Abstract
Cellular programming of naïve T cells can improve the efficacy of adoptive T-cell therapy. However, the current ex vivo engineering of T cells requires the pre-activation of T cells, which causes them to lose their naïve state. In this study, cationic-polymer-functionalized nanowires were used to pre-program the fate of primary naïve CD8+ T cells to achieve a therapeutic response in vivo. This was done by delivering single or multiple microRNAs to primary naïve mouse and human CD8+ T cells without pre-activation. The use of nanowires further allowed for the delivery of large, whole lentiviral particles with potential for long-term integration. The combination of deletion and overexpression of miR-29 and miR-130 impacted the ex vivo T-cell differentiation fate from the naïve state. The programming of CD8+ T cells using nanowire-delivered co-delivery of microRNAs resulted in the modulation of T-cell fitness by altering the T-cell proliferation, phenotypic and transcriptional regulation, and secretion of effector molecules. Moreover, the in vivo adoptive transfer of murine CD8+ T cells programmed through the nanowire-mediated dual delivery of microRNAs provided enhanced immune protection against different types of intracellular pathogen (influenza and Listeria monocytogenes). In vivo analyses demonstrated that the simultaneous alteration of miR-29 and miR-130 levels in naïve CD8+ T cells reduces the persistence of canonical memory T cells whereas increases the population of short-lived effector T cells. Nanowires could potentially be used to modulate CD8+ T-cell differentiation and achieve a therapeutic response in vivo without the need for pre-activation.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The raw RNA-seq data on nanowires are available in the GEO database under accession code GSE255468. The naïve dataset used for comparison was the expression profiles of adult naïve CD8+ T cells from GSE97795 (ref. 57). The remaining data are available within the Article and its Supplementary Information. Due to the very large file sizes and volume of data, the remaining raw data supporting the findings of this study are available from the corresponding authors on reasonable request. Source data are provided with this paper.
References
Kaech, S. M. & Cui, W. Transcriptional control of effector and memory CD8+ T cell differentiation. Nat. Rev. Immunol. 12, 749–761 (2012).
Joshi, N. S. et al. Inflammation directs memory precursor and short-lived effector CD8+ T cell fates via the graded expression of T-bet transcription factor. Immunity 27, 281–295 (2007).
Yee Mon, K. J. et al. MicroRNA-29 specifies age-related differences in the CD8+ T cell immune response. Cell Rep. 37, 109969 (2021).
Maus, M. V. et al. Adoptive immunotherapy for cancer or viruses. Annu. Rev. Immunol. 32, 189–225 (2014).
Morotti, M. et al. Promises and challenges of adoptive T-cell therapies for solid tumours. Br. J. Cancer 124, 1759–1776 (2021).
Cappell, K. M. & Kochenderfer, J. N. Long-term outcomes following CAR T cell therapy: what we know so far. Nat. Rev. Clin. Oncol. 20, 359–371 (2023).
Chow, A., Perica, K., Klebanoff, C. A. & Wolchok, J. D. Clinical implications of T cell exhaustion for cancer immunotherapy. Nat. Rev. Clin. Oncol. 19, 775–790 (2022).
Hinrichs, C. S. et al. Human effector CD8+ T cells derived from naive rather than memory subsets possess superior traits for adoptive immunotherapy. Blood 117, 808–814 (2011).
Klebanoff, C. A. et al. Memory T cell-driven differentiation of naive cells impairs adoptive immunotherapy. J. Clin. Invest. 126, 318–334 (2016).
Trifari, S. et al. MicroRNA-directed program of cytotoxic CD8+ T-cell differentiation. Proc. Natl Acad. Sci. USA 110, 18608–18613 (2013).
Hinrichs, C. S. et al. Adoptively transferred effector cells derived from naïve rather than central memory CD8+ T cells mediate superior antitumor immunity. Proc. Natl Acad. Sci. USA 106, 17469–17474 (2009).
Bronevetsky, Y. et al. T cell activation induces proteasomal degradation of Argonaute and rapid remodeling of the microRNA repertoire. J. Exp. Med. 210, 417–432 (2013).
Zhang, N. & Bevan, M. J. Dicer controls CD8+ T-cell activation, migration, and survival. Proc. Natl Acad. Sci. USA 107, 21629–21634 (2010).
Smith, N. L., Wissink, E. M., Grimson, A. & Rudd, B. D. miR-150 regulates differentiation and cytolytic effector function in CD8+ T cells. Sci. Rep. 5, 16399 (2015).
Muljo, S. A. et al. Aberrant T cell differentiation in the absence of Dicer. J. Exp. Med. 202, 261–269 (2005).
Wissink, E. M., Smith, N. L., Spektor, R., Rudd, B. D. & Grimson, A. MicroRNAs and their targets are differentially regulated in adult and neonatal mouse CD8+ T cells. Genetics 201, 1017–1030 (2015).
Friedman, R. C., Farh, K. K.-H., Burge, C. B. & Bartel, D. P. Most mammalian mRNAs are conserved targets of microRNAs. Genome Res. 19, 92–105 (2009).
Liang, Y., Pan, H. F. & Ye, D. Q. microRNAs function in CD8+ T cell biology. J. Leukoc. Biol. 97, 487–497 (2015).
Ji, Y. et al. miR-155 augments CD8+ T-cell antitumor activity in lymphoreplete hosts by enhancing responsiveness to homeostatic γc cytokines. Proc. Natl Acad. Sci. USA 112, 476–481 (2015).
Tsai, C. Y., Allie, S. R., Zhang, W. & Usherwood, E. J. MicroRNA miR-155 affects antiviral effector and effector memory CD8 T cell differentiation. J. Virol. 87, 2348–2351 (2013).
Lind, E. F., Elford, A. R. & Ohashi, P. S. Micro-RNA 155 is required for optimal CD8+ T cell responses to acute viral and intracellular bacterial challenges. J. Immunol. 190, 1210–1216 (2013).
Wu, T. et al. Temporal expression of microRNA cluster miR-17-92 regulates effector and memory CD8+ T-cell differentiation. Proc. Natl Acad. Sci. USA 109, 9965–9970 (2012).
Boldin, M. P. et al. miR-146a is a significant brake on autoimmunity, myeloproliferation, and cancer in mice. J. Exp. Med. 208, 1189–1201 (2011).
Huffaker, T. B. et al. Epistasis between microRNAs 155 and 146a during T cell-mediated antitumor immunity. Cell Rep. 2, 1697–1709 (2012).
Xu, Y. & Dotti, G. Selection bias: maintaining less-differentiated T cells for adoptive immunotherapy. J. Clin. Invest. 126, 35–37 (2016).
Tumeh, P. C. et al. The impact of ex vivo clinical grade activation protocols on human T-cell phenotype and function for the generation of genetically modified cells for adoptive cell transfer therapy. J. Immunother. 33, 759–768 (2010).
Shalek, A. K. et al. Nanowire-mediated delivery enables functional interrogation of primary immune cells: application to the analysis of chronic lymphocytic leukemia. Nano Lett. 12, 6498–6504 (2012).
Zhang, Z., Qiu, S., Zhang, X. & Chen, W. Optimized DNA electroporation for primary human T cell engineering. BMC Biotechnol. 18, 4 (2018).
Chevrier, N. et al. Systematic discovery of TLR signaling components delineates viral-sensing circuits. Cell 147, 853–867 (2011).
Shalek, A. K. et al. Vertical silicon nanowires as a universal platform for delivering biomolecules into living cells. Proc. Natl Acad. Sci. USA 107, 1870–1875 (2010).
Robinson, J. T. et al. Vertical nanowire electrode arrays as a scalable platform for intracellular interfacing to neuronal circuits. Nat. Nanotechnol. 7, 180–184 (2012).
Chiappini, C. et al. Biodegradable silicon nanoneedles delivering nucleic acids intracellularly induce localized in vivo neovascularization. Nat. Mater. 14, 532–539 (2015).
Chen, Y. et al. Emerging roles of 1D vertical nanostructures in orchestrating immune cell functions. Adv. Mater. 32, e2001668 (2020).
Choi, M. et al. Intracellular delivery of bioactive cargos to hard-to-transfect cells using carbon nanosyringe arrays under an applied centrifugal g-force. Adv. Health. Mater. 5, 101–107 (2016).
Yosef, N. et al. Dynamic regulatory network controlling TH17 cell differentiation. Nature 496, 461–468 (2013).
Pop, M. A. & Almquist, B. D. Controlled delivery of microRNAs into primary cells using nanostraw technology. Adv. NanoBiomed Res. 1, 2000061 (2021).
Bhingardive, V. et al. Antibody-functionalized nanowires: a tuner for the activation of T cells. Nano Lett. 21, 4241–4248 (2021).
Dixit, H. G. et al. Massively-parallelized, deterministic mechanoporation for intracellular delivery. Nano Lett. 20, 860–867 (2020).
Stuchbury, G. & Munch, G. Optimizing the generation of stable neuronal cell lines via pre-transfection restriction enzyme digestion of plasmid DNA. Cytotechnology 62, 189–194 (2010).
Shokouhi, A. R. et al. Engineering efficient CAR-T cells via electroactive nanoinjection. Adv. Mater. 35, e2304122 (2023).
Chen, Y. et al. Cellular deformations induced by conical silicon nanowire arrays facilitate gene delivery. Small 15, e1904819 (2019).
Singh, A. et al. Efficient modulation of T-cell response by dual-mode, single-carrier delivery of cytokine-targeted siRNA and DNA vaccine to antigen-presenting cells. Mol. Ther. 16, 2011–2021 (2008).
Singh, A. et al. An injectable synthetic immune-priming center mediates efficient T-cell class switching and T-helper 1 response against B cell lymphoma. J. Control. Release 155, 184–192 (2011).
Singh, A., Suri, S. & Roy, K. In-situ crosslinking hydrogels for combinatorial delivery of chemokines and siRNA-DNA carrying microparticles to dendritic cells. Biomaterials 30, 5187–5200 (2009).
Ma, F. et al. The microRNA miR-29 controls innate and adaptive immune responses to intracellular bacterial infection by targeting interferon-γ. Nat. Immunol. 12, 861–869 (2011).
Liu, X. et al. Genome-wide analysis identifies NR4A1 as a key mediator of T cell dysfunction. Nature 567, 525–529 (2019).
Hansel, C. S. et al. Nanoneedle-mediated stimulation of cell mechanotransduction machinery. ACS Nano 13, 2913–2926 (2019).
Olson, J. A., McDonald-Hyman, C., Jameson, S. C. & Hamilton, S. E. Effector-like CD8+ T cells in the memory population mediate potent protective immunity. Immunity 38, 1250–1260 (2013).
Jameson, S. C. & Masopust, D. Understanding subset diversity in T cell memory. Immunity 48, 214–226 (2018).
Huster, K. M. et al. Unidirectional development of CD8+ central memory T cells into protective Listeria-specific effector memory T cells. Eur. J. Immunol. 36, 1453–1464 (2006).
Higgins, S. G. et al. High-aspect-ratio nanostructured surfaces as biological metamaterials. Adv. Mater. 32, e1903862 (2020).
Kim, D., Paggi, J. M., Park, C., Bennett, C. & Salzberg, S. L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 37, 907–915 (2019).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
McCarthy, D. J., Chen, Y. & Smyth, G. K. Differential expression analysis of multifactor RNA-Seq experiments with respect to biological variation. Nucleic Acids Res. 40, 4288–4297 (2012).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Smith, N. L. et al. Developmental origin governs CD8+ T cell fate decisions during infection. Cell 174, 117–130 (2018).
Shah, S. B. et al. Combinatorial treatment rescues tumour-microenvironment-mediated attenuation of MALT1 inhibitors in B-cell lymphomas. Nat. Mater. 22, 511–523 (2023).
Moeller, T. D. et al. Profiling germinal center-like B cell responses to conjugate vaccines using synthetic immune organoids. ACS Cent. Sci. 9, 787–804 (2023).
Purwada, A. et al. Ex vivo synthetic immune tissues with T cell signals for differentiating antigen-specific, high affinity germinal center B cells. Biomaterials 198, 27–36 (2019).
Mosquera, M. J. et al. Extracellular matrix in synthetic hydrogel-based prostate cancer organoids regulate therapeutic response to EZH2 and DRD2 inhibitors. Adv. Mater. 34, e2100096 (2022).
Mosquera, M. J. et al. Immunomodulatory nanogels overcome restricted immunity in a murine model of gut microbiome-mediated metabolic syndrome. Sci. Adv. 5, eaav9788 (2019).
Acknowledgements
We acknowledge financial support from the Curci Foundation Award (A.S.); the National Science Foundation (EEC-1648035, seed funding awarded to A.S.); US National Institutes of Health NIH 5R01AI132738-06, 1R01CA266052-01, 1R01CA238745-01A1 and U01CA280984-01 (awarded to A.S.); and NIH R01AI110613 and U01AI131348 (awarded to B.D.R.). This work was performed in part at the Georgia Institute of Technology, Institute for Electronics and Nanotechnology, a member of the National Nanotechnology Coordinated Infrastructure (NNCI), which is supported by the National Science Foundation (ECCS-2025462). We further acknowledge support from the Cornell Nanoscale Science and Technology Facility under the NSF Grant ECCS-1542081 for the use of equipment. We acknowledge technical support on STEM from M. Tian at Georgia Institute of Technology, Institute for Electronics and Nanotechnology, IEN/IMAT material characterization facilities. We acknowledge technical support on Zetasizer from A. J. Heiler and S. N. Thomas at Georgia Institute of Technology. Figures 1a, 2i, 4a, 5a and 6a,e, and Extended Fig. 2c were created with BioRender.com.
Author information
Authors and Affiliations
Contributions
A.S. conceived the nanowire fabrication, and B.D.R. and A.S. conceived and designed the T-cell studies. K.J.Y.M. and S.K. designed and performed the original experiments and analysis. Z.D. performed all the revision experiments and analysis with A.S. K.J.Y.M., S.K., B.D.R., Z.D. and A.S. wrote the manuscript. A.G. and J.D.W. generated the miRNA. A.G. and H.Z. performed RNA-seq analysis. R.J. generated the eGFP-encoding lentiviruses. All authors provided critical feedback on the research, analysis and manuscript.
Corresponding authors
Ethics declarations
Competing interests
A.S. receives research support from Genentech, Inc. A.S. has an intellectual property disclosure filed on the technology with US Patent and Trademark Office (USPTO) assigned U.S. Application No. 63/591,553. The other authors declare no competing interests.
Peer review
Peer review information
Nature Nanotechnology thanks Mark Schvartzman and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 T cell characterization on nanowires.
a) Effect of Centrifugal g-Force on the survival of primary naïve CD8+ T cells on PEI-functionalized nanowires with 10 wt/v% PEI. Data are presented as mean ± s.e.m. One-way ANOVA with Tukey’s test (n = 3, each dot represents an independent nanowire chip). b-d) Representative SEM images of primary naïve murine CD8+ T cells seeded on the nanowires (B) and FIB-SEM etched T cells on the nanowires (C-D). e) Representative FIB-SEM image of melted nanowire inside a T cell.
Extended Data Fig. 2 Nanowire functionalization and characterization of T cell response.
a) Representative pseudocolor plots of small molecular weight dextran delivery into primary naïve murine CD8+ T cells using nanowires. b) Representative flow cytometry histograms of tbet and emoes proteins in naïve mouse CD8+ T cells incubated with no-nanowire (2D) and no-PEI coated nanowire delivery of miR-29-mimic to murine CD8+ T cells. c) Schematic of functionalization process of nanowires via covalent conjugation of PEI on nanowire surface. d) Representative flow cytometry gating of cell viability (top) and delivery efficacy (bottom) of 6-FAM-miR-29-mimic in primary naïve murine CD8+ T cells using 2D control, unmodified nanowires, and covalently conjugated (CC) or surface coated (SC) nanowires with PEI. T cells were incubated on nanowires for 96 hrs. e) Representative STEM images of blank nanowires showing the absence of a PEI layer. f) Representative EDX images of blank nanowires showing the minimal presence of Nitrogen (N), Carbon (C), and Oxygen (O) on Silicon (Si). g) Zeta potential measurement of bare silicon nanowires, PEI-functionalized nanowires, and miRNA-loaded PEI-functionalized nanowires. Data are presented as mean ± s.e.m. One-way ANOVA with Tukey’s test (n = 3, each dot represents an independent nanowire chip). h) Representative flow cytometry gating of cell viability (top) and delivery efficacy (bottom) of 6-FAM-miR-29-mimic in primary naïve murine CD8+ T cells at various miRNA doses and fixed PEI wt/v%. T cells were incubated on nanowires for 96 hrs.
Extended Data Fig. 3 Nanowire penetration in T cells.
a) 3D projection of confocal microscopy image of nanowires in T cells. PEI-functionalized nanowires were loaded with FITC-BSA (green), naïve mouse CD8+ T cells were seeded on nanowires, centrifuged, and cultured for 12 hrs. Fixed samples were stained with phalloidin (actin, orange) and DAPI (nucleus, blue). Data representative of 40 cells from 4 independent technical replicates, with 10 cells from each replicate. b) Orthogonal projection with nanowires (FITC), DAPI, and actin (orange) from the same sample as in (A). The inset on the right represents the maximum intensity projection (MIP) of the representative sample. Data representative of 40 cells from 4 independent technical replicates, with 10 cells from each replicate. c) 3D projection of representative confocal microscopy image of nanowires in T cells comparing PEI-functionalized nanowires to blank nanowires. Data representative of 40 cells from 4 technical replicates for each condition, with 10 cells from each replicate. d) Measurement of the height of nanowire penetration inside a naïve mouse CD8+ T cell. Data represents 10 samples randomly selected from 40 cells from 4 independent technical replicates. Data are presented as mean ± s.e.m. A two-tailed, unpaired t-test with Welch’s correction was performed. e) Quantification of the number of nanowires per cell from maximum intensity projection of confocal microscopy images. Each dot represents a cell and a total of 40 cells were analyzed across 4 independent technical replicates. Data are presented as mean ± s.e.m. A two-tailed, unpaired t-test with Welch’s correction was performed.
Extended Data Fig. 4 miRNA delivery and release in T cells.
a-b) Histogram overlays (A) and bar graph of 6-FAM-miR-29-mimic uptake or release in primary naïve mouse CD8+ T cells (B). Data are presented as mean ± s.e.m. One-way ANOVA with Tukey’s test (n = 3, each dot represents an independent nanowire chip). c) Representative confocal microscopy image of murine CD8+ T cell with delivered miR-29-mimic using PEI-functionalized nanowires. T cells were incubated on nanowires for 96 hrs. d) Bar graph of miR-29 delivery efficacy of CD8+ T cells upon 6-FAM-miR-29-mimic delivery of primary naïve murine CD8+ T cells with different nanowire tip sizes and covalently conjugated with 10 wt/v% PEI. T cells were incubated on nanowires for 96 hrs. Data are presented as mean ± s.e.m. One-way ANOVA with Tukey’s test (n = 3, each dot represents an independent nanowire chip).
Extended Data Fig. 5 Benchmarking PEI-functionalized nanowires against conventional methods of miRNA delivery.
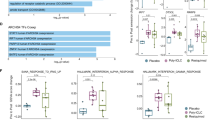
a) Bar graph representing % viability of primary naïve murine CD8+ T cells exposed to PEI-functionalized nanowires, Lipofectamine, PEI complexation, lentivirus, and nucleofection. 2D soluble miRNA delivery was used as a positive control. T cells were incubated on nanowires for 96 hrs. b) Bar graph representing % viability of primary, pre-activated murine CD8+ T cells exposed to PEI-functionalized nanowires, Lipofectamine, PEI complexation, lentivirus, and nucleofection. 2D soluble miRNA delivery was used as a positive control. T cells were pre-activated using anti-CD28/anti-CD3 beads. T cells were incubated on nanowires for 96 hrs. c) Bar graph representing % viability of primary naïve human CD8+ T cells exposed to PEI-functionalized nanowires, Lipofectamine, PEI complexation, lentivirus, and nucleofection. 2D soluble miRNA delivery was used as a positive control. T cells were incubated on nanowires for 96 hrs. d) Bar graph representing % viability of primary, pre-activated human CD8+ T cells exposed to PEI-functionalized nanowires, Lipofectamine, PEI complexation, lentivirus, and nucleofection. 2D soluble miRNA delivery was used as a positive control. T cells were pre-activated using anti-CD28/anti-CD3 beads. T cells were incubated on nanowires for 96 hrs. e) Bar graph representing % naïvity after 96 hrs when naïve murine CD8+ T cells are exposed to PEI-functionalized nanowires, Lipofectamine, PEI complexation, lentivirus, and nucleofection. 2D soluble miRNA delivery was used as a positive control. T cells were incubated on nanowires for 96 hrs. f) Bar graph representing % naïvity after 96 hrs when naïve human CD8+ T cells are exposed to PEI-functionalized nanowires, Lipofectamine, PEI complexation, lentivirus, and nucleofection. 2D soluble miRNA delivery was used as a positive control. T cells were incubated on nanowires for 96 hrs. In all figures, data are presented as mean ± s.e.m. One-way ANOVA with Tukey’s test (n = 3, each dot represents an independent nanowire chip).
Extended Data Fig. 6 Delivery of eGFP-encoding lentivirus to primary naïve human CD8+ T cells using nanowires.
a-b) Standard curve of concentration versus absorbance of ELISA against p24 antigen with lentiviral particulates (A) and bar graph representing % loading efficiency of lentivirus on nanowires (B) at various concentrations. Data are presented as mean ± s.e.m. (n = 3, each dot represents an independent nanowire chip). c) Histograms of fold-change in CD8+ GFP+ T cells upon eGFP-encoding lentiviral delivery using nanowires and without nanowires, in the presence or absence of reverse transcriptase inhibitor Efavirenz. T cells were incubated on lentiviruses on nanowires or without nanowires for 24 hr to minimize toxicity with lentiviruses, followed by culture in hIL-2 till day 4.
Extended Data Fig. 7 Co-delivery of miRNA antisense and mimic oligonucleotides by single functionalized nanowire platform toggles human CD8+ T cell fate switches.
a) % CD8+ CD62L- T cells in human CD8+ T cells. Primary naïve human CD8+ T cells were incubated on PEI-functionalized nanowires, loaded with miR-29-ASO, miR-130-mimic, or coloaded with miR-29-ASO and miR-130-mimic for 96 hrs, followed by TCR stimulation for the next 48 hrs. 2D culture of Primary naïve human CD8+ T cells with soluble miR-29-ASO and miR-130-mimic, followed by TCR stimulation for the next 48 hrs was used as a control. b) % viable CD8+ T cells in human CD8+ T cells in conditions defined in (A). c-d) % activated T cells expressing CD69 (C) and CD28 (D) markers in human CD8+ T cells in conditions defined in (A). e) % effector T cells in human CD8+ T cells in conditions defined in (A). f) % PD1+ T cells in human CD8+ T cells. Primary naïve human CD8+ T cells were incubated on PEI-functionalized nanowires, followed by TCR stimulation for the next 3 or 6 days. 2D culture of Primary naïve human CD8+ T cells with soluble miR-29-ASO and miR-130-mimic, followed by TCR stimulation for the next 48 hrs was used as a control. In all figures, data are presented as mean ± s.e.m. One-way ANOVA with Tukey’s test (n = 3 for A-D, F; n = 6 for E; each dot represents an independent nanowire chip).
Extended Data Fig. 8 Delivery of two miRNAs by functionalized nanowire enhances murine CD8+ T-cell effector function in vitro.
a-b) Representative histograms (A) and quantification (B) of cell-death markers Fas and AnnexinV expression in murine CD8+ T cells post 72 hrs of gB peptide stimulation. Primary naïve murine CD8+ T cells from gBT-1Tg mice were incubated on nanowires for 96 hrs, and gB peptide stimulation was done for the next 72 hrs. Data are presented as mean ± s.e.m. Two-way ANOVA with Tukey’s test, (n = 4, each dot represents an independent nanowire chip). c) Representative histograms (left) and bar graph (right) representing the expression of CD62L (MFI) on CD8+ T cells post 72 hrs of gB peptide stimulation. Primary naïve murine CD8+ T cells from gBT-1Tg mice were incubated on nanowires for 96 hrs, and gB peptide stimulation was done for the next 72 hrs. Data are presented as mean ± s.e.m. One-way ANOVA with Tukey’s test, (n = 4, each dot represents an independent nanowire chip). d) Bar graph representing the % CD8+ CD62L- cells in CD8+ T cells post 72 hr of gB peptide stimulation. Primary naïve murine CD8+ T cells from gBT-1Tg mice were incubated on nanowires for 96 hrs, and gB peptide stimulation was done for the next 72 hrs. Data are presented as mean ± s.e.m. One-way ANOVA with Tukey’s test, (n = 4, each dot represents an independent nanowire chip). e) Representative activation markers CD44, CD69, and CD28 expression histograms post 72 hr of gB peptide stimulation. Primary naïve murine CD8+ T cells from gBT-1Tg mice were incubated on nanowires for 96 hrs, and gB peptide stimulation was done for the next 72 hrs. Data are presented as mean ± s.e.m. Two-way ANOVA with Tukey’s test, (n = 4, each dot represents an independent nanowire chip). f) Representative histograms of cytokine TNFα and perforin-producing CD8+ T cells post 72 hr of gB peptide stimulation. Primary naïve murine CD8+ T cells from gBT-1Tg mice were incubated on nanowires for 96 hrs, and gB peptide stimulation was done for the next 72 hrs. Data are presented as mean ± s.e.m. Two-way ANOVA with Tukey’s test, (n = 4, each dot represents an independent nanowire chip).
Extended Data Fig. 9 Functionalized nanowire delivery of dual miRNA-related oligos influence the naïve CD8 T cell transcriptome.
a). Principal component analysis of naïve CD8+ T cells incubated for 96 hrs on nanowires coated with negative control miRNA (salmon) relative to miR-29-ASO/miR-130-mimic miRNA (blue). Data (each dot) represents a biological replicate (mice). b). Volcano plot of statistically significant, differentially expressed T cell-related genes comparing nanowires coated with negative control miRNA relative to miR-29-ASO/miR-130-mimic miRNA. The red dots represent select labeled genes that are upregulated, the blue dots represent select labeled genes that are downregulated. The colors green and magenta represent select genes from other biological processes. Data represent an average of two biological replicates (mice). c). Volcano plot of statistically significant, differentially expressed mechanoregulatory genes in naïve CD8+ T cells on nanowires coated with negative control miRNA (blue) relative to no nanowire control (red). Data represent an average of two biological replicates (mice).
Extended Data Fig. 10 Delivery of dual miRNA components by functionalized nanowire enhances murine CD8+ T cell effector memory function in vivo in influenza A.
a-b) Representative contour plots (A) and quantification (B) of effector memory T cell (Tem) and central memory T cell (Tcm) phenotype of Lung and spleen CD8+ T cells at Day 8 Flu gB primary infection. Data are presented as mean ± s.e.m. Two-way ANOVA with Tukey’s multiple comparison test, (n = 6 mice).
Supplementary information
Supplementary Information
Supplementary Figs. 1–5.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 9
Statistical source data.
Source Data Extended Data Fig. 10
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yee Mon, K.J., Kim, S., Dai, Z. et al. Functionalized nanowires for miRNA-mediated therapeutic programming of naïve T cells. Nat. Nanotechnol. (2024). https://doi.org/10.1038/s41565-024-01649-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41565-024-01649-7