Abstract
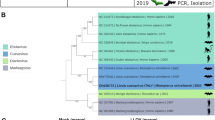
Filoviruses, especially Ebola virus (EBOV) and Marburg virus (MARV), are notoriously pathogenic and capable of causing severe haemorrhagic fever diseases in humans with high lethality1,2. The risk of future outbreaks is exacerbated by the discovery of other bat-borne filoviruses of wide genetic diversity globally3,4,5. Here we report the characterization of a phylogenetically distinct bat filovirus, named Měnglà virus (MLAV). The coding-complete genome of MLAV shares 32–54% nucleotide sequence identity with known filoviruses. Phylogenetic analysis places this new virus between EBOV and MARV, suggesting the need for a new genus taxon. Importantly, despite the low amino acid sequence identity (22–39%) of the glycoprotein with other filoviruses, MLAV is capable of using the Niemann–Pick C1 (NPC1) as entry receptor. MLAV is also replication-competent with chimeric MLAV mini-genomes containing EBOV or MARV leader and trailer sequences, indicating that these viruses are evolutionally and functionally closely related. Finally, MLAV glycoprotein-typed pseudo-types transduced cell lines derived from humans, monkeys, dogs, hamsters and bats, implying a broad species cell tropism with a high risk of interspecies spillover transmission.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
The data that support the findings of this study are available from the corresponding author upon request. The GenBank accession number for the genome sequence of MLAV is KX371887.
Change history
08 February 2019
In the version of this Letter originally published, in the ‘Phylogenetic analysis’ section of the Methods, the authors mistakenly stated that the GenBank accession number for the Ravn virus genome sequence was FJ750958. The correct accession number is DQ447649 for Ravn virus, Kenya, 1987. Accordingly, the label ‘RAVN2007’ in Fig. 1b should have been ‘RAVV1987’. This mistake does not change any conclusions in this study. This statement and figure have now been amended in all versions of the Letter, and the Supplementary Information file has been updated accordingly.
References
Feldmann, H., Sanchez, A. & Geisbert, T. W. in Fields Virology (eds Knipe, D. M. & Howley, P. M.) Ch. Filoviridae: Marburg and Ebola viruses, 923–956 (Lippincott Williams & Wilkins, Philadelphia, 2013).
Radoshitzky, S. R., Bavari, S., Jahrling, P. B. & Kuhn, J. H. in Medical Aspects of Biological Warfare (ed. Garr, J. H.) Ch. Filoviruses, 569–614 (Borden Institute, Fort Sam Houston, 2018).
He, B. et al. Filovirus RNA in fruit bats, China. Emerg. Infect. Dis. 21, 1675–1677 (2015).
Yang, X. L. et al. Genetically diverse filoviruses in rousettus and eonycteris spp. bats, China, 2009 and 2015. Emerg. Infect. Dis. 23, 482–486 (2017).
Kemenesi, G. et al. Re-emergence of Lloviu virus in Miniopterus schreibersii bats, Hungary, 2016. Emerg. Microbes Infect. 7, 66 (2018).
Kuhn, J. H. et al. in Classification and Nomenclature of Viruses. Virus Taxonomy: Ninth Report of the International Committee on Taxonomy of Viruses (eds King, A. M. Q., Adams, M. J., Carstens, E. B. & Lefkowitz, E. J.) Ch. Filoviridae, 665–671 (Academic Press, London, 2012).
Bao, Y. et al. Implementation of objective PASC-derived taxon demarcation criteria for official classification of filoviruses. Viruses 9, 105 (2017).
Amarasinghe, G. K. et al. Taxonomy of the order Mononegavirales: update 2018. Arch. Virol. 163, 2283–2294 (2018).
Shi, M. et al. The evolutionary history of vertebrate RNA viruses. Nature 556, 197–202 (2018).
Miller, E. H. et al. Ebola virus entry requires the host-programmed recognition of an intracellular receptor. EMBO J. 31, 1947–1960 (2012).
Hoffmann, M., Gonzalez Hernandez, M., Berger, E., Marzi, A. & Pohlmann, S. The glycoproteins of all filovirus species use the same host factors for entry into bat and human cells but entry efficiency is species dependent. PLoS. ONE 11, e0149651 (2016).
Carette, J. E. et al. Ebola virus entry requires the cholesterol transporter Niemann–Pick C1. Nature 477, 340–343 (2011).
Côté, M. et al. Small molecule inhibitors reveal Niemann–Pick C1 is essential for Ebola virus infection. Nature 477, 344–348 (2011).
Towner, J. S. et al. Isolation of genetically diverse Marburg viruses from Egyptian fruit bats. PLoS Path. 5, e1000536 (2009).
Schuh, A. J., Amman, B. R. & Towner, J. S. Filoviruses and bats. Microbiol. Aus. 38, 12–16 (2017).
Olival, K. J. & Hayman, D. T. Filoviruses in bats: current knowledge and future directions. Viruses 6, 1759–1788 (2014).
Laing, E. D. et al. Serologic evidence of fruit bat exposure to filoviruses, Singapore, 2011-2016. Emerg. Infect. Dis. 24, 114–117 (2018).
Leroy, E. M. et al. Fruit bats as reservoirs of Ebola virus. Nature 438, 575–576 (2005).
Negredo, A. et al. Discovery of an ebolavirus-like filovirus in europe. PLoS Pathog. 7, e1002304 (2011).
Jayme, S. I. et al. Molecular evidence of Ebola Reston virus infection in Philippine bats. Virol. J. 12, 107 (2015).
Yuan, J., Zhang, Y., Li, J., Wang, L. F. & Shi, Z. Serological evidence of ebolavirus infection in bats, China. Virol. J. 9, 236 (2012).
Mühlberger, E. Filovirus replication and transcription. Future Virol. 2, 205–215 (2007).
Hartman, A. L., Towner, J. S. & Nichol, S. T. A C-terminal basic amino acid motif of Zaire ebolavirus VP35 is essential for type I interferon antagonism and displays high identity with the RNA-binding domain of another interferon antagonist, the NS1 protein of influenza A virus. Virology 328, 177–184 (2004).
Edwards, M. R. et al. Differential regulation of interferon responses by ebola and marburg virus VP35 proteins. Cell Rep. 14, 1632–1640 (2016).
Valmas, C. et al. Marburg virus evades interferon responses by a mechanism distinct from Ebola virus. PLoS Pathog. 6, e1000721 (2010).
Kuhn, J. H. et al. Conserved receptor-binding domains of Lake Victoria marburgvirus and Zaire ebolavirus bind a common receptor. J. Biol. Chem. 281, 15951–15958 (2006).
Wang, H. et al. Ebola viral glycoprotein bound to its endosomal receptor niemann-pick C1. Cell 164, 258–268 (2016).
Thomas, H. & Groseth, A. (eds) Ebolaviruses, 53–78 (Humana Press, New York, 2017).
Theriault, S., Groseth, A., Neumann, G., Kawaoka, Y. & Feldmann, H. Rescue of Ebola virus from cDNA using heterologous support proteins. Virus Res. 106, 43–50 (2004).
Watt, A. et al. A novel life cycle modeling system for Ebola virus shows a genome length-dependent role of VP24 in virus infectivity. J. Virol. 88, 10511–10524 (2014).
Kumar, S., Stecher, G. & Tamura, K. MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 33, 1870–1874 (2016).
Li, Y. et al. Host range, prevalence, and genetic diversity of adenoviruses in bats. J. Virol. 84, 3889–3897 (2010).
Albariño, C. G. et al. Development of a reverse genetics system to generate recombinant Marburg virus derived from a bat isolate. Virology 446, 230–237 (2013).
Brouillette, R. B. & Maury, W. Production of filovirus glycoprotein-pseudotyped vesicular stomatitis virus for study of filovirus entry mechanisms. Methods Mol. Biol. 1628, 53–63 (2017).
Ran, F. A. et al. Genome engineering using the CRISPR-Cas9 system. Nat. Protoc. 8, 2281–2308 (2013).
Acknowledgements
This study was supported in part by the strategic priority research programme of the Chinese Academy of Sciences (no. XDB29010000 to Z-L.S.), Singapore National Research Foundation grants (nos. NRF2012NRF-CRP001-056 and NRF2016NRF-NSFC002-013 to L.-F.W.) and a Singapore MINDEF grant (no. MINDEF-NUS-DIRP/2015/04 to L.-F.W. and D.E.A.). X.L.Y. is supported by the Visiting Scientist Fellowship from the Chinese Academy of Sciences. We thank T. Hoenen and H. Feldmann for providing plasmids from which the relevant EBOV genes were derived for this study.
Author information
Authors and Affiliations
Contributions
X.-L.Y., L.-F.W. and Z.-L.S. conceived the study. X.-L.Y., C.W.T., R.-D.J., B.L., W.Z., Y.Z., X.F.L., P.Z., X.-L.L., L.Z., S.-Y.L. and Y.-Z.Z. conducted the experiments. W.G., Z.-L.S. and L.-F.W. provided resources and reagents. X.-L.Y., C.W.T. and D.E.A. analysed the data. X.-L.Y., C.W.T., D.E.A., Z.-L.S. and L.-F.W. prepared the manuscript. All authors read, edited and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figures 1–5, and Supplementary Tables 1 and 2.
Rights and permissions
About this article
Cite this article
Yang, XL., Tan, C.W., Anderson, D.E. et al. Characterization of a filovirus (Měnglà virus) from Rousettus bats in China. Nat Microbiol 4, 390–395 (2019). https://doi.org/10.1038/s41564-018-0328-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-018-0328-y
This article is cited by
-
Isolation and genome characterization of Lloviu virus from Italian Schreibers’s bats
Scientific Reports (2023)
-
Structure of the Ebola virus polymerase complex
Nature (2022)
-
Isolation of infectious Lloviu virus from Schreiber’s bats in Hungary
Nature Communications (2022)
-
Functional comparison of SARS-CoV-2 with closely related pangolin and bat coronaviruses
Cell Discovery (2021)
-
Viral genomics in Ebola virus research
Nature Reviews Microbiology (2020)