Abstract
Microbial interactions are expected to be major determinants of microbiome structure and function. Although fungi are found in diverse microbiomes, their interactions with bacteria remain largely uncharacterized. In this work, we characterize interactions in 16 different bacterial–fungal pairs, examining the impacts of 8 different fungi isolated from cheese rind microbiomes on 2 bacteria (Escherichia coli and a cheese-isolated Pseudomonas psychrophila). Using random barcode transposon-site sequencing with an analysis pipeline that allows statistical comparisons between different conditions, we observed that fungal partners caused widespread changes in the fitness of bacterial mutants compared to growth alone. We found that all fungal species modulated the availability of iron and biotin to bacterial species, which suggests that these may be conserved drivers of bacterial–fungal interactions. Species-specific interactions were also uncovered, a subset of which suggested fungal antibiotic production. Changes in both conserved and species-specific interactions resulted from the deletion of a global regulator of fungal specialized metabolite production. This work highlights the potential for broad impacts of fungi on bacterial species within microbiomes.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Sequence data that support the findings of this study (RB-TnSeq and RNA-seq) have been deposited in the NCBI SRA database with SRA accession codes SRR11514793–SRR11514872 and BioProject code PRJNA624168. MS data are available in the MassIVE database under accession numbers MSV000085070 and MSV000085054. The GNPS molecular network is available at https://gnps.ucsd.edu/ProteoSAFe/status.jsp?task=464b331ef9d54de9957d23b4f9b9db14. The E. coli annotation database used for GO functional enrichment is available at http://bioconductor.org/packages/release/data/annotation/html/org.EcK12.eg.db.html. The Whole Genome Shotgun project for Penicillium sp. str. 12, including reads, genome assembly and annotation has been deposited at DDBJ/ENA/GenBank under the accession JAASRZ000000000 in BioProject PRJNA612335 (BioSample SAMN14369290 and SRA SRR11536435). In addition to these sources, the data used to create Figs. 2, 3, 5 and 6 are available in the Supplementary Tables provided with the paper. Uncropped Southern blots associated with Supplementary Fig. 3 are provided with the manuscript as Supplementary Data. Source data are provided with this paper.
Code availability
The R scripts developed for processing RB-TnSeq data described in this manuscript are available at https://github.com/DuttonLab/RB-TnSeq-Microbial-interactions along with usage instructions. The perl scripts needed for initial processing of RB-TnSeq data published in Wetmore et al.18 are available at https://bitbucket.org/berkeleylab/feba/src/master/.
References
Laforest-Lapointe, I. & Arrieta, M.-C. Microbial eukaryotes: a missing link in gut microbiome studies. mSystems 3, e00201-17 (2018).
Huseyin, C. E., O’Toole, P. W., Cotter, P. D. & Scanlan, P. D. Forgotten fungi—the gut mycobiome in human health and disease. FEMS Microbiol. Rev. 41, 479–511 (2017).
Bergelson, J., Mittelstrass, J. & Horton, M. W. Characterizing both bacteria and fungi improves understanding of the Arabidopsis root microbiome. Sci. Rep. 9, 24 (2019).
Huffnagle, G. B. & Noverr, M. C. The emerging world of the fungal microbiome. Trends Microbiol. 21, 334–341 (2013).
Bradford, L. L. & Ravel, J. The vaginal mycobiome: a contemporary perspective on fungi in women’s health and diseases. Virulence 8, 342–351 (2017).
de Phillips, F., Laiola, M., Blaiotta, G. & Ercolini, D. Different amplicon targets for sequencing-based studies of fungal diversity. Appl. Environ. Microbiol. 83, e00905-17 (2017).
Jiang, T. T. et al. Commensal fungi recapitulate the protective benefits of intestinal bacteria. Cell Host Microbe 22, 809–816 (2017).
Wagg, C., Schlaeppi, K., Banerjee, S., Kuramae, E. E. & van der Heijden, M. G. A. Fungal–bacterial diversity and microbiome complexity predict ecosystem functioning. Nat. Commun. 10, 4841 (2019).
Durán, P. et al. Microbial interkingdom interactions in roots promote Arabidopsis survival. Cell 175, 973–983 (2018).
Tourneroche, A. et al. Bacterial–fungal interactions in the kelp endomicrobiota drive autoinducer-2 quorum sensing. Front. Microbiol. 10, 1693 (2019).
Lindsay, A. K. & Hogan, D. A. Candida albicans: molecular interactions with Pseudomonas aeruginosa and Staphylococcus aureus. Fungal Biol. Rev. 28, 85–96 (2014).
Xu, X.-L. et al. Bacterial peptidoglycan triggers Candida albicans hyphal growth by directly activating the adenylyl cyclase Cyr1p. Cell Host Microbe 4, 28–39 (2008).
Spraker, J. E. et al. Conserved responses in a war of small molecules between a plant-pathogenic bacterium and fungi. mBio 9, e00820-18 (2018).
Khalid, S. et al. NRPS-derived isoquinolines and lipopetides mediate antagonism between plant pathogenic fungi and bacteria. ACS Chem. Biol. 13, 171–179 (2018).
Wolfe, B. E., Button, J. E., Santarelli, M. & Dutton, R. J. Cheese rind communities provide tractable systems for in situ and in vitro studies of microbial diversity. Cell 158, 422–433 (2014).
Morin, M., Pierce, E. C. & Dutton, R. J. Changes in the genetic requirements for microbial interactions with increasing community complexity. eLife 7, e37072 (2018).
Zhang, Y., Kastman, E. K., Guasto, J. S. & Wolfe, B. E. Fungal networks shape dynamics of bacterial dispersal and community assembly in cheese rind microbiomes. Nat. Commun. 9, 336 (2018).
Wetmore, K. M. et al. Rapid quantification of mutant fitness in diverse bacteria by sequencing randomly bar-coded transposons. mBio 6, e00306-15 (2015).
Hallen-Adams, H. E. & Suhr, M. J. Fungi in the healthy human gastrointestinal tract. Virulence 8, 352–358 (2017).
Frąc, M., Hannula, S. E., Bełka, M. & Jędryczka, M. Fungal biodiversity and their role in soil health. Front. Microbiol. 9, 707 (2018).
Dukare, A. S. et al. Exploitation of microbial antagonists for the control of postharvest diseases of fruits: a review. Crit. Rev. Food Sci. Nutr. 59, 1498–1513 (2019).
Richards, T. A., Jones, M. D. M., Leonard, G. & Bass, D. Marine fungi: their ecology and molecular diversity. Ann. Rev. Mar. Sci. 4, 495–522 (2012).
Choi, K.-H., Lee, H., Lee, S., Kim, S. & Yoon, Y. Cheese microbial risk assessments—a review. Asian-Australas. J. Anim. Sci. 29, 307–314 (2016).
Perrin, F. et al. Quantitative risk assessment of haemolytic and uremic syndrome linked to O157:H7 and non-O157:H7 shiga-toxin producing Escherichia coli strains in raw milk soft cheeses: quantitative risk assessment of HUS linked to pathogenic STEC in cheese. Risk Anal. 35, 109–128 (2015).
Cosetta, C. M. & Wolfe, B. E. Deconstructing and reconstructing cheese rind microbiomes for experiments in microbial ecology and evolution. Curr. Protoc. Microbiol. 56, e95 (2020).
Calvo, A. M., Wilson, R. A., Bok, J. W. & Keller, N. P. Relationship between secondary metabolism and fungal development. Microbiol. Mol. Biol. Rev. 66, 447–459 (2002).
Nonejuie, P., Burkart, M., Pogliano, K. & Pogliano, J. Bacterial cytological profiling rapidly identifies the cellular pathways targeted by antibacterial molecules. Proc. Natl Acad. Sci. USA 110, 16169–16174 (2013).
Bok, J. W. & Keller, N. P. LaeA, a regulator of secondary metabolism in Aspergillus spp. Eukaryot. Cell 3, 527–535 (2004).
Kosalková, K. et al. The global regulator LaeA controls penicillin biosynthesis, pigmentation and sporulation, but not roquefortine C synthesis in Penicillium chrysogenum. Biochimie 91, 214–225 (2009).
Laich, F., Fierro, F. & Martín, J. F. Production of penicillin by fungi growing on food products: identification of a complete penicillin gene cluster in Penicillium griseofulvum and a truncated cluster in Penicillium verrucosum. Appl. Environ. Microbiol. 68, 1211–1219 (2002).
Streit, W. R. & Entcheva, P. Biotin in microbes, the genes involved in its biosynthesis, its biochemical role and perspectives for biotechnological production. Appl. Microbiol. Biotechnol. 61, 21–31 (2003).
Kastman, E. K. et al. Biotic interactions shape the ecological distributions of Staphylococcus species. mBio 7, e01157-16 (2016).
Bonham, K. S., Wolfe, B. E. & Dutton, R. J. Extensive horizontal gene transfer in cheese-associated bacteria. eLife 6, e22144 (2017).
Dean, C. R. & Poole, K. Expression of the ferric enterobactin receptor (PfeA) of Pseudomonas aeruginosa: involvement of a two-component regulatory system. Mol. Microbiol. 8, 1095–1103 (1993).
Schalk, I. J., Rigouin, C. & Godet, J. An overview of siderophore biosynthesis among fluorescent Pseudomonads and new insights into their complex cellular organization. Environ. Microbiol. 22, 1447–1466 (2020).
Fecker, L. & Braun, V. Cloning and expression of the fhu genes involved in iron(III)-hydroxamate uptake by Escherichia coli. J. Bacteriol. 156, 1301–1314 (1983).
Sauer, M., Hantke, K. & Braun, V. Ferric-coprogen receptor FhuE of Escherichia coli: processing and sequence common to all TonB-dependent outer membrane receptor proteins. J. Bacteriol. 169, 2044–2049 (1987).
Blin, K. et al. antiSMASH 5.0: updates to the secondary metabolite genome mining pipeline. Nucleic Acids Res. 47, W81–W87 (2019).
Triana, S. et al. Draft genome sequence of the animal and human pathogen malassezia pachydermatis strain CBS 1879. Genome Announc. 3, e01197-15 (2015).
Sarkar, S. K., Chowdhury, C. & Ghosh, A. S. Deletion of penicillin-binding protein 5 (PBP5) sensitises Escherichia coli cells to β-lactam agents. Int. J. Antimicrob. Agents 35, 244–249 (2010).
Perrin, R. M. et al. Transcriptional regulation of chemical diversity in Aspergillus fumigatus by LaeA. PLoS Pathog. 3, e50 (2007).
Haas, H. Fungal siderophore metabolism with a focus on Aspergillus fumigatus. Nat. Prod. Rep. 31, 1266–1276 (2014).
Luckner, M. [On the synthesis of quinoline alkaloids in plants. 2. Fermentativ conversion of the penicillin alkaloids cyclopenin and cyclopenol to viridicatin and viridicatol]. Eur. J. Biochem. 2, 74–78 (1967).
Peters, B. M., Jabra-Rizk, M. A., O’May, G. A., Costerton, J. W. & Shirtliff, M. E. Polymicrobial interactions: impact on pathogenesis and human disease. Clin. Microbiol. Rev. 25, 193–213 (2012).
Scherlach, K., Graupner, K. & Hertweck, C. Molecular bacteria–fungi interactions: effects on environment, food, and medicine. Annu. Rev. Microbiol. 67, 375–397 (2013).
de Boer, W., Folman, L. B., Summerbell, R. C. & Boddy, L. Living in a fungal world: impact of fungi on soil bacterial niche development. FEMS Microbiol. Rev. 29, 795–811 (2005).
Johansson, J. F., Paul, L. R. & Finlay, R. D. Microbial interactions in the mycorrhizosphere and their significance for sustainable agriculture. FEMS Microbiol. Ecol. 48, 1–13 (2004).
Tarkka, M. T., Sarniguet, A. & Frey-Klett, P. Inter-kingdom encounters: recent advances in molecular bacterium–fungus interactions. Curr. Genet. 55, 233–243 (2009).
Taga, M. E. & Walker, G. C. Sinorhizobium meliloti requires a cobalamin-dependent ribonucleotide reductase for symbiosis with its plant host. Mol. Plant. Microbe Interact. 23, 1643–1654 (2010).
Deveau, A. et al. Role of fungal trehalose and bacterial thiamine in the improved survival and growth of the ectomycorrhizal fungus Laccaria bicolor S238N and the helper bacterium Pseudomonas fluorescens BBc6R8. Environ. Microbiol. Rep. 2, 560–568 (2010).
Hantke, K. Identification of an iron uptake system specific for coprogen and rhodotorulic acid in Escherichia coli K12. Mol. Gen. Genet. 191, 301–306 (1983).
Arias, A. A. et al. Growth of desferrioxamine-deficient Streptomyces mutants through xenosiderophore piracy of airborne fungal contaminations. FEMS Microbiol. Rev. 91, fiv080 (2015).
Haas, H., Eisendle, M. & Turgeon, B. G. Siderophores in fungal physiology and virulence. Annu. Rev. Phytopathol. 46, 149–187 (2008).
Park, M., Cho, Y.-J., Lee, Y. W. & Jung, W. H. Understanding the mechanism of action of the anti-dandruff agent zinc pyrithione against Malassezia restricta. Sci. Rep. 8, 12086 (2018).
Gründlinger, M. et al. Fungal siderophore biosynthesis is partially localized in peroxisomes. Mol. Microbiol. 88, 862–875 (2013).
Heymann, P., Ernst, J. F. & Winkelmann, G. A gene of the major facilitator superfamily encodes a transporter for enterobactin (Enb1p) in Saccharomyces cerevisiae. Biometals 13, 65–72 (2000).
Sass, G. et al. Studies of Pseudomonas aeruginosa mutants indicate pyoverdine as the central factor in inhibition of Aspergillus fumigatus biofilm. J. Bacteriol. 200, e00345-17 (2017).
Briard, B. et al. Pseudomonas aeruginosa manipulates redox and iron homeostasis of its microbiota partner Aspergillus fumigatus via phenazines. Sci. Rep. 5, 8220 (2015).
Clancy, A. et al. Evidence for siderophore-dependent iron acquisition in group B streptococcus. Mol. Microbiol. 59, 707–721 (2006).
Jin, B. et al. Iron acquisition systems for ferric hydroxamates, haemin and haemoglobin in Listeria monocytogenes. Mol. Microbiol. 59, 1185–1198 (2006).
Mishra, R. P. N. et al. Staphylococcus aureus FhuD2 is involved in the early phase of staphylococcal dissemination and generates protective immunity in mice. J. Infect. Dis. 206, 1041–1049 (2012).
Rocha, E. R. & Krykunivsky, A. S. Anaerobic utilization of Fe(III)-xenosiderophores among Bacteroides species and the distinct assimilation of Fe(III)-ferrichrome by Bacteroides fragilis within the genus. MicrobiologyOpen 6, e00479 (2017).
Li, H. et al. The outer mucus layer hosts a distinct intestinal microbial niche. Nat. Commun. 6, 8292 (2015).
Ong, S. A. & Neilands, J. B. Siderophores in microbially processed cheese. J. Agric. Food Chem. 27, 990–995 (1979).
David, L. A. et al. Diet rapidly and reproducibly alters the human gut microbiome. Nature 505, 559–563 (2014).
Rehner, S. A. & Samuels, G. J. Molecular systematics of the Hypocreales: a teleomorph gene phylogeny and the status of their anamorphs. Can. J. Bot. 73, 816–823 (1995).
Glass, N. L. & Donaldson, G. C. Development of primer sets designed for use with the PCR to amplify conserved genes from filamentous ascomycetes. Appl. Environ. Microbiol. 61, 1323–1330 (1995).
Ronquist, F. & Huelsenbeck, J. P. MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19, 1572–1574 (2003).
Dunnett, C. W. A multiple comparison procedure for comparing several treatments with a Control. J. Am. Stat. Assoc. 50, 1096–1121 (1955).
Wickham, H. ggplot2: Elegant Graphics for Data Analysis (Springer Science+Business Media, 2009).
Cleary, J. L., Luu, G. T., Pierce, E. C., Dutton, R. J. & Sanchez, L. M. BLANKA: an algorithm for blank subtraction in mass spectrometry of complex biological samples. J. Am. Soc. Mass. Spectrom. 30, 1426–1434 (2019).
Mohimani, H. et al. Dereplication of microbial metabolites through database search of mass spectra. Nat. Commun. 9, 4035 (2018).
Mohimani, H. et al. Dereplication of peptidic natural products through database search of mass spectra. Nat. Chem. Biol. 13, 30–37 (2017).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B Stat. Methodol. 57, 289–300 (1995).
Tang, Y., Horikoshi, M. & Li, W. ggfortify: unified interface to visualize statistical result of popular R packages. R Journal 8, 474–485 (2016).
Huerta-Cepas, J. et al. eggNOG 5.0: a hierarchical, functionally and phylogenetically annotated orthology resource based on 5090 organisms and 2502 viruses. Nucleic Acids Res. 47, D309–D314 (2019).
Yu, G., Wang, L.-G., Han, Y. & He, Q.-Y. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16, 284–287 (2012).
Carlson, M. org.EcK12.eg.db: Genome wide annotation for E. coli strain K12. R package version 3.8.2. (Bioconductor, 2019).
Carlson, M. & Pagès, H. AnnotationForge: tools for building SQLite-based annotation data packages. R package version 1.26.0 (Bioconductor, 2019).
Baba, T. et al. Construction of Escherichia coli K-12 in-frame, single-gene knockout mutants: the Keio collection. Mol. Syst. Biol. 2, 2006.0008 (2006).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Schwyn, B. & Neilands, J. B. Universal chemical assay for the detection and determination of siderophores. Anal. Biochem. 160, 47–56 (1987).
Payne, S. M. Detection, isolation, and characterization of siderophores. Methods Enzymol. 235, 329–344 (1994).
Grenier, F., Matteau, D., Baby, V. & Rodrigue, S. Complete genome sequence of Escherichia coli BW25113. Genome Announc. 2, e01038-14 (2014).
Lawrence, M. et al. Software for computing and annotating genomic ranges. PLoS Comput. Biol. 9, e1003118 (2013).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Datsenko, K. A. & Wanner, B. L. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc. Natl Acad. Sci. USA 97, 6640–6645 (2000).
Koren, S. et al. Canu: scalable and accurate long-read assembly via adaptive k-mer weighting and repeat separation. Genome Res. 27, 722–736 (2017).
Vaser, R., Sović, I., Nagarajan, N. & Šikić, M. Fast and accurate de novo genome assembly from long uncorrected reads. Genome Res. 27, 737–746 (2017).
Walker, B. J. et al. Pilon: an integrated tool for comprehensive microbial variant detection and genome assembly improvement. PLoS ONE 9, e112963 (2014).
Min, B., Grigoriev, I. V. & Choi, I.-G. FunGAP: fungal genome annotation pipeline using evidence-based gene model evaluation. Bioinformatics 33, 2936–2937 (2017).
Jones, P. et al. InterProScan 5: genome-scale protein function classification. Bioinformatics 30, 1236–1240 (2014).
Lim, F. Y., Sanchez, J. F., Wang, C. C. C. & Keller, N. P. Toward awakening cryptic secondary metabolite gene clusters in filamentous fungi. Methods Enzymol. 517, 303–324 (2012).
Xie, C. et al. KOBAS 2.0: a web server for annotation and identification of enriched pathways and diseases. Nucleic Acids Res. 39, W316–W322 (2011).
The Gene Ontology Consortium. The Gene Ontology Resource: 20 years and still GOing strong. Nucleic Acids Res. 47, D330–D338 (2019).
Ashburner, M. et al. Gene Ontology: tool for the unification of biology. Nat. Genet. 25, 25–29 (2000).
Jukes, T. H. & Cantor, C. R. in Mammalian Protein Metabolism Vol. 3 (ed. Munro, H. N.) 21–132 (Academic Press, 1969).
Conway, J. R., Lex, A. & Gehlenborg, N. UpSetR: an R package for the visualization of intersecting sets and their properties. Bioinformatics 33, 2938–2940 (2017).
Patti, G. J. et al. A view from above: cloud plots to visualize global metabolomic data. Anal. Chem. 85, 798–804 (2013).
Acknowledgements
The authors would like to thank the following people and groups: the Arkin Lab and the Deutschbauer Lab at UC Berkeley for the E. coli Keio_ML9 RB-TnSeq library; K. Jepsen at the IGM Genomics Center at the University of California, San Diego for assistance with sequencing; S. Kryazhimskiy (UCSD) for his input on RB-TnSeq data processing; C. Dinh (UCSD) for assistance with fungal genome assembly; W. Bushnell (UCSD) for assistance with fitness validation experiments; S. Beyhan (JCVI) for advice on fungal genome annotation; L. Marotz (UCSD) for assistance with non-cheese yeasts; and members of the Dutton Lab, especially B. Anderson and C. Saak, for constructive comments on the manuscript. This work was supported by the National Institutes of Health grant nos. T32-AT007533 (to J.C.L.) and F31-AT010418 (to J.C.L.), the National Institutes of Health grant no. R01-AI117712 (to R.B.L), National Science Foundation grant no. MCB-1817955 (to L.M.S.), National Science Foundation grant no. MCB-1817887 (to R.J.D. and L.M.S.), National Science Foundation grant no. MCB-1715553 (to B.E.W.), the UCSD Center for Microbiome Innovation (to E.C.P.), the UCSD Ruth Stern Award (to E.C.P.), NIH Institutional Training grant no. 5 T32 GM 7240-40 (to E.C.P.) and the National Institutes of Health grant no. R01GM112739-01 (to N.P.K.).
Author information
Authors and Affiliations
Contributions
R.J.D., L.M.S. and B.E.W. conceptualized the study. E.C.P., M.M., J.T., R.B.L. and J.C.L. performed the experiments. The R data processing pipeline was written by M.M. Data analyses were performed by E.C.P., J.C.L. and M.M. The article was written by E.C.P., M.M., L.M.S., J.C.L. and R.J.D. and revised with input from all authors. The figures were made by E.C.P. with input from L.M.S., J.C.L., R.J.D. and M.M., except for Fig. 4a and Extended Data Fig. 7 (R.B.L.), Fig. 6d (J.C.L.), Extended Data Figs. 1 and 2 (M.M.) and Supplementary Fig. 3 (J.T.). The study was supervised by R.J.D. and L.M.S.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 RB-TnSeq R data processing pipeline for gene fitness comparison across multiple conditions.
The pipeline is divided into three main scripts. Script 1 calculates the normalized gene fitness for each replicate of an RB-TnSeq experiment. This script has to be run for each replicate independently. Then, the .Rdata files from Script 1 are loaded in Script 2. Script 2 calculates for each RB-TnSeq condition the average gene fitness across replicates (inverse-variance weighted average). Finally, Script 3 compares gene fitness values of each RB-TnSeq condition against a chosen reference condition.
Extended Data Fig. 2 RB-TnSeq assay for fungal impacts on bacterial gene fitness.
Characterized pooled bacterial mutant libraries were grown in a biofilm either alone or with a fungal partner. After seven days of growth, mutant abundances were compared to the starting library abundances for each condition. Changes in barcode abundances were used to calculate gene fitness values. Genes with fitness values that differed significantly between co-culture and alone conditions (significant interaction fitness) were identified as potentially relevant to fungal interaction.
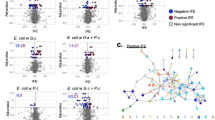
Extended Data Fig. 3 Comparison of E. coli and P. psychrophila interaction fitness values for the 874 genes found in both bacteria.
BLAST comparison (e-value cutoff of 1e-2) of protein sequences from P. psychrophila to those from E. coli and comparison of eggNOG gene assignments were used to find the best cross-species gene match for all genes with significant interaction fitness for at least one of the two bacterial species. A match was found for 874 genes. For each fungal condition, the fitness value of these genes with E. coli is on the x-axis and with P. psychrophila on the y-axis. In each condition, the genes are colored according to whether they have significant interaction fitness in the condition for E. coli (red), P. psychrophila (blue), both (purple), or neither (white).
Extended Data Fig. 4 Network of E. coli (left) or P. psychrophila (right) genes with positive and negative RB-TnSeq interaction fitness.
Each purple node represents a fungal partner and is labeled as follows: fungal partner (number of genes with interaction fitness; number of genes with interaction fitness unique to this condition). Each blue or orange node represents a bacterial gene. Nodes are colored by whether the average interaction fitness is positive (blue) or negative (orange) as shown in the legend below and are sized by average strength of interaction fitness across partners.
Extended Data Fig. 5 Principal Component Analysis of RB-TnSeq data.
Analysis was done on the fitness values in each fungal condition for all E. coli (left) or P. psychrophila (right) genes with an interaction fitness in at least one fungal condition. Each colored circle represents a fungal condition.
Extended Data Fig. 6 Clusters of Orthologous Genes (COG) categories of genes with interaction fitness.
Charts display the number of genes with interaction fitness that fall into each COG category for E. coli (left) or P. psychrophila (right).
Extended Data Fig. 7 Bacterial Cytological Profiling of ΔtolC E.coli treated with known antibiotic compounds on cheese curd agar.
DAPI dye stains DNA and FM4-64 dye stains bacterial membranes. SYTOX green stains nucleic acids but cannot penetrate live cells. Scale bars represent 2 µm. Testing of each antibiotic at four concentrations was performed once, and cells from the edges of zones of clearing were imaged for at least 5 fields from each condition to ensure consistency in phenotype.
Extended Data Fig. 8 Siderophore production by filamentous fungi.
Liquid CAS assay was performed on filtered and concentrated fungal supernatants from three replicates grown in 2% liquid cheese pH 7 for 12 days. Row A) 1-3: Liquid cheese control 4-6: Penicillium SAM3. Row B) 1-3: Debaryomyces 135B 4-6: Penicillium #12. Row C) 1-3: Candida 135E. 4-6: Penicillium RS17. Row D) 1-3: Scopulariopsis 165-5 Row E) 1-3: Scopulariopsis JB370. Row F) 1-3: Fusarium 554A. % Siderophore units calculated as [(Ar - As)/(Ar)]*100, where Ar is the absorbance of the cheese curd agar supernatant blank and As is the absorbance of the sample. N=3 biologically independent samples, error bars show standard deviation and black point is the mean.
Extended Data Fig. 9 Fitness defect of Δfep mutants on iron-limiting CCA.
Visual assays of E. coli mutant growth spotted alone on CCA pH 7 supplemented with tetrazolium chloride, an indicator of respiration.
Extended Data Fig. 10 Comparison of Penicillium sp. str. 12 WT and laeA deletion mutant growth on CCA.
Radial growth assay, including quantification, of Penicillium sp. str. 12 WT and laeA deletion mutant grown alone or with E. coli on CCA pH 7 (N=3 biologically independent experiments, error bars show standard deviation and black point is the mean). Spore counts from Penicillium sp. str. 12 WT and laeA deletion mutant grown alone or with E. coli for 7 days on CCA are also shown (N=3 biologically independent samples, error bars show standard deviation and black point is the mean).
Supplementary information
Supplementary Information
Supplementary Figs. 1–3 and Supplementary Method 1.
Supplementary Tables
Supplementary Tables 1–19.
Supplementary Data
Uncropped Southern blots related to Supplementary Fig. 3.
Rights and permissions
About this article
Cite this article
Pierce, E.C., Morin, M., Little, J.C. et al. Bacterial–fungal interactions revealed by genome-wide analysis of bacterial mutant fitness. Nat Microbiol 6, 87–102 (2021). https://doi.org/10.1038/s41564-020-00800-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-020-00800-z
This article is cited by
-
Acidification suppresses the natural capacity of soil microbiome to fight pathogenic Fusarium infections
Nature Communications (2023)
-
Mutualism reduces the severity of gene disruptions in predictable ways across microbial communities
The ISME Journal (2023)
-
Mycobiota and diet-derived fungal xenosiderophores promote Salmonella gastrointestinal colonization
Nature Microbiology (2022)
-
Multi-kingdom microbiota analyses identify bacterial–fungal interactions and biomarkers of colorectal cancer across cohorts
Nature Microbiology (2022)
-
Higher-order interactions shape microbial interactions as microbial community complexity increases
Scientific Reports (2022)