Abstract
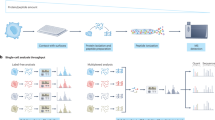
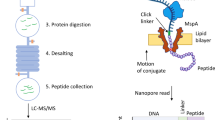
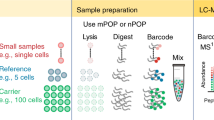
Proteins are major building blocks of life. The protein content of a cell and an organism provides key information for the understanding of biological processes and disease. Despite the importance of protein analysis, only a handful of techniques are available to determine protein sequences, and these methods face limitations, for example, requiring a sizable amount of sample. Single-molecule techniques would revolutionize proteomics research, providing ultimate sensitivity for the detection of low-abundance proteins and the realization of single-cell proteomics. In recent years, novel single-molecule protein sequencing schemes that use fluorescence, tunnelling currents and nanopores have been proposed. Here, we present a review of these approaches, together with the first experimental efforts towards their realization. We discuss their advantages and drawbacks, and present our perspective on the development of single-molecule protein sequencing techniques.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Miyashita, M. et al. Attomole level protein sequencing by Edman degradation coupled with accelerator mass spectrometry. Proc. Natl Acad. Sci. USA 98, 4403–8 (2001).
Shimonishi, Y. et al. Sequencing of peptide mixtures by Edman degradation and field‐desorption mass spectrometry. Eur. J. Biochem. 112, 251–264 (1980).
Bradley, C. V., Williams, D. H. & Hanley, M. R. Peptide sequencing using the combination of edman degradation, carboxypeptidase digestion and fast atom bombardment mass spectrometry. Biochem. Biophys. Res. Commun. 104, 1223–30 (1982).
Steen, H. & Mann, M. The abc’s (and xyz’s) of peptide sequencing. Nat. Rev. Mol. Cell Biol. 5, 699–711 (2004).
Yates, J. R. III A century of mass spectrometry: from atoms to proteomes. Nat. Methods 8, 633–637 (2011).
Aebersold, R. & Mann, M. Mass-spectrometric exploration of proteome structure and function. Nature 537, 347–355 (2016).
Walther, T. C. & Mann, M. Mass spectrometry-based proteomics in cell biology. J. Cell Biol. 190, 491–500 (2010).
Domon, B. & Aebersold, R. Options and considerations when selecting a quantitative proteomics strategy. Nat. Biotechnol. 28, 710–721 (2010).
A cast of thousands. Nat. Biotechnol. 21, 213 (2003).
Anderson, N. L. The human plasma proteome: History, character, and diagnostic prospects. Mol. Cell. Proteomics 1, 845–867 (2002).
Zubarev, R. A. The challenge of the proteome dynamic range and its implications for in-depth proteomics. Proteomics 13, 723–726 (2013).
Huang, B. et al. Counting low-copy number proteins in a single cell. Science 315, 81–84 (2007).
Ham, B. M. & MaHam, A. Analytical Chemistry: A Chemist and Laboratory Technician’s Toolkit (Wiley, Hoboken, 2015).
Hawkridge, A. M. in Quantitative Proteomics (eds Eyers, C. E. & Gaskell, S.) 1–25 (RSC, Cambridge, 2014).
Pagel, O., Loroch, S., Sickmann, A. & Zahedi, R. P. Current strategies and findings in clinically relevant post-translational modification-specific proteomics. Expert Rev. Proteomics 12, 235–253 (2015).
Heath, J. R., Ribas, A. & Mischel, P. S. Single-cell analysis tools for drug discovery and development. Nat. Rev. Drug Discov. 15, 204–216 (2015).
Prakadan, S. M., Shalek, A. K. & Weitz, D. A. Scaling by shrinking: empowering single-cell ‘omics’ with microfluidic devices. Nat. Rev. Genet. 18, 345–361 (2017).
Su, Y., Shi, Q. & Wei, W. Single cell proteomics in biomedicine: High-dimensional data acquisition, visualization,and analysis. Proteomics 17, 1600267 (2017).
Lu, Y., Yang, L., Wei, W. & Shi, Q. Microchip-based single-cell functional proteomics for biomedical applications. Lab Chip 17, 1250–1263 (2017).
Spitzer, M. H. & Nolan, G. P. Mass Cytometry: Single Cells, Many Features. Cell 165, 780–791 (2016).
Bentley, D. R. et al. Accurate whole human genome sequencing using reversible terminator chemistry. - Supplement. Nature 456, 53–9 (2008).
Eid, J. et al. Real-Time DNA Sequencing from Single Polymerase Molecules. Science 323, 133–138 (2009).
Braslavsky, I., Hebert, B., Kartalov, E. & Quake, S. R. Sequence information can be obtained from single DNA molecules. Proc. Natl Acad. Sci.USA 100, 3960–3964 (2003).
Hernandez, E. T., Swaminathan, J., Marcotte, E. M. & Anslyn, E. V. Solution-phase and solid-phase sequential, selective modification of side chains in KDYWEC and KDYWE as models for usage in single-molecule protein sequencing. New J. Chem. 41, 462–469 (2017).
Yao, Y., Docter, M., van Ginkel, J., de Ridder, D. & Joo, C. Single-molecule protein sequencing through fingerprinting: computational assessment. Phys. Biol. 12, 055003 (2015).
Swaminathan, J., Boulgakov, A. A. & Marcotte, E. M. A Theoretical justification for single molecule peptide sequencing. PLOS Comput. Biol. 11, e1004080 (2015).
Müller, V. & Westerlund, F. Optical DNA mapping in nanofluidic devices: principles and applications. Lab Chip 17, 579–590 (2017).
van Ginkel, J. et al. Single-molecule peptide fingerprinting. Proc. Natl Acad. Sci.USA 115, 3338–3343 (2018).
Preminger, M. & Smilansky, Z. Methods for evaluating ribonucleotide sequences. US patent 9,012,150 (2009).
Stevens, B. et al. Fret-based identification of mRNAs undergoing translation. PLoS One 7, e38344 (2012).
Swaminathan, J. Single Molecule Peptide Sequencing. PhD thesis, University of Texas at Austin (2015).
Borgo, B. & Havranek, J. J. Computer-aided design of a catalyst for Edman degradation utilizing substrate-assisted catalysis. Protein Sci. 24, 571–579 (2015).
Aviram, A. & Ratner, M. A. Molecular rectifiers. Chem. Phys. Lett. 29, 277–283 (1974).
Dekker, C., Tans, S. J., Oberndorff, B., Meyer, R. & Venema, L. C. STM imaging and spectroscopy of single copperphthalocyanine molecules. Synth. Met. 84, 853–854 (1997).
Reed, M. A. Conductance of a Molecular Junction. Science 278, 252–254 (1997).
Ratner, M. A brief history of molecular electronics. Nat. Nanotech. 8, 378–381 (2013).
Tsutsui, M., Taniguchi, M., Yokota, K. & Kawai, T. Identifying single nucleotides by tunnelling current. Nat. Nanotech. 5, 286–290 (2010).
Tanaka, H. & Kawai, T. Partial sequencing of a single DNA molecule with a scanning tunnelling microscope. Nat. Nanotech. 4, 518–522 (2009).
Shapir, E. et al. Electronic structure of single DNA molecules resolved by transverse scanning tunnelling spectroscopy. Nat. Mater. 7, 68–74 (2008).
Chang, S. et al. Electronic signatures of all four DNA nucleosides in a tunneling gap. Nano Lett. 10, 1070–1075 (2010).
Feng, J. et al. Identification of single nucleotides in MoS2 nanopores. Nat. Nanotech. 11, 117–126 (2015).
Lindsay, S. et al. Recognition tunneling. Nanotechnology 21, 262001 (2010).
Zhao, Y. et al. Single-molecule spectroscopy of amino acids and peptides by recognition tunnelling. Nat. Nanotech. 9, 466–73 (2014).
Ohshiro, T. et al. Detection of post-translational modifications in single peptides using electron tunnelling currents. Nat. Nanotech. 9, 835–840 (2014).
Morikawa, T., Yokota, K., Tsutsui, M. & Taniguchi, M. Fast and low-noise tunnelling current measurements forsingle-molecule detection in an electrolyte solution using insulator-protected nanoelectrodes. Nanoscale 9, 4076–4081 (2017).
Morikawa, T., Yokota, K., Tanimoto, S., Tsutsui, M. & Taniguchi, M. Detecting single-nucleotides by tunneling current measurements at sub-MHz temporal resolution. Sensors 17, 885–893 (2017).
Taniguchi, M., Tsutsui, M., Yokota, K. & Kawai, T. Fabrication of the gating nanopore device. Appl. Phys. Lett. 95, 123701 (2009).
Yokota, K., Tsutsui, M. & Taniguchi, M. Electrode-embedded nanopores for label-free single-moleculesequencing by electric currents. RSC Adv. 4, 15886–15899 (2014).
Heerema, S. J. & Dekker, C. Graphene nanodevices for DNA sequencing. Nat. Nanotech. 11, 127–136 (2016).
Ivanov, A. P. et al. DNA tunneling detector embedded in a nanopore. Nano Lett. 11, 279–285 (2011).
Lu, H., Giordano, F. & Ning, Z. Oxford nanopore MinION sequencing and genome assembly. Genomics Proteomics Bioinformatics 14, 265–279 (2016).
Jain, M. et al. Improved data analysis for the MinION nanopore sequencer. Nat. Methods 12, 351–356 (2015).
Jain, M., Olsen, H. E., Paten, B. & Akeson, M. The Oxford Nanopore MinION: delivery of nanopore sequencing to the genomics community. Genome Biol. 17, 239–249 (2016).
Deamer, D., Akeson, M. & Branton, D. Three decades of nanopore sequencing. Nat. Biotechnol. 34, 518–524 (2016).
Gaskill, M. First DNA sequencing in space a game changer. NASA (2016); https://www.nasa.gov/mission_pages/station/research/news/dna_sequencing
Waduge, P. et al. Nanopore-based measurements of protein size, fluctuations, and conformational changes. ACS Nano. 11, 5706–5716 (2017).
Bell, N. A. W. & Keyser, U. F. Digitally encoded DNA nanostructures for multiplexed, single-molecule protein sensing with nanopores. Nat. Nanotech. 11, 645–651 (2016).
Plesa, C., Ruitenberg, J. W., Witteveen, M. J. & Dekker, C. Detection of individual proteins bound along DNA using solid-state nanopores. Nano Lett. 15, 3153–3158 (2015).
Venkatesan, B. M. & Bashir, R. Nanopore sensors for nucleic acid analysis. Nat. Nanotech. 6, 615–624 (2011).
Stefureac, R., Long, Y.-T., Kraatz, H.-B., Howard, P. & Lee, J. S. Transport of α-helical peptides through α-jemolysin and aerolysin pores. Biochemistry 45, 9172–9179 (2006).
Sutherland, T. C. et al. Structure of peptides investigated by nanopore analysis. Nano Lett. 4, 1273–1277 (2004).
Movileanu, L., Schmittschmitt, J. P., Martin Scholtz, J. & Bayley, H. Interactions of peptides with a protein pore. Biophys. J. 89, 1030–1045 (2005).
Goodrich, C. P. et al. Single-molecule electrophoresis of β-hairpin peptides by electrical recordings and langevin dynamics simulations. J. Phys. Chem. B 111, 3332–3335 (2007).
Mohammad, M. M. & Movileanu, L. Excursion of a single polypeptide into a protein pore: simple physics, but complicated biology. Eur. Biophys. J. 37, 913–925 (2008).
Mahendran, K. R., Romero-Ruiz, M., Schlösinger, A., Winterhalter, M. & Nussberger, S. Protein translocation through Tom40: Kinetics of peptide release. Biophys. J. 102, 39–47 (2012).
Ji, Z. et al. Fingerprinting of peptides with a large channel of bacteriophage Phi29 DNA packaging motor. Small 12, 4572–4578 (2016).
Talaga, D. S & Li, J. Single-molecule protein unfolding in solid state nanopores. J. Am. Chem. Soc. 131, 9287–9297 (2009).
Li, J., Fologea, D., Rollings, R. & Ledden, B. Characterization of protein unfolding with solid-state nanopores. Protein Pept. Lett. 21, 256–265 (2014).
Restrepo-Pérez, L., John, S., Aksimentiev, A., Joo, C. & Dekker, C. SDS-assisted protein transport through solid-state nanopores. Nanoscale 9, 11685–11693 (2017).
Oukhaled, G. et al. Unfolding of proteins and long transient conformations detected by single nanopore recording. Phys. Rev. Lett. 98, 158101 (2007).
Pastoriza-Gallego, M. et al. Dynamics of unfolded protein transport through an aerolysin pore. J. Am. Chem.Soc. 133, 2923–2931 (2011).
Merstorf, C. et al. Wild type, mutant protein unfolding and phase transition detected by single-nanopore recording. ACS Chem. Biol. 7, 652–658 (2012).
Pastoriza-Gallego, M. et al. Urea denaturation of α-hemolysin pore inserted in planar lipid bilayer detected bysingle nanopore recording: Loss of structural asymmetry. FEBS Lett. 581, 3371–3376 (2007).
Freedman, K. J. et al. Chemical, thermal, and electric field induced unfolding of single protein molecules studiedusing nanopores. Anal. Chem. 83, 5137–5144 (2011).
Payet, L. et al. Thermal unfolding of proteins probed at the single molecule level using nanopores. Anal. Chem. 84, 4071–4076 (2012).
Cressiot, B. et al. Protein transport through a narrow solid-state nanopore at high voltage: Experiments andtheory. ACS Nano 6, 6236–6243 (2012).
Oukhaled, A. et al. Dynamics of completely unfolded and native proteins through solid-state nanopores as afunction of electric driving force. ACS Nano 5, 3628–3638 (2011).
Freedman, K. J., Haq, S. R., Edel, J. B., Jemth, P. & Kim, M. J. Single molecule unfolding and stretching of protein domains inside a solid-state nanopore by electric field. Sci. Rep. 3, 1638 (2013).
Firnkes, M., Pedone, D., Knezevic, J., Döblinger, M. & Rant, U. Electrically facilitated translocations of proteins through silicon nitride nanopores: Conjoint and competitive action of diffusion, electrophoresis, and electroosmosis. Nano Lett. 10, 2162–2167 (2010).
Huang, G., Willems, K., Soskine, M., Wloka, C. & Maglia, G. Electro-osmotic capture and ionic discrimination of peptide and protein biomarkers with FraC nanopores. Nat. Commun. 8, 935 (2017).
Kennedy, E., Dong, Z., Tennant, C. & Timp, G. Reading the primary structure of a protein with 0.07 nm3 resolution using a subnanometre-diameter pore. Nat. Nanotech. 11, 968–976 (2016).
Dong, Z., Kennedy, E., Hokmabadi, M. & Timp, G. Discriminating residue substitutions in a single protein molecule using a sub-nanopore. ACS Nano 11, 5440–5452 (2017).
Rodriguez-Larrea, D. & Bayley, H. Multistep protein unfolding during nanopore translocation. Nat. Nanotech. 8, 288–295 (2013).
Rodriguez-Larrea, D. & Bayley, H. Protein co-translocational unfolding depends on the direction of pulling. Nat. Commun. 5, 4841 (2014).
Rosen, C. B., Rodriguez-Larrea, D. & Bayley, H. Single-molecule site-specific detection of protein phosphorylation with a nanopore. Nat. Biotechnol. 32, 179–181 (2014).
Biswas, S., Song, W., Borges, C., Lindsay, S. & Zhang, P. Click addition of a DNA thread to the N-termini of peptides for their translocation through solid-state nanopores. ACS Nano 9, 9652–9664 (2015).
Pastoriza-Gallego, M. et al. Evidence of unfolded protein translocation through a protein nanopore. ACS Nano 8, 11350–11360 (2014).
Plesa, C. et al. Fast translocation of proteins through solid state nanopores. Nano Lett. 13, 658–63 (2013).
Nivala, J., Marks, D. B. & Akeson, M. Unfoldase-mediated protein translocation through an α-hemolysin nanopore. Nat. Biotechnol. 31, 247–250 (2013).
Nivala, J., Mulroney, L., LiG., Schreiber, J. & Akeson, M. Discrimination among protein variants using an unfoldase-coupled nanopore. ACS Nano 8, 12365–12375 (2014).
Aubin-Tam, M.-E., Olivares, A. O., Sauer, R. T., Baker, T. A. & Lang, M. J. Single-molecule protein unfolding and translocation by an ATP-fueled proteolytic machine. Cell 145, 257–67 (2011).
Sampath, G. Amino acid discrimination in a nanopore and the feasibility of sequencing peptides with a tandem cell and exopeptidase. RSC Adv. 5, 30694–30700 (2015).
Boynton, P. & Di Ventra, M. Sequencing proteins with transverse ionic transport in nanochannels. Sci. Rep. 6, 25232 (2016).
Wilson, J., Sloman, L., He, Z. & Aksimentiev, A. Graphene nanopores for protein sequencing. Adv. Funct.Mater. 26, 4830–4838 (2016).
Maulbetsch, W., Wiener, B., Poole, W., Bush, J. & Stein, D. Preserving the sequence of a biopolymer’s monomers as they enter an electrospray mass spectrometer. Phys. Rev. Appl. 6, 054006 (2016).
Keifer, D. Z. & Jarrold, M. F. Single-molecule mass spectrometry. Mass Spec. Rev. 36, 715–733 (2016).
Manrao, E. A. et al. Reading DNA at single-nucleotide resolution with a mutant MspA nanopore and phi29 DNA polymerase. Nat. Biotechnol. 30, 349–353 (2012).
Millioni, R. et al. High abundance proteins depletion vs low abundance proteins enrichment: Comparison of methods to reduce the plasma proteome complexity. PLoS One 6, e19603 (2011).
Baker, M. S. et al. Accelerating the search for the missing proteins in the human proteome. Nat. Commun. 8, 14271 (2017).
Wetterstrand, K. DNA Sequencing Costs: Data from the NHGRI Genome Sequencing Program (GSP). Available at: www.genome.gov/sequencingcostsdata (Accessed 2 July 2018).
Goodwin, S., McPherson, J. D. & McCombie, W. R. Coming of age: ten years of next-generation sequencing technologies. Nat. Rev. Genet. 17, 333–351 (2016).
Haider, S. & Pal, R. Integrated analysis of transcriptomic and proteomic data. Curr. Genomics 14, 91–110 (2013).
Pandey, A. & Mann, M. Proteomics to study genes and genomes. Nature 405, 837–846 (2000).
Tyers, M. & Mann, M. From genomics to proteomics. Nature 422, 193–197 (2003).
Edman, P. Method for determination of the amino acid sequence in peptides. Acta Chemica Scandinavica 4, 283–293 (1950).
Li, K. W. & Geraerts, W. P. M. in Neuropeptide Protocols (eds Irvine, G. B. & Williams, C. H.) 17–26 (Humana Press, New York, 1997).
McCormack, A. L. et al. Direct analysis and identification of proteins in mixtures by LC/MS/MS and database searching at the low-femtomole level. Anal. Chem. 69, 767–776 (1997).
Acknowledgements
We thank S. Pud, S. Schmid, S. Caneva, J. van Ginkel and M. Filius for discussions. We acknowledge funding received from the Netherlands Organisation for Scientific Research (NWO/OCW) as a part of the Frontiers of Nanoscience programme. The C.D. lab was further supported by the ERC Advanced Grant SynDiv (No. 669598) and by the National Human Genome Research Institute of the National Institute of Health under Award Number R01-HG007406. C.J. was funded by the Foundation for Fundamental Research on Matter (12PR3029 and SMPS).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Restrepo-Pérez, L., Joo, C. & Dekker, C. Paving the way to single-molecule protein sequencing. Nature Nanotech 13, 786–796 (2018). https://doi.org/10.1038/s41565-018-0236-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41565-018-0236-6
This article is cited by
-
Nanopore DNA sequencing technologies and their applications towards single-molecule proteomics
Nature Chemistry (2024)
-
Peptide sequencing based on host–guest interaction-assisted nanopore sensing
Nature Methods (2024)
-
Real-time detection of 20 amino acids and discrimination of pathologically relevant peptides with functionalized nanopore
Nature Methods (2024)
-
Optical sequencing of single synthetic polymers
Nature Chemistry (2024)
-
Full-length single-molecule protein fingerprinting
Nature Nanotechnology (2024)