Abstract
In the past decade, an exciting realization has been that diverse liver diseases — ranging from nonalcoholic steatohepatitis, alcoholic steatohepatitis and cirrhosis to hepatocellular carcinoma — fall along a spectrum. Work on the biology of the gut–liver axis has assisted in understanding the basic biology of both alcoholic fatty liver disease and nonalcoholic fatty liver disease (NAFLD). Of immense importance is the advancement in understanding the role of the microbiome, driven by high-throughput DNA sequencing and improved computational techniques that enable the complexity of the microbiome to be interrogated, together with improved experimental designs. Here, we review gut–liver communications in liver disease, exploring the molecular, genetic and microbiome relationships and discussing prospects for exploiting the microbiome to determine liver disease stage and to predict the effects of pharmaceutical, dietary and other interventions at a population and individual level. Although much work remains to be done in understanding the relationship between the microbiome and liver disease, rapid progress towards clinical applications is being made, especially in study designs that complement human intervention studies with mechanistic work in mice that have been humanized in multiple respects, including the genetic, immunological and microbiome characteristics of individual patients. These ‘avatar mice’ could be especially useful for guiding new microbiome-based or microbiome-informed therapies.
Key points
-
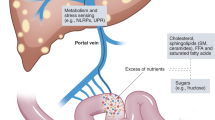
The liver and intestine communicate extensively through the biliary tract, portal vein and systemic mediators.
-
Liver products primarily influence the gut microbiota composition and gut barrier integrity, whereas intestinal factors regulate bile acid synthesis, glucose and lipid metabolism in the liver.
-
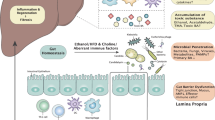
Diverse liver diseases (including nonalcoholic fatty liver disease and alcoholic liver disease) are not unrelated but converge along a common path of progression; pro-inflammatory changes in the liver and intestine mediate development of fibrosis, cirrhosis and, ultimately, hepatocellular carcinoma.
-
Alcoholic and nonalcoholic fatty liver diseases share key characteristics, such as intestinal dysbiosis, gut permeability and shifts in levels of bile acids, ethanol and choline metabolites.
-
Precise contributions of the microbiome to liver diseases could differ based on aetiology; improvements in experimental design and development of animal models are rapidly elucidating causal mechanisms.
-
Advances in understanding the gut–liver axis could encourage research into microbiome-based, diagnostic, prognostic and therapeutic modalities to improve management of liver diseases.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Change history
21 May 2018
In the original version of Table 1 published online, upward arrows to indicate increased translocation of PAMPs were missing from the row entitled ‘Translocation’ for both the column on alcoholic liver disease and nonalcoholic fatty liver disease. This error has now been updated in the PDF and HTML version of the article.
References
Schnabl, B. & Brenner, D. A. Interactions between the intestinal microbiome and liver diseases. Gastroenterology 146, 1513–1524 (2014).
Hartmann, P., Seebauer, C. T. & Schnabl, B. Alcoholic liver disease: the gut microbiome and liver cross talk. Alcohol. Clin. Exp. Res. 39, 763–775 (2015).
Younossi, Z. M. et al. The economic and clinical burden of nonalcoholic fatty liver disease in the United States and Europe. Hepatology 64, 1577–1586 (2016).
Bertola, A., Mathews, S., Ki, S. H., Wang, H. & Gao, B. Mouse model of chronic and binge ethanol feeding (the NIAAA model). Nat. Protoc. 8, 627–637 (2013).
Pelz, S., Stock, P., Brückner, S. & Christ, B. A methionine-choline-deficient diet elicits NASH in the immunodeficient mouse featuring a model for hepatic cell transplantation. Exp. Cell Res. 318, 276–287 (2012).
Itagaki, H., Shimizu, K., Morikawa, S., Ogawa, K. & Ezaki, T. Morphological and functional characterization of non-alcoholic fatty liver disease induced by a methionine-choline-deficient diet in C57BL/6 mice. Int. J. Clin. Exp. Pathol. 6, 2683–2696 (2013).
Loomba, R. et al. Gut microbiome-based metagenomic signature for non-invasive detection of advanced fibrosis in human nonalcoholic fatty liver disease. Cell Metab. 25, 1054–1062.e5 (2017).
Yang, A.-M. et al. Intestinal fungi contribute to development of alcoholic liver disease. J. Clin. Invest. 127, 2829–2841 (2017). This is the first study to implicate the mycobiome in ALD.
Chen, Y.-M. et al. Associations of gut-flora-dependent metabolite trimethylamine-N-oxide, betaine and choline with non-alcoholic fatty liver disease in adults. Sci. Rep. 6, 19076 (2016).
Csak, T. et al. Fatty acid and endotoxin activate inflammasomes in mouse hepatocytes that release danger signals to stimulate immune cells. Hepatology 54, 133–144 (2011).
Uesugi, T., Froh, M., Arteel, G. E., Bradford, B. U. & Thurman, R. G. Toll-like receptor 4 is involved in the mechanism of early alcohol-induced liver injury in mice. Hepatology 34, 101–108 (2001).
Anand, G., Zarrinpar, A. & Loomba, R. Targeting dysbiosis for the treatment of liver disease. Semin. Liver Dis. 36, 37–47 (2016).
Seki, E. & Schnabl, B. Role of innate immunity and the microbiota in liver fibrosis: crosstalk between the liver and gut. J. Physiol. 590, 447–458 (2012).
Jemal, A. et al. Global cancer statistics. CA Cancer J. Clin. 61, 69–90 (2011).
Schwabe, R. F. & Jobin, C. The microbiome and cancer. Nat. Rev. Cancer 13, 800–812 (2013).
Garrett, W. S. Cancer and the microbiota. Science 348, 80–86 (2015).
Koppel, N., Maini Rekdal, V. & Balskus, E. P. Chemical transformation of xenobiotics by the human gut microbiota. Science 356, eaag2770 (2017).
Tolba, R., Kraus, T., Liedtke, C., Schwarz, M. & Weiskirchen, R. Diethylnitrosamine (DEN)-induced carcinogenic liver injury in mice. Lab. Anim. 49, 59–69 (2015).
Stärkel, P. & Schnabl, B. Bidirectional communication between liver and gut during alcoholic liver disease. Semin. Liver Dis. 36, 331–339 (2016).
Chiang, J. Y. L. Bile acid metabolism and signaling. Compr. Physiol. 3, 1191–1212 (2013).
Wahlström, A., Sayin, S. I., Marschall, H. U. & Bäckhed, F. Intestinal crosstalk between bile acids and microbiota and its impact on host metabolism. Cell Metab. 24, 41–50 (2016).
Arab, J. P., Karpen, S. J., Dawson, P. A., Arrese, M. & Trauner, M. Bile acids and nonalcoholic fatty liver disease: molecular insights and therapeutic perspectives. Hepatology 65, 350–362 (2017).
Zarrinpar, A. & Loomba, R. Review article: The emerging interplay among the gastrointestinal tract, bile acids and incretins in the pathogenesis of diabetes and non-alcoholic fatty liver disease. Aliment. Pharmacol. Ther. 36, 909–921 (2012).
Copple, B. L. & Li, T. Pharmacology of bile acid receptors: evolution of bile acids from simple detergents to complex signaling molecules. Pharmacol. Res. 104, 9–21 (2016).
Sinal, C. J. et al. Targeted disruption of the nuclear receptor FXR/BAR impairs bile acid and lipid homeostasis. Cell 102, 731–744 (2000).
Pols, T. W. H., Noriega, L. G., Nomura, M., Auwerx, J. & Schoonjans, K. The bile acid membrane receptor TGR5 as an emerging target in metabolism and inflammation. J. Hepatol. 54, 1263–1272 (2011).
Broeders, E. P. M. et al. The bile acid chenodeoxycholic acid increases human brown adipose tissue activity. Cell Metab. 22, 418–426 (2015).
Thomas, C. et al. TGR5-mediated bile acid sensing controls glucose homeostasis. Cell Metab. 10, 167–177 (2009).
Perino, A. & Schoonjans, K. TGR5 and immunometabolism: insights from physiology and pharmacology. Trends Pharmacol. Sci. 36, 847–857 (2015).
Schaap, F. G., Trauner, M. & Jansen, P. L. M. Bile acid receptors as targets for drug development. Nat. Rev. Gastroenterol. Hepatol. 11, 55–67 (2014).
Inagaki, T. et al. Regulation of antibacterial defense in the small intestine by the nuclear bile acid receptor. Proc. Natl Acad. Sci. USA 103, 3920–3925 (2006).
Parséus, A. et al. Microbiota-induced obesity requires farnesoid X receptor. Gut 66, 429–437 (2017).
Jiang, C. et al. Intestinal farnesoid X receptor signaling promotes nonalcoholic fatty liver disease. J. Clin. Invest. 125, 386–402 (2015).
Ridlon, J. M., Kang, D. J., Hylemon, P. B. & Bajaj, J. S. Bile acids and the gut microbiome. Curr. Opin. Gastroenterol. 30, 332–338 (2014).
Mouzaki, M. et al. Bile acids and dysbiosis in non-alcoholic fatty liver disease. PLoS ONE 11, e0151829 (2016).
Odenwald, M. A. & Turner, J. R. The intestinal epithelial barrier: a therapeutic target? Nat. Rev. Gastroenterol. Hepatol. 14, 9–21 (2017).
Turner, J. R. Intestinal mucosal barrier function in health and disease. Nat. Rev. Immunol. 9, 799–809 (2009).
Abreu, M. T. Toll-like receptor signalling in the intestinal epithelium: how bacterial recognition shapes intestinal function. Nat. Rev. Immunol. 10, 131–144 (2010).
Gallo, R. L. & Hooper, L. V. Epithelial antimicrobial defence of the skin and intestine. Nat. Rev. Immunol. 12, 503–516 (2012).
Mantis, N. J., Rol, N. & Corthésy, B. Secretory IgA’s complex roles in immunity and mucosal homeostasis in the gut. Mucosal Immunol. 4, 603–611 (2011).
Rakoff-Nahoum, S., Paglino, J., Eslami-Varzaneh, F., Edberg, S. & Medzhitov, R. Recognition of commensal microflora by toll-like receptors is required for intestinal homeostasis. Cell 118, 229–241 (2004).
Yaku, K. et al. The enhancement of phase 2 enzyme activities by sodium butyrate in normal intestinal epithelial cells is associated with Nrf2 and p53. Mol. Cell. Biochem. 370, 7–14 (2012).
Wächtershäuser, A. & Stein, J. Rationale for the luminal provision of butyrate in intestinal diseases. Eur. J. Nutr. 39, 164–171 (2000).
Ziegler, K., Kerimi, A., Poquet, L. & Williamson, G. Butyric acid increases transepithelial transport of ferulic acid through upregulation of the monocarboxylate transporters SLC16A1 (MCT1) and SLC16A3 (MCT4). Arch. Biochem. Biophys. 599, 3–12 (2016).
Lobos, O., Barrera, A. & Padilla, C. Microorganisms of the intestinal microbiota of oncorhynchus mykiss produce antagonistic substances against bacteria contaminating food and causing disease in humans. Ital. J. Food Saf. 6, 6240 (2017).
Walsh, C. J., Guinane, C. M., O’ Toole, P. W. & Cotter, P. D. A Profile Hidden Markov Model to investigate the distribution and frequency of LanB-encoding lantibiotic modification genes in the human oral and gut microbiome. PeerJ 5, e3254 (2017).
Graham, C. E., Cruz, M. R., Garsin, D. A. & Lorenz, M. C. Enterococcus faecalis bacteriocin EntV inhibits hyphal morphogenesis, biofilm formation, and virulence of Candida albicans. Proc. Natl Acad. Sci. USA 114, 4507–4512 (2017).
Leclercq, S. et al. Role of intestinal permeability and inflammation in the biological and behavioral control of alcohol-dependent subjects. Brain. Behav. Immun. 26, 911–918 (2012). This paper demonstrates the role of inflammation in ALD and reversibility on abstinence in humans.
Henao-Mejia, J. et al. Inflammasome-mediated dysbiosis regulates progression of NAFLD and obesity. Nature 482, 179–185 (2012). This paper shows the role of inflammation in NAFLD and transferability of symptoms by co-housing mice.
Martinez-Medina, M. et al. Western diet induces dysbiosis with increased E. coli in CEABAC10 mice, alters host barrier function favouring AIEC colonisation. Gut 63, 116–124 (2014).
Serino, M. et al. Metabolic adaptation to a high-fat diet is associated with a change in the gut microbiota. Gut 61, 543–553 (2012).
Pendyala, S., Walker, J. M. & Holt, P. R. A high-fat diet is associated with endotoxemia that originates from the gut. Gastroenterology 142, 1100–1101.e2 (2012).
Wang, Y. et al. Effects of alcohol on intestinal epithelial barrier permeability and expression of tight junction-associated proteins. Mol. Med. Rep. 9, 2352–2356 (2014).
Fukui, H., Brauner, B., Bode, J. C. & Bode, C. Plasma endotoxin concentrations in patients with alcoholic and non-alcoholic liver disease: reevaluation with an improved chromogenic assay. J. Hepatol. 12, 162–169 (1991).
Schäfer, C., Parlesak, A., Schütt, C., Bode, J. C. & Bode, C. Concentrations of lipopolysaccharide-binding protein, bactericidal/permeability-increasing protein, soluble CD14 and plasma lipids in relation to endotoxaemia in patients with alcoholic liver disease. Alcohol Alcohol. 37, 81–86 (2002).
Tulstrup, M. V.-L. et al. Antibiotic treatment affects intestinal permeability and gut microbial composition in wistar rats dependent on antibiotic class. PLoS ONE 10, e0144854 (2015).
Forbes, J. D., Van Domselaar, G. & Bernstein, C. N. The gut microbiota in immune-mediated inflammatory diseases. Front. Microbiol. 7, 1081 (2016).
Grander, C. et al. Recovery of ethanol-induced Akkermansia muciniphila depletion ameliorates alcoholic liver disease. Gut https://doi.org/10.1136/gutjnl-2016-313432 (2017).
Everard, A. et al. Cross-talk between Akkermansia muciniphila and intestinal epithelium controls diet-induced obesity. Proc. Nati. Acad. Sci. USA 110, 9066–9071 (2013).
Elamin, E. E., Masclee, A. A., Dekker, J. & Jonkers, D. M. Ethanol metabolism and its effects on the intestinal epithelial barrier. Nutr. Rev. 71, 483–499 (2013).
Filliol, A. et al. RIPK1 protects hepatocytes from Kupffer cells-mediated TNF-induced apoptosis in mouse models of PAMP-induced hepatitis. J. Hepatol. 66, 1205–1213 (2017).
Ni, Y. H., Huo, L. J. & Li, T. T. Effect of interleukin-22 on proliferation and activation of hepatic stellate cells induced by acetaldehyde and related mechanism [Chinese]. Zhonghua Gan Zang Bing Za Zhi 25, 9–14 (2017).
Wu, X., Wang, Y., Wang, S., Xu, R. & Lv, X. Purinergic P2X7 receptor mediates acetaldehyde-induced hepatic stellate cells activation via PKC-dependent GSK3β pathway. Int Immunopharmacol. 43, 164–171 (2017).
López-Lázaro, M. A local mechanism by which alcohol consumption causes cancer. Oral Oncol. 62, 149–152 (2016).
Zhu, L. et al. Characterization of gut microbiomes in nonalcoholic steatohepatitis (NASH) patients: A connection between endogenous alcohol and NASH. Hepatology 57, 601–609 (2013). This study reports elevated ethanol production by gut microbiota in paediatric patients with NASH.
Miele, L. et al. Increased intestinal permeability and tight junction alterations in nonalcoholic fatty liver disease. Hepatology 49, 1877–1887 (2009).
Pascual, S. et al. Intestinal permeability is increased in patients with advanced cirrhosis. Hepatogastroenterology 50, 1482–1486 (2003).
Philips, C. A. et al. Healthy donor fecal microbiota transplantation in steroid-ineligible severe alcoholic hepatitis: a pilot study. Clin. Gastroenterol. Hepatol. 15, 600–602 (2017).
Seki, E. et al. TLR4 enhances TGF-β signaling and hepatic fibrosis. Nat. Med. 13, 1324–1332 (2007).
Isayama, F. et al. LPS signaling enhances hepatic fibrogenesis caused by experimental cholestasis in mice. Am. J. Physiol. Gastrointest. Liver Physiol. 290, G1318–G1328 (2006).
Gäbele, E. et al. Role of TLR9 in hepatic stellate cells and experimental liver fibrosis. Biochem. Biophys. Res. Commun. 376, 271–276 (2008).
Hartmann, P., Haimerl, M., Mazagova, M., Brenner, D. A. & Schnabl, B. Toll-Like receptor 2-mediated intestinal injury and enteric tumor necrosis factor receptor i contribute to liver fibrosis in mice. Gastroenterology 143, 1330–1340.e1 (2012).
Lebeaupin, C. et al. ER stress induces NLRP3 inflammasome activation and hepatocyte death. Cell Death Dis. 6, e1879 (2015).
Zeisel, S. H. & da Costa, K.-A. Choline: an essential nutrient for public health. Nutr. Rev. 67, 615–623 (2009).
Han, J. et al. Metabolomic profiling distinction of human nonalcoholic fatty liver disease progression from a common rat model. Obesity 25, 1069–1076 (2017).
Muraki, Y., Makita, Y., Yamasaki, M., Amano, Y. & Matsuo, T. Elevation of liver endoplasmic reticulum stress in a modified choline-deficient l -amino acid-defined diet-fed non-alcoholic steatohepatitis mouse model. Biochem. Biophys. Res. Commun. 486, 632–638 (2017).
Rutenburg, A. M. et al. The role of intestinal bacteria in the development of dietary cirrhosis in rats. J. Exp. Med. 106, 1–14 (1957).
Mehedint, M. G. & Zeisel, S. H. Choline’s role in maintaining liver function: new evidence for epigenetic mechanisms. Curr. Opin. Clin. Nutr. Metab. Care 16, 339–345 (2013).
Velasquez, M., Ramezani, A., Manal, A. & Raj, D. Trimethylamine N-oxide: the good, the bad and the unknown. Toxins 8, 326 (2016).
Del Rio, D. et al. The Gut microbial metabolite trimethylamine-N-oxide is present in human cerebrospinal fluid. Nutrients 9, 1053 (2017).
Spencer, M. D. et al. Association between composition of the human gastrointestinal microbiome and development of fatty liver with choline deficiency. Gastroenterology 140, 976–986 (2011).
Gogiashvili, M. et al. Metabolic profiling of ob/ob mouse fatty liver using HR-MAS 1H-NMR combined with gene expression analysis reveals alterations in betaine metabolism and the transsulfuration pathway. Anal. Bioanal. Chem. 409, 1591–1606 (2017).
Sherriff, J. L., O.Sullivan, T. A., Properzi, C., Oddo, J.-L. & Adams, L. A. Choline, its potential role in nonalcoholic fatty liver disease, and the case for human and bacterial genes. Adv. Nutr. An. Int. Rev. J. 7, 5–13 (2016).
Chen, P. et al. Supplementation of saturated long-chain fatty acids maintains intestinal eubiosis and reduces ethanol-induced liver injury in mice. Gastroenterology 148, 203–214.e16 (2015).
Cresci, G. A. et al. Prophylactic tributyrin treatment mitigates chronic-binge alcohol-induced intestinal barrier and liver injury. J. Gastroenterol. Hepatol. 32, 1587–1597 (2017). This paper describes an example of microbiome-based therapeutics for liver disease management.
Shi, X. et al. Hepatic and fecal metabolomic analysis of the effects of Lactobacillus rhamnosus gg on alcoholic fatty liver disease in mice. J. Proteome Res. 14, 1174–1182 (2015).
Kim, D.-H. et al. Dual function of Lactobacillus kefiri DH5 in preventing high-fat-diet-induced obesity: direct reduction of cholesterol and upregulation of PPARα in adipose tissue. Mol. Nutr. Food Res. 61, 1700252 (2017).
Nanji, A. A., Khettry, U. & Sadrzadeh, S. M. Lactobacillus feeding reduces endotoxemia and severity of experimental alcoholic liver (disease). Proc. Soc. Exp. Biol. Med. 205, 243–247 (1994).
Forsyth, C. B. et al. Lactobacillus GG treatment ameliorates alcohol-induced intestinal oxidative stress, gut leakiness, and liver injury in a rat model of alcoholic steatohepatitis. Alcohol 43, 163–172 (2009).
Loguercio, C. et al. Beneficial effects of a probiotic VSL#3 on parameters of liver dysfunction in chronic liver diseases. J. Clin. Gastroenterol. 39, 540–543 (2005).
Kirpich, I. A. et al. Probiotics restore bowel flora and improve liver enzymes in human alcohol-induced liver injury: a pilot study. Alcohol 42, 675–682 (2008).
Stadlbauer, V. et al. Effect of probiotic treatment on deranged neutrophil function and cytokine responses in patients with compensated alcoholic cirrhosis. J. Hepatol. 48, 945–951 (2008).
Chen, R.-C. et al. Lactobacillus rhamnosus GG supernatant promotes intestinal barrier function, balances Treg and TH17 cells and ameliorates hepatic injury in a mouse model of chronic-binge alcohol feeding. Toxicol. Lett. 241, 103–110 (2016).
Levitt, M. D. et al. Use of measurements of ethanol absorption from stomach and intestine to assess human ethanol metabolism. Am. J. Physiol. 273, G951–G957 (1997).
Norberg, A., Jones, A. W., Hahn, R. G. & Gabrielsson, J. L. Role of variability in explaining ethanol pharmacokinetics. Clin. Pharmacokinet. 42, 1–31 (2003).
Hamarneh, S. R. et al. Intestinal alkaline phosphatase attenuates alcohol-induced hepatosteatosis in mice. Dig. Dis. Sci. 62, 2021–2034 (2017).
Chen, P. et al. Microbiota protects mice against acute alcohol-induced liver injury. Alcohol. Clin. Exp. Res. 39, 2313–2323 (2015).
Ansari, R., Husain, K. & Rizvi, S. Role of transcription factors in steatohepatitis and hypertension after ethanol: the epicenter of metabolism. Biomolecules 6, 29 (2016).
Setshedi, M., Wands, J. R. & de la Monte, S. M. Acetaldehyde adducts in alcoholic liver disease. Oxid. Med. Cell. Longev. 3, 178–185 (2010).
Rao, R. K. Acetaldehyde-induced barrier disruption and paracellular permeability in caco-2 cell monolayer. Methods Mol. Biol. 447, 171–183 (2008).
Mir, H. et al. Occludin deficiency promotes ethanol-induced disruption of colonic epithelial junctions, gut barrier dysfunction and liver damage in mice. Biochim. Biophys. Acta 1860, 765–774 (2016).
Chaudhry, K. K. et al. Glutamine supplementation attenuates ethanol-induced disruption of apical junctional complexes in colonic epithelium and ameliorates gut barrier dysfunction and fatty liver in mice. J. Nutr. Biochem. 27, 16–26 (2016).
Chen, P., Stärkel, P., Turner, J. R., Ho, S. B. & Schnabl, B. Dysbiosis-induced intestinal inflammation activates tumor necrosis factor receptor I and mediates alcoholic liver disease in mice. Hepatology 61, 883–894 (2015).
Forsyth, C. B., Voigt, R. M., Burgess, H. J., Swanson, G. R. & Keshavarzian, A. Circadian rhythms, alcohol and gut interactions. Alcohol 49, 389–398 (2015).
Yan, A. W. & Schnabl, B. Bacterial translocation and changes in the intestinal microbiome associated with alcoholic liver disease. World J. Hepatol. 4, 110–118 (2012).
Yan, A. W. et al. Enteric dysbiosis associated with a mouse model of alcoholic liver disease. Hepatology 53, 96–105 (2011).
Hartmann, P. et al. Deficiency of intestinal mucin-2 ameliorates experimental alcoholic liver disease in mice. Hepatology 58, 108–119 (2013).
Park, B., Lee, H.-R. & Lee, Y.-J. Alcoholic liver disease: focus on prodromal gut health. J. Dig. Dis. 17, 493–500 (2016).
Wang, H., Lafdil, F., Kong, X. & Gao, B. Signal transducer and activator of transcription 3 in liver diseases: a novel therapeutic target. Int. J. Biol. Sci. 7, 536–550 (2011).
Mottaran, E. et al. Lipid peroxidation contributes to immune reactions associated with alcoholic liver disease. Free Radic. Biol. Med. 32, 38–45 (2002).
Xie, G. et al. Chronic Ethanol consumption alters mammalian gastrointestinal content metabolites. J. Proteome Res. 12, 3297–3306 (2013).
Couch, R. D. et al. Alcohol induced alterations to the human fecal VOC metabolome. PLoS ONE 10, e0119362 (2015).
Leclercq, S. et al. Intestinal permeability, gut-bacterial dysbiosis, and behavioral markers of alcohol-dependence severity. Proc. Natl Acad. Sci. USA 111, E4485–E4493 (2014).
Arroyo, V. et al. Acute-on-chronic liver failure in cirrhosis. Nat. Rev. Dis. Primers 2, 16041 (2016).
Cresci, G. A., Bush, K. & Nagy, L. E. Tributyrin supplementation protects mice from acute ethanol-induced gut injury. Alcohol. Clin. Exp. Res. 38, 1489–1501 (2014).
Spengler, E. K. & Loomba, R. Recommendations for diagnosis, referral for liver biopsy, and treatment of nonalcoholic fatty liver disease and nonalcoholic steatohepatitis. Mayo Clin. Proc. 90, 1233–1246 (2015).
Loomba, R., Abraham, M. & Unalp, A. Association between diabetes, family history of diabetes and risk of nonalcoholic steatohepatitis and fibrosis. Hepatology 56, 943–951 (2012).
Doycheva, I. et al. Non-invasive screening of diabetics in primary care for NAFLD and advanced fibrosis by MRI and MRE. Aliment. Pharmacol. Ther. 43, 83–95 (2016).
Loomba, R. et al. Heritability of hepatic fibrosis and steatosis based on a prospective twin study. Gastroenterology 149, 1784–1793 (2015).
Cui, J. et al. Shared genetic effects between hepatic steatosis and fibrosis: a prospective twin study. Hepatology 64, 1547–1558 (2016).
Caussy, C. et al. Nonalcoholic fatty liver disease with cirrhosis increases familial risk for advanced fibrosis. J. Clin. Invest. 127, 2697–2704 (2017).
Gao, B. & Bataller, R. Alcoholic liver disease: pathogenesis and new therapeutic targets. Gastroenterology 141, 1572–1585 (2011).
Wieland, A., Frank, D. N., Harnke, B. & Bambha, K. Systematic review: microbial dysbiosis and nonalcoholic fatty liver disease. Aliment. Pharmacol. Ther. 42, 1051–1063 (2015).
Kapil, S. et al. Small intestinal bacterial overgrowth and toll-like receptor signaling in patients with non-alcoholic fatty liver disease. J. Gastroenterol. Hepatol. 31, 213–221 (2016).
Boursier, J. et al. The severity of nonalcoholic fatty liver disease is associated with gut dysbiosis and shift in the metabolic function of the gut microbiota. Hepatology 63, 764–775 (2016).
Bajaj, J. S. et al. Salivary microbiota reflects changes in gut microbiota in cirrhosis with hepatic encephalopathy. Hepatology 62, 1260–1271 (2015).
Rahman, K. et al. Loss of Junctional adhesion molecule a promotes severe steatohepatitis in mice on a diet high in saturated fat, fructose, and cholesterol. Gastroenterology 151, 733–746.e12 (2016).
Arendt, B. M. et al. Nonalcoholic fatty liver disease is associated with lower hepatic and erythrocyte ratios of phosphatidylcholine to phosphatidylethanolamine. Appl. Physiol. Nutr. Metab. 38, 334–340 (2013).
Rao, R. K., Seth, A. & Sheth, P. Recent advances in alcoholic liver disease I. Role of intestinal permeability and endotoxemia in alcoholic liver disease. Am. J. Physiol. Gastrointest. Liver Physiol. 286, G881–G884 (2004).
Ferrier, L. et al. Impairment of the intestinal barrier by ethanol involves enteric microflora and mast cell activation in rodents. Am. J. Pathol. 168, 1148–1154 (2006).
Ferrere, G. et al. Fecal microbiota manipulation prevents dysbiosis and alcohol-induced liver injury in mice. J. Hepatol. 66, 806–815 (2017). This study shows that FMT could prevent alcohol-induced liver damage.
Mutlu, E. A. et al. Colonic microbiome is altered in alcoholism. Am. J. Physiol. Gastrointest. Liver Physiol. 302, G966–G978 (2012).
Tuomisto, S. et al. Changes in gut bacterial populations and their translocation into liver and ascites in alcoholic liver cirrhotics. BMC Gastroenterol. 14, 40 (2014).
Chen, Y. et al. Characterization of fecal microbial communities in patients with liver cirrhosis. Hepatology 54, 562–572 (2011).
Wang, L. et al. Intestinal REG3 lectins protect against alcoholic steatohepatitis by reducing mucosa-associated microbiota and preventing bacterial translocation. Cell Host Microbe 19, 227–239 (2016).
Inamine, T. et al. Genetic loss of immunoglobulin A does not influence development of alcoholic steatohepatitis in mice. Alcohol. Clin. Exp. Res. 40, 2604–2613 (2016).
Adachi, Y., Bradford, B. U., Gao, W., Bojes, H. K. & Thurman, R. G. Inactivation of Kupffer cells prevents early alcohol-induced liver injury. Hepatology 20, 453–460 (1994).
Seo, W. & Jeong, W. Il. Hepatic non-parenchymal cells: Master regulators of alcoholic liver disease? World J. Gastroenterol. 22, 1348–1356 (2016).
Ju, C. & Mandrekar, P. Macrophages and alcohol-related liver inflammation. Alcohol Res. 37, 251–262 (2015).
Tilg, H., Moschen, A. R. & Szabo, G. Interleukin-1 and inflammasomes in alcoholic liver disease/acute alcoholic hepatitis and nonalcoholic fatty liver disease/nonalcoholic steatohepatitis. Hepatology 64, 955–965 (2016).
Axelson, M., Mörk, B. & Sjövall, J. Ethanol has an acute effect on bile acid biosynthesis in man. FEBS Lett. 281, 155–159 (1991).
Xie, G. et al. Alteration of bile acid metabolism in the rat induced by chronic ethanol consumption. FASEB J. 27, 3583–3593 (2013).
Wu, W.-B. et al. Excessive bile acid activated NF-kappa B and promoted the development of alcoholic steatohepatitis in farnesoid X receptor deficient mice. Biochimie 115, 86–92 (2015).
Wu, W.-B. et al. Agonist of farnesoid X receptor protects against bile acid induced damage and oxidative stress in mouse placenta — a study on maternal cholestasis model. Placenta 36, 545–551 (2015).
Bhat, M. et al. Implication of the intestinal microbiome in complications of cirrhosis. World J. Hepatol. 8, 1128–1136 (2016).
National Institute of Diabetes and Digestive and Kidney Diseases. Cirrhosis. NIDDK https://www.niddk.nih.gov/health-information/liver-disease/cirrhosis (2014).
Mells, G. F. et al. Genome-wide association study identifies 12 new susceptibility loci for primary biliary cirrhosis. Nat. Genet. 43, 329–332 (2011).
Charlton, M. R. et al. Frequency and outcomes of liver transplantation for nonalcoholic steatohepatitis in the United States. Gastroenterology 141, 1249–1253 (2011).
Yang, J. D. et al. Diabetes mellitus heightens the risk of hepatocellular carcinoma except in patients with hepatitis C cirrhosis. Am. J. Gastroenterol. 111, 1573–1580 (2016).
Bajaj, J. S. et al. Gut microbiota alterations can predict hospitalizations in cirrhosis independent of diabetes mellitus. Sci. Rep. 5, 18559 (2016).
Jun, D. W. et al. Association between small intestinal bacterial overgrowth and peripheral bacterial dna in cirrhotic patients. Dig. Dis. Sci. 55, 1465–1471 (2010).
Yao, J., Chang, L., Yuan, L. & Duan, Z. Nutrition status and small intestinal bacterial overgrowth in patients with virus-related cirrhosis. Asia Pac. J. Clin. Nutr. 25, 283–291 (2016).
Chen, Y. et al. Dysbiosis of small intestinal microbiota in liver cirrhosis and its association with etiology. Sci. Rep. 6, 34055 (2016).
Mas, A. et al. Comparison of rifaximin and lactitol in the treatment of acute hepatic encephalopathy: results of a randomized, double-blind, double-dummy, controlled clinical trial. J. Hepatol. 38, 51–58 (2003).
Bajaj, J. S. et al. Rifaximin improves driving simulator performance in a randomized trial of patients with minimal hepatic encephalopathy. Gastroenterology 140, 478–487.e1 (2011).
Vlachogiannakos, J. et al. Long-term administration of rifaximin improves the prognosis of patients with decompensated alcoholic cirrhosis. J. Gastroenterol. Hepatol. 28, 450–455 (2013).
Qin, N. et al. Alterations of the human gut microbiome in liver cirrhosis. Nature 513, 59–64 (2014).
Bajaj, J. S. et al. Fungal dysbiosis in cirrhosis. Gut https://doi.org/10.1136/gutjnl-2016-313170 (2017). This article discusses the role of the mycobiome in cirrhosis.
Fouts, D. E., Torralba, M., Nelson, K. E., Brenner, D. A. & Schnabl, B. Bacterial translocation and changes in the intestinal microbiome in mouse models of liver disease. J. Hepatol. 56, 1283–1292 (2012).
Yoshimoto, S. et al. Obesity-induced gut microbial metabolite promotes liver cancer through senescence secretome. Nature 499, 97–101 (2013). In this study, deoxycholic acid, a gut-microbiota-derived bile acid, is shown to promote HCC.
Xie, G. et al. Distinctly altered gut microbiota in the progression of liver disease. Oncotarget 7, 19355–19366 (2016).
Grat, M. et al. Relevance of pre-transplant α-fetoprotein dynamics in liver transplantation for hepatocellular cancer. Ann. Transplant. 21, 115–124 (2016).
Fox, J. G. et al. Gut microbes define liver cancer risk in mice exposed to chemical and viral transgenic hepatocarcinogens. Gut 59, 88–97 (2010).
Rogers, A. B. Distance burning. Gut Microbes 2, 52–57 (2011).
Huang, Y. et al. Identification of helicobacter species in human liver samples from patients with primary hepatocellular carcinoma. J. Clin. Pathol. 57, 1273–1277 (2004).
Krüttgen, A. et al. Study on the association of helicobacter species with viral hepatitis-induced hepatocellular carcinoma. Gut Microbes 3, 228–233 (2012).
Ward, J. M. et al. Chronic active hepatitis and associated liver tumors in mice caused by a persistent bacterial infection with a novel Helicobacter species. J. Natl Cancer Inst. 86, 1222–1227 (1994).
Mima, K. et al. The microbiome and hepatobiliary-pancreatic cancers. Cancer Lett. 402, 9–15 (2017).
Brandtzaeg, P. Secretory IgA: designed for anti-microbial defense. Front. Immunol. 4, 222 (2013).
Shalapour, S. et al. Inflammation-induced IgA+ cells dismantle anti-liver cancer immunity. Nature 551, 340–345 (2017).
Dapito, D. H. et al. Promotion of hepatocellular carcinoma by the intestinal microbiota and TLR4. Cancer Cell 21, 504–516 (2012). This paper demonstrates the role of the gut microbiota in HCC development.
Xie, G. et al. Dysregulated hepatic bile acids collaboratively promote liver carcinogenesis. Int. J. Cancer 139, 1764–1775 (2016).
Ruhland, M. K. et al. Stromal senescence establishes an immunosuppressive microenvironment that drives tumorigenesis. Nat. Commun. 7, 11762 (2016).
Demaria, M. et al. Cellular senescence promotes adverse effects of chemotherapy and cancer relapse. Cancer Discov. 7, 165–176 (2017).
Gomes, A. L. et al. Metabolic inflammation-associated IL-17A causes non-alcoholic steatohepatitis and hepatocellular carcinoma. Cancer Cell 30, 161–175 (2016).
Li, J. et al. Interleukin 17A promotes hepatocellular carcinoma metastasis via NF-kB induced matrix metalloproteinases 2 and 9 expression. PLoS ONE 6, e21816 (2011).
Hammerich, L., Heymann, F. & Tacke, F. Role of IL-17 and Th17 Cells in liver diseases. Clin. Dev. Immunol. 2011, 1–12 (2011).
Viaud, S. et al. The intestinal microbiota modulates the anticancer immune effects of cyclophosphamide. Science 342, 971–976 (2013).
Neuschwander-Tetri, B. A. et al. Farnesoid X nuclear receptor ligand obeticholic acid for non-cirrhotic, non-alcoholic steatohepatitis (FLINT): a multicentre, randomised, placebo-controlled trial. Lancet 385, 956–965 (2015).
Thiele, M., Wiest, R., Gluud, L. L., Albillos, A. & Krag, A. Can non-selective beta-blockers prevent hepatocellular carcinoma in patients with cirrhosis? Med. Hypotheses 81, 871–874 (2013).
Li, J. et al. Probiotics modulated gut microbiota suppresses hepatocellular carcinoma growth in mice. Proc. Natl Acad. Sci. USA 113, E1306–E1315 (2016).
Forslund, K. et al. Disentangling type 2 diabetes and metformin treatment signatures in the human gut microbiota. Nature 528, 262–266 (2015).
Caporaso, J. G. et al. Moving pictures of the human microbiome. Genome Biol. 12, R50 (2011).
Halfvarson, J. et al. Dynamics of the human gut microbiome in inflammatory bowel disease. Nat. Microbiol. 2, 17004 (2017).
Wu, G. D. et al. Linking long-term dietary patterns with gut microbial enterotypes. Science 334, 105–108 (2011).
Noguera-Julian, M. et al. Gut microbiota linked to sexual preference and HIV infection. EBioMedicine 5, 135–146 (2016).
Jackson, M. A. et al. Proton pump inhibitors alter the composition of the gut microbiota. Gut 65, 749–756 (2016).
Goodrich, J. K. et al. Human genetics shape the gut microbiome. Cell 159, 789–799 (2014).
Debelius, J. et al. Tiny microbes, enormous impacts: what matters in gut microbiome studies? Genome Biol. 17, 217 (2016).
Lozupone, C. A. et al. Meta-analyses of studies of the human microbiota. Genome Res. 23, 1704–1714 (2013).
Sinha, R., Abnet, C. C., White, O., Knight, R. & Huttenhower, C. The microbiome quality control project: baseline study design and future directions. Genome Biol. 16, 276 (2015).
van Dongen, J., Slagboom, P. E., Draisma, H. H. M., Martin, N. G. & Boomsma, D. I. The continuing value of twin studies in the omics era. Nat. Rev. Genet. 13, 640–653 (2012).
Smith, M. I. et al. Gut microbes of Malawian twin pairs discordant for kwashiorkor. Science 339, 548–554 (2013).
Goodrich, J. K. et al. Genetic determinants of the gut microbiome in UK twins. Cell Host Microbe 19, 731–743 (2016).
Sookoian, S. & Pirola, C. J. Genetic predisposition in nonalcoholic fatty liver disease. Clin. Mol. Hepatol. 23, 1–12 (2017).
Weingarden, A. et al. Dynamic changes in short- and long-term bacterial composition following fecal microbiota transplantation for recurrent Clostridium difficile infection. Microbiome 3, 10 (2015).
Zaneveld, J. R., McMinds, R. & Vega Thurber, R. Stress and stability: applying the Anna Karenina principle to animal microbiomes. Nat. Microbiol. 2, 17121 (2017). This article explains how instability as a community characteristic might be important to understand inflammatory disease.
Piscaglia, F. et al. Clinical patterns of hepatocellular carcinoma in nonalcoholic fatty liver disease: a multicenter prospective study. Hepatology 63, 827–838 (2016).
Kostic, A. D. et al. The dynamics of the human infant gut microbiome in development and in progression toward type 1 diabetes. Cell Host Microbe 17, 260–273 (2015).
Nguyen, T. L. A., Vieira-Silva, S., Liston, A. & Raes, J. How informative is the mouse for human gut microbiota research? Dis. Model. Mech. 8, 1–16 (2015).
Manichanh, C. et al. Reshaping the gut microbiome with bacterial transplantation and antibiotic intake. Genome Res. 20, 1411–1419 (2010).
Kiraly, D. D. et al. Alterations of the host microbiome affect behavioral responses to cocaine. Sci. Rep. 6, 35455 (2016).
Sampson, T. R. et al. Gut microbiota regulate motor deficits and neuroinflammation in a model of Parkinson’s disease. Cell 167, 1469–1480.e12 (2016).
Markle, J. G. M. et al. Sex differences in the gut microbiome drive hormone-dependent regulation of autoimmunity. Science 339, 1084–1088 (2013).
Llopis, M. et al. Intestinal microbiota contributes to individual susceptibility to alcoholic liver disease. Gut 65, 830–839 (2016).
Byrne, A. T. et al. Interrogating open issues in cancer precision medicine with patient-derived xenografts. Nat. Rev. Cancer 17, 254–268 (2017).
Hidalgo, M. et al. Patient-derived xenograft models: an emerging platform for translational cancer research. Cancer Discov. 4, 998–1013 (2014).
Mattner, J. Impact of microbes on the pathogenesis of primary biliary cirrhosis (PBC) and primary sclerosing cholangitis (PSC). Int. J. Mol. Sci. 17, 1864 (2016).
Verdier, J., Luedde, T. & Sellge, G. Biliary mucosal barrier and microbiome. Viszeralmedizin 31, 156–161 (2015).
Miyake, Y. & Yamamoto, K. Role of gut microbiota in liver diseases. Hepatol. Res. 43, 139–146 (2013).
Pflughoeft, K. J. & Versalovic, J. Human microbiome in health and disease. Annu. Rev. Pathol. Mech. Dis. 7, 99–122 (2012).
Bogdanos, D.-P. et al. Primary biliary cirrhosis is characterized by IgG3 antibodies cross-reactive with the major mitochondrial autoepitope and its Lactobacillus mimic. Hepatology 42, 458–465 (2005).
Padgett, K. et al. Phylogenetic and immunological definition of four lipoylated proteins from, implications for primary biliary cirrhosis. J. Autoimmun. 24, 209–219 (2005).
Mohammed, J. P. et al. Identification of Cd101 as a susceptibility gene for Novosphingobium aromaticivorans-induced liver autoimmunity. J. Immunol. 187, 337–349 (2011).
Lee, J.-Y. et al. Contribution of the 7β-hydroxysteroid dehydrogenase from Ruminococcus gnavus N53 to ursodeoxycholic acid formation in the human colon. J. Lipid Res. 54, 3062–3069 (2013).
Olsson, R. et al. Bile duct bacterial isolates in primary sclerosing cholangitis: a study of explanted livers. J. Hepatol. 28, 426–432 (1998).
Pollheimer, M. J., Halilbasic, E., Fickert, P. & Trauner, M. Pathogenesis of primary sclerosing cholangitis. Best Pract. Res. Clin. Gastroenterol. 25, 727–739 (2011).
Toyoki, Y. et al. Semiquantitative evaluation of hepatic fibrosis by measuring tissue hydroxyproline. Hepatogastroenterology 45, 2261–2264 (1998).
Karrar, A. et al. Biliary epithelial cell antibodies link adaptive and innate immune responses in primary sclerosing cholangitis. Gastroenterology 132, 1504–1514 (2007).
Katt, J. et al. Increased T helper type 17 response to pathogen stimulation in patients with primary sclerosing cholangitis. Hepatology 58, 1084–1093 (2013).
Loftus, E. V., Sandborn, W. J., Lindor, K. D. & Larusso, N. F. Interactions between chronic liver disease and inflammatory bowel disease. Inflamm. Bowel Dis. 3, 288–302 (1997).
Bode, J. C., Bode, C., Heidelbach, R., Dürr, H. K. & Martini, G. A. Jejunal microflora in patients with chronic alcohol abuse. Hepatogastroenterology 31, 30–34 (1984).
Bull-Otterson, L. et al. Metagenomic analyses of alcohol induced pathogenic alterations in the intestinal microbiome and the effect of Lactobacillus rhamnosus GG treatment. PLoS ONE 8, e53028 (2013).
Jiang, W. et al. Dysbiosis gut microbiota associated with inflammation and impaired mucosal immune function in intestine of humans with non-alcoholic fatty liver disease. Sci. Rep. 5, 8096 (2015).
Raman, M. et al. Fecal microbiome and volatile organic compound metabolome in obese humans with nonalcoholic fatty liver disease. Clin. Gastroenterol. Hepatol. 11, 868–875.e3 (2013).
Bäckhed, F. et al. The gut microbiota as an environmental factor that regulates fat storage. Proc. Natl Acad. Sci. USA 101, 15718–15723 (2004).
Bäckhed, F., Manchester, J. K., Semenkovich, C. F. & Gordon, J. I. Mechanisms underlying the resistance to diet-induced obesity in germ-free mice. Proc. Natl Acad. Sci. USA 104, 979–984 (2007).
Achur, R. N., Freeman, W. M. & Vrana, K. E. Circulating cytokines as biomarkers of alcohol abuse and alcoholism. J. Neuroimmune Pharmacol. 5, 83–91 (2010).
Luck, H. et al. Regulation of obesity-related insulin resistance with gut anti-inflammatory agents. Cell Metab. 21, 527–542 (2015).
Luther, J. et al. Hepatic Injury in nonalcoholic steatohepatitis contributes to altered intestinal permeability. Cell. Mol. Gastroenterol. Hepatol. 1, 222–232 (2015).
Le Roy, T. et al. Intestinal microbiota determines development of non-alcoholic fatty liver disease in mice. Gut 62, 1787–1794 (2013).
Bala, S., Marcos, M., Gattu, A., Catalano, D. & Szabo, G. Acute binge drinking increases serum endotoxin and bacterial DNA levels in healthy individuals. PLoS ONE 9, e96864 (2014).
Bode, C., Kugler, V. & Bode, J. C. Endotoxemia in patients with alcoholic and non-alcoholic cirrhosis and in subjects with no evidence of chronic liver disease following acute alcohol excess. J. Hepatol. 4, 8–14 (1987).
Parlesak, A., Schäfer, C., Schütz, T., Bode, J. C. & Bode, C. Increased intestinal permeability to macromolecules and endotoxemia in patients with chronic alcohol abuse in different stages of alcohol-induced liver disease. J. Hepatol. 32, 742–747 (2000).
Roh, Y. S., Zhang, B., Loomba, R. & Seki, E. TLR2 and TLR9 contribute to alcohol-mediated liver injury through induction of CXCL1 and neutrophil infiltration. Am. J. Physiol. Gastrointest. Liver Physiol. 309, G30–G41 (2015).
Jin, R. et al. Fructose induced endotoxemia in pediatric nonalcoholic fatty liver disease. Int. J. Hepatol. 2014, 560620 (2014).
Mridha, A. R. et al. TLR9 is up-regulated in human and murine NASH: pivotal role in inflammatory recruitment and cell survival. Clin. Sci. 131, 2145–2159 (2017).
Alm, R., Carlson, J. & Eriksson, S. Fasting serum bile acids in liver disease. A comparison with histological features. Scand. J. Gastroenterol. 17, 213–218 (1982).
Ferslew, B. C. et al. Altered bile acid metabolome in patients with nonalcoholic steatohepatitis. Dig. Dis. Sci. 60, 3318–3328 (2015).
Fernando, H., Bhopale, K. K., Kondraganti, S., Kaphalia, B. S. & Shakeel Ansari, G. A. Lipidomic changes in rat liver after long-term exposure to ethanol. Toxicol. Appl. Pharmacol. 255, 127–137 (2011).
Fernando, H. et al. 1H and 31P NMR lipidome of ethanol-induced fatty liver. Alcohol. Clin. Exp. Res. 34, 1937–1947 (2010).
Dumas, M.-E. et al. Metabolic profiling reveals a contribution of gut microbiota to fatty liver phenotype in insulin-resistant mice. Proc. Natl Acad. Sci. USA 103, 12511–12516 (2006).
Liu, J., Han, L., Zhu, L. & Yu, Y. Free fatty acids, not triglycerides, are associated with non-alcoholic liver injury progression in high fat diet induced obese rats. Lipids Health Dis. 15, 27 (2016).
Volynets, V. et al. Nutrition, intestinal permeability, and blood ethanol levels are altered in patients with nonalcoholic fatty liver disease (NAFLD). Dig. Dis. Sci. 57, 1932–1941 (2012).
Engstler, A. J. et al. Insulin resistance alters hepatic ethanol metabolism: studies in mice and children with non-alcoholic fatty liver disease. Gut 65, 1564–1571 (2016).
Nakamura, A. & Terauchi, Y. Lessons from mouse models of high-fat diet-induced NAFLD. Int. J. Mol. Sci. 14, 21240–21257 (2013).
Ishioka, M., Miura, K., Minami, S., Shimura, Y. & Ohnishi, H. Altered gut microbiota composition and immune response in experimental steatohepatitis mouse models. Dig. Dis. Sci. 62, 396–406 (2017).
Lieber, C. S. & DeCarli, L. M. The feeding of alcohol in liquid diets: two decades of applications and 1982 update. Alcohol. Clin. Exp. Res. 6, 523–531 (1982).
Tsukamoto, H., Reidelberger, R. D., French, S. W. & Largman, C. Long-term cannulation model for blood sampling and intragastric infusion in the rat. Am. J. Physiol. 247, R595–R599 (1984).
Ronis, M. J. J. et al. Increased 4-hydroxynonenal protein adducts in male GSTA4-4/PPAR-α double knockout mice enhance injury during early stages of alcoholic liver disease. Am. J. Physiol. Gastrointest. Liver Physiol. 308, G403–G415 (2015).
Ericsson, A. C. et al. Effects of vendor and genetic background on the composition of the fecal microbiota of inbred mice. PLoS ONE 10, e0116704 (2015).
Stappenbeck, T. S. & Virgin, H. W. Accounting for reciprocal host-microbiome interactions in experimental science. Nature 534, 191–199 (2016).
Nakagawa, H. et al. Loss of liver E-cadherin induces sclerosing cholangitis and promotes carcinogenesis. Proc. Natl Acad. Sci. USA 111, 1090–1095 (2014).
Etienne-Mesmin, L., Vijay-Kumar, M., Gewirtz, A. T. & Chassaing, B. Hepatocyte Toll-like receptor 5 promotes bacterial clearance and protects mice against high-fat diet-induced liver disease. Cell. Mol. Gastroenterol. Hepatol. 2, 584–604 (2016). This well-designed mouse study shows the importance of both inflammatory and tolerizing bacteria in regulating liver inflammation.
Wu, T. et al. Multimodal imaging of a humanized orthotopic model of hepatocellular carcinoma in immunodeficient mice. Sci. Rep. 6, 35230 (2016).
Round, J. L. & Mazmanian, S. K. The gut microbiota shapes intestinal immune responses during health and disease. Nat. Rev. Immunol. 9, 313–323 (2009).
Acknowledgements
The authors thank D. McDonald, T. Kosciółek, Z. Xu and A. Plymoth for their helpful discussions. M.K. is supported by NIH grants R01 AI043477 and R01 CA118165. R.L. is supported in part by grant R01-DK106419-03. Research reported in this publication was supported in part by the National Institute of Environmental Health Sciences of the NIH under award number P42ES010337. B.S. is supported by NIH grants R01 AA020703, U01 AA021856 and U01AA24726 and by award number I01BX002213 from the Biomedical Laboratory Research and Development Service of the VA Office of Research and Development. J.D. is supported by the Robert Wood Johnson Foundation. The content is solely the responsibility of the authors and does not necessarily represent the official views of the NIH.
Author information
Authors and Affiliations
Contributions
A.T., J.D., D.A.B., M.K., R.L., B.S. and R.K. researched data for the article. A.T., J.D., D.A.B., M.K., R.L., B.S. and R.K. made substantial contributions to discussion of content. A.T., D.A.B., M.K., R.L., B.S. and R.K. reviewed and edited the manuscript before submission. A.T., J.D. and R.K. wrote the article.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Qiita: http://qiita.microbio.me
Rights and permissions
About this article
Cite this article
Tripathi, A., Debelius, J., Brenner, D.A. et al. The gut–liver axis and the intersection with the microbiome. Nat Rev Gastroenterol Hepatol 15, 397–411 (2018). https://doi.org/10.1038/s41575-018-0011-z
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41575-018-0011-z
This article is cited by
-
Jian Gan powder ameliorates immunological liver injury in mice by modulating the gut microbiota and metabolic profiles
European Journal of Medical Research (2024)
-
Advances in the study of the interaction between schistosome infections and the host's intestinal microorganisms
Parasites & Vectors (2024)
-
From Cirrhosis to the Dysbiosis (A Loop of Cure or Complications?)
Indian Journal of Microbiology (2024)
-
A Novel Bifidobacterium/Klebsiella Ratio in Characterization Analysis of the Gut and Bile Microbiota of CCA Patients
Microbial Ecology (2024)
-
Splenectomy ameliorates liver cirrhosis by restoring the gut microbiota balance
Cellular and Molecular Life Sciences (2024)