum, A., Rubin, E., Orr, A. & Michalopoulos, G. K. Expression of hepatocyte epidermal growth factor receptor, FAS and glypican 3 in EpCAM-positive regenerative clusters of hepatocytes, cholangiocytes, and progenitor cells in human liver failure. Hum. Pathol. 44, 743–749 (2013).
Stueck, A. E. & Wanless, I. R. Hepatocyte buds derived from progenitor cells repopulate regions of parenchymal extinction in human cirrhosis. Hepatology 61, 1696–1707 (2015).
Bhushan, B. & Apte, U. Liver regeneration after acetaminophen hepatotoxicity: mechanisms and therapeutic opportunities. Am. J. Pathol. 189, 719–729 (2019).
Bhushan, B. et al. Pro-regenerative signaling after acetaminophen-induced acute liver injury in mice identified using a novel incremental dose model. Am. J. Pathol. 184, 3013–3025 (2014).
Bhushan, B. et al. Dual role of epidermal growth factor receptor in liver injury and regeneration after acetaminophen overdose in mice. Toxicol. Sci. 155, 363–378 (2017).
Hughes, R. D., Zhang, L., Tsubouchi, H., Daikuhara, Y. & Williams, R. Plasma hepatocyte growth factor and biliprotein levels and outcome in fulminant hepatic failure. J. Hepatol. 20, 106–111 (1994).
James, L. P., Kurten, R. C., Lamps, L. W., McCullough, S. & Hinson, J. A. Tumour necrosis factor receptor 1 and hepatocyte regeneration in acetaminophen toxicity: a kinetic study of proliferating cell nuclear antigen and cytokine expression. Basic Clin. Pharmacol. Toxicol. 97, 8–14 (2005).
James, L. P., Lamps, L. W., McCullough, S. & Hinson, J. A. Interleukin 6 and hepatocyte regeneration in acetaminophen toxicity in the mouse. Biochem. Biophys. Res. Commun. 309, 857–863 (2003).
Donahower, B. et al. Vascular endothelial growth factor and hepatocyte regeneration in acetaminophen toxicity. Am. J. Physiol. Gastrointest. Liver Physiol. 291, G102–G109 (2006).
Kato, T. et al. Vascular endothelial growth factor receptor-1 signaling promotes liver repair through restoration of liver microvasculature after acetaminophen hepatotoxicity. Toxicol. Sci. 120, 218–229 (2011).
Bhushan, B. et al. Role of bile acids in liver injury and regeneration following acetaminophen overdose. Am. J. Pathol. 183, 1518–1526 (2013).
Bhushan, B., Poudel, S., Manley, M. W. Jr. Roy, N. & Apte, U. Inhibition of glycogen synthase kinase 3 accelerated liver regeneration after acetaminophen-induced hepatotoxicity in mice. Am. J. Pathol. 187, 543–552 (2017).
Alvarez-Sola, G. et al. Engineered fibroblast growth factor 19 protects from acetaminophen-induced liver injury and stimulates aged liver regeneration in mice. Cell Death Dis. 8, e3083 (2017).
Bird, T. G. et al. TGFβ inhibition restores a regenerative response in acute liver injury by suppressing paracrine senescence. Sci. Transl Med. 10, eaan1230 (2018).
Borude, P., Bhushan, B. & Apte, U. DNA damage response regulates initiation of liver regeneration following acetaminophen overdose. Gene Expr. 18, 115–123 (2018).
Borude, P. et al. Pleiotropic role of p53 in injury and liver regeneration after acetaminophen overdose. Am. J. Pathol. 188, 1406–1418 (2018).
Overturf, K. et al. Hepatocytes corrected by gene therapy are selected in vivo in a murine model of hereditary tyrosinaemia type I. Nat. Genet. 12, 266–273 (1996).
Overturf, K., al-Dhalimy, M., Ou, C. N., Finegold, M. & Grompe, M. Serial transplantation reveals the stem-cell-like regenerative potential of adult mouse hepatocytes. Am. J. Pathol. 151, 1273–1280 (1997).
Monga, S. P. Updates on hepatic homeostasis and the many tiers of hepatobiliary repair. Nat. Rev. Gastroenterol. Hepatol. 16, 84–86 (2019).
Wang, B., Zhao, L., Fish, M., Logan, C. Y. & Nusse, R. Self-renewing diploid Axin2+ cells fuel homeostatic renewal of the liver. Nature 524, 180–185 (2015).
Kennedy, S., Rettinger, S., Flye, M. W. & Ponder, K. P. Experiments in transgenic mice show that hepatocytes are the source for postnatal liver growth and do not stream. Hepatology 22, 160–168 (1995).
Lin, S. et al. Distributed hepatocytes expressing telomerase repopulate the liver in homeostasis and injury. Nature 556, 244–248 (2018).
Chen, F. et al. Broad distribution of hepatocyte proliferation in liver homeostasis and regeneration. Cell Stem Cell 26, 27–33.e4 (2020).
Sun, T. et al. AXIN2+ pericentral hepatocytes have limited contributions to liver homeostasis and regeneration. Cell Stem Cell 26, 97–107.e6 (2019).
Monga, S. P. No zones left behind: democratic hepatocytes contribute to liver homeostasis and repair. Cell Stem Cell 26, 2–3 (2020).
Michalopoulos, G. K. Hepatostat: liver regeneration and normal liver tissue maintenance. Hepatology 65, 1384–1392 (2017).
Klaas, M. et al. The alterations in the extracellular matrix composition guide the repair of damaged liver tissue. Sci. Rep. 6, 27398 (2016).
Canbay, A. et al. Apoptotic body engulfment by a human stellate cell line is profibrogenic. Lab. Invest. 83, 655–663 (2003).
Duncan, A. W. et al. The ploidy conveyor of mature hepatocytes as a source of genetic variation. Nature 467, 707–710 (2010).
Anti, M. et al. DNA ploidy pattern in human chronic liver diseases and hepatic nodular lesions. Flow cytometric analysis on echo-guided needle liver biopsy. Cancer 73, 281–288 (1994).
Duncan, A. W. et al. Frequent aneuploidy among normal human hepatocytes. Gastroenterology 142, 25–28 (2012).
Boege, Y. et al. A dual role of caspase-8 in triggering and sensing proliferation-associated DNA damage, a key determinant of liver cancer development. Cancer Cell 32, 342–359 (2017).
Zhu, M. et al. Somatic mutations increase hepatic clonal fitness and regeneration in chronic liver disease. Cell 177, 608–621 (2019).
Luo, J. H. et al. Transcriptomic and genomic analysis of human hepatocellular carcinomas and hepatoblastomas. Hepatology 44, 1012–1024 (2006).
Author information
Authors and Affiliations
Contributions
G.K.M. and B.B. wrote the article. G.K.M. reviewed/edited the manuscript before submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information
Nature Reviews Gastroenterology & Hepatology thanks S. Dooley, S. Huppert, H. Jaeschke and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Michalopoulos, G.K., Bhushan, B. Liver regeneration: biological and pathological mechanisms and implications. Nat Rev Gastroenterol Hepatol 18, 40–55 (2021). https://doi.org/10.1038/s41575-020-0342-4
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41575-020-0342-4
This article is cited by
-
Identification and validation of a novel predictive signature based on hepatocyte-specific genes in hepatocellular carcinoma by integrated analysis of single-cell and bulk RNA sequencing
BMC Medical Genomics (2024)
-
The dual function of cGAS-STING signaling axis in liver diseases
Acta Pharmacologica Sinica (2024)
-
Commentary on the challenges of modeling hepatoblastoma in-vivo
Pediatric Research (2024)
-
Heterogeneity of hepatocyte dynamics restores liver architecture after chemical, physical or viral damage
Nature Communications (2024)
-
Three Dimensional Bioprinting for Hepatic Tissue Engineering: From In Vitro Models to Clinical Applications
Tissue Engineering and Regenerative Medicine (2024)