Abstract
Osteoporosis is a highly prevalent disorder characterized by low bone mineral density and an increased risk of fracture, termed osteoporotic fracture. Notably, bone mineral density, osteoporosis and osteoporotic fracture are highly heritable; however, determining the genetic architecture, and especially the underlying genomic and molecular mechanisms, of osteoporosis in vivo in humans is still challenging. In addition to susceptibility loci identified in genome-wide association studies, advances in various omics technologies, including genomics, transcriptomics, epigenomics, proteomics and metabolomics, have all been applied to dissect the pathogenesis of osteoporosis. However, each technology individually cannot capture the entire view of the disease pathology and thus fails to comprehensively identify the underlying pathological molecular mechanisms, especially the regulatory and signalling mechanisms. A change to the status quo calls for integrative multi-omics and inter-omics analyses with approaches in ‘systems genetics and genomics’. In this Review, we highlight findings from genome-wide association studies and studies using various omics technologies individually to identify mechanisms of osteoporosis. Furthermore, we summarize current studies of data integration to understand, diagnose and inform the treatment of osteoporosis. The integration of multiple technologies will provide a road map to illuminate the complex pathogenesis of osteoporosis, especially from molecular functional aspects, in vivo in humans.
Key points
-
Osteoporosis, which is the most common bone disorder worldwide, and its related traits (low bone mineral density and osteoporotic fracture) are highly heritable.
-
Multiple omics technologies, including genomics, transcriptomics, epigenomics, proteomics and metabolomics, have been applied to identify the molecular factors contributing to the pathogenesis of osteoporosis.
-
Building upon the success in single-omics discovery research, studies have integrated data from different omics levels to better elucidate the molecular and functional mechanisms for osteoporosis.
-
Integration of omics approaches can provide a holistic road map to comprehensively illuminate the complex pathogenesis of osteoporosis and fulfil the potential of personalized disease risk prediction, intervention and treatment as well as drug development or re-purposing.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Kanis, J. A. Diagnosis of osteoporosis. Osteoporos. Int. 7, S108–S116 (1997).
World Health Organization. Assessment of fracture risk and its application to screening for postmenopausal osteoporosis : report of a WHO study group [meeting held in Rome from 22 to 25 June 1992] (WHO, 1994).
Johnell, O. et al. Predictive value of BMD for hip and other fractures. J. Bone Miner. Res. 20, 1185–1194 (2005).
Peacock, M., Turner, C. H., Econs, M. J. & Foroud, T. Genetics of osteoporosis. Endocr. Rev. 23, 303–326 (2002).
Deng, H. W. et al. Relevance of the genes for bone mass variation to susceptibility to osteoporotic fractures and its implications to gene search for complex human diseases. Genet. Epidemiol. 22, 12–25 (2002).
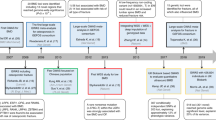
Morris, J. A. et al. An atlas of genetic influences on osteoporosis in humans and mice. Nat. Genet. 51, 258–266 (2019). As the largest GWAS of BMD until 2019, this paper performed analyses in over 426,000 individuals from the UK Biobank and identified 518 loci associated with BMD, 301 of which were novel.
Sun, Y. V. & Hu, Y. J. Integrative analysis of multi-omics data for discovery and functional studies of complex human diseases. Adv. Genet. 93, 147–190 (2016).
Hasin, Y., Seldin, M. & Lusis, A. Multi-omics approaches to disease. Genome Biol. 18, 83 (2017).
Raisz, L. G. Pathogenesis of osteoporosis: concepts, conflicts, and prospects. J. Clin. Invest. 115, 3318–3325 (2005).
Pittenger, M. F. et al. Multilineage potential of adult human mesenchymal stem cells. Science 284, 143–147 (1999).
Long, F. Building strong bones: molecular regulation of the osteoblast lineage. Nat. Rev. Mol. Cell Biol. 13, 27–38 (2011).
Udagawa, N. et al. Origin of osteoclasts: mature monocytes and macrophages are capable of differentiating into osteoclasts under a suitable microenvironment prepared by bone marrow-derived stromal cells. Proc. Natl Acad. Sci. USA 87, 7260–7264 (1990).
Boyle, W. J., Simonet, W. S. & Lacey, D. L. Osteoclast differentiation and activation. Nature 423, 337–342 (2003).
Teitelbaum, S. L. Bone resorption by osteoclasts. Science 289, 1504–1508 (2000).
Knothe Tate, M. L., Adamson, J. R., Tami, A. E. & Bauer, T. W. The osteocyte. Int. J. Biochem. Cell Biol. 36, 1–8 (2004).
Dallas, S. L., Prideaux, M. & Bonewald, L. F. The osteocyte: an endocrine cell and more. Endocr. Rev. 34, 658–690 (2013).
Sims, N. A. & Walsh, N. C. Intercellular cross-talk among bone cells: new factors and pathways. Curr. Osteoporos. Rep. 10, 109–117 (2012).
Sabik, O. L. & Farber, C. R. Using GWAS to identify novel therapeutic targets for osteoporosis. Transl. Res. 181, 15–26 (2017).
Richards, J. B., Zheng, H. F. & Spector, T. D. Genetics of osteoporosis from genome-wide association studies: advances and challenges. Nat. Rev. Genet. 13, 576–588 (2012).
Liu, Y. J., Zhang, L., Papasian, C. J. & Deng, H. W. Genome-wide association studies for osteoporosis: a 2013 update. J. Bone Metab. 21, 99–116 (2014).
Styrkarsdottir, U. et al. Multiple genetic loci for bone mineral density and fractures. N. Engl. J. Med. 358, 2355–2365 (2008).
Richards, J. B. et al. Bone mineral density, osteoporosis, and osteoporotic fractures: a genome-wide association study. Lancet 371, 1505–1512 (2008).
Styrkarsdottir, U. et al. New sequence variants associated with bone mineral density. Nat. Genet. 41, 15–17 (2009).
Xiong, D. H. et al. Genome-wide association and follow-up replication studies identified ADAMTS18 and TGFBR3 as bone mass candidate genes in different ethnic groups. Am. J. Hum. Genet. 84, 388–398 (2009).
Zheng, H. F. et al. Whole-genome sequencing identifies EN1 as a determinant of bone density and fracture. Nature 526, 112–117 (2015). By using whole-genome sequencing, this study identified a low-frequency non-coding variant near the novel locus (EN1) associated with lumbar spine BMD and the risk of fracture.
Rivadeneira, F. et al. Twenty bone-mineral-density loci identified by large-scale meta-analysis of genome-wide association studies. Nat. Genet. 41, 1199–1206 (2009).
Richards, J. B. et al. Collaborative meta-analysis: associations of 150 candidate genes with osteoporosis and osteoporotic fracture. Ann. Intern. Med. 151, 528–537 (2009).
Estrada, K. et al. Genome-wide meta-analysis identifies 56 bone mineral density loci and reveals 14 loci associated with risk of fracture. Nat. Genet. 44, 491–501 (2012).
Moayyeri, A. et al. Genetic determinants of heel bone properties: genome-wide association meta-analysis and replication in the GEFOS/GENOMOS consortium. Hum. Mol. Genet. 23, 3054–3068 (2014).
Medina-Gomez, C. et al. Life-course genome-wide association study meta-analysis of total body bmd and assessment of age-specific effects. Am. J. Hum. Genet. 102, 88–102 (2018).
Parikka, V. et al. Estrogen responsiveness of bone formation in vitro and altered bone phenotype in aged estrogen receptor-α-deficient male and female mice. Eur. J. Endocrinol. 152, 301–314 (2005).
Maatta, J. A. et al. Inactivation of estrogen receptor α in bone-forming cells induces bone loss in female mice. FASEB J. 27, 478–488 (2013).
Crockett, J. C., Rogers, M. J., Coxon, F. P., Hocking, L. J. & Helfrich, M. H. Bone remodelling at a glance. J. Cell Sci. 124, 991–998 (2011).
Collins, R. What makes UK Biobank special? Lancet 379, 1173–1174 (2012).
Kemp, J. P. et al. Identification of 153 new loci associated with heel bone mineral density and functional involvement of GPC6 in osteoporosis. Nat. Genet. 49, 1468–1475 (2017).
Riggs, B. L., Khosla, S. & Melton, L. J. 3rd. A unitary model for involutional osteoporosis: estrogen deficiency causes both type I and type II osteoporosis in postmenopausal women and contributes to bone loss in aging men. J. Bone Miner. Res. 13, 763–773 (1998).
Koller, D. L. et al. Meta-analysis of genome-wide studies identifies WNT16 and ESR1 SNPs associated with bone mineral density in premenopausal women. J. Bone Miner. Res. 28, 547–558 (2013).
Wang, C. et al. Susceptibility genes for osteoporotic fracture in postmenopausal Chinese women. J. Bone Miner. Res. 27, 2582–2591 (2012).
Zheng, H. F. et al. WNT16 influences bone mineral density, cortical bone thickness, bone strength, and osteoporotic fracture risk. PLOS Genet. 8, e1002745 (2012).
Simonet, W. S. et al. Osteoprotegerin: a novel secreted protein involved in the regulation of bone density. Cell 89, 309–319 (1997).
Xiong, L. et al. Lrp4 in osteoblasts suppresses bone formation and promotes osteoclastogenesis and bone resorption. Proc. Natl Acad. Sci. USA 112, 3487–3492 (2015).
Kaprio, J. Twins and the mystery of missing heritability: the contribution of gene-environment interactions. J. Intern. Med. 272, 440–448 (2012).
Yang, T. L. et al. Genome-wide copy-number-variation study identified a susceptibility gene, UGT2B17, for osteoporosis. Am. J. Hum. Genet. 83, 663–674 (2008). The first study to investigate the contribution of copy number variation to osteoporosis identified a deletion variant of UGT2B17 associated with osteoporotic fracture.
Zhu, W. et al. Gene-based GWAS analysis for consecutive studies of GEFOS. Osteoporos. Int. 29, 2645–2658 (2018).
Zhang, L. S. et al. Pathway-based genome-wide association analysis identified the importance of regulation-of-autophagy pathway for ultradistal radius BMD. J. Bone Miner. Res. 25, 1572–1580 (2010).
Huang, H., Chanda, P., Alonso, A., Bader, J. S. & Arking, D. E. Gene-based tests of association. PLOS Genet. 7, e1002177 (2011).
Liu, Y. Z. et al. Powerful bivariate genome-wide association analyses suggest the Sox6 gene influencing both obesity and osteoporosis phenotypes in males. PLOS ONE 4, e6827 (2009).
Medina-Gomez, C. et al. Bivariate genome-wide association meta-analysis of pediatric musculoskeletal traits reveals pleiotropic effects at the SREBF1/TOM1L2 locus. Nat. Commun. 8, 121 (2017).
Pei, Y. F. et al. Joint association analysis identified 18 new loci for bone mineral density. J. Bone Miner. Res. 34, 1086–1094 (2019).
Greenbaum, J. et al. Increased detection of genetic loci associated with risk predictors of osteoporotic fracture using a pleiotropic cFDR method. Bone 99, 62–68 (2017).
Calabrese, G. M. et al. Integrating GWAS and co-expression network data identifies bone mineral density genes SPTBN1 and MARK3 and an osteoblast functional module. Cell Syst. 4, 46–59.e4 (2017).
Boyle, E. A., Li, Y. I. & Pritchard, J. K. An expanded view of complex traits: from polygenic to omnigenic. Cell 169, 1177–1186 (2017).
Lee, S. H. et al. Prediction of future osteoporotic fracture occurrence by genetic profiling: a 6-year follow-up observational study. J. Clin. Endocrinol. Metab. 101, 1215–1224 (2016).
Ho-Le, T. P., Center, J. R., Eisman, J. A., Nguyen, H. T. & Nguyen, T. V. Prediction of bone mineral density and fragility fracture by genetic profiling. J. Bone Miner. Res. 32, 285–293 (2017).
Kim, S. K. Identification of 613 new loci associated with heel bone mineral density and a polygenic risk score for bone mineral density, osteoporosis and fracture. PLOS ONE 13, e0200785 (2018).
Nelson, M. R. et al. The support of human genetic evidence for approved drug indications. Nat. Genet. 47, 856–860 (2015).
Xia, W. F. et al. Swedish mutant APP suppresses osteoblast differentiation and causes osteoporotic deficit, which are ameliorated by N-acetyl-L-cysteine. J. Bone Miner. Res. 28, 2122–2135 (2013).
Cornelius, C. et al. Osteoporosis and Alzheimer pathology: role of cellular stress response and hormetic redox signaling in aging and bone remodeling. Front. Pharmacol. 5, 120 (2014).
Sato, T. et al. Donepezil prevents RANK-induced bone loss via inhibition of osteoclast differentiation by downregulating acetylcholinesterase. Heliyon 1, e00013 (2015).
Karasik, D. et al. Heritability and genetic correlations for bone microarchitecture: the framingham study families. J. Bone Miner. Res. 32, 106–114 (2017).
Trajanoska, K. et al. Assessment of the genetic and clinical determinants of fracture risk: genome wide association and mendelian randomisation study. BMJ 362, k3225 (2018).
Nielson, C. M. et al. Novel genetic variants associated with increased vertebral volumetric BMD, reduced vertebral fracture risk, and increased expression of SLC1A3 and EPHB2. J. Bone Miner. Res. 31, 2085–2097 (2016).
Guo, Y. et al. Genome-wide association study identifies ALDH7A1 as a novel susceptibility gene for osteoporosis. PLOS Genet. 6, e1000806 (2010).
Greene, C. S., Penrod, N. M., Williams, S. M. & Moore, J. H. Failure to replicate a genetic association may provide important clues about genetic architecture. PLOS ONE 4, e5639 (2009).
Kraft, P., Zeggini, E. & Ioannidis, J. P. Replication in genome-wide association studies. Stat. Sci. 24, 561–573 (2009).
Liu, Y. J., Zhang, L., Pei, Y. F., Papasian, C. J. & Deng, H. W. On genome-wide association studies and their meta-analyses: lessons learned from osteoporosis studies. J. Clin. Endocrinol. Metab. 98, E1278–E1282 (2013).
Kichaev, G. et al. Integrating functional data to prioritize causal variants in statistical fine-mapping studies. PLOS Genet. 10, e1004722 (2014).
Yang, J., Lee, S. H., Goddard, M. E. & Visscher, P. M. GCTA: a tool for genome-wide complex trait analysis. Am. J. Hum. Genet. 88, 76–82 (2011).
Benner, C. et al. FINEMAP: efficient variable selection using summary data from genome-wide association studies. Bioinformatics 32, 1493–1501 (2016).
Greenbaum, J. & Deng, H. W. A statistical approach to fine mapping for the identification of potential causal variants related to bone mineral density. J. Bone Miner. Res. 32, 1651–1658 (2017).
Benner, C. et al. Prospects of fine-mapping trait-associated genomic regions by using summary statistics from genome-wide association studies. Am. J. Hum. Genet. 101, 539–551 (2017).
Liu, Y. Z. et al. A novel pathophysiological mechanism for osteoporosis suggested by an in vivo gene expression study of circulating monocytes. J. Biol. Chem. 280, 29011–29016 (2005).
Mohan, S., Hu, Y. & Edderkaoui, B. Chemokine receptor 3 is a negative regulator of trabecular bone mass in female mice. J. Cell. Biochem. 120, 13974–13984 (2019).
Fitzpatrick, L. A. et al. Targeted deletion of histidine decarboxylase gene in mice increases bone formation and protects against ovariectomy-induced bone loss. Proc. Natl Acad. Sci. USA 100, 6027–6032 (2003).
Takuma, A. et al. Dexamethasone enhances osteoclast formation synergistically with transforming growth factor-β by stimulating the priming of osteoclast progenitors for differentiation into osteoclasts. J. Biol. Chem. 278, 44667–44674 (2003).
Eskildsen, T. et al. MicroRNA-138 regulates osteogenic differentiation of human stromal (mesenchymal) stem cells in vivo. Proc. Natl Acad. Sci. USA 108, 6139–6144 (2011).
Yang, L. et al. The long non-coding RNA-ORLNC1 regulates bone mass by directing mesenchymal stem cell fate. Mol. Ther. 27, 394–410 (2019).
Zhao, K. et al. Hsa_Circ_0001275: a potential novel diagnostic biomarker for postmenopausal osteoporosis. Cell Physiol. Biochem. 46, 2508–2516 (2018).
Vidal, C., Cachia, A. & Xuereb-Anastasi, A. Effects of a synonymous variant in exon 9 of the CD44 gene on pre-mRNA splicing in a family with osteoporosis. Bone 45, 736–742 (2009).
Scotti, M. M. & Swanson, M. S. RNA mis-splicing in disease. Nat. Rev. Genet. 17, 19–32 (2016).
Marini, F., Cianferotti, L. & Brandi, M. L. Epigenetic mechanisms in bone biology and osteoporosis: can they drive therapeutic choices? Int. J. Mol. Sci. 17, E1329 (2016).
van Meurs, J. B., Boer, C. G., Lopez-Delgado, L. & Riancho, J. A. Role of epigenomics in bone and cartilage disease. J. Bone Miner. Res. 34, 215–230 (2019).
Reppe, S. et al. Methylation of bone SOST, its mRNA, and serum sclerostin levels correlate strongly with fracture risk in postmenopausal women. J. Bone Miner. Res. 30, 249–256 (2015).
Li, X. et al. Sclerostin binds to LRP5/6 and antagonizes canonical Wnt signaling. J. Biol. Chem. 280, 19883–19887 (2005).
Cao, Y. & Wang, B. Expression of sclerostin in osteoporotic fracture patients is associated with DNA methylation in the CpG island of the SOST gene. Int. J. Genomics 2019, 7076513 (2019).
Delgado-Calle, J. et al. Genome-wide profiling of bone reveals differentially methylated regions in osteoporosis and osteoarthritis. Arthritis Rheum. 65, 197–205 (2013). The first epigenome-wide association study of human bone detected 241 differentially methylated CpG sites in femoral head trabecular bone specimens between 27 patients with osteoporotic hip fractures and 26 patients with hip osteoarthritis.
Reppe, S. et al. Distinct DNA methylation profiles in bone and blood of osteoporotic and healthy postmenopausal women. Epigenetics 12, 674–687 (2017).
Morris, J. A. et al. Epigenome-wide association of DNA methylation in whole blood with bone mineral density. J. Bone Miner. Res. 32, 1644–1650 (2017).
Fernandez-Rebollo, E. et al. Primary osteoporosis is not reflected by disease-specific DNA methylation or accelerated epigenetic age in blood. J. Bone Miner. Res. 33, 356–361 (2018).
ENCODE Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Roadmap Epigenomics Consortium et al. Integrative analysis of 111 reference human epigenomes. Nature 518, 317–330 (2015).
Deng, F. Y. et al. Proteomic analysis of circulating monocytes in Chinese premenopausal females with extremely discordant bone mineral density. Proteomics 8, 4259–4272 (2008).
Deng, F. Y. et al. Peripheral blood monocyte-expressed ANXA2 gene is involved in pathogenesis of osteoporosis in humans. Mol. Cell Proteom. 10, M111.011700 (2011).
Deng, F. Y. et al. Is GSN significant for hip BMD in female Caucasians? Bone 63, 69–75 (2014).
Zhou, X. et al. Anxa2 attenuates osteoblast growth and is associated with hip BMD and osteoporotic fracture in Chinese elderly. PLOS ONE 13, e0194781 (2018).
Zeng, Y. et al. Mass spectrometry based proteomics profiling of human monocytes. Protein Cell 8, 123–133 (2017). The first comprehensive proteome knowledgebase for human monocytes was developed in 2017; it involves a total of 2,237 unique protein-encoding genes and provides a reference map for future in-depth research on monocyte biology and osteoporosis.
Bhattacharyya, S., Siegel, E. R., Achenbach, S. J., Khosla, S. & Suva, L. J. Serum biomarker profile associated with high bone turnover and BMD in postmenopausal women. J. Bone Miner. Res. 23, 1106–1117 (2008).
Deng, F. Y. et al. An integrative study ascertained SOD2 as a susceptibility gene for osteoporosis in Chinese. J. Bone Miner. Res. 26, 2695–2701 (2011).
Zhang, L. et al. Network-based proteomic analysis for postmenopausal osteoporosis in Caucasian females. Proteomics 16, 12–28 (2016).
Zeng, Y. et al. Quantitative proteomics and integrative network analysis identified novel genes and pathways related to osteoporosis. J. Proteomics 142, 45–52 (2016).
Arasu, A. et al. Serum sclerostin and risk of hip fracture in older Caucasian women. J. Clin. Endocrinol. Metab. 97, 2027–2032 (2012).
Johnson, C. H., Ivanisevic, J. & Siuzdak, G. Metabolomics: beyond biomarkers and towards mechanisms. Nat. Rev. Mol. Cell Biol. 17, 451–459 (2016).
Wu, Q. Y. et al. Long non-coding RNAs: a new regulatory code for osteoporosis. Front. Endocrinol. 9, 587 (2018).
Ma, B. et al. Metabolomic profiles delineate signature metabolic shifts during estrogen deficiency-induced bone loss in rat by GC-TOF/MS. PLOS ONE 8, e54965 (2013).
You, Y. S. et al. Association between the metabolome and low bone mineral density in Taiwanese women determined by 1H NMR spectroscopy. J. Bone Miner. Res. 29, 212–222 (2014). The first metabolomics study of osteoporosis in humans compared high and low BMD groups and reported four distinguishing metabolites: lactate, acetone, acetate and glutamine.
Miyamoto, T. et al. Metabolomics-based profiles predictive of low bone mass in menopausal women. Bone Rep. 9, 11–18 (2018).
Zhao, Q. et al. Metabolomic profiles associated with bone mineral density in US Caucasian women. Nutr. Metab. 15, 57 (2018).
Yu, L. et al. Association between metabolic profiles in urine and bone mineral density of pre- and postmenopausal Chinese women. Menopause 26, 94–102 (2019).
Cabrera, D. et al. Association of plasma lipids and polar metabolites with low bone mineral density in Singaporean-Chinese menopausal women: a pilot study. Int. J. Environ. Res. Public Health 15, E1045 (2018).
Moayyeri, A. et al. Metabolomic pathways to osteoporosis in middle-aged women: a genome-metabolome-wide Mendelian randomization study. J. Bone Miner. Res. 33, 643–650 (2018).
Liu, L. et al. Assessing the associations of blood metabolites with osteoporosis: a Mendelian randomization study. J. Clin. Endocrinol. Metab. 103, 1850–1855 (2018).
Schrimpe-Rutledge, A. C., Codreanu, S. G., Sherrod, S. D. & McLean, J. A. Untargeted metabolomics strategies — challenges and emerging directions. J. Am. Soc. Mass Spectrom. 27, 1897–1905 (2016).
Ding, C., Cicuttini, F. & Jones, G. Tibial subchondral bone size and knee cartilage defects: relevance to knee osteoarthritis. Osteoarthritis Cartilage 15, 479–486 (2007).
Chou, C. H. et al. Genome-wide expression profiles of subchondral bone in osteoarthritis. Arthritis Res. Ther. 15, R190 (2013).
Jaffe, A. E. & Irizarry, R. A. Accounting for cellular heterogeneity is critical in epigenome-wide association studies. Genome Biol. 15, R31 (2014).
Teschendorff, A. E. & Zheng, S. C. Cell-type deconvolution in epigenome-wide association studies: a review and recommendations. Epigenomics 9, 757–768 (2017).
Maurano, M. T. et al. Systematic localization of common disease-associated variation in regulatory DNA. Science 337, 1190–1195 (2012).
Schaid, D. J., Chen, W. & Larson, N. B. From genome-wide associations to candidate causal variants by statistical fine-mapping. Nat. Rev. Genet. 19, 491–504 (2018).
Boyle, A. P. et al. Annotation of functional variation in personal genomes using regulomeDB. Genome Res. 22, 1790–1797 (2012).
Guo, Y. et al. Integrating epigenomic elements and gwass identifies bdnf gene affecting bone mineral density and osteoporotic fracture risk. Sci. Rep. 6, 30558 (2016).
Yao, S. et al. Regulatory element-based prediction identifies new susceptibility regulatory variants for osteoporosis. Hum. Genet. 136, 963–974 (2017).
Grundberg, E. et al. Population genomics in a disease targeted primary cell model. Genome Res. 19, 1942–1952 (2009).
Mullin, B. H. et al. Expression quantitative trait locus study of bone mineral density gwas variants in human osteoclasts. J. Bone Miner. Res. 33, 1044–1051 (2018).
Chen, X. F. et al. An osteoporosis risk SNP at 1p36.12 acts as an allele-specific enhancer to modulate LINC00339 expression via long-range loop formation. Am. J. Hum. Genet. 102, 776–793 (2018). This paper provides a mechanistic explanation for the influence of the 1p36.12 locus on human osteoporosis, using multiple omics technologies to bridge GWAS results to physiology.
Meng, X. H. et al. Integration of summary data from GWAS and eQTL studies identified novel causal BMD genes with functional predictions. Bone 113, 41–48 (2018).
Giambartolomei, C. et al. Bayesian test for colocalisation between pairs of genetic association studies using summary statistics. PLOS Genet. 10, e1004383 (2014).
Hormozdiari, F. et al. Colocalization of GWAS and eQTL signals detects target genes. Am. J. Hum. Genet. 99, 1245–1260 (2016).
Chesi, A. et al. Genome-scale Capture C promoter interactions implicate effector genes at GWAS loci for bone mineral density. Nat. Commun. 10, 1260 (2019).
Zhu, D. L. et al. Multiple functional variants at 13q14 Risk locus for osteoporosis regulate RANKL expression through long-range super-enhancer. J. Bone Miner. Res. 33, 1335–1346 (2018).
Wang, Q. et al. A Bayesian framework that integrates multi-omics data and gene networks predicts risk genes from schizophrenia GWAS data. Nat. Neurosci. 22, 691–699 (2019).
Langfelder, P. & Horvath, S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinformatics 9, 559 (2008).
Lusis, A. J. et al. The Hybrid Mouse Diversity Panel: a resource for systems genetics analyses of metabolic and cardiovascular traits. J. Lipid Res. 57, 925–942 (2016).
Calabrese, G. et al. Systems genetic analysis of osteoblast-lineage cells. PLOS Genet. 8, e1003150 (2012).
Chen, Y. C. et al. Integrative analysis of genomics and transcriptome data to identify potential functional genes of BMDs in females. J. Bone Miner. Res. 31, 1041–1049 (2016).
Chenu, C., Serre, C. M., Raynal, C., Burt-Pichat, B. & Delmas, P. D. Glutamate receptors are expressed by bone cells and are involved in bone resorption. Bone 22, 295–299 (1998).
Zhang, J. G. et al. Integrative analysis of transcriptomic and epigenomic data to reveal regulation patterns for BMD variation. PLOS ONE 10, e0138524 (2015).
Al-Barghouthi, B. M. & Farber, C. R. Dissecting the genetics of osteoporosis using systems approaches. Trends Genet. 35, 55–67 (2019).
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012).
Duan, X. et al. Deficiency of ATP6V1H causes bone loss by inhibiting bone resorption and bone formation through the TGF-β1 pathway. Theranostics 6, 2183–2195 (2016).
Saito, A. et al. Targeted reversion of induced pluripotent stem cells from patients with human cleidocranial dysplasia improves bone regeneration in a rat calvarial bone defect model. Stem Cell Res. Ther. 9, 12 (2018).
Freudenthal, B. et al. Rapid phenotyping of knockout mice to identify genetic determinants of bone strength. J. Endocrinol. 231, R31–R46 (2016).
Brommage, R. et al. High-throughput screening of mouse gene knockouts identifies established and novel skeletal phenotypes. Bone Res. 2, 14034 (2014).
Chappell, L., Russell, A. J. C. & Voet, T. Single-cell (multi)omics technologies. Annu. Rev. Genomics Hum. Genet. 19, 15–41 (2018).
Chan, C. K. F. et al. Identification of the human skeletal stem cell. Cell 175, 43–56.e21 (2018).
Santiago-Algarra, D., Dao, L. T. M., Pradel, L., Espana, A. & Spicuglia, S. Recent advances in high-throughput approaches to dissect enhancer function. F1000Res 6, 939 (2017).
Chen, Y. C., Greenbaum, J., Shen, H. & Deng, H. W. Association between gut microbiota and bone health: potential mechanisms and prospective. J. Clin. Endocrinol. Metab. 102, 3635–3646 (2017).
Jones, R. M., Mulle, J. G. & Pacifici, R. Osteomicrobiology: the influence of gut microbiota on bone in health and disease. Bone 115, 59–67 (2018).
Wang, J. et al. Diversity analysis of gut microbiota in osteoporosis and osteopenia patients. PeerJ 5, e3450 (2017).
Goodrich, J. K., Davenport, E. R., Clark, A. G. & Ley, R. E. The relationship between the human genome and microbiome comes into view. Annu. Rev. Genet. 51, 413–433 (2017).
Timpson, N. J., Greenwood, C. M. T., Soranzo, N., Lawson, D. J. & Richards, J. B. Genetic architecture: the shape of the genetic contribution to human traits and disease. Nat. Rev. Genet. 19, 110–124 (2018).
Price, A. L., Zaitlen, N. A., Reich, D. & Patterson, N. New approaches to population stratification in genome-wide association studies. Nat. Rev. Genet. 11, 459–463 (2010).
Zhang, L. et al. A new method for estimating effect size distribution and heritability from genome-wide association summary results. Hum. Genet. 135, 171–184 (2016).
Wright, N. C. et al. The recent prevalence of osteoporosis and low bone mass in the United States based on bone mineral density at the femoral neck or lumbar spine. J. Bone Miner. Res. 29, 2520–2526 (2014).
International Osteoporosis Foundation. Broken bones, broken lives: a roadmap to solve the fragility fracture crisis in Europe (IOF, 2018).
Chen, P., Li, Z. & Hu, Y. Prevalence of osteoporosis in China: a meta-analysis and systematic review. BMC Public Health 16, 1039 (2016).
Lee, K. S., Bae, S. H., Lee, S. H., Lee, J. & Lee, D. R. New reference data on bone mineral density and the prevalence of osteoporosis in Korean adults aged 50 years or older: the Korea National Health and Nutrition Examination Survey 2008-2010. J. Korean Med. Sci. 29, 1514–1522 (2014).
Wade, S. W., Strader, C., Fitzpatrick, L. A., Anthony, M. S. & O’Malley, C. D. Estimating prevalence of osteoporosis: examples from industrialized countries. Arch. Osteoporos. 9, 182 (2014).
Acknowledgements
T.L.Y. and S.S.D. acknowledge the support of the National Natural Science Foundation of China (31771399, 31970569 and 81573241), the Innovative Talent Promotion Plan of Shaanxi Province for Young Sci-Tech New Star (2018KJXX-010) and the special guidance funds for the construction of world-class universities (disciplines) and characteristic development in central universities. H.S. and H.W.D. acknowledge the support of grants from the National Institutes of Health (R01AR059781, P20GM109036, R01MH107354, R01MH104680, R01GM109068, R01AR069055, U19AG055373, R01DK115679), the Edward G. Schlieder Endowment and the Drs. W. C. Tsai and P. T. Kung Professorship in Biostatistics from Tulane University.
Author information
Authors and Affiliations
Contributions
H-W.D., T-L.Y., H.S., S-S.D., L.Z., F-Y.D and Q.Z. researched data for the article, made substantial contributions to the discussion of content, and contributed to the writing and review/editing of the manuscript before submission. A.L. researched data for the article and contributed to the writing and review/editing of the manuscript before submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information
Nature Reviews Endocrinology thanks D. Karasik, J. Tobias and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
FINEMAP: http://www.christianbenner.com/
GCTA-COJO: https://cnsgenomics.com/software/gcta/#COJO
International Mouse Phenotyping Consortium: http://www.mousephenotype.org/
Mouse Genome Informatics: http://www.informatics.jax.org/
PAINTOR: https://github.com/gkichaev/PAINTOR_V3.0
The Origins of Bone and Cartilage Disease project: http://www.boneandcartilage.com/
UK Biobank: https://www.ukbiobank.ac.uk/
Supplementary information
41574_2019_282_MOESM2_ESM.xlsx
Supplementary Table 2. Drug target information for GWAS genes (drugs for osteoporosis, other diseases or disease pathways).
41574_2019_282_MOESM5_ESM.xlsx
Supplementary Table 5 and 6. Summary of potential diagnostic protein biomarkers of osteoporotic fracture predictive protein biomarkers of osteoporotic fracture.
Rights and permissions
About this article
Cite this article
Yang, TL., Shen, H., Liu, A. et al. A road map for understanding molecular and genetic determinants of osteoporosis. Nat Rev Endocrinol 16, 91–103 (2020). https://doi.org/10.1038/s41574-019-0282-7
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41574-019-0282-7