Abstract
The food-borne bacterial pathogen Salmonella Typhimurium uses a type III protein secretion system to deliver multiple proteins into host cells. These secreted effectors modulate the functions of host cells and activate specific signalling cascades that result in the production of pro-inflammatory cytokines and intestinal inflammation. Some of the Salmonella-encoded effectors counteract this inflammatory response and help to preserve host homeostasis. Here, we demonstrate that the Salmonella effector protein SopD, which is required for pathogenesis, functions to both activate and inhibit the inflammatory response by targeting the Rab8 GTPase, which is a negative regulator of inflammation. We show that SopD has GTPase-activating protein activity for Rab8 and, therefore, inhibits this GTPase and stimulates inflammation. We also show that SopD activates Rab8 by displacing it from its cognate guanosine dissociation inhibitor, resulting in the stimulation of a signalling cascade that suppresses inflammation. We solved the crystal structure of SopD in association with Rab8 to a resolution of 2.3 Å, which reveals a unique contact interface that underlies these complex interactions. These findings show the remarkable evolution of a bacterial effector protein to exert both agonistic and antagonistic activities towards the same host cellular target to modulate the inflammatory response.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The atomic coordinates and structure factors generated in this study have been deposited in the PDB under the accession code 7BWT. Source data are provided with this paper.
References
Chen, L. M., Hobbie, S. & Galan, J. E. Requirement of CDC42 for Salmonella-induced cytoskeletal and nuclear responses. Science 274, 2115–2118 (1996).
Bruno, V. M. et al. Salmonella Typhimurium type III secretion effectors stimulate innate immune responses in cultured epithelial cells. PLoS Pathog. 5, e1000538 (2009).
Patel, J. C. & Galan, J. E. Differential activation and function of Rho GTPases during Salmonella-host cell interactions. J. Cell Biol. 175, 453–463 (2006).
Sun, H., Kamanova, J., Lara-Tejero, M. & Galán, J. E. Salmonella stimulates pro-inflammatory signalling through p21-activated kinases bypassing innate immune receptors. Nat. Microbiol. 3, 1122–1130 (2018).
Hobbie, S., Chen, L. M., Davis, R. & Galán, J. E. Involvement of the mitogen-activated protein kinase pathways in the nuclear responses and cytokine production induced by Salmonella Typhimurium in cultured intestinal cells. J. Immunol. 159, 5550–5559 (1997).
Kamanova, J., Sun, H., Lara-Tejero, M. & Galán, J. The Salmonella effector protein SopA modulates innate immune responses by targeting TRIM E3 ligase family members. PLoS Pathog. 12, e1005552 (2016).
Keestra, A. et al. A Salmonella virulence factor activates the NOD1/NOD2 signaling pathway. mBio 2, e00266-11 (2011).
Haraga, A. & Miller, S. I. A Salmonella type III secretion effector interacts with the mammalian serine/threonine protein kinase PKN1. Cell Microbiol. 8, 837–846 (2006).
Stecher, B. et al. Salmonella enterica serovar Typhimurium exploits inflammation to compete with the intestinal microbiota. PLoS Biol. 5, 2177–2189 (2007).
Winter, S. et al. Gut inflammation provides a respiratory electron acceptor for Salmonella. Nature 467, 426–429 (2010).
Fu, Y. & Galan, J. E. A Salmonella protein antagonizes Rac-1 and Cdc42 to mediate host-cell recovery after bacterial invasion. Nature 401, 293–297 (1999).
Sun, H., Kamanova, J., Lara-Tejero, M. & Galán, J. A family of Salmonella type III secretion effector proteins selectively targets the NF-κB signaling pathway to preserve host homeostasis. PLoS Pathog. 12, e1005484 (2016).
Jones, M. A. et al. Secreted effector proteins of Salmonella dublin act in concert to induce enteritis. Infect. Immun. 66, 5799–5804 (1998).
Zhang, S. et al. The Salmonella enterica serotype Typhimurium effector proteins SipA, SopA, SopB, SopD, and SopE2 act in concert to induce diarrhea in calves. Infect. Immun. 70, 3843–3855 (2002).
Wall, A. et al. Small GTPase Rab8a-recruited phosphatidylinositol 3-kinase γ regulates signaling and cytokine outputs from endosomal Toll-like receptors. J. Biol. Chem. 292, 4411–4422 (2017).
Luo, L. et al. TLR crosstalk activates LRP1 to recruit Rab8a and PI3Kγ for suppression of inflammatory responses. Cell Rep. 24, 3033–3044 (2018).
Tong, S., Wall, A., Hung, Y., Luo, L. & Stow, J. Guanine nucleotide exchange factors activate Rab8a for Toll-like receptor signalling. Small GTPases 12, 27–43 (2021).
Brumell, J. et al. SopD2 is a novel type III secreted effector of Salmonella Typhimurium that targets late endocytic compartments upon delivery into host cells. Traffic 4, 36–48 (2003).
Spanò, S., Gao, X., Hannemann, S., Lara-Tejero, M. & Galán, J. A bacterial pathogen targets a host Rab-Family GTPase defense pathway with a GAP. Cell Host Microbe 19, 216–226 (2016).
Creagh, E. & O’Neill, L. TLRs, NLRs and RLRs: a trinity of pathogen sensors that co-operate in innate immunity. Trends Immunol. 27, 352–357 (2006).
Lee, J., Mo, J., Shen, C., Rucker, A. & Raz, E. Toll-like receptor signaling in intestinal epithelial cells contributes to colonic homoeostasis. Curr. Opin. Gastroenterol. 23, 27–31 (2007).
Kelly, D., Conway, S. & Aminov, R. Commensal gut bacteria: mechanisms of immune modulation. Trends Immunol. 26, 326–333 (2005).
Eckmann, L. Sensor molecules in intestinal innate immunity against bacterial infections. Curr. Opin. Gastroenterol. 22, 95–101 (2006).
Shibolet, O. & Podolsky, D. TLRs in the Gut. IV. Negative regulation of Toll-like receptors and intestinal homeostasis: addition by subtraction. Am. J. Physiol. Gastrointest. Liver Physiol. 292, G1469–G1473 (2007).
Lang, T. & Mansell, A. The negative regulation of Toll-like receptor and associated pathways. Immunol. Cell Biol. 85, 425–434 (2007).
Hardt, W.-D., Chen, L.-M., Schuebel, K. E., Bustelo, X. R. & Galán, J. E. Salmonella Typhimurium encodes an activator of Rho GTPases that induces membrane ruffling and nuclear responses in host cells. Cell 93, 815–826 (1998).
Zhou, D., Chen, L. M., Hernandez, L., Shears, S. B. & Galan, J. E. A Salmonella inositol polyphosphatase acts in conjunction with other bacterial effectors to promote host cell actin cytoskeleton rearrangements and bacterial internalization. Mol. Microbiol. 39, 248–259 (2001).
Sato, T. et al. Rab8a and Rab8b are essential for several apical transport pathways but insufficient for ciliogenesis. J. Cell Sci. 127, 422–431 (2014).
Steele-Mortimer, O. et al. Activation of Akt/protein kinase B in epithelial cells by the Salmonella Typhimurium effector sigD. J. Biol. Chem. 275, 37718–37724 (2000).
Marcus, S., Wenk, M., Steele-Mortimer, O. & Finlay, B. A synaptojanin-homologous region of Salmonella Typhimurium SigD is essential for inositol phosphatase activity and Akt activation. FEBS Lett. 494, 201–207 (2001).
Pereira-Leal, J. B. & Seabra, M. C. The mammalian Rab family of small GTPases: definition of family and subfamily sequence motifs suggests a mechanism for functional specificity in the Ras superfamily. J. Mol. Biol. 301, 1077–1087 (2000).
Pereira-Leal, J. B. & Seabra, M. C. Evolution of the Rab family of small GTP-binding proteins. J. Mol. Biol. 313, 889–901 (2001).
D’Costa, V. et al. Salmonella disrupts host endocytic trafficking by SopD2-mediated inhibition of Rab7. Cell Rep. 12, 1508–1518 (2015).
Guo, Z., Hou, X., Goody, R. S. & Itzen, A. Intermediates in the guanine nucleotide exchange reaction of Rab8 protein catalyzed by guanine nucleotide exchange factors rabin8 and GRAB. J.Biol.Chem. 288, 32466–32474 (2013).
Hattula, K., Furuhjelm, J., Arffman, A. & Peränen, J. A Rab8-specific GDP/GTP exchange factor is involved in actin remodeling and polarized membrane transport. Mol. Biol. Cell 13, 3268–3280 (2002).
Homma, Y. & Fukuda, M. Rabin8 regulates neurite outgrowth in both GEF activity-dependent and -independent manners. Mol. Biol. Cell 27, 2107–2118 (2016).
Müller, M. & Goody, R. Molecular control of Rab activity by GEFs, GAPs and GDI. Small GTPases 9, 5–21 (2018).
Cherfils, J. & Zeghouf, M. Regulation of small GTPases by GEFs, GAPs, and GDIs. Physiol. Rev. 93, 269–309 (2013).
Collins, R. “Getting it on”—GDI displacement and small GTPase membrane recruitment. Mol. Cell 12, 1064–1066 (2003).
Sivars, U., Aivazian, D. & Pfeffer, S. Yip3 catalyses the dissociation of endosomal Rab–GDI complexes. Nature 425, 856–859 (2003).
Yamashita, T. & Tohyama, M. The p75 receptor acts as a displacement factor that releases Rho from Rho-GDI. Nat. Neurosci. 6, 461–467 (2003).
Hoiseth, S. K. & Stocker, B. A. Aromatic-dependent Salmonella Typhimurium are non-virulent and effective as live vaccines. Nature 291, 238–239 (1981).
Kaniga, K., Bossio, J. C. & Galan, J. E. The Salmonella Typhimurium invasion genes invF and invG encode homologues of the AraC and PulD family of proteins. Mol. Microbiol. 13, 555–568 (1994).
Demarre, G. et al. A new family of mobilizable suicide plasmids based on broad host range R388 plasmid (IncW) and RP4 plasmid (IncPα) conjugative machineries and their cognate Escherichia coli host strains. Res. Microbiol. 156, 245–255 (2005).
Gibson, D. et al. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat. Methods 6, 343–345 (2009).
Kabsch, W. XDS. Acta Crystallogr. D 66, 125–132 (2010).
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Adams, P. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
DeLano, W. L. The PyMOL Molecular Graphics System (2002); http://www.pymol.org
Itzen, A., Rak, A. & Goody, R. Sec2 is a highly efficient exchange factor for the Rab protein Sec4. J. Mol. Biol. 365, 1359–1367 (2007).
Galán, J. E. & Curtiss III, R. Cloning and molecular characterization of genes whose products allow Salmonella Typhimurium to penetrate tissue culture cells. Proc. Natl Acad. Sci. USA 86, 6383–6387 (1989).
Chang, S., Song, J. & Galán, J. Receptor-mediated sorting of typhoid toxin during its export from Salmonella Typhi-infected cells. Cell Host Microbe 20, 682–689 (2016).
Ran, F. A. et al. Genome engineering using the CRISPR–Cas9 system. Nat. Protoc. 8, 2281–2308 (2013).
Lara-Tejero, M. et al. Role of the caspase-1 inflammasome in Salmonella Typhimurium pathogenesis. J. Exp. Med. 203, 1407–1412 (2006).
Acknowledgements
We thank J. Wang (Tsinghua University, China) for suggestions on structure data processing; the staff of the BL17U1 and BL19U1 beamlines of the National Facility for Protein Science Shanghai (NFPS) at Shanghai Synchrotron Radiation Facility (SSRF) for assistance during data collection; and X. Li (Shandong University, Core facilities for life and environmental sciences) for help with the XRD. Work in X.G.’s laboratory was supported by the National Key R&D Program of China (no. 2018YFE0113000), the National Natural Science Foundation of China (nos. 31770143 and 31901943), the Major Basic Program of Natural Science Foundation of Shandong Province (no. ZR2019ZD21), the Youth Interdisciplinary Innovative Research Group of Shandong University (no. 2020QNQT009) and the Taishan Young Scholars Program (no. tsqn20161005). Work in J.E.G.’s laboratory was supported by NIH grants (nos. R01AI055472 and R01AI079022).
Author information
Authors and Affiliations
Contributions
H.L. performed all of the functional analyses, animal and cell biological experiments. K.J., M.T. and Z.C. carried out the biochemical characterization of SopD and solved its crystal structure bound to Rab8. J.E.G. and X.G. supervised the study. J.E.G. and X.G. wrote the paper with comments from all of the authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information: Nature Microbiology thanks Samuel Miller, Laurent Terradot and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Size-exclusion chromatography analyses of Rab8 in the presence of SopD or SopD2.
Purified Rab81–183 preloaded with GTPγS was incubated with purified SopD or SopD2 and subjected to size-exclusion chromatography in a Superdex 75 increase column. Elution profiles along with SDS-PAGE analyses of the elution fractions are shown. This experiment was conducted at least three times with equivalent results.
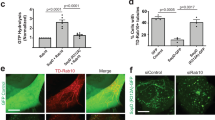
Extended Data Fig. 2 Rab8 is recruited to the Salmonella-induced membrane ruffles.
Henle-407 cells were transiently transfected with a plasmid expressing GFP-Rab8A and subsequently infected with wild-type S. Typhimurium, its ∆sopD isogeneic mutant, or left uninfected (mock). Fifteen minutes after infection, cells were fixed and stained with anti-GFP antibody, to visualize Rab8A (green), rhodamine-labelled Phalloidin, to visualize the actin cytoskeleton (red), and 4’,6-diamidino-2-phenylindole (DAPI) to visualize nuclear and bacteria DNA (blue). Scale bar: 10 µm. This experiment was conducted at least three times with equivalent results.
Extended Data Fig. 3 Rab8 negatively modulates Salmonella-induced pro-inflammatory signaling.
Effects of Rab8a (a and b), Rab8b (c), or Rab8ab (d) deficiency on S. Typhimurium-induced AKT activation. Control or deficient Raw264.7 or HT29 (as indicated) cells were infected with wild-type S. Typhimurium with a multiplicity of infection of 2 and 10, respectively, and at the indicated times after infection, the levels of phosphorylated Akt were determined by immunoblotting analyses as indicated in Materials and Methods. Values are normalized to the β-actin signal, which served as a loading control. The western blots for this experiment are shown in Fig. 2.
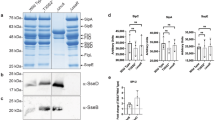
Extended Data Fig. 4 SopD enhances pro-inflammatory signaling by antagonizing Rab8 through its GAP activity.
(a) Expression levels of SopD and its catalytic mutant SopDR312A in stable cell lines (Raw264.7). Stable cell lines (Raw264.7) expressing HA-tagged SopD or SopDR312A were lysed before immunoblotting analysis with the antibodies directed to the HA tag and to β-actin (as a loading control). (b) Effect of the expression of SopD or its catalytic mutant SopDR312A on LPS-induced activation of AKT, p70S6K, Erk1/2, and p38 MAP, and NF-κB signaling pathways. Raw264.7 cells stably expressing HA-tagged SopD or its GAP-deficient mutant SopDR312A were treated with LPS (100 ng/ml) for the indicated times, lysed, and analyzed by immunoblotting with antibodies specific for the phosphorylated state of AKT, p70S6K, p38, and Erk1/2, as well as an antibody to I-κBα and β-actin (as a loading control). The quantification of the western blot analyses is shown on Fig. 2e.
Extended Data Fig. 5 PI3-kinase is required for S. Typhimurium-induced signaling.
(a) Effects of the PI3-kinase inhibitor Wortmannin on S. Typhimurium-induced phosphorylation of AKT. Raw264.7 (MOI=2) or HT29 cells (MOI=10) were pre-treated with increasing concentrations of Wortmannin (10 nM to 1000 nM) for 30 min. Cells were then infected with wild-type S. Typhimurium for 30 min and cell lysates were analyzed by immunoblotting with antibodies directed to the phosphorylated (activated) form of Akt or β-actin (as a loading control). Controls included cells left uninfected or infected with the ∆sopB S. Typhimurium mutant strain, which is defective for the activation of AKT. Quantification of the immunoblots is shown on the right panels. (b) Effects of the PI3-kinase inhibitor Wortmannin on S. Typhimurium-induced transcription of cytokine genes. Raw264.7 cells (MOI=5) and HT29 cells (MOI=20) were pre-treated with Wortmannin (100 nM) for 30 min. Cells were left uninfected or infected with wild-type S. Typhimurium for 4 hs and the cytokine mRNA levels were measured by qPCR assay. Data are the mean ± standard deviation of three independent experiments. ** P < 0.01, *** P < 0.001 (unpaired two-sided t test).
Extended Data Fig. 6 Size-exclusion chromatography profile of SopD/GDP-bound Rab8 complex.
Rab81–183 preloaded with GDP was incubated with SopD, and subjected to size-exclusion chromatography with Superdex 200 increase column. The elution profile along with the SDS-PAGE analyses of the elution fractions of the SopD/Rab8 complex are shown. This experiment was conducted at least three times with equivalent results.
Extended Data Fig. 7 SDS-PAGE analyses of the elution fractions of the size-exclusion chromatography analyses of SopD carrying mutations in amino acids defining its interface with Rab8.
Purified Rab8a1–183 preloaded with GDP was incubated with purified SopD or the indicated SopD mutants and subjected to size-exclusion chromatography (see the chromatographic profile in Fig. 3e). Fractions were collected, separated on an SDS-PAGE, and stained with Coomassie blue. This experiment was conducted at least three times with equivalent results.
Extended Data Fig. 8 The SopDE293A mutation disrupts the ability of SopD to form a complex with Rab8.
FLAG-epitope tagged SopD or SopDE293A were co-expressed with GFP-Rab8 or empty vector in HEK-293T cells. The cell lysates were co-immunoprecipitated with anti-FLAG M2 beads followed by immunoblotting with anti-GFP and anti-Flag antibodies. The quantification of the intensity of the bands is shown in the right panel. This experiment was conducted at least three times with equivalent results.
Extended Data Fig. 9 SopD lacks guanine-nucleotide exchange activity.
Mant-GDP-loaded Rab8a (2 μM) was incubated with Rabin8 (0.4 μM) (positive control), SopD (2 μM), 2 μM SopDR312A (2 μM), SopDE293A (2 μM) or buffer only (negative control), in the presence of 5 μM GTPγS. The decreased fluorescence as a result of the mant-GDP to GTPγS exchange was monitored over time. Data are expressed as the mean ± SD from three independent experiments.
Extended Data Fig. 10 Effect of SopD or its mutants on cytokine production in S. Typhimurium-infected cells.
Raw264.7 cells were infected with the indicated S. Typhimurium strains (MOI=5) and 18 hours after infection, the levels of the indicated cytokines in cell supernatants were quantified by ELISA. Values are the mean ± SD of three independent measurements. ** P < 0.01, *** P < 0.001, ns: not significant (unpaired two-sided t test).
Supplementary information
Supplementary Information
Supplementary Figs. 1–10 and Tables 1–4.
Supplementary Data 1
Statistical source data.
Supplementary Data 2
Unprocessed western blots.
Supplementary Data 3
Statistical source data.
Supplementary Data 4
Unprocessed western blots.
Supplementary Data 5
Statistical source data.
Supplementary Data 6
Unprocessed western blots.
Supplementary Data 7
Statistical source data.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 1
Unprocessed western blots.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 2
Unprocessed western blots.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 3
Unprocessed western blots.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 4
Unprocessed western blots.
Source Data Fig. 6
Statistical source data.
Source Data Fig. 6
Unprocessed western blots.
Source Data Extended Data Fig. 1
Unmodified gels.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 4
Unprocessed western blots.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 5
Unprocessed western blots.
Source Data Extended Data Fig. 6
Unmodified gels.
Source Data Extended Data Fig. 7
Unmodified gels.
Source Data Extended Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 8
Unprocessed western blots.
Source Data Extended Data Fig. 9
Statistical source data.
Source Data Extended Data Fig. 10
Statistical source data.
Rights and permissions
About this article
Cite this article
Lian, H., Jiang, K., Tong, M. et al. The Salmonella effector protein SopD targets Rab8 to positively and negatively modulate the inflammatory response. Nat Microbiol 6, 658–671 (2021). https://doi.org/10.1038/s41564-021-00866-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-021-00866-3
This article is cited by
-
Bacteroides fragilis ubiquitin homologue drives intraspecies bacterial competition in the gut microbiome
Nature Microbiology (2023)
-
Salmonella Typhimurium and inflammation: a pathogen-centric affair
Nature Reviews Microbiology (2021)
-
Salmonella effector SopD promotes plasma membrane scission by inhibiting Rab10
Nature Communications (2021)