Abstract
CRISPR–Cas systems provide heritable immunity against viruses and other mobile genetic elements by incorporating fragments of invader DNA into the host CRISPR array as spacers. Integration of new spacers is localized to the 5′ end of the array, and in certain Gram-negative Bacteria this polarized localization is accomplished by the integration host factor. For most other Bacteria and Archaea, the mechanism for 5′ end localization is unknown. Here we show that archaeal histones play a key role in directing integration of CRISPR spacers. In Pyrococcus furiosus, deletion of either histone A or B impairs integration. In vitro, purified histones are sufficient to direct integration to the 5′ end of the CRISPR array. Archaeal histone tetramers and bacterial integration host factor induce similar U-turn bends in bound DNA. These findings indicate a co-evolution of CRISPR arrays with chromosomal DNA binding proteins and a widespread role for binding and bending of DNA to facilitate accurate spacer integration.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Sequencing data generated for this study have been deposited in the NCBI Sequence Read Archive (SRA) under the BioProjectID PRJNA901420. Source data are provided with this paper.
Code availability
Custom python scripts were generated to quantify read coverage patterns and dinucleotide frequencies, and to parse and reformat alignment output files for display on the UCSC Genome Browser. The codes are available from the corresponding authors upon request or S.C.G. at garrett@uchc.edu.
References
Barrangou, R. et al. CRISPR provides acquired resistance against viruses in prokaryotes. Science 315, 1709–1712 (2007).
Alkhnbashi, O. S. et al. Characterizing leader sequences of CRISPR loci. Bioinformatics 32, i576–i585 (2016).
Wei, Y., Chesne, M. T., Terns, R. M. & Terns, M. P. Sequences spanning the leader–repeat junction mediate CRISPR adaptation to phage in Streptococcus thermophilus. Nucleic Acids Res. 43, 1749–1758 (2015).
Yosef, I., Goren, M. G. & Qimron, U. Proteins and DNA elements essential for the CRISPR adaptation process in Escherichia coli. Nucleic Acids Res. 40, 5569–5576 (2012).
Hale, C. R. et al. Essential features and rational design of CRISPR RNAs that function with the Cas RAMP module complex to cleave RNAs. Mol. Cell 45, 292–302 (2012).
Carte, J., Wang, R. Y., Li, H., Terns, R. M. & Terns, M. P. Cas6 is an endoribonuclease that generates guide RNAs for invader defense in prokaryotes. Genes Dev. 22, 3489–3496 (2008).
Makarova, K. S. et al. Evolutionary classification of CRISPR–Cas systems: a burst of class 2 and derived variants. Nat. Rev. Microbiol., https://doi.org/10.1038/s41579-019-0299-x (2019).
Brouns, S. J. et al. Small CRISPR RNAs guide antiviral defense in prokaryotes. Science 321, 960–964 (2008).
Hale, C. R. et al. RNA-guided RNA cleavage by a CRISPR RNA-Cas protein complex. Cell 139, 945–956 (2009).
Nussenzweig, P. M. & Marraffini, L. A. Molecular mechanisms of CRISPR–Cas immunity in bacteria. Annu. Rev. Genet. 54, 93–120 (2020).
McGinn, J. & Marraffini, L. A. Molecular mechanisms of CRISPR–Cas spacer acquisition. Nat. Rev. Microbiol. 17, 7–12 (2019).
Xiao, Y., Ng, S., Nam, K. H. & Ke, A. How type II CRISPR–Cas establish immunity through Cas1–Cas2-mediated spacer integration. Nature 550, 137–141 (2017).
Nunez, J. K. et al. Cas1–Cas2 complex formation mediates spacer acquisition during CRISPR–Cas adaptive immunity. Nat. Struct. Mol. Biol. 21, 528–534 (2014).
Shiimori, M., Garrett, S. C., Graveley, B. R. & Terns, M. P. Cas4 nucleases define the PAM, length, and orientation of DNA fragments integrated at CRISPR loci. Mol. Cell 70, 814–824 e816 (2018).
Hu, C. et al. Mechanism for Cas4-assisted directional spacer acquisition in CRISPR–Cas. Nature 598, 515–520 (2021).
Heler, R. et al. Cas9 specifies functional viral targets during CRISPR–Cas adaptation. Nature 519, 199–202 (2015).
Wei, Y. Z., Terns, R. M. & Terns, M. P. Cas9 function and host genome sampling in type II-A CRISPR–Cas adaptation. Genes Dev. 29, 356–361 (2015).
Levy, A. et al. CRISPR adaptation biases explain preference for acquisition of foreign DNA. Nature 520, 505–510 (2015).
Jakhanwal, S. et al. A CRISPR–Cas9–integrase complex generates precise DNA fragments for genome integration. Nucleic Acids Res. 49, 3546–3556 (2021).
Ivancic-Bace, I., Cass, S. D., Wearne, S. J. & Bolt, E. L. Different genome stability proteins underpin primed and naive adaptation in E. coli CRISPR–Cas immunity. Nucleic Acids Res. 43, 10821–10830 (2015).
Nunez, J. K., Lee, A. S., Engelman, A. & Doudna, J. A. Integrase-mediated spacer acquisition during CRISPR–Cas adaptive immunity. Nature 519, 193–198 (2015).
Rollie, C., Schneider, S., Brinkmann, A. S., Bolt, E. L. & White, M. F. Intrinsic sequence specificity of the Cas1 integrase directs new spacer acquisition. eLife 4, https://doi.org/10.7554/eLife.08716 (2015).
Barrangou, R. & Dudley, E. G. CRISPR-based typing and next-generation tracking technologies. Annu. Rev. Food Sci. Technol. 7, 395–411 (2016).
Kim, J. G., Garrett, S., Wei, Y., Graveley, B. R. & Terns, M. P. CRISPR DNA elements controlling site-specific spacer integration and proper repeat length by a type II CRISPR–Cas system. Nucleic Acids Res. 47, 8632–8648 (2019).
Wright, A. V. & Doudna, J. A. Protecting genome integrity during CRISPR immune adaptation. Nat. Struct. Mol. Biol. 23, 876–883 (2016).
Yoganand, K. N., Sivathanu, R., Nimkar, S. & Anand, B. Asymmetric positioning of Cas1–2 complex and Integration Host Factor induced DNA bending guide the unidirectional homing of protospacer in CRISPR–Cas type I-E system. Nucleic Acids Res. 45, 367–381 (2017).
Nunez, J. K., Bai, L., Harrington, L. B., Hinder, T. L. & Doudna, J. A. CRISPR immunological memory requires a host factor for specificity. Mol. Cell 62, 824–833 (2016).
Santiago-Frangos, A., Buyukyoruk, M., Wiegand, T., Krishna, P. & Wiedenheft, B. Distribution and phasing of sequence motifs that facilitate CRISPR adaptation. Curr. Biol. 31, 3515–3524 e3516 (2021).
Fagerlund, R. D. et al. Spacer capture and integration by a type I-F Cas1–Cas2-3 CRISPR adaptation complex. Proc. Natl Acad. Sci. USA 114, E5122–E5128 (2017).
Wright, A. V. et al. Structures of the CRISPR genome integration complex. Science 357, 1113–1118 (2017).
Rice, P. A., Yang, S., Mizuuchi, K. & Nash, H. A. Crystal structure of an IHF-DNA complex: a protein-induced DNA U-turn. Cell 87, 1295–1306 (1996).
Rollie, C., Graham, S., Rouillon, C. & White, M. F. Prespacer processing and specific integration in a type I-A CRISPR system. Nucleic Acids Res. 46, 1007–1020 (2018).
Garrett, S. et al. Primed CRISPR DNA uptake in Pyrococcus furiosus. Nucleic Acids Res. 48, 6120–6135 (2020).
Shiimori, M. et al. Role of free DNA ends and protospacer adjacent motifs for CRISPR DNA uptake in Pyrococcus furiosus. Nucleic Acids Res. https://doi.org/10.1093/nar/gkx839 (2017).
Terns, R. M. & Terns, M. P. The RNA- and DNA-targeting CRISPR–Cas immune systems of Pyrococcus furiosus. Biochem. Soc. Trans. 41, 1416–1421 (2013).
Tang, D. et al. A distinct structure of Cas1–Cas2 complex provides insights into the mechanism for the longer spacer acquisition in Pyrococcus furiosus. Int. J. Biol. Macromol. 183, 379–386 (2021).
Grainy, J., Garrett, S., Graveley, B. R., & Terns, M. P. CRISPR repeat sequences and relative spacing specify DNA integration by Pyrococcus furiosus Cas1 and Cas2. Nucleic Acids Res. https://doi.org/10.1093/nar/gkz548 (2019).
Laursen, S. P., Bowerman, S. & Luger, K. Archaea: the final frontier of chromatin. J. Mol. Biol. 433, 166791 (2021).
Henneman, B., van Emmerik, C., van Ingen, H. & Dame, R. T. Structure and function of archaeal histones. PLoS Genet. 14, e1007582 (2018).
Hocher, A. et al. Growth temperature and chromatinization in Archaea. Nat. Microbiol. https://doi.org/10.1038/s41564-022-01245-2 (2022).
Mattiroli, F. et al. Structure of histone-based chromatin in Archaea. Science 357, 609–612 (2017).
Maruyama, H. et al. An alternative beads-on-a-string chromatin architecture in Thermococcus kodakarensis. EMBO Rep. 14, 711–717 (2013).
Sandman, K., Krzycki, J. A., Dobrinski, B., Lurz, R. & Reeve, J. N. HMf, a DNA-binding protein isolated from the hyperthermophilic archaeon Methanothermus fervidus, is most closely related to histones. Proc. Natl Acad. Sci. USA 87, 5788–5791 (1990).
Stevens, K. M., Hocher, A. & Warnecke, T. Deep conservation of histone variants in Thermococcales Archaea. Genome Biol. Evol. 14, https://doi.org/10.1093/gbe/evab274 (2022).
Nalabothula, N. et al. Archaeal nucleosome positioning in vivo and in vitro is directed by primary sequence motifs. BMC Genom. 14, 391 (2013).
Bailey, K. A., Pereira, S. L., Widom, J. & Reeve, J. N. Archaeal histone selection of nucleosome positioning sequences and the procaryotic origin of histone-dependent genome evolution. J. Mol. Biol. 303, 25–34 (2000).
Watts, E. A. et al. Hyper-stimulation of Pyrococcus furiosus CRISPR DNA uptake by a self-transmissible plasmid. Extremophiles 26, 36 (2022).
Cubonovaa, L. et al. An archaeal histone is required for transformation of Thermococcus kodakarensis. J. Bacteriol. 194, 6864–6874 (2012).
Efremov, A. K. et al. Transcriptional repressor TrmBL2 from Thermococcus kodakarensis forms filamentous nucleoprotein structures and competes with histones for DNA binding in a salt- and DNA supercoiling-dependent manner. J. Biol. Chem. 290, 15770–15784 (2015).
Crnigoj, M. et al. Interactions of archaeal chromatin proteins Alba1 and Alba2 with nucleic acids. PLoS ONE 8, e58237 (2013).
Garrett, S. C. Pruning and tending immune memories: spacer dynamics in the CRISPR array. Front. Microbiol. 12, 664299 (2021).
Adam, P. S., Borrel, G., Brochier-Armanet, C. & Gribaldo, S. The growing tree of Archaea: new perspectives on their diversity, evolution and ecology. ISME J. 11, 2407–2425 (2017).
McGinn, J. & Marraffini, L. A. CRISPR–Cas systems optimize their immune response by specifying the site of spacer integration. Mol. Cell 64, 616–623 (2016).
Shipman, S. L., Nivala, J., Macklis, J. D. & Church, G. M. CRISPR–Cas encoding of a digital movie into the genomes of a population of living bacteria. Nature 547, 345–349 (2017).
Shipman, S. L., Nivala, J., Macklis, J. D. & Church, G. M. Molecular recordings by directed CRISPR spacer acquisition. Science 353, aaf1175 (2016).
Bhattarai-Kline, S. et al. Recording gene expression order in DNA by CRISPR addition of retron barcodes. Nature 608, 217–225 (2022).
Schmidt, F., Cherepkova, M. Y. & Platt, R. J. Transcriptional recording by CRISPR spacer acquisition from RNA. Nature 562, 380–385 (2018).
Schmidt, F. et al. Noninvasive assessment of gut function using transcriptional recording sentinel cells. Science 376, eabm6038 (2022).
Bridger, S. L., Lancaster, W. A., Poole, F. L. 2nd, Schut, G. J. & Adams, M. W. Genome sequencing of a genetically tractable Pyrococcus furiosus strain reveals a highly dynamic genome. J. Bacteriol. 194, 4097–4106 (2012).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Adams, M. W. et al. Key role for sulfur in peptide metabolism and in regulation of three hydrogenases in the hyperthermophilic archaeon Pyrococcus furiosus. J. Bacteriol. 183, 716–724 (2001).
Elmore, J., Deighan, T., Westpheling, J., Terns, R. M. & Terns, M. P. DNA targeting by the type I-G and type I-A CRISPR–Cas systems of Pyrococcus furiosus. Nucleic Acids Res. 43, 10353–10363 (2015).
Liman, G. L. S., Stettler, M. E. & Santangelo, T. J. Transformation techniques for the anaerobic hyperthermophile Thermococcus kodakarensis. Methods Mol. Biol. 2522, 87–104 (2022).
Sanders, T. J. et al. TFS and Spt4/5 accelerate transcription through archaeal histone-based chromatin. Mol. Microbiol. 111, 784–797 (2019).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Lynch, T. W., Read, E. K., Mattis, A. N., Gardner, J. F. & Rice, P. A. Integration host factor: putting a twist on protein-DNA recognition. J. Mol. Biol. 330, 493–502 (2003).
Acknowledgements
We thank the members of the Terns, Graveley and Santangelo labs for the helpful discussions. M.P.T. discloses support for this work from the National Institutes of Health (R35GM118160). B.R.G. discloses support for this work from the National Institutes of Health (R35GM118140). T.J. Santangelo discloses support for this work from the National Institutes of Health (R35GM143963 and RO1GM100329).
Author information
Authors and Affiliations
Contributions
E.A.W., S.C.G. and R.J.C. conducted experiments, analysed and interpreted results and wrote the manuscript. L.M.C., T.J. Sanders., C.J.M., B.R.W., R.L.V., R.F. and B.R. conducted experiments and interpreted results. T.J. Santangelo., B.R.G. and M.P.T. supervised experiments, interpreted results and wrote the manuscript and/or guided its conception and completion.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Microbiology thanks Tobias Warnecke, Jun-Jie Liu and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Steps of integration.
Cartoon diagram shows the steps by which a spacer is integrated into the CRISPR array at the leader-adjacent repeat.
Extended Data Fig. 2 Distributions of aligned DNA sequences from MNase protection assay.
MNase protected DNA (Fig. 1a) was sequenced by HTS, aligned, and genome browser tracks were generated to examine the genome-wide distribution of aligned DNA fragments. Overlapping subtracks are color-coded according to the size of the DNA fragment (red: 30 bp +/- 5; golden yellow: 60 bp +/- 5; green: 90 bp +/- 5; blue 120 bp +/- 5). Positions of CRISPR loci are indicated (leader in pale yellow, repeat array in black). Approximately 10 kb of genome space upstream and downstream from each CRISPR locus is shown. Six replicates (three stationary growth, three exponential) were sequenced and yielded similar patterns; a representative replicate is shown here.
Extended Data Fig. 3 Positions of MNase protected DNA fragments and promoter elements within leaders of the seven CRISPR loci in P. furiosus.
Genome browser tracks were examined to determine the distribution of aligned MNase-protected DNA fragments within CRISPR leaders. Overlapping subtracks are color-coded according to the size of the DNA fragment (red: 30 bp +/- 5; golden yellow: 60 bp +/- 5; green: 90 bp +/- 5; blue 120 bp +/- 5). The x-axis indicates the cumulative depth of read coverage. Six replicates (three stationary growth (St), three exponential (Ex)) are shown. We noted that the 60 bp read coverage (peak height) over L1 and LR was unusually high compared to 60 bp coverage elsewhere in the genome. To quantify this, we analyzed sliding 60 bp windows across the genome, determined the density of 60 bp reads in those sliding windows, then binned and tallied the results to determine the distribution of 60 bp read densities. The values found under the label “60mer peak height” indicate the percentile of the L1 or LR peaks for each replicate; for example, the LR peak in CR1 exponential replicate 1 is higher than 99.92% of all 60 bp windows in the genome. We also noted that L1 and LR peaks usually had more 60mer coverage than 90mer coverage, even though 90mer reads were more abundant elsewhere in the genome. We used the sliding windows to quantify the 60mer to 90mer read density ratios across the genome. The values found under the label “60/90 peak ratio” indicate how biased towards 60mer read covereager the L1 or LR peak is; for example, the LR peak in CR1 exponential replicate 1 is more biased towards 60mer read coverage than 97.68% of all 60 bp windows in the genome.
Extended Data Fig. 4 Nucleotide alignment of seven CRISPR array leaders with MNase protected DNA peaks, promoter elements, and integration sites mapped.
Nucleotide sequences of seven leaders were aligned; conserved nucleotide regions are highlighted in gray. Peaks of MNase protected DNA are highlighted in yellow; the purple star and line at the leader-repeat junction show the position of typical, in vivo spacer integration. Promoter elements in the leader are labeled (BRE and TATA).
Extended Data Fig. 5 Growth, RNA expression, and histone abundance characteristics in wildtype, ∆histone A, and ∆histone B P. furiosus strains used in this study.
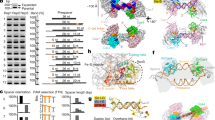
Total, ribo-depleted RNA was sequenced for wildtype, ∆histone A, and ∆histone B strains (five replicates each) and analyzed using the DESeq2 package to identify annotated transcripts with differential expression between (a) wildtype and ∆histone A and (b) wildtype and ∆histone B. Volcano plots show raw p-values against log2-fold changes in abundance for each annotated transcript. A difference in transcript abundance was considered significant if the log2-fold difference was equal or greater than 0.58 and the adjusted p-value was equal or less than 0.1; significantly different points are shown in blue; all other points for transcripts not meeting this threshold are colored gray. (c) Wildtype, ∆histone A, and ∆histone B strains were grown in liquid medium supplemented with pyruvate and Na2S at 95 °C and optical density (OD600nm) was measured every hour as a proxy for cell growth. Blank sample contained medium but no inoculum. Error bars show standard error of the mean for three biological replicates. (d) Western blots, employing polyclonal antibodies raised against HTkA (that recognize HTkA, HTkB, HPfA, and HPfB), were done on total protein from each strain harvested either from exponential (E, 10 hour time point) or stationary (S, 30 hour time point) phase cells. Recombinantly purified HPfA and HPfB were resolved on the gels to provide size references. Six biological replicates were run with similar results; a representative blot image is shown here. Although samples were boiled in 2% SDS before gel loading, histones retained dimer, tetramer, and larger multimeric complexes, giving rise to the multi-band ladder appearance visible on the gel.
Extended Data Fig. 6 Conservation of histones in Euryarchaeota and purification of recombinant proteins.
(a) Clustal Omega Multiple Sequence Alignment of histones from P. furiosus (Pfu A and Pfu B), Thermococcus kodakarensis (Tko A and Tko B), and Methanothermus fervidus (HMf A and HMf B). * = completely conserved residues,: = conservation between groups with strongly similar properties,. = conservation between groups with weakly similar properties. (b) SDS PAGE gels with the purified proteins used in the in vitro integration assays. Cas1 = 37.5 kDa, Cas2 = 10 kDa, P. furiosus histones A = 7.4 kDa and B = 7.3 kDa, T. kodakarensis histones A = 7.3 kDa and B = 7.1 kDa, TrmBL2 = 30.6 kDa.
Extended Data Fig. 7 In vitro evaluation of spacer integration into four CRISPR arrays in the presence or absence of P. furiosus histones.
(a) Gel images show representative results from PCR (carried out as in Fig. 5d) with primers targeting four CRISPR arrays: CRISPR5, CRISPR6, CRISPR7, and CRISPR8. The expected sizes for PCR products resulting from integrations at repeats 1 - 5 are marked with R1 – R5, respectively. Red asterisks identify the band corresponding to integrations at repeat 1, which is the natural, preferred point of integration in vivo. (b) Intensity of all PCR bands was quantified using ImageJ and the proportion of integration events at the five repeats was determined. Single biological replicates were evaluated with two different primer pairs producing with similar outcomes; one PCR reaction is shown and quantitated here.
Extended Data Fig. 8 MNase protection assay to characterize binding patterns for P. furiosus histones incubated with the pCR7-long plasmid.
(a) Gel image of DNA fragments generated by micrococcal nuclease digestion of pCR7-long plasmid incubated with purified recombinant P. furiosus histones A and B. Colored boxes highlight major bands of protected DNA. The gel image is for a digest containing 100 units of MNase; a 500 unit digestion was also done and resulting DNA bands were much fainter. (b) DNA bands highlighted in A were sequenced and genome browser tracks were generated. Overlapping subtracks are color-coded according to the size of the DNA fragment (red: 30 bp +/- 5; golden yellow: 60 bp +/- 5; green: 90 bp +/- 5; blue 120 bp +/- 5). The full CRISPR7 and part of one repeat are shown, with the purple line indicating the position where integration would typically occur in vivo. The x-axis indicates the cumulative depth of read coverage. One replicate was done for each treatment (100 and 500 units MNase).
Extended Data Fig. 9 In vitro evaluation of spacer integration into pCR7-long in the presence or absence of DNA-binding proteins.
(a) Diagrammatric representation of the PCR used to assess in vitro integration into pCR7-long. (b) Gel images show representative results from the PCR in (a) when recombinant purified histones, Alba, or Trmbl2 were added to the reaction at the indicated concentrations. The expected sizes for PCR products resulting from integrations at repeats 1 - 4 are marked with R1 - R4, respectively. Red asterisks identify the band corresponding to integrations at repeat 1, which is the natural, preferred point of integration in vivo. (c) Intensity of all PCR bands was quantified using ImageJ and the proportion of integration events at the five repeats was determined. Three replicates were assessed with similar outcomes, representative gels and quantifications are shown here.
Extended Data Fig. 10 Unbiased HTS sequencing of in vitro integration of a spacer into pCR7-long.
The in vitro integration assay with and without histones (a) or alba (b) was carried out, as before, and integration events were then sequenced using a two-step, semi-degenerate PCR protocol which targets all DNA fragments bearing the spacer. Sequencing results were used to generate genome browser tracks; the amplitude of the black peaks in these tracks indicate the total number of reads supporting an integration event at that nucleotide position. The positions of the leader (pale yellow) and repeats (black) are shown. Assay products with histones were sequenced for six biological replicates yielding very similar patterns; with alba one biological replicate was sequenced.
Supplementary information
Source data
Source Data Fig. 1
Uncropped agarose gel images and MNase digested DNA.
Source Data Fig. 3
Uncropped agarose gel images and PCR products.
Source Data Fig. 4
Uncropped agarose gel images and PCR products.
Source Data Fig. 5
Uncropped agarose gel images and PCR products.
Source Data Extended Data Fig. 5
Uncropped gel image and western blot.
Source Data Extended Data Fig. 6
Uncropped gel images and western blot.
Source Data Extended Data Fig. 7
Uncropped agarose gel images and PCR products.
Source Data Extended Data Fig. 8
Uncropped agarose gel images and MNase digested DNA.
Source Data Extended Data Fig. 9
Uncropped agarose gel images and PCR products.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Watts, E.A., Garrett, S.C., Catchpole, R.J. et al. Histones direct site-specific CRISPR spacer acquisition in model archaeon. Nat Microbiol 8, 1682–1694 (2023). https://doi.org/10.1038/s41564-023-01446-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-023-01446-3