Abstract
Pharmaceuticals have extensive reciprocal interactions with the microbiome, but whether bacterial drug sensitivity and metabolism is driven by pathways conserved in host cells remains unclear. Here we show that anti-cancer fluoropyrimidine drugs inhibit the growth of gut bacterial strains from 6 phyla. In both Escherichia coli and mammalian cells, fluoropyrimidines disrupt pyrimidine metabolism. Proteobacteria and Firmicutes metabolized 5-fluorouracil to its inactive metabolite dihydrofluorouracil, mimicking the major host mechanism for drug clearance. The preTA operon was necessary and sufficient for 5-fluorouracil inactivation by E. coli, exhibited high catalytic efficiency for the reductive reaction, decreased the bioavailability and efficacy of oral fluoropyrimidine treatment in mice and was prevalent in the gut microbiomes of colorectal cancer patients. The conservation of both the targets and enzymes for metabolism of therapeutics across domains highlights the need to distinguish the relative contributions of human and microbial cells to drug efficacy and side-effect profiles.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The raw data for our mouse experiments as well as uncropped gel images and tumour photographs can be found in the source data section. The Genome Taxonomy Database95, Kyoto Encyclopedia of Genes and Genomes (KEGG) database96 and MIDAS v1.0 database65 are publicly available. Sequencing data have been deposited under the NCBI BioProjects PRJNA576932 (RNA-seq, 16S rRNA gene sequences and isolate genomes) and PRJNA720145 (GO Study metagenomic data). Source data are provided with this paper.
Code availability
Source code for our analyses of bacterial preTA can be found at https://bitbucket.org/pbradz/preta/.
References
Thorn, C. F., Klein, T. E. & Altman, R. B. PharmGKB: the pharmacogenomics knowledge base. Methods Mol. Biol. 1015, 311–320 (2013).
Spanogiannopoulos, P., Bess, E. N., Carmody, R. N. & Turnbaugh, P. J. The microbial pharmacists within us: a metagenomic view of xenobiotic metabolism. Nat. Rev. Microbiol. 14, 273–287 (2016).
Lam, K. N., Alexander, M. & Turnbaugh, P. J. Precision medicine goes microscopic: engineering the microbiome to improve drug outcomes. Cell Host Microbe 26, 22–34 (2019).
Zimmermann, M., Zimmermann-Kogadeeva, M., Wegmann, R. & Goodman, A. L. Mapping human microbiome drug metabolism by gut bacteria and their genes. Nature https://doi.org/10.1038/s41586-019-1291-3 (2019).
Javdan, B. et al. Personalized mapping of drug metabolism by the human gut microbiome. Cell 181, 1661–1679.e22 (2020).
Wallace, B. D. et al. Alleviating cancer drug toxicity by inhibiting a bacterial enzyme. Science 330, 831–835 (2010).
Biernat, K. A. et al. Structure, function, and inhibition of drug reactivating human gut microbial β-glucuronidases. Sci. Rep. 9, 825 (2019).
Iida, N. et al. Commensal bacteria control cancer response to therapy by modulating the tumor microenvironment. Science 342, 967–970 (2013).
Sivan, A. et al. Commensal Bifidobacterium promotes antitumor immunity and facilitates anti-PD-L1 efficacy. Science 350, 1084–1089 (2015).
Vétizou, M. et al. Anticancer immunotherapy by CTLA-4 blockade relies on the gut microbiota. Science 350, 1079–1084 (2015).
Viaud, S. et al. The intestinal microbiota modulates the anticancer immune effects of cyclophosphamide. Science 342, 971–976 (2013).
Haiser, H. J. et al. Predicting and manipulating cardiac drug inactivation by the human gut bacterium Eggerthella lenta. Science 341, 295–298 (2013).
Maurice, C. F., Haiser, H. J. & Turnbaugh, P. J. Xenobiotics shape the physiology and gene expression of the active human gut microbiome. Cell 152, 39–50 (2013).
Maini Rekdal, V., Bess, E. N., Bisanz, J. E., Turnbaugh, P. J. & Balskus, E. P. Discovery and inhibition of an interspecies gut bacterial pathway for Levodopa metabolism. Science 364, eaau6323 (2019).
Nayak, R. R. et al. Methotrexate impacts conserved pathways in diverse human gut bacteria leading to decreased host immune activation. Cell Host Microbe https://doi.org/10.1016/j.chom.2020.12.008 (2021).
Artacho, A. et al. The pretreatment gut microbiome is associated with lack of response to methotrexate in new-onset rheumatoid arthritis. Arthritis Rheumatol. 73, 931–942 (2021).
Bisanz, J. E., Spanogiannopoulos, P., Pieper, L. M., Bustion, A. E. & Turnbaugh, P. J. How to determine the role of the microbiome in drug disposition. Drug Metab. Dispos. 46, 1588–1595 (2018).
Zimmermann, M., Zimmermann-Kogadeeva, M., Wegmann, R. & Goodman, A. L. Separating host and microbiome contributions to drug pharmacokinetics and toxicity. Science 363, eaat9931 (2019).
Arruebo, M. et al. Assessment of the evolution of cancer treatment therapies. Cancers 3, 3279–3330 (2011).
Gadiko, C. et al. Comparative bioavailability study of capecitabine tablets of 500 mg in metastatic breast cancer and colorectal cancer patients under fed condition. Clin. Res. Regul. Aff. 29, 72–76 (2012).
Haller, D. G. et al. Potential regional differences for the tolerability profiles of fluoropyrimidines. J. Clin. Oncol. 26, 2118–2123 (2008).
Jennings, B. A. et al. Evaluating predictive pharmacogenetic signatures of adverse events in colorectal cancer patients treated with fluoropyrimidines. PLoS ONE 8, e78053 (2013).
Saif, M. W., Syrigos, K., Mehra, R., Mattison, L. K. & Diasio, R. B. Dihydropyrimidine dehydrogenase deficiency (DPD) in GI malignancies: experience of 4-years. Pak. J. Med. Sci. Q. 23, 832–839 (2007).
Leonard, R., Hennessy, B. T., Blum, J. L. & O’Shaughnessy, J. Dose-adjusting capecitabine minimizes adverse effects while maintaining efficacy: a retrospective review of capecitabine for metastatic breast cancer. Clin. Breast Cancer 11, 349–356 (2011).
Horowitz, J., Saukkonen, J. J. & Chargaff, E. Effects of fluoropyrimidines on the synthesis of bacterial proteins and nucleic acids. J. Biol. Chem. 235, 3266–3272 (1960).
Bloch, A. & Hutchison, D. J. A mechanism of resistance to fluoropyrimidines. Cancer Res. 24, 433–439 (1964).
Stringer, A. M. et al. Gastrointestinal microflora and mucins may play a critical role in the development of 5-Fluorouracil-induced gastrointestinal mucositis. Exp. Biol. Med. 234, 430–441 (2009).
Von Bültzingslöwen, I., Adlerberth, I., Wold, A. E., Dahlén, G. & Jontell, M. Oral and intestinal microflora in 5-fluorouracil treated rats, translocation to cervical and mesenteric lymph nodes and effects of probiotic bacteria. Oral. Microbiol. Immunol. 18, 278–284 (2003).
Stringer, A. M. et al. Chemotherapy-induced diarrhea is associated with changes in the luminal environment in the DA rat. Exp. Biol. Med. 232, 96–106 (2007).
Zwielehner, J. et al. Changes in human fecal microbiota due to chemotherapy analyzed by TaqMan-PCR, 454 sequencing and PCR-DGGE fingerprinting. PLoS ONE 6, e28654 (2011).
van Vliet, M. J. et al. Chemotherapy treatment in pediatric patients with acute myeloid leukemia receiving antimicrobial prophylaxis leads to a relative increase of colonization with potentially pathogenic bacteria in the gut. Clin. Infect. Dis. 49, 262–270 (2009).
Stringer, A. M. et al. Biomarkers of chemotherapy-induced diarrhoea: a clinical study of intestinal microbiome alterations, inflammation and circulating matrix metalloproteinases. Support. Care Cancer 21, 1843–1852 (2013).
Montassier, E. et al. Chemotherapy-driven dysbiosis in the intestinal microbiome. Aliment. Pharmacol. Ther. 42, 515–528 (2015).
Scott, T. A. et al. Host-microbe co-metabolism dictates cancer drug efficacy in C. elegans. Cell 169, 442–456.e18 (2017).
García-González, A. P. et al. Bacterial metabolism affects the C. elegans response to cancer chemotherapeutics. Cell 169, 431–441.e8 (2017).
Rosener, B. et al. Evolved bacterial resistance against fluoropyrimidines can lower chemotherapy impact in the Caenorhabditis elegans host. eLife 9, e59831 (2020).
Grothey, A. et al. Duration of adjuvant chemotherapy for stage III colon cancer. N. Engl. J. Med. 378, 1177–1188 (2018).
Walko, C. M. & Lindley, C. Capecitabine: a review. Clin. Ther. 27, 23–44 (2005).
Zhang, L., Zhang, Y. D., Strong, J. M., Reynolds, K. S. & Huang, S.-M. A regulatory viewpoint on transporter-based drug interactions. Xenobiotica 38, 709–724 (2008).
Longley, D. B., Harkin, D. P. & Johnston, P. G. 5-fluorouracil: mechanisms of action and clinical strategies. Nat. Rev. Cancer 3, 330–338 (2003).
Islam, Z. et al. Bacterial versus human thymidylate synthase: kinetics and functionality. PLoS ONE 13, e0196506 (2018).
Pinedo, H. M. & Peters, G. F. Fluorouracil: biochemistry and pharmacology. J. Clin. Oncol. 6, 1653–1664 (1988).
O’Donovan, G. A. & Neuhard, J. Pyrimidine metabolism in microorganisms. Bacteriol. Rev. 34, 278–343 (1970).
Lewis, K. Platforms for antibiotic discovery. Nat. Rev. Drug Discov. 12, 371–387 (2013).
Spanogiannopoulos, P., Waglechner, N., Koteva, K. & Wright, G. D. A rifamycin inactivating phosphotransferase family shared by environmental and pathogenic bacteria. Proc. Natl Acad. Sci. USA 111, 7102–7107 (2014).
Lunenburg, C. A. T. C. et al. Prospective DPYD genotyping to reduce the risk of fluoropyrimidine-induced severe toxicity: ready for prime time. Eur. J. Cancer 54, 40–48 (2016).
Wei, X., McLeod, H. L., McMurrough, J., Gonzalez, F. J. & Fernandez-Salguero, P. Molecular basis of the human dihydropyrimidine dehydrogenase deficiency and 5-fluorouracil toxicity. J. Clin. Invest. 98, 610–615 (1996).
Vreken, P., Van Kuilenburg, A. B., Meinsma, R. & van Gennip, A. H. Dihydropyrimidine dehydrogenase (DPD) deficiency: identification and expression of missense mutations C29R, R886H and R235W. Hum. Genet. 101, 333–338 (1997).
Hidese, R., Mihara, H., Kurihara, T. & Esaki, N. Escherichia coli dihydropyrimidine dehydrogenase is a novel NAD-dependent heterotetramer essential for the production of 5,6-dihydrouracil. J. Bacteriol. 193, 989–993 (2011).
Smith, A. E. & Yamada, E. W. Dihydrouracil dehydrogenase of rat liver. J. Biol. Chem. 246, 3610–3617 (1971).
Podschun, B., Jahnke, K., Schnackerz, K. D. & Cook, P. F. Acid base catalytic mechanism of the dihydropyrimidine dehydrogenase from pH studies. J. Biol. Chem. 268, 3407–3413 (1993).
Porter, D. J. Dehalogenating and NADPH-modifying activities of dihydropyrimidine dehydrogenase. J. Biol. Chem. 269, 24177–24182 (1994).
Porter, D. J., Harrington, J. A., Almond, M. R., Lowen, G. T. & Spector, T. (R)-5-fluoro-5,6-dihydrouracil: kinetics of oxidation by dihydropyrimidine dehydrogenase and hydrolysis by dihydropyrimidine aminohydrolase. Biochem. Pharmacol. 48, 775–779 (1994).
Beaupre, B. A., Forouzesh, D. C. & Moran, G. R. Transient-state analysis of porcine dihydropyrimidine dehydrogenase reveals reductive activation by NADPH. Biochemistry 59, 2419–2431 (2020).
Lam, K. N. et al. Phage-delivered CRISPR-Cas9 for strain-specific depletion and genomic deletions in the gut microbiome. Cell Rep. 37, 109930 (2021).
Barba, M., Dutoit, R., Legrain, C. & Labedan, B. Identifying reaction modules in metabolic pathways: bioinformatic deduction and experimental validation of a new putative route in purine catabolism. BMC Syst. Biol. 7, 99 (2013).
Bradley, P. H., Nayfach, S. & Pollard, K. S. Phylogeny-corrected identification of microbial gene families relevant to human gut colonization. PLoS Comput. Biol. 14, e1006242 (2018).
Nayfach, S., Shi, Z. J., Seshadri, R., Pollard, K. S. & Kyrpides, N. C. New insights from uncultivated genomes of the global human gut microbiome. Nature 568, 505–510 (2019).
Zeller, G. et al. Potential of fecal microbiota for early-stage detection of colorectal cancer. Mol. Syst. Biol. 10, 766 (2014).
Ai, D. et al. Identifying gut microbiota associated with colorectal cancer using a zero-inflated lognormal model. Front. Microbiol. 10, 826 (2019).
Geller, L. T. et al. Potential role of intratumor bacteria in mediating tumor resistance to the chemotherapeutic drug gemcitabine. Science 357, 1156–1160 (2017).
Maier, L. et al. Extensive impact of non-antibiotic drugs on human gut bacteria. Nature 555, 623–628 (2018).
Guo, C.-J. et al. Depletion of microbiome-derived molecules in the host using Clostridium genetics. Science 366, eaav1282 (2019).
Nayak, R. R. & Turnbaugh, P. J. Mirror, mirror on the wall: which microbiomes will help heal them all? BMC Med. 14, 72 (2016).
Nayfach, S., Rodriguez-Mueller, B., Garud, N. & Pollard, K. S. An integrated metagenomics pipeline for strain profiling reveals novel patterns of bacterial transmission and biogeography. Genome Res. 26, 1612–1625 (2016).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Anders, S., Pyl, P. T. & Huber, W. HTSeq–a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol. 11, R106 (2010).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Seemann, T. snippy. (GitHub) (2020).
Titz, B., Häuser, R., Engelbrecher, A. & Uetz, P. The Escherichia coli protein YjjG is a house-cleaning nucleotidase in vivo. FEMS Microbiol. Lett. 270, 49–57 (2007).
Sharan, S. K., Thomason, L. C., Kuznetsov, S. G. & Court, D. L. Recombineering: a homologous recombination-based method of genetic engineering. Nat. Protoc. 4, 206–223 (2009).
Jensen, S. I., Lennen, R. M., Herrgård, M. J. & Nielsen, A. T. Seven gene deletions in seven days: Fast generation of Escherichia coli strains tolerant to acetate and osmotic stress. Sci. Rep. 5, 17874 (2015).
Baba, T. et al. Construction of Escherichia coli K-12 in-frame, single-gene knockout mutants: the Keio collection. Mol. Syst. Biol. 2, 2006.0008 (2006).
Datsenko, K. A. & Wanner, B. L. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc. Natl Acad. Sci. USA 97, 6640–6645 (2000).
Cox, G. et al. A common platform for antibiotic dereplication and adjuvant discovery. Cell Chem. Biol. 24, 98–109 (2017).
Funatsu, G. & Wittmann, H. G. Ribosomal proteins. 33. Location of amino-acid replacements in protein S12 isolated from Escherichia coli mutants resistant to streptomycin. J. Mol. Biol. 68, 547–550 (1972).
Cicchillo, R. M. et al. Lipoyl synthase requires two equivalents ofS-Adenosyl-l-methionine to synthesize one equivalent of lipoic acid. Biochemistry 43, 6378–6386 (2004).
Beaupre, B. A., Roman, J. V. & Moran, G. R. An improved method for the expression and purification of porcine dihydropyrimidine dehydrogenase. Protein Expr. Purif. 171, 105610 (2020).
Myhal, M. L., Laux, D. C. & Cohen, P. S. Relative colonizing abilities of human fecal and K 12 strains of Escherichia coli in the large intestines of streptomycin-treated mice. Eur. J. Clin. Microbiol. 1, 186–192 (1982).
Measuring treatment response in Patient Derived Xenograft (PDX) models at The Jackson Laboratory. The Jackson Laboratory http://tumor.informatics.jax.org/mtbwi/live/www/html/SOCHelp.html (2017).
Gohl, D. M. et al. Systematic improvement of amplicon marker gene methods for increased accuracy in microbiome studies. Nat. Biotechnol. 34, 942–949 (2016).
McMurdie, P. J. & Holmes, S. phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLoS ONE 8, e61217 (2013).
Haft, D. H. et al. RefSeq: an update on prokaryotic genome annotation and curation. Nucleic Acids Res. 46, D851–D860 (2018).
Notredame, C., Higgins, D. G. & Heringa, J. T-Coffee: a novel method for fast and accurate multiple sequence alignment. J. Mol. Biol. 302, 205–217 (2000).
Madeira, F. et al. The EMBL-EBI search and sequence analysis tools APIs in 2019. Nucleic Acids Res. 47, W636–W641 (2019).
Capella-Gutiérrez, S., Silla-Martínez, J. M. & Gabaldón, T. trimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 25, 1972–1973 (2009).
Finn, R. D., Clements, J. & Eddy, S. R. HMMER web server: interactive sequence similarity searching. Nucleic Acids Res. 39, W29–W37 (2011).
Wu, D., Jospin, G. & Eisen, J. A. Systematic identification of gene families for use as ‘Markers’ for phylogenetic and phylogeny-driven ecological studies of bacteria and archaea and their major subgroups. PLoS ONE 8, e77033 (2013).
Nguyen, L.-T., Schmidt, H. A., von Haeseler, A. & Minh, B. Q. IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 32, 268–274 (2015).
Sievers, F. et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 7, 539 (2011).
Madden, T. The BLAST Sequence Analysis Tool (National Center for Biotechnology Information (US), 2003).
Rognes, T., Flouri, T., Nichols, B., Quince, C. & Mahé, F. VSEARCH: a versatile open source tool for metagenomics. PeerJ 4, e2584 (2016).
Nayfach, S., Fischbach, M. A. & Pollard, K. S. MetaQuery: a web server for rapid annotation and quantitative analysis of specific genes in the human gut microbiome. Bioinformatics 31, 3368–3370 (2015).
Parks, D. H. et al. A complete domain-to-species taxonomy for Bacteria and Archaea. Nat. Biotechnol. 38, 1079–1086 (2020).
Minoru, K. et al. KEGG: integrating viruses and cellular organisms. Nucleic Acids Res. 49, D545–D551 (2021).
Guindon, S., Delsuc, F., Dufayard, J.-F. & Gascuel, O. Estimating maximum likelihood phylogenies with PhyML. Methods Mol. Biol. 537, 113–137 (2009).
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Yu, G., Wang, L.-G., Han, Y. & He, Q.-Y. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16, 284–287 (2012).
Waterhouse, A. et al. SWISS-MODEL: homology modelling of protein structures and complexes. Nucleic Acids Res. 46, W296–W303 (2018).
Acknowledgements
We thank the other members of the Gerona, Goga, Pollard and Turnbaugh labs for their helpful suggestions during the preparation of this manuscript; B. Yu, M. Tan and R. Sit from the Chan Zuckerberg Biohub for assistance with DNA sequencing; G. Wright and L. Ejim (McMaster University) for providing pINT1; J. Bisanz for advice with computational methods; K. Spitler in the UCSF Quantitative Metabolite Analysis Center for help with analytical methods; and the GO Study clinical research coordinators (D. Stanfield, P. Steiding and J. Whitman). This work was supported by the National Institutes of Health (R01HL122593; R21CA227232; R01CA223817), the Searle Scholars Program (SSP-2016-1352) and CDMRP W81XWH-18-1-0713. P.J.T. is a Chan Zuckerberg Biohub Investigator and was a Nadia’s Gift Foundation Innovator who was supported in part by the Damon Runyon Cancer Research Foundation (DRR-42-16). A.G. was supported in part by a MARK Foundation Endeavor Award. Fellowship support was provided by the Canadian Institutes of Health Research (P.S. and K.N.L.) and the NIH (J.V.L. - F32CA243548, T32CA108462; T.S.K. - F30CA257378). B.G.H.G. is a Connie and Bob Lurie Fellow of the Damon Runyon Cancer Research Foundation (DRG-2450-21).
Author information
Authors and Affiliations
Contributions
P.J.T. conceived the project and was the primary supervisor for the study. R.R.G., A.G. and K.S.P. also supervised components of this work. P.S. led the in vitro screens and E. coli strain construction and established protocols for the pharmacokinetics and xenograft experiments. T.S.K. led the final pharmacokinetics, xenograft, transcriptomics and amplicon sequencing data generation and analysis. B.G.H.G. led the biochemical characterization of PreTA. P.H.B. led the bioinformatic analysis of preTA operons across genomes and microbiomes. J.M., Y.N.A.M., T.S.K., B.G.H.G. and M.S. performed mass spectrometry. K.N.L. sequenced and analysed the drug-resistant E. coli mutants. J.V.L. assisted with the tumour xenograft measurements. C.E.A., A.V. and W.K. (GO Study PI) oversaw the conception and design of the GO Study and contributed patient samples. E.L.V.B. contributed to developing the study protocol and supervision of data collection for the GO Study. D.G. designed the GO Study specimen collection kits and managed biospecimen collection, storage and retrieval. P.S. wrote the initial draft. T.S.K. and P.J.T. revised the manuscript with input from all authors.
Corresponding author
Ethics declarations
Competing interests
P.J.T. is on the scientific advisory boards of Pendulum, Seed and SNIPRbiome; there is no direct overlap between the current study and these consulting duties. K.S.P. is on the scientific advisory board of Phylagen; there is no direct overlap between the current study and these consulting duties. C.E.A serves on the scientific advisory board of Pionyr Immunotherapeutics and has received research funding (institution) from Bristol Meyer Squibb, Guardant Health, Kura Oncology, Merck and Novartis; there is no direct overlap with the current study. All other authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Bacterial taxa vary in their sensitivity to fluoropyrimidines despite the rapid emergence of resistance during in vitro growth.
(a) Simplified metabolic pathway for the bioactivation of the oral prodrug capecitabine (CAP) to 5-fluorouracil (5-FU), 5-fluorodeoxyuridine (FUDR), and downstream metabolites1. Red indicates chemical groups hydrolyzed during the conversion of CAP to 5-FU. Sequential reactions are indicated by multiple arrows. (b,c) Variation in (b) 5-FU and (c) FUDR minimal inhibitory concentration (MIC) at the phylum level. Bacterial strains where no MIC could be determined were set to the maximum tested concentration. Each dot represents a bacterial isolate. Red lines indicate the median. p-values, Kruskal-Wallis test with Dunn’s correction for multiple comparisons. (d) 5-FU MICs of parent and 5-FU-resistant strains of E. coli MG1655, B. fragilis DSM2151, and B. ovatus DSM1896 (see Supplementary Table 2). The type of selection is indicated by the strain identifier: A = agar; L = liquid. (e) Wild-type E. coli BW25113 was assayed for conversion of CAP to 5-FU by LC-MS/MS (n = 3 biological replicates per group). Open circles represent individual values, filled circles represent mean values. p-values, two-tailed paired t test comparing final vs. baseline values for each analyte.
Extended Data Fig. 2 Multiple fluoropyrimidines disrupt the pyrimidine metabolism pathway in E. coli.
(a) A working model for uracil and uridine import and metabolism in bacterial and mammalian cells. Underlined genes are essential for E. coli growth and metabolites with red circles indicate putative 5-FU metabolites. (b-d) Uracil rescues the growth of E. coli BW25113 in the presence of (b) 5-FU, (c) FUDR, and (d) CAP in a dose-dependent manner. M9MM plus glucose was used as the base media with added uracil from 0.1–10 µg/ml. (e-g) A loss-of-function mutation in uracil phosphoribosyltransferase gene (Δupp) rescues the growth of E. coli BW25113 in the presence of (e) 5-FU, (f) FUDR, and (g) CAP. MIC assays performed in M9MM plus glucose. Values in panels b-g are mean±stdev (n = 4 biological replicates per group).
Extended Data Fig. 3 The preTA operon is necessary and sufficient for the inactivation of 5-fluorouracil in an independent E. coli background and has a modest impact on growth.
(a) Wild-type (preTA+), deletion (ΔpreTA), complemented (preTA++), and empty vector E. coli BW25113 strains were assayed for residual 5-FU using disk diffusion (0–48 hours incubation) and LC-QTOF/MS (48 hours). Complementation and empty vector were on the ΔpreTA background. (b) Wild-type, single gene deletion (ΔpreT, ΔpreA) and operon deletion (ΔpreTA) strains of E. coli BW25113 were assayed for conversion of 5-FU to DHFU by LC-MS/MS. Open circles represent individual values, filled circles represent means [n = 3; *p-value<0.05; 5-FU: 52 hr, p = 0.041; DHFU: 30 hr, p = 0.011; 45 hr, p = 0.004; 52 hr, p = 0.009, 2-way ANOVA with Tukey’s correction relative to baseline of the same analyte]. (c) 5-FU MIC determination of the E. coli strains shown in panel a and Fig. 1d in minimal (M9MM) media. Values are normalized to the growth control (no 5-FU) with darker colors indicating growth inhibition. Sterile media is shown in the final row.
Extended Data Fig. 4 Purification and biochemical characterization of E. coli PreTA.
(a) Purification workflow for E. coli PreTA including immobilized metal affinity chromatography (M: marker, I: insoluble fraction, S: soluble fraction, FT: flowthrough, W: wash, numbers across the top indicate imidazole concentration), TEV cleavage (M: marker, L: loaded, FT: flowthrough, W: wash, E: elute), size exclusion chromatography (M: marker, L: loaded, lanes are labeled by fraction from a 96 well plate collected via FPLC), and by UV–visible absorption spectroscopy where normalizing protein levels showed a ratio of greater than 0.35 for A280/A377 indicated holoenzyme. In all gels, numbers down the side indicate protein molecular weight in kDa and solid lines indicate lanes that were carried forward in the preparation. (b-c) Analytical size exclusion chromatography (SEC) traces (b) and analysis (c), characterizing the main peak as a heterotetramer. Grey shaded region indicates 95% confidence intervals of the linear model. (d) High pressure liquid chromatography (HPLC) of NADH, uracil (Ura), dihydrouracil (DHU), NAD+, 5-fluorouracil (5-FU), and dihydrofluorouracil (DHFU). (e) Liquid Chromatography Mass Spectrometry (LC-MS/MS) of enzymatic reaction with 5-FU confirms the presence of the exact mass of DHFU.
Extended Data Fig. 5 PreTA decreases the efficacy and oral bioavailability of capecitabine (CAP).
(a, g) Xenograft experiment (expt) 1 and 2 design, respectively. (b, h) Pre-treatment tumor volumes: (b) expt1, (h) expt2 (n = 8 mice ΔpreTA-CAP; n = 10 mice preTA++-Veh; n = 9 mice/group remainder expt1; n = 8 mice/group expt2; lines represent medians; 2-way ANOVA with Tukey’s correction). (c, i) Colonization levels of streptomycin-resistant (SmR) E. coli in colony-forming units (CFU)/gram stool in (c) expt1 and (i) expt2 (n = 6 mice/group expt1; n = 8 mice/group expt2; opaque dots and lines represent mean±SEM; 2-way ANOVA with Tukey’s correction). Zero values at baseline replaced with our limit of detection (1000 CFU/g). (d, j) Percentage of starting tumor volumes over time in (d) expt1 and (j) expt2 (n = 8 mice ΔpreTA-CAP group; n = 10 mice preTA++-Veh; n = 9 mice/group remainder expt1; n = 8 mice/group expt2; opaque dots and lines represent mean±SEM; 2-way ANOVA with Tukey’s correction did not reach significance). (e, k) Percentage of starting tumor volumes on (e) day 15 expt1 and (k) day 22 expt2 (n = 8 mice ΔpreTA-CAP group; n = 10 mice preTA++-Veh; n = 9 mice/group remainder expt1; n = 8 mice/group expt2; timepoints selected to capture all mice prior to euthanasia; lines represent medians; 2-way ANOVA with Tukey’s correction). (f, l) Percentage of mice reaching the humane endpoint in (f) expt1 and (l) expt2 (expt1: n = 8 mice ΔpreTA-CAP group; n = 10 mice preTA++-Veh; n = 9 mice/group remainder; expt2: n = 3 preTA++ groups, n = 4 ΔpreTA groups; log-rank Mantel-Cox test comparing ΔpreTA-CAP to all other groups. Eight mice were censored (black boxes) as they did not reach the endpoint in expt1. (m) LC-QTOF/MS quantification of 5-FU from pooled plasma samples following 500 mg/kg CAP administration in mice (same design as Fig. 4g; n = 5 mice/group pooled into n = 1 sample/group).
Extended Data Fig. 6 Consistent shifts in the gut microbiota across groups.
(a) Quantification of streptomycin-resistant (SmR) E. coli in feces across time (n = 11 mice for for preTA++-Veh group and n = 10 mice/group for the rest; 2-way ANOVA test with Tukey’s correction; lines and ribbons represent mean±SEM). Zero values at baseline replaced with our limit of detection (1000 CFU/g). (b) Principal coordinate analysis of fecal microbiota from mice treated with CAP or vehicle (Veh) and colonized with ΔpreTA or preTA++ E. coli across time (Bray-Curtis distance matrix, permutational multivariate analysis of variance test with Benjamini-Hochberg correction using ADONIS statistical package). (c) Microbial community composition at the phylum level. Each bar represents stool from each mouse. Short horizontal lines within bars represent different amplicon sequence variants (ASV). (d) Number of ASVs over time for each treatment group (exact n values indicated in panel c; 2-way ANOVA with Tukey’s correction; lines and ribbons represent mean±SEM). (e, f) Proportion of (e) Bacteroidota and (f) E. coli over time for each treatment group (exact n values indicated in panel c; 2-way ANOVA with Tukey’s correction; lines and ribbons represent mean±SEM). (g) Volcano plots of differentially abundant sequence variants at day 17 (FDR < 0.1, |log2 fold-change|>1, Wald test with Benjamini-Hochberg correction, significance limits are marked with dash lines). Blue and red texts represent enriched and depleted taxa, respectively.
Extended Data Fig. 7 Distribution of preTA operons across gut metagenome-assembled genomes (MAGs).
A phylogenetic tree of species from IGGdb, made using a concatenated alignment of single-copy marker genes, is shown in black. Species in which a MAG contains a putative preTA operon are identified with colored circles, where the color of the circle corresponds to prevalence in the gut (blue: low prevalence; orange: high prevalence). All species displayed were detected at least once from 3,810 gut metagenome samples. Phylum-level annotations are shown as colored ring segments surrounding the tree for the ten phyla with the most gut MAGs. Data can be found in Supplementary Table 9.
Extended Data Fig. 8 Bacterial preTA operons from diverse strains, preTA encoding natural strains, and pooled isolates from patient samples are capable of 5-FU inactivation.
(a) preTA operons from Salmonella enterica LT2 (DSM17058), Oxalobacter formigenes ATCC35274, and Lactobacillus reuteri DSM20016 were amplified and integrated into the chromosome of E. coli MG1655 ∆preTA. Strains were incubated with 20 μg/ml 5-FU and assayed for residual drug using disk diffusion (0–48 hours incubation) and LC-QTOF/MS (48 hours). E. coli MG1655 preTA complementation and empty vector controls are also included on the ΔpreTA background. (b) We also tested strains predicted to encode the preTA operon: Salmonella enterica LT2, Anaerostipes caccae DSM14662, and Clostridium sporogenes DSM795. Residual 5-FU was assayed as in panel a. Of note, we did not detect either compound from A. caccae, suggesting that this strain may further metabolize DHFU. (c) Bacterial isolates from 22 CRC patient stool samples in the GO Study (Supplementary Table 12) were isolated on McConkey agar, pooled, and incubated in BHI+ with 5-FU for 4 days. Residual 5-FU levels were assessed using a disk diffusion assay. Colors indicate the treatment cohort: (A) CAP (yellow); (B) TAS-102 (blue); and (C) CAP plus immunotherapy (green). Red boxes in panels a-c indicate the first timepoint without a clear zone of inhibition.
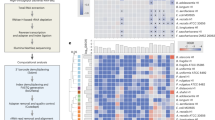
Extended Data Fig. 9 preTA sources and abundance in CRC patients during fluoropyrimidine treatment.
(a) Fraction of total preTA reads mapping to different species in metagenomic samples from GO Study patients undergoing CAP treatment with or without pembrolizumab immunotherapy (IO). Heatmap values were linearly interpolated between samples (filled circles in panel b). (b) Total abundance, as log10 RPKG, of the preTA operon in the gut microbiome prior to and during treatment with the oral fluoropyrimidine CAP with or without IO, as shown in Fig. 6d. Lines connect measurements (filled circles) for the same patient. One zero RPKG value was replaced with half the minimum non-zero value prior to taking the logarithm. The first day of treatment is defined as day 1. Days are the same between panels.
Supplementary information
Supplementary Table 1
Supplementary Table 1.
Source data
Source Data Fig. 4
Statistical source data.
Source Data Fig. 4
Unprocessed tumour images.
Source Data Extended Data Fig. 4
Unprocessed protein blots
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Spanogiannopoulos, P., Kyaw, T.S., Guthrie, B.G.H. et al. Host and gut bacteria share metabolic pathways for anti-cancer drug metabolism. Nat Microbiol 7, 1605–1620 (2022). https://doi.org/10.1038/s41564-022-01226-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-022-01226-5
This article is cited by
-
High-throughput transcriptomics of 409 bacteria–drug pairs reveals drivers of gut microbiota perturbation
Nature Microbiology (2024)
-
Gut bugs disrupt cancer drugs
Nature Reviews Microbiology (2022)