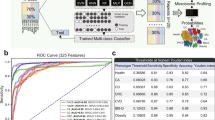
Abstract
The gut microbiome has been implicated in cancer in several ways, as specific microbial signatures are known to promote cancer development and influence safety, tolerability and efficacy of therapies. The ‘omics’ technologies used for microbiome analysis continuously evolve and, although much of the research is still at an early stage, large-scale datasets of ever increasing size and complexity are being produced. However, there are varying levels of difficulty in realizing the full potential of these new tools, which limit our ability to critically analyse much of the available data. In this Perspective, we provide a brief overview on the role of gut microbiome in cancer and focus on the need, role and limitations of a machine learning-driven approach to analyse large amounts of complex health-care information in the era of big data. We also discuss the potential application of microbiome-based big data aimed at promoting precision medicine in cancer.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Marchesi, J. R. et al. The gut microbiota and host health: a new clinical frontier. Gut 65, 330–339 (2016).
Yue, B. et al. Inflammatory bowel disease: a potential result from the collusion between gut microbiota and mucosal immune system. Microorganisms 7, E440 (2019).
Zhang, Z. et al. Impact of fecal microbiota transplantation on obesity and metabolic syndrome — a systematic review. Nutrients 11, E2291 (2019).
Thaiss, C. A. et al. Persistent microbiome alterations modulate the rate of post-dieting weight regain. Nature 540, 544–551 (2016).
Mullish, B. H. & Williams, H. R. Clostridium difficile infection and antibiotic-associated diarrhoea. Clin. Med. 18, 237–241 (2018).
van der Giessen, J. et al. Modulation of cytokine patterns and microbiome during pregnancy in IBD. Gut 69, 473–486 (2020).
Konstantinov, S. R., van der Woude, C. J. & Peppelenbosch, M. P. Do pregnancy-related changes in the microbiome stimulate innate immunity? Trends Mol. Med. 19, 454–459 (2013).
Maguire, M. & Maguire, G. Gut dysbiosis, leaky gut, and intestinal epithelial proliferation in neurological disorders: towards the development of a new therapeutic using amino acids, prebiotics, probiotics, and postbiotics. Rev. Neurosci. 30, 179–201 (2019).
Tang, W. H. et al. Intestinal microbial metabolism of phosphatidylcholine and cardiovascular risk. N. Engl. J. Med. 368, 1575–1584 (2013).
Vivarelli, S. et al. Gut microbiota and cancer: from pathogenesis to therapy. Cancers 11, 38 (2019).
Bi, J. H. et al. ClickGene: an open cloud-based platform for big pan-cancer data genome-wide association study, visualization and exploration. BioData Min. 12, 12 (2019).
Rodriguez-Martin, B. et al. Pan-cancer analysis of whole genomes identifies driver rearrangements promoted by LINE-1 retrotransposition. Nat. Genet. 52, 306–319 (2020).
Zhang, J. et al. The International Cancer Genome Consortium data portal. Nat. Biotechnol. 37, 367–369 (2019).
Brown, J. A., Ni Chonghaile, T., Matchett, K. B., Lynam-Lennon, N. & Kiely, P. A. Big data-led cancer research, application, and insights. Cancer Res. 76, 6167–6170 (2016).
Evans, B. J. & Krumholz, H. M. People-powered data collaboratives: fueling data science with the health-related experiences of individuals. J. Am. Med. Inform. Assoc. 26, 159–161 (2019).
Provost, F. & Fawcett, T. Data science and its relationship to big data and data-driven decision making. Big Data 1, 51–59 (2013).
Sanchez-Pinto, L. N., Luo, Y. & Churpek, M. M. Big data and data science in critical care. Chest 154, 1239–1248 (2018).
Gruson, D., Helleputte, T., Rousseau, P. & Gruson, D. Data science, artificial intelligence, and machine learning: opportunities for laboratory medicine and the value of positive regulation. Clin. Biochem. 69, 1–7 (2019).
Rajkomar, A., Dean, J. & Kohane, I. Machine learning in medicine. N. Engl. J. Med. 380, 1347–1358 (2019).
Sender, R., Fuchs, S. & Milo, R. Are we really vastly outnumbered? Revisiting the ratio of bacterial to host cells in humans. Cell 164, 337–340 (2016).
Lozupone, C. A. et al. Diversity, stability and resilience of the human gut microbiota. Nature 489, 220–230 (2012).
Conlon, M. A. & Bird, A. R. The impact of diet and lifestyle on gut microbiota and human health. Nutrients 7, 17–44 (2014).
Imhann, F. et al. The influence of proton pump inhibitors and other commonly used medication on the gut microbiota. Gut Microbes 8, 351–358 (2017).
Thomas, S. et al. The host microbiome regulates and maintains human health: a primer and perspective for non-microbiologists. Cancer Res. 77, 1783–1812 (2017).
Fessler, J., Matson, V. & Gajewski, T. F. Exploring the emerging role of the microbiome in cancer immunotherapy. J. Immunother. Cancer 7, 108 (2019).
Scott, A. J. et al. International Cancer Microbiome Consortium consensus statement on the role of the human microbiome in carcinogenesis. Gut 68, 1624–1632 (2019).
Lazar, V. et al. Aspects of gut microbiota and immune system interactions in infectious diseases, immunopathology, and cancer. Front. Immunol. 9, 1830 (2018).
Pagliari, D. et al. Gut microbiota–immune system crosstalk and pancreatic disorders. Mediators Inflamm. 2018, 7946431 (2018).
Bingula, R. et al. Desired turbulence? Gut–lung axis, immunity, and lung cancer. J. Oncol. 2017, 5035371 (2017).
Gopalakrishnan, V. et al. The influence of the gut microbiome on cancer, immunity, and cancer immunotherapy. Cancer Cell 33, 570–580 (2018).
Rugge, M. et al. Gastric cancer as preventable disease. Clin. Gastroenterol. Hepatol. 15, 1833–1843 (2017).
Parsonnet, J. et al. Helicobacter pylori infection and the risk of gastric carcinoma. N. Engl. J. Med. 325, 1127–1131 (1991).
Garrett, W. S. Cancer and the microbiota. Science 348, 80–86 (2015).
Wu, S. et al. A human colonic commensal promotes colon tumorigenesis via activation of T helper type 17 T cell responses. Nat. Med. 15, 1016–1022 (2009).
Raza, M. H. et al. Microbiota in cancer development and treatment. J. Cancer Res. Clin. Oncol. 145, 49–63 (2019).
Pushalkar, S. et al. The pancreatic cancer microbiome promotes oncogenesis by induction of innate and adaptive immune suppression. Cancer Discov. 8, 403–416 (2018).
Kostic, A. D. et al. Fusobacterium nucleatum potentiates intestinal tumorigenesis and modulates the tumor-immune microenvironment. Cell Host Microbe 14, 207–215 (2013).
Rubinstein, M. R. et al. Fusobacterium nucleatum promotes colorectal carcinogenesis by modulating E-cadherin/beta-catenin signaling via its FadA adhesin. Cell Host Microbe 14, 195–206 (2013).
Yoshimoto, S. et al. Obesity-induced gut microbial metabolite promotes liver cancer through senescence secretome. Nature 499, 97–101 (2013).
Raskov, H., Burcharth, J. & Pommergaard, H. C. Linking gut microbiota to colorectal cancer. J. Cancer 8, 3378–3395 (2017).
Li, S., Peppelenboscha, M. P. & Smits, R. Bacterial biofilms as a potential contributor to mucinous colorectal cancer formation. Biochim. Biophys. Acta Rev. Cancer 1872, 74–79 (2019).
Belkaid, Y. & Hand, T. W. Role of the microbiota in immunity and inflammation. Cell 157, 121–141 (2014).
Yu, J. et al. Metagenomic analysis of faecal microbiome as a tool towards targeted non-invasive biomarkers for colorectal cancer. Gut 66, 70–78 (2017).
Zackular, J. P. et al. The gut microbiome modulates colon tumorigenesis. mBio 4, e00692-13 (2013).
Wirbel, J. et al. Meta-analysis of fecal metagenomes reveals global microbial signatures that are specific for colorectal cancer. Nat. Med. 25, 679–689 (2019).
Sobhani, I. et al. Microbial dysbiosis in colorectal cancer (CRC) patients. PLoS One 6, e16393 (2011).
Ren, Z. et al. Gut microbial profile analysis by MiSeq sequencing of pancreatic carcinoma patients in China. Oncotarget 8, 95176–95191 (2017).
Ren, Z. et al. Gut microbiome analysis as a tool towards targeted non-invasive biomarkers for early hepatocellular carcinoma. Gut 58, 1014–1023 (2019).
Pouncey, A. L. et al. Gut microbiota, chemotherapy and the host: the influence of the gut microbiota on cancer treatment. Ecancermedicalscience 12, 868 (2018).
Alexander, J. L. et al. Gut microbiota modulation of chemotherapy efficacy and toxicity. Nat. Rev. Gastroenterol. Hepatol. 14, 356–365 (2017).
Touchefeu, Y. et al. Systematic review: the role of the gut microbiota in chemotherapy- or radiation-induced gastrointestinal mucositis — current evidence and potential clinical applications. Aliment. Pharmacol. Ther. 40, 409–421 (2014).
Mathijssen, R. H. et al. Clinical pharmacokinetics and metabolism of irinotecan (CPT-11). Clin. Cancer Res. 7, 2182–2194 (2001).
Ma, M. K. & McLeod, H. L. Lessons learned from the irinotecan metabolic pathway. Curr. Med. Chem. 10, 41–49 (2003).
Wallace, B. D. et al. Alleviating cancer drug toxicity by inhibiting a bacterial enzyme. Science 330, 831–835 (2010).
Kodawara, T. et al. The inhibitory effect of ciprofloxacin on the beta-glucuronidase-mediated deconjugation of the irinotecan metabolite SN-38-G. Basic Clin. Pharmacol. Toxicol. 118, 333–337 (2016).
Frank, M. et al. TLR signaling modulates side effects of anticancer therapy in the small intestine. J. Immunol. 194, 1983–1995 (2015).
Hooper, L. V. & Macpherson, A. J. Immune adaptations that maintain homeostasis with the intestinal microbiota. Nat. Rev. Immunol. 10, 159–169 (2010).
Quince, C. et al. Shotgun metagenomics, from sampling to analysis. Nat. Biotechnol. 35, 833–844 (2017).
Vetizou, M. et al. Anticancer immunotherapy by CTLA-4 blockade relies on the gut microbiota. Science 350, 1079–1084 (2015).
Frankel, A. E. et al. Metagenomic shotgun sequencing and unbiased metabolomic profiling identify specific human gut microbiota and metabolites associated with immune checkpoint therapy efficacy in melanoma patients. Neoplasia 19, 848–855 (2017).
Gopalakrishnan, V. et al. Gut microbiome modulates response to anti-PD-1 immunotherapy in melanoma patients. Science 359, 97–103 (2018).
Routy, B. et al. Gut microbiome influences efficacy of PD-1-based immunotherapy against epithelial tumors. Science 359, 91–97 (2018).
Sivan, A. et al. Commensal Bifidobacterium promotes antitumor immunity and facilitates anti-PD-L1 efficacy. Science 350, 1084–1089 (2015).
Gerassy-Vainberg, S. et al. Radiation induces proinflammatory dysbiosis: transmission of inflammatory susceptibility by host cytokine induction. Gut 67, 97–107 (2018).
Kumagai, T., Rahman, F. & Smith, A. M. The microbiome and radiation induced-bowel injury: evidence for potential mechanistic role in disease pathogenesis. Nutrients 10, E1405 (2018).
Cui, M. et al. Faecal microbiota transplantation protects against radiation-induced toxicity. EMBO Mol. Med. 9, 448–461 (2017).
Manichanh, C. et al. The gut microbiota predispose to the pathophysiology of acute postradiotherapy diarrhea. Am. J. Gastroenterol. 103, 1754–1761 (2008).
Nam, Y. D. et al. Impact of pelvic radiotherapy on gut microbiota of gynecological cancer patients revealed by massive pyrosequencing. PLoS One 8, e82659 (2013).
Wang, A. et al. Gut microbial dysbiosis may predict diarrhea and fatigue in patients undergoing pelvic cancer radiotherapy: a pilot study. PLoS One 10, e0126312 (2015).
Reis Ferreira, M. et al. Microbiota- and radiotherapy-induced gastrointestinal side-effects (MARS) study: a large pilot study of the microbiome in acute and late-radiation enteropathy. Clin. Cancer Res. 25, 6487–6500 (2019).
Lam, S. Y., Peppelenbosch, M. P. & Fuhler, G. M. Prediction and treatment of radiation enteropathy: can intestinal bugs lead the way? Clin. Cancer Res. 25, 6280–6282 (2019).
Roy, S. & Trinchieri, G. Microbiota: a key orchestrator of cancer therapy. Nat. Rev. Cancer. 17, 271–285 (2017).
Lehouritis, P. et al. Local bacteria affect the efficacy of chemotherapeutic drugs. Sci. Rep. 5, 14554 (2015).
Viaud, S. et al. The intestinal microbiota modulates the anticancer immune effects of cyclophosphamide. Science 342, 971–976 (2013).
Ghiringhelli, F. et al. Activation of the NLRP3 inflammasome in dendritic cells induces IL-1beta-dependent adaptive immunity against tumors. Nat. Med. 15, 1170–1178 (2009).
Ozben, T. Oxidative stress and apoptosis: impact on cancer therapy. J. Pharm. Sci. 96, 2181–2196 (2007).
Iida, N. et al. Commensal bacteria control cancer response to therapy by modulating the tumor microenvironment. Science 342, 967–970 (2013).
Daillere, R. et al. Enterococcus hirae and Barnesiella intestinihominis facilitate cyclophosphamide-induced therapeutic immunomodulatory effects. Immunity 45, 931–943 (2016).
Fyza, Y., Gills, J. & Sears, C. L. Impact of the microbiome on checkpoint inhibitor treatment in patients with non-small cell lung cancer and melanoma. EBioMedicine 48, 642–647 (2019).
Seidel, J. A., Otsuka, A. & Kabashima, K. Anti-PD-1 and anti-CTLA-4 therapies in cancer: mechanisms of action, efficacy, and limitations. Front. Oncol. 8, 86 (2018).
Darvin, P., Toor, S. M., Sasidharan Nair, V. & Elkord, E. Immune checkpoint inhibitors: recent progress and potential biomarkers. Exp. Mol. Med. 50, 1–11 (2018).
Yang, B. et al. Progresses and perspectives of anti-PD-1/PD-L1 antibody therapy in head and neck cancers. Front. Oncol. 8, 563 (2018).
Matson, V. et al. The commensal microbiome is associated with anti-PD-1 efficacy in metastatic melanoma patients. Science 359, 104–108 (2018).
Peled, J. U. et al. Microbiota predictor of mortality in allogeneic hematopoietic-cell transplantation. N. Engl. J. Med. 382, 822–834 (2020).
Schwabe, R. F. & Jobin, C. The microbiome and cancer. Nat. Rev. Cancer 13, 800–812 (2013).
Elinav, E. et al. The cancer microbiome. Nat. Rev. Cancer. 19, 371–376 (2018).
de Martel, C. et al. Global burden of cancers attributable to infections in 2008: a review and synthetic analysis. Lancet Oncol. 13, 607–615 (2012).
Fais, T. et al. Targeting colorectal cancer-associated bacteria: a new area of research for personalized treatments. Gut Microbes 7, 329–333 (2016).
Shah, M. S. et al. Leveraging sequence-based faecal microbial community survey data to identify a composite biomarker for colorectal cancer. Gut 67, 882–891 (2018).
Armour, C. R., Nayfach, S., Pollard, K. S. & Sharpton, T. J. A metagenomic meta-analysis reveals functional signatures of health and disease in the human gut microbiome. mSystems 4, e00332-18 (2019).
Segata, N. et al. Metagenomic biomarker discovery and explanation. Genome Biol. 12, R60 (2011).
Bhatt, A. S. et al. Sequence-based discovery of Bradyrhizobium enterica in cord colitis syndrome. N. Engl. J. Med. 369, 517–528 (2013).
Kostic, A. D. et al. Genomic analysis identifies association of Fusobacterium with colorectal carcinoma. Genome Res. 22, 292–298 (2012).
Drewes, J. L. et al. High-resolution bacterial 16S rRNA gene profile meta-analysis and biofilm status reveal common colorectal cancer consortia. NPJ Biofilms Microbiomes 3, 34 (2017).
Thomas, A. M. et al. Metagenomic analysis of colorectal cancer datasets identifies cross-cohort microbial diagnostic signatures and a link with choline degradation. Nat. Med. 25, 667–678 (2019).
Esteban-Gil, A. et al. ColPortal, an integrative multiomic platform for analysing epigenetic interactions in colorectal cancer. Sci. Data 6, 255 (2019).
Derosa, L. et al. Negative association of antibiotics on clinical activity of immune checkpoint inhibitors in patients with advanced renal cell and non-small-cell lung cancer. Ann. Oncol. 29, 1437–1444 (2018).
Li, Y., Wu, F. X. & Ngom, A. A review on machine learning principles for multi-view biological data integration. Brief. Bioinform. 19, 325–340 (2018).
Alanazi, H. O., Abdullah, A. H. & Qureshi, K. N. A critical review for developing accurate and dynamic predictive models using machine learning methods in medicine and health care. J. Med. Syst. 41, 69 (2017).
Camacho, D. M., Collins, K. M., Powers, R. K., Costello, J. C. & Collins, J. J. Next-generation machine learning for biological networks. Cell 173, 1581–1592 (2018).
Zhang, Y. et al. Machine learning performance in a microbial molecular autopsy context: a cross-sectional postmortem human population study. PLoS One 14, e0213829 (2019).
Ruffle, J. K., Farmer, A. D. & Aziz, Q. Artificial intelligence-assisted gastroenterology — promises and pitfalls. Am. J. Gastroenterol. 114, 422–428 (2019).
Saito, H. et al. Automatic detection and classification of protruding lesions in wireless capsule endoscopy images based on a deep convolutional neural network. Gastrointest. Endosc. 92, 144–151 (2020).
Lui, T. K., Guo, C. G. & Leung, W. K. Accuracy of artificial intelligence on histology prediction and detection of colorectal polyps: a systematic review and meta-analysis. Gastrointest. Endosc. 92, 11–22 (2020).
Seyed Tabib, N. S. et al. Big data in IBD: big progress for clinical practice. Gut https://doi.org/10.1136/gutjnl-2019-320065 (2020).
Olivera, P., Danese, S., Jay, N., Natoli, G. & Peyrin-Biroulet, L. Big data in IBD: a look into the future. Nat. Rev. Gastroenterol. Hepatol. 16, 312–321 (2019).
Noor, E., Cherkaoui, S. & Sauer, U. Biological insights through omics data integration. Curr. Opin. Syst. Biol. 15, 39–47 (2019).
Lopez, C., Tcker, S., Salameh, T. & Tucker, C. An unsupervised machine learning method for discovering patient clusters based on genetic signatures. J. Biomed. Inform. 85, 30–39 (2018).
Shomorony, I. et al. An unsupervised learning approach to identify novel signatures of health and disease from multimodal data. Genome Med. 12, 7 (2020).
Price, N. D. et al. A wellness study of 108 individuals using personal, dense, dynamic data clouds. Nat. Biotechnol. 35, 747–756 (2017).
Argelaguet, R. et al. Multi-omics factor analysis — a framework for unsupervised integration of multi-omics data sets. Mol. Syst. Biol. 14, e8124 (2018).
Bisikirska, B. et al. Elucidation and pharmacological targeting of novel molecular drivers of follicular lymphoma progression. Cancer Res. 76, 664–674 (2016).
Costello, J. C. et al. A community effort to assess and improve drug sensitivity prediction algorithms. Nat. Biotechnol. 32, 1202–1212 (2014).
Mezlini, A. M. & Goldenberg, A. Incorporating networks in a probabilistic graphical model to find drivers for complex human disease. PLoS Comput. Biol. 13, e1005580 (2017).
Fabris, F., Magalhaes, J. P. & Freitas, A. A. A review of supervised machine learning applied to ageing research. Biogerontology 18, 171–188 (2017).
Yu, Z. et al. Progressive semisupervised learning of multiple classifiers. IEEE Trans. Cybern. 48, 689–702 (2018).
Huang, H., Vangay, P., McKinlay, C. E. & Knights, D. Multi-omics analysis of inflammatory bowel disease. Immunol. Lett. 162, 62–68 (2014).
Lio, W. et al. Guidelines for developing and reporting machine learning predictive models in biomedical research: a multidisciplinary view. J. Med. Internet Res. 18, e323 (2016).
Doostparast Torshizi, A. & Petzold, L. R. Graph-based semi-supervised learning with genomic data integration using condition-responsive genes applied to phenotype classification. J. Am. Med. Inform. Assoc. 25, 99–108 (2018).
Lin, Y. et al. Computer-aided biomarker discovery for precision medicine: data resources, models and applications. Brief. Bioinform 20, 952–975 (2019).
Tang, B., Pan, Z., Yin, K. & Khateeb, A. Recent advances of deep learning in bioinformatics and computational biology. Front. Genet. 10, 214 (2019).
Londhe, V. Y. & Bhasin, B. Artificial intelligence and its potential in oncology. Drug. Discov. Today 24, 228–232 (2019).
Ngiam, K. Y. & Khor, I. W. Big data and machine learning algorithms for health-care delivery. Lancet Oncol. 20, e262–e273 (2019).
Babarenda Gamage, T. P. et al. An automated computational biomechanics workflow for improving breast cancer diagnosis and treatment. Interface Focus. 9, 20190034 (2019).
Tseng, Y. J. et al. Predicting breast cancer metastasis by using serum biomarkers and clinicopathological data with machine learning technologies. Int. J. Med. Inform. 128, 79–86 (2019).
Goldenberg, S. L., Nir, G. & Salcudean, S. E. A new era: artificial intelligence and machine learning in prostate cancer. Nat. Rev. Urol. 16, 391–403 (2019).
Paik, E. S. et al. Prediction of survival outcomes in patients with epithelial ovarian cancer using machine learning methods. J. Gynecol. Oncol. 30, e65 (2019).
Kouznetsova, V. L. et al. Recognition of early and late stages of bladder cancer using metabolites and machine learning. Metabolomics 15, 94 (2019).
Jin, Y. et al. The diversity of gut microbiome is associated with favorable responses to anti-PD-1 immunotherapy in Chinese non-small cell lung cancer patients. J. Thorac. Oncol. 14, 1378–1389 (2019).
Qian, Z. et al. Differentiation of glioblastoma from solitary brain metastases using radiomic machine-learning classifiers. Cancer Lett. 451, 128–135 (2019).
Leatherdale, S. T. & Lee, J. Artificial intelligence (AI) and cancer prevention: the potential application of AI in cancer control programming needs to be explored in population laboratories such as COMPASS. Cancer Causes Control. 30, 671–675 (2019).
Veselkov, K. et al. HyperFoods: machine intelligent mapping of cancer-beating molecules in foods. Sci. Rep. 9, 9237 (2019).
Zhao, W. et al. Toward automatic prediction of EGFR mutation status in pulmonary adenocarcinoma with 3D deep learning. Cancer Med. 8, 3532–3543 (2019).
Sato, M. et al. Machine-learning approach for the development of a novel predictive model for the diagnosis of hepatocellular carcinoma. Sci. Rep. 9, 7704 (2019).
Feng, Q. X. et al. An intelligent clinical decision support system for preoperative prediction of lymph node metastasis in gastric cancer. J. Am. Coll. Radiol. 16, 952–960 (2019).
You, J., McLeod, R. D. & Hu, P. Predicting drug–target interaction network using deep learning model. Comput. Biol. Chem. 80, 90–101 (2019).
Kessler, R. C., Bossarte, R. M., Luedtke, A., Zaslavsky, A. M. & Zubizarreta, J. R. Machine learning methods for developing precision treatment rules with observational data. Behav. Res. Ther. 120, 103412 (2019).
Mottini, C., Napolitano, F., Li, Z., Gao, X. & Cardone, L. Computer-aided drug repurposing for cancer therapy: approaches and opportunities to challenge anticancer targets. Semin. Cancer Biol. https://doi.org/10.1016/j.semcancer.2019.09.023 (2019).
Bera, K., Schalper, K. A., Rimm, D. L., Velcheti, V. & Madabhushi, A. Artificial intelligence in digital pathology — new tools for diagnosis and precision oncology. Nat. Rev. Clin. Oncol. 16, 703–715 (2019).
Penson, A. et al. Development of genome-derived tumor type prediction to inform clinical cancer care. JAMA Oncol. https://doi.org/10.1001/jamaoncol.2019.3985 (2019).
Grewal, J. K. et al. Application of a neural network whole transcriptome-based pan-cancer method for diagnosis of primary and metastatic cancers. JAMA Netw. Open 2, e192597 (2019).
Subramanian, S. et al. Persistent gut microbiota immaturity in malnourished Bangladeshi children. Nature 510, 417–421 (2014).
Vervier, K. et al. Large-scale machine learning for metagenomics sequence classification. Bioinformatics 32, 1023–1032 (2016).
Fernandez-Navarro, T. et al. Exploring the interactions between serum free fatty acids and fecal microbiota in obesity through a machine learning algorithm. Food Res. Int. 121, 533–541 (2019).
Thompson, J. et al. Machine learning to predict microbial community functions: an analysis of dissolved organic carbon from litter decomposition. PLoS One 14, e0215502 (2019).
Shinn, L. et al. Applying machine-learning to human gastrointestinal microbial species to predict dietary intake. Curr. Dev. Nutr. 3, https://doi.org/10.1093/cdn/nzz040.P20-040-19 (2019).
Turnbaugh, P. J. et al. The human microbiome project. Nature 449, 804–810 (2007).
Integrative HMP (iHMP) Research Network Consortium. The Integrative Human Microbiome Project. Nature 569, 641-648.
Vangav, P., Hillmann, B. M. & Knights, D. Microbiome learning repo (ML Repo): a public repository of microbiome regression and classification tasks. Gigascience 8, giz042 (2019).
Mallick., H. et al. Experimental design and quantitative analysis of microbial community multiomics. Genome Biol. 18, 228 (2017).
Knight, R. et al. Best practices for analysing microbiomes. Nat. Rev. Microbiol. 16, 410–422 (2018).
Cani, P. D. Human gut microbiome: hopes, threats and promises. Gut 67, 1716–1725 (2018).
Knights, D., Costello, E. K. & Knight, R. Supervised classification of human microbiota. FEMS Microbiol. Rev. 35, 343–359 (2011).
Moitinho-Silva, L. et al. Predicting the HMA–LMA status in marine sponges by machine learning. Front. Microbiol. 8, 752 (2017).
Bockulich, N. A. et al. Antibiotics, birth mode, and diet shape microbiome maturation during early life. Sci. Transl Med. 8, 343ra82 (2016).
Pasolli, E., Truong, D. T., Malik, F., Waldron, L. & Segata, N. Machine learning meta-analysis of large metagenomic datasets: tools and biological insights. PLoS Comput. Biol. 12, e1004977 (2016).
Heshiki, Y. et al. Predictable modulation of cancer treatment outcomes by the gut microbiota. Microbiome 8, 28 (2020).
Zuo, T. et al. Gut mucosal virome alterations in ulcerative colitis. Gut 68, 1169–1179 (2019).
Larsen, P. E. & Dai, Y. Metabolome of human gut microbiome is predictive of host dysbiosis. Gigascience 4, 42 (2015).
Bokulich, N. et al. q2-sample-classifier: machine-learning tools for microbiome classification and regression. J. Open Source Softw. 3, 934 (2018).
Dhariwal, A. et al. MicrobiomeAnalyst: a web-based tool for comprehensive statistical, visual and meta-analysis of microbiome data. Nucleic Acids Res. 45, W180–W188 (2017).
Edgar, R. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26, 2460–2461 (2010).
Prifti, E. et al. Interpretable and accurate prediction models for metagenomics data. Gigascience 9, 1–11 (2020).
Zhou, Y.-H. & Gallins, P. A review and tutorial of machine learning methods for microbiome host trait prediction. Front. Genet. 10, 579 (2019).
Ananthakrishnan, A. et al. Gut microbiome function predicts response to anti-integrin biologic therapy in inflammatory bowel diseases. Cell Host Microbe 21, 603–610 (2017).
Zhu, Q., Jiang, X., Zhu, Q., Pan, M. & He, T. Graph embedding deep learning guides microbial biomarkers’ identification. Front. Genet. 10, 1182 (2019).
Angermueller, C., Parnamaa, T., Parts, L. & Stegle, O. Deep learning for computational biology. Mol. Syst. Biol. 12, 878 (2016).
Eraslan, G., Avsec, Z., Gagneur, J. & Theis, F. J. Deep learning: new computational modelling techniques for genomics. Nat. Rev. Genet. 20, 389–403 (2019).
LaPierre, N., Ju, C. J., Zhou, G. & Wang, W. MetaPheno: a critical evaluation of deep learning and machine learning in metagenome-based disease prediction. Methods 166, 74–82 (2019).
Lloyd-Price, J. et al. Multi-omics of the gut microbial ecosystem in inflammatory bowel diseases. Nature 569, 655–662 (2019).
Stols-Goncalves, D. et al. Epigenetic markers and mMicrobiota/mMetabolite-induced epigenetic modifications in the pathogenesis of obesity, metabolic syndrome, type 2 diabetes, and non-alcoholic fatty liver disease. Curr. Diab. Rep. 19, 31 (2019).
Palsson, B. & Zengler, K. The challenges of integrating multi-omic data sets. Nat. Chem. Biol. 6, 787–789 (2010).
Zitnik, M. et al. Machine learning for integrating data in biology and medicine: principles, practice, and opportunities. Inf. Fusion. 50, 71–91 (2019).
Bycroft, C. et al. The UK Biobank resource with deep phenotyping and genomic data. Nature 562, 203–209 (2018).
Perkins, B. A. et al. Precision medicine screening using whole-genome sequencing and advanced imaging to identify disease risk in adults. Proc. Natl Acad. Sci. USA 115, 3685–3691 (2018).
Hollister, E. B. et al. Leveraging human microbiome features to diagnose and stratify children with irritable bowel syndrome. J. Mol. Diagn. 21, 449–461 (2019).
Kreznar, J. H. et al. Host genotype and gut microbiome modulate insulin secretion and diet-induced metabolic phenotypes. Cell Rep. 18, 1739–1750 (2017).
Schussler-Fiorenza Rose, S. M. et al. A longitudinal big data approach for precision health. Nat. Med. 25, 792–804 (2019).
Flemer, B. et al. The oral microbiota in colorectal cancer is distinctive and predictive. Gut 67, 1454–1463 (2018).
Zackular, J. P. et al. The human gut microbiome as a screening tool for colorectal cancer. Cancer Prev. Res. 7, 1112–1121 (2014).
Imhann, F. et al. The 1000IBD project: multi-omics data of 1000 inflammatory bowel disease patients; data release 1. BMC Gastroenterol. 19, 5 (2019).
Casals-Pascual, C. et al. Microbial diversity in clinical microbiome studies: sample size and statistical power considerations. Gastroenterology 158, 1524–1528 (2020).
Shenoi, S.J., Ly, V., Soni, S. & Roberts, K. Developing a serach engine for precision medicine. AMIA Jt. Summits Transl Sci. Proc. 2020, 579–588 (2020).
Goecks, J., Jalili, V., Heiser, L. M. & Gray, J. W. How machine learning will transform biomedicine. Cell 181, 92–101 (2020).
Zheng, Y. et al. Specific gut microbiome signature predicts the early-stage lung cancer. Gut Microbes 11, 1–12 (2020).
Zeevi, D. et al. Personalized nutrition by prediction of glycemic responses. Cell 163, 1079–1094 (2015).
Gevers, D. et al. The treatment-naive microbiome in new-onset Crohn’s disease. Cell Host Microbe 15, 382–392 (2014).
Franzosa, E. A. et al. Gut microbiome structure and metabolic activity in inflammatory bowel disease. Nat. Microbiol. 4, 293–305 (2019).
Coburn, B. et al. Lung microbiota across age and disease stage in cystic fibrosis. Sci. Rep. 5, 10241 (2015).
Alekseyenko, A. V. et al. Community differentiation of the cutaneous microbiota in psoriasis. Microbiome 1, 31 (2013).
Wu, H. et al. Metformin alters the gut microbiome of individuals with treatment-naive type 2 diabetes, contributing to the therapeutic effects of the drug. Nat. Med. 23, 850–858 (2017).
Forslund, K. et al. Disentangling type 2 diabetes and metformin treatment signatures in the human gut microbiota. Nature 528, 262–266 (2015).
Haiser, H. J. et al. Predicting and manipulating cardiac drug inactivation by the human gut bacterium Eggerthella lenta. Science 341, 295–298 (2013).
Cussotto, S., Clarke, G., Dinan, T. G. & Cryan, J. F. Psychotropics and the microbiome: a chamber of secrets. Psychopharmacology 236, 1411–1432 (2019).
Claesson, M. J. et al. Composition, variability, and temporal stability of the intestinal microbiota of the elderly. Proc. Natl Acad. Sci. USA 108, 4586–4591 (2011).
Bibbo, S. et al. The role of diet on gut microbiota composition. Eur. Rev. Med. Pharmacol. Sci. 20, 4742–4749 (2016).
Qiu, G. et al. The significance of probiotics in preventing radiotherapy-induced diarrhea in patients with cervical cancer: a systematic review and meta-analysis. Int. J. Surg. 65, 61–69 (2019).
Liu, M. M. et al. Probiotics for prevention of radiation-induced diarrhea: a meta-analysis of randomized controlled trials. PLoS One 12, e0178870 (2017).
Wang, Y. H. et al. The efficacy and safety of probiotics for prevention of chemoradiotherapy-induced diarrhea in people with abdominal and pelvic cancer: a systematic review and meta-analysis. Eur. J. Clin. Nutr. 70, 1246–1253 (2016).
Delia, P. et al. Use of probiotics for prevention of radiation-induced diarrhea. World J. Gastroenterol. 13, 912–915 (2007).
Henson, C. C. et al. Nutritional interventions for reducing gastrointestinal toxicity in adults undergoing radical pelvic radiotherapy. Cochrane Database Syst. Rev. 11, CD009896 (2013).
Reyna-Figueroa, J. et al. Probiotic supplementation decreases chemotherapy-induced gastrointestinal side effects in patients with acute leukemia. J. Pediatr. Hematol. Oncol. 41, 468–472 (2019).
Osterlund, P. et al. Lactobacillus supplementation for diarrhoea related to chemotherapy of colorectal cancer: a randomised study. Br. J. Cancer 97, 1028–1034 (2007).
Wada, M. et al. Effects of the enteral administration of Bifidobacterium breve on patients undergoing chemotherapy for pediatric malignancies. Support. Care Cancer 18, 751–759 (2010).
Tian, Y. et al. Effects of probiotics on chemotherapy in patients with lung cancer. Oncol. Lett. 17, 2836–2848 (2019).
Mego, M. et al. Prevention of irinotecan induced diarrhea by probiotics: a randomized double blind, placebo controlled pilot study. Complement. Ther. Med. 23, 356–432 (2015).
Chitapanarux, I. et al. Randomized controlled trial of live Lactobacillus acidophilus plus Bifidobacterium bifidum in prophylaxis of diarrhea during radiotherapy in cervical cancer patients. Radiat. Oncol. 5, 31 (2010).
Wei, D. et al. Probiotics for the prevention or treatment of chemotherapy- or radiotherapy-related diarrhoea in people with cancer. Cochrane Database Syst. Rev. 8, CD008831 (2018).
Jonasch, E. et al. Phase II study of two weeks on, one week off sunitinib scheduling in patients with metastatic renal cell carcinoma. J. Clin. Oncol. 36, 1588–1593 (2018).
Andreyev, J. et al. Guidance on the management of diarrhoea during cancer chemotherapy. Lancet Oncol. 15, e447–e460 (2014).
Wang, Y. et al. Fecal microbiota transplantation for refractory immune checkpoint inhibitor-associated colitis. Nat. Med. 24, 1804–1808 (2018).
Cammarota, G. et al. Fecal microbiota transplantation: a new old kid on the block for the management of gut microbiota-related disease. J. Clin. Gastroenterol. 48, S80–S84 (2014).
O’Toole, P. W., Marchesi, J. R. & Hill, C. Next-generation probiotics: the spectrum from probiotics to live biotherapeutics. Nat. Microbiol. 2, 17057 (2017).
Song, H. et al. Synthetic microbial consortia: from systematic analysis to construction and applications. Chem. Soc. Rev. 43, 6954–6981 (2014).
Yuvaraj, S. et al. E. coli-produced BMP-2 as a chemopreventive strategy for colon cancer: a proof-of-concept study. Gastroenterol. Res. Pract. 2012, 895462 (2012).
Huibregtse, I. L. et al. Genetically modified Lactococcus lactis for delivery of human interleukin-10 to dendritic cells. Gastroenterol. Res. Pract. 2012, 639291 (2012).
Pellegrini, M. et al. Gut microbiota composition after diet and probiotics in overweight breast cancer survivors: a randomized open-label pilot intervention trial. Nutrition 74, 110749 (2020).
Moore, J. H. et al. Preparing next-generation scientists for biomedical big data: artificial intelligence approaches. Per. Med. 16, 247–257 (2019).
Buruk, B., Ekmekci, P. E. & Arda, B. A critical perspective on guidelines for responsible and trustworthy artificial intelligence. Med. Health Care Philos. https://doi.org/10.1007/s11019-020-09948-1 (2020).
Price, W. N. II & Cohen, I. G. Privacy in the age of medical big data. Nat. Med. 25, 37–43 (2019).
van den Bogert, B., Boekhorst, J., Provano, W. & May, A. On the role of bioinformatics and data science in industrial applications. Front. Genet. 10, 721 (2019).
Wang, Y. & Qian, P. Y. Conservative fragments in bacterial 16S rRNA genes and primer design for 16S ribosomal DNA amplicons in metagenomic studies. PLoS One 4, e7401 (2009).
Budding, A. E. et al. Automated broad-range molecular detection of bacteria in clinical samples. J. Clin. Microbiol. 54, 934–943 (2016).
Franzosa, E. A. et al. Relating the metatranscriptome and metagenome of the human gut. Proc. Natl Acad. Sci. USA 111, e2329–e2338 (2014).
Jin, P. et al. Mining the fecal proteome: from biomarkers to personalised medicine. Expert. Rev. Proteom. 14, 445–459 (2017).
Daliri, E. B. et al. The human microbiome and metabolomics: current concepts and applications. Crit. Rev. Food Sci. Nutr. 57, 3565–3576 (2017).
Ranjan, R., Rani, A., Metwally, A., McGee, H. S. & Perkins, D. L. Analysis of the microbiome: advantages of whole genome shotgun versus 16S amplicon sequencing. Biochem. Biophys. Res. Commun. 469, 967–977 (2016).
Vuik, F. et al. Composition of the mucosa-associated microbiota along the entire gastrointestinal tract of human individuals. United European Gastroenterol. J. 7, 897–907 (2019).
Li, S. et al. Pancreatic cyst fluid harbors a unique microbiome. Microbiome 5, 147 (2016).
Acknowledgements
This publication has in part emanated from research conducted with the financial support of AIRC Foundation for Cancer Research (AIRC IG grant number 18599, MFAG grant number 23681) and Science Foundation Ireland (grant number SFI/12/RC/2273).
Author information
Authors and Affiliations
Contributions
The authors contributed equally to all aspects of the article.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information
Nature Reviews Gastroenterology & Hepatology thanks M. Peppelenbosch and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
ColPortal: https://colportal.imib.es
GenBank and Gene Expression Omnibus: https://www.ncbi.nlm.nih.gov/gds
IBD Multi-omics Database: http://ibdmdb.org/
Microbiome Learning Repo: https://knights-lab.github.io/MLRepo/
ML4Microbiome COST Action: https://www.cost.eu/actions/CA18131
ONCOBIOME Project: https://cordis.europa.eu/project/id/825410/it
Supplementary information
Rights and permissions
About this article
Cite this article
Cammarota, G., Ianiro, G., Ahern, A. et al. Gut microbiome, big data and machine learning to promote precision medicine for cancer. Nat Rev Gastroenterol Hepatol 17, 635–648 (2020). https://doi.org/10.1038/s41575-020-0327-3
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41575-020-0327-3
This article is cited by
-
Microbiome as a biomarker and therapeutic target in pancreatic cancer
BMC Microbiology (2024)
-
Anesthesia decision analysis using a cloud-based big data platform
European Journal of Medical Research (2024)
-
Gut microbiome features and metabolites in non-alcoholic fatty liver disease among community-dwelling middle-aged and older adults
BMC Medicine (2024)
-
Utilization of the microbiome in personalized medicine
Nature Reviews Microbiology (2024)
-
The transition from genomics to phenomics in personalized population health
Nature Reviews Genetics (2024)