Abstract
Despite the wide availability of antibiotics, infectious diseases remain a leading cause of death worldwide1. In the absence of new therapies, mortality rates due to untreatable infections are predicted to rise more than tenfold by 2050. Natural products (NPs) made by cultured bacteria have been a major source of clinically useful antibiotics. In spite of decades of productivity, the use of bacteria in the search for new antibiotics was largely abandoned due to high rediscovery rates2,3. As only a fraction of bacterial diversity is regularly cultivated in the laboratory and just a fraction of the chemistries encoded by cultured bacteria are detected in fermentation experiments, most bacterial NPs remain hidden in the global microbiome. In an effort to access these hidden NPs, we have developed a culture-independent NP discovery platform that involves sequencing, bioinformatic analysis and heterologous expression of biosynthetic gene clusters captured on DNA extracted from environmental samples. Here, we describe the application of this platform to the discovery of the malacidins, a distinctive class of antibiotics that are commonly encoded in soil microbiomes but have never been reported in culture-based NP discovery efforts. The malacidins are active against multidrug-resistant pathogens, sterilize methicillin-resistant Staphylococcus aureus skin infections in an animal wound model and did not select for resistance under our laboratory conditions.
Similar content being viewed by others
Main
Calcium-dependent antibiotics are a small family of N-acylated cyclic peptides that require calcium for antibacterial activity. Known members of this family contain a conserved Asp-X-Asp-Gly motif that is thought to facilitate calcium binding4,5,6. One example, daptomycin, has proved useful clinically in the treatment of multidrug-resistant bacteremia. Calcium-dependent antibiotics were of particular interest to us because individual family members have been shown to have discrete modes of action, targeting either cell wall biosynthesis or cell membrane integrity. Given these differences in modes of action, we hypothesized that the conserved Asp-X-Asp-Gly calcium-binding motif might be indicative of a broader collection of uncharacterized, bacterially encoded antibiotics with diverse mechanisms of action and therapeutic potential.
To test this hypothesis, we undertook a sequence-guided screen of diverse soils for biosynthetic gene clusters (BGCs) that encode calcium-binding motifs. Due to the complexity of soil metagenomes, it remains challenging to shotgun sequence deep enough to generate data that are broadly useful for BGC discovery7. We have developed sequencing strategies that rely on the barcoding of biosynthetic genes using degenerate polymerase chain reaction (PCR) primers to parse mixtures of BGCs present in environmental samples7,8. In this approach, primers targeting conserved NP biosynthetic genes are used to generate PCR amplicon pools containing homologous genes from BGCs present in an environmental DNA (eDNA) sample (Fig. 1a). Individual next-generation sequencing reads derived from these amplicons (NP sequence tags, NPSTs) are used to predict BGCs present in a sample by comparing them to a database of sequences from characterized BGCs. This analysis is carried out using the bioinformatics platform eSNaPD (environmental Surveyor of Natural Product Diversity: http://esnapd2.rockefeller.edu) that was developed to evaluate metagenome-derived NPSTs8,9.
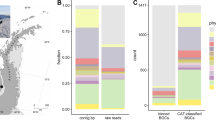
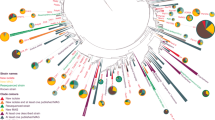
a, (i) Degenerate PCR primers targeting the conserved regions of adenylation domains found in non-ribosomal peptide synthetase genes were used to generate amplicons from an arrayed collection of eDNA isolated from 2,000 unique soils. The reads from these next-generation sequenced amplicons (NPSTs) were analysed by eSNaPD. (ii) A desert soil rich in AD NPSTs from the previously unknown malacidin clade was used to build an arrayed cosmid library. Cosmids harbouring all fragments of a targeted BGC were assembled and integrated into a heterologous host for production, extraction and characterization. b, AD NPSTs identified by the eSNaPD analysis to be evolutionarily related to the conserved Asp4 ADs of known calcium-dependent antibiotics were used to phylogenetically map the unexplored clades of this larger family across all tested soil microbiomes. The subfamilies of calcium-dependent antibiotics and their relative abundance are illustrated on the phylogenetic tree by colour and percentage. Across all sampled soil metagenomes, the malacidin antibiotic-clade represents 19% of the NPSTs, and 59% of calcium-dependent antibiotic tags originate from unexplored branches. c, Geospatial distribution of calcium-dependent antibiotics across sampled US soil metagenomes. States containing at least one soil sample with AD NPSTs from the malacidin clade are indicated in orange. States lacking malacidin tags but still containing calcium-dependent-antibiotic NPSTs are indicated in blue. States with at least one sampled soil but no detected calcium-dependent-antibiotic NPSTs are highlighted in dark grey.
Known calcium-dependent antibiotics are biosynthesized by non-ribosomal peptide synthetases (NRPS). Accordingly, we used primers targeting NRPS adenylation domains (ADs) to track this family of NPs across diverse soil microbiomes. For this study, and as part of our ongoing soil metagenome-driven NP discovery efforts, we expanded our soil collection to more than 2,000 soils from ecologically and geographically diverse environments8. Even using a conservative estimate of 103 unique bacterial species per gram of soil2, we expect the diversity of bacteria present in this collection to rival that of the largest culture collections. Initially, primers targeting NRPS ADs were used to screen eDNA isolated from small aliquots of each soil to identify environments predicted to contain gene clusters that encode for unidentified calcium-dependent antibiotics.
Three-quarters of sequenced soils had NPSTs that mapped to at least one AD from a known calcium-dependent antibiotic BGC (Fig. 1b,c and Supplementary Fig. 1). Only 13% of these identified NPSTs cluster at ≥95% nucleotide identity to ADs found in characterized calcium-dependent antibiotics and less than 30% of them are found in more than one soil metagenome. Taken together, this indicates that the majority of lipopeptides encoded by the global soil metagenome are probably uncharacterized and that even within our large soil collection, we have captured only a fraction of the biosynthetic diversity that exists within the calcium-dependent antibiotic family.
Phylogenetic analysis of AD sequences from characterized calcium-dependent antibiotics indicated that the domain responsible for incorporating the first aspartic acid (Asp4) in the conserved Asp-X-Asp-Gly motif most closely mapped to functional divergence of BGCs in this family (Supplementary Fig. 1b). We, therefore, focused on eSNaPD data for this domain to track calcium-dependent antibiotic BGCs. A phylogenetic tree derived from tags associated with this domain showed numerous clades not associated with known BGCs, indicating the existence of uncharacterized calcium-dependent antibiotics in soil microbiomes. One distinct, eDNA-specific clade was found in 19% of metagenomes (Fig. 1b), suggesting that the BGCs associated with these tags belong to an abundant and yet uncharacterized family of antibiotics, which we have called the malacidins (metagenomic acidic lipopeptide antibiotic-cidins).
To recover a complete malacidin BGC for use in heterologous expression studies, we retrieved from our soil archive a desert soil (DFD0097) that was rich in NPSTs from the malacidin branch of the AD phylogenetic tree (Fig. 1c). Employing standard soil metagenome cloning methods, a saturating cosmid library was constructed using DNA extracted from DFD0097 soil10. This 20-million-membered library was archived as purified cosmid DNA and Escherichia coli glycerol stocks arrayed in 96-well format, with each library well containing ~20,000 unique clones10,11. To expedite the recovery of BGCs, each well of the library was individually screened by PCR using the barcoded AD-targeting primers used to profile soils. Library-derived NPST data were analysed by eSNaPD to generate a map of BGC information across the arrayed library.
Using this BGC prediction map, overlapping cosmid clones predicted to contain the malacidin BGC were recovered from the library. Sequencing and in silico analysis of these clones suggested that the malacidin BGC spanned 72 kilobases across 3 cosmids (DFD0097-644, DFD0097-735 and DFD0097-388) (Fig. 2a and Supplementary Table 1, GenBank Accession KY654519). For the purposes of heterologous expression, these three overlapping cosmids were assembled into a contiguous fragment of DNA using transformation-associated recombination in yeast and the E. coli:yeast:Streptomyces shuttle vector, pTARa (Fig. 2b)11. The resulting bacterial artificial chromosome (DFD0097-644:735:388) and the empty pTARa vector were separately conjugated into Streptomyces albus J1074. Extracts from cultures of S. albus harbouring DFD0097-644:735:388 were found to exhibit antibacterial activity against Staphylococcus aureus and contain clone-specific metabolites (Fig. 2c,d). The major clone-specific metabolites, malacidin A and B, were isolated from cultures of S. albus DFD0097-644:735:388 and their structures were elucidated using a combination of mass spectrometry and NMR data. The malacidin structures were supported by a detailed bioinformatic analysis of the BGC (Fig. 2e, Supplementary Figs. 2–25, Supplementary Tables 2 and 3 and Supplementary Discussion).
a,b, The malacidin BGC was recovered on three overlapping cosmid clones (a) and assembled from these three overlapping clones in yeast using transformation-associated recombination (TAR) (b). The resulting BAC was integrated into the S. albus genome for heterologous expression studies. c, A representative HPLC analysis of crude extracts derived from cultures of S. albus transformed with the malacidin BGC shows the presence of BGC-specific small molecules. The two primary malacidin peaks are highlighted with an asterisk. d, Unlike crude extracts of the S. albus host strain alone, only extracts from the S. albus harbouring the malacidin BGC showed antibacterial activity when applied to a lawn of S. aureus USA300. Both the HPLC analysis (c) and antibacterial activity (d) are representative of four independent fermentations. e, Malacidin A and B are cyclic lipopeptides containing eight-amino-acid macrocycles and polyunsaturated lipids. The malacidins do not contain the conserved DXDG motif seen in all known calcium-dependent antibiotics—incorporating a rare 3-hydroxyl aspartic acid (HyAsp, highlighted in violet) and lacking the spacer residue. Biosynthetic enzymes predicted to be involved in the production of non-proteinogenic amino acid (3-methyl aspartic acid, 4-methyl proline and 2,3-diamino 3-methyl propanoic acid) and fatty acid substrates required for the biosynthesis of the maladicins are shown and colour-coded according to their activities. Stereocentres in malacidin that were predicted bioinformatically, as opposed to through chemical and spectroscopic analysis, are denoted with an asterisk.
The malacidins are 10-membered cyclic lipopeptides that differ only by a methylene on the branch at the terminus of their lipid tails. Their peptide cores include four non-proteinogenic amino acids (Fig. 2e). Calcium-dependent antibiotics characterized from culture-based discovery programmes contain larger, 11- to 13-amino-acid rings and completely distinct peptide sequences (Supplementary Fig. 24). The malacidins do not contain the canonical Asp-X-Asp-Gly calcium-binding motif found in known calcium-dependent antibiotics4. They lack the variable spacer residue found in this canonical motif and contain an ASP-OH, suggesting that they either no longer bind calcium or may represent a different calcium-binding motif4,5,6. To determine the requirement of calcium for the antibacterial activity of the malacidins, we tested for antibiosis against methicillin-resistant Staphylococcus aureus (MRSA) across a range of calcium concentrations. In these assays, we saw a clear dependence on calcium for antibiosis, indicating that although the malacidins do not contain a canonical Asp-X-Asp-Gly motif, their antibacterial activity remains calcium-dependent (Fig. 3a). Similar experiments using cations other than calcium showed no antibiosis (Supplementary Fig. 26). The malacidins are broadly active against Gram-positive bacteria including multidrug-resistant pathogens and bacteria resistant to mechanistically diverse, clinically used antibiotics (Table 1 and Supplementary Table 4). As the most common form of Staphylococcus infection occurs on the skin, we elected to test the in vivo efficacy of the malacidins using an animal wound model. Topical administration of malacidin A was successful in sterilizing MRSA-infected wounds in a rat model (Fig. 3b). At 24 and 72 h post infection, malacidin A treatment resulted in no observed bacterial burdens in the wounds. The vehicle-treated controls had an average of 5.5 log and 7.0 log of MRSA at 24 h and 72 h, respectively (Kruskal–Wallis P value <0.0001). Likewise, the malacidins showed no significant toxicity or haemolytic activity against mammalian cells at the highest concentrations tested (100–250 µg ml−1, >100 minimal inhibitory concentration, MIC) (Supplementary Fig. 27 and Supplementary Table 3). Unlike daptomycin, which is unable to treat severe community-acquired pneumonia due to loss of activity in the presence of pulmonary surfactants12, malacidin A does not share this liability (Fig. 3c). Our experimental efforts to induce resistance to malacidin in the laboratory have so far been unsuccessful. Even after 20 days of exposure to sub-lethal levels of malacidin A, we did not detect any malacidin-resistant S. aureus (Fig. 3d). Whether resistance can arise through horizontal gene transfer from environmental bacteria remains to be seen.
a, The MIC of malacidin A against MRSA was assessed at various concentrations of calcium and the antibiosis of malacidin A was found to be calcium-dependent. The error bars represent the standard deviation across two replicates over three independent experiments (n = 6). b, Malacidin A is an effective treatment against MRSA in rat cutaneous wound infections. The error bars represent the standard deviation across replicate wounds (n = 4). c, Unlike daptomycin, malacidin A activity against S. pneumoniae is largely unaffected by the presence of pulmonary surfactants. The error bars represent the standard deviation across two replicates over three independent experiments (n = 6). d, After 20 days of repeated exposure to 0.5 × MIC of malacidin A (Mal.), we did not detect any malacidin-resistant S. aureus. Vancomycin (Van.), daptomycin (Dap.) and rifamycin (Rif.) were used as controls in this assay. The error bars represent the standard deviation across three replicates for MIC determination (n = 3).
Characterized calcium-dependent antibiotics function by one of two distinct modes of action (Fig. 4a). Daptomycin displays rapid bactericidal activity by binding cytoplasmic membrane phosopholipids and oligomerizing in the membrane13,14. This affects phospholipid synthesis and overall membrane fluidity, ultimately leading to decreased membrane integrity and cell death15. The potent antibacterial activity of friulimicin and its structural relatives is due to inhibition of bacterial cell wall biosynthesis through binding of the lipid II precursor, undecaprenyl phosphate (C55-P)16,17. As the malacidins are structurally distinct from other calcium-dependent antibiotics, we sought to determine whether they function by one of these known mechanisms or a third distinct mode of action. We first assessed the effect of malacidin on membrane integrity (Fig. 4b and Supplementary Fig. 28). No membrane leakage was observed when S. aureus cells pretreated with SYTOX green or DiBAC4 were exposed to either daptomycin or malacidin in the absence of calcium supplementation. With the addition of calcium, daptomycin-treated S. aureus showed a rapid increase in fluorescence, which is indicative of a loss of membrane integrity. Malacidin, however, did not demonstrate the same effect, indicating that malacidin and daptomycin have distinct modes of action.
a, Schematic diagram showing modes of action of daptomycin, friulimicin and malacidin. b, In contrast to daptomycin, malacidin A does not cause MRSA membrane leakage in a SYTOX green fluorescent assay. The error bars represent the standard deviation across three biological replicates (n = 3). c, As seen with the cell wall biosynthesis inhibitor vancomycin, exposing MRSA to malacidin A results in the accumulation of the cell wall intermediate UDP-MurNAc-pentapeptide. The UDP-MurNAc-pentapeptide peak ([M–H]– = 1148.35) is indicated with a red asterisk on the UPLC-MS trace. The chromatograms are representative of at least three independent experiments. d, Interaction of malacidin A and daptomycin with purified cell wall precursors. An interaction is indicated by a reduction of the amount of free antibiotic (visible on the TLC by ultraviolet light). e, The interaction of malacidin A with cell wall precursor, lipid II, is calcium-dependent. Both TLCs in d and e are representative of at least two replicate experiments.
As seen with friulimicin and other cell-wall-intermediate binding antibiotics, S. aureus treated with malacidin accumulates the cell wall precursor undecaprenyl-N-acetylmuramic acid-pentapeptide (UDP-MurNAc-pentapeptide) (Fig. 4c)16,17. This signalled that the target of malacidin, like that of friulimicin, lies downstream of UDP-MurNAc-pentapeptide formation. Surprisingly, in a thin-layer chromatography (TLC) mobility shift assay, malacidin did not sequester C55-P, the target of friulimicin (Fig. 4d)16,17. Lipid II is the key downstream intermediate of MurNAc-pentapeptide. We therefore tested malacidin for lipid-II-binding activity. In this TLC-based mobility shift assay, we observed lipid-II-dependent disappearance of the malacidin band (Fig. 4d,e). Unlike previously characterized calcium-dependent antibiotics, malacidin neither depolarizes the membrane nor binds C55-P but instead appears to interact with lipid II in a calcium-dependent manner. Fortuitously, despite the fact that vancomycin also binds lipid II, the malacidins are active against both vancomycin-intermediate- and vancomycin-resistant pathogens.
The malacidins exhibited potent antibacterial activity against Gram-positive pathogens resistant to clinically used antibiotics, including the antibiotic of last resort vancomycin, and did not select for resistance in the laboratory under the conditions of our experiments. The discovery of the malacidins supports our hypothesis that the calcium-dependent antibiotics are a larger than previously thought family of NPs with low susceptibility to resistance and diverse modes of action. Environmental microbes are in a continuous antibiotic arms race that is likely to select for antibiotic variants capable of circumventing existing resistance mechanisms. The sequence-guided metagenomic discovery pipeline outlined here provides a means to interrogate complex environmental metagenomes for these uncharacterized antibiotics by tracking NPSTs that differ from those associated with known antibiotic BGCs. While metagenome-based antibiotic discovery methods are still in their infancy, the scaling and automation of the pipeline described here should permit the systematic discovery of NP antibiotics that have until now remained hidden in the global metagenome, providing a potentially powerful approach for combating antibiotic resistance.
Methods
NPST generation and sequencing
To add to the diversity of the 185 previously collected soil samples8,18, an additional 1,800 soils were collected for this study from sites throughout the United States. Crude eDNA was extracted from each of these following established protocols10,18. Briefly, 25 g of soil was heated (70 °C) in lysis buffer (100 mM Tris-HCl, 100 mM EDTA, 1.5 M NaCl, 1% (w/v) CTAB, 2% (w/v) SDS, pH 8.0) for 2 h. Soil particulates were removed from the crude lysate by centrifugation, and eDNA was precipitated from the resulting supernatant with the addition of 0.7 volumes of isopropanol. Crude eDNA was collected by centrifugation, washed with 70% ethanol and resuspended in TE. Crude eDNA was then spin-column-purified (PowerMax soil DNA kit) and employed as a template in PCR experiments targeting ADs as follows: AD fragments (~795 bp) were amplified using primers: 5′-GCSTACSYSATSTACACSTCSGG-3′ and 5′-SASGTCVCCSGTSCGGTA-3′. These primers are designed to recognize the conserved regions in NRPS ADs8,18. The 5′ ends of the primers were augmented with MiSeq sequencing adapters followed by unique 8 bp barcode sequences identifying the soil metagenome from which they were amplified. PCR conditions: 12 μl reaction, 1 × Buffer G (Epicentre), 50 pmol of each primer, 2.5 units Omni Klentaq polymerase (DNA Polymerase Technology) and 100 ng eDNA. Cycle conditions for AD amplification: 95 °C 4 min, (95 °C 30 s, 63.5 °C 30 s, 72 °C 45 s) x 34 cycles, 72 °C 5 min. First-round amplicons contained incomplete Illumina adaptors and therefore required a second round of PCR to append the remainder of the adaptor sequence19. Amplicons were pooled as collections of 96 samples and cleaned using Agencourt Ampure XP magnetic beads (Beckman Coulter). Cleaned, pooled amplicons were used as a template in a second 20-μl PCR using the following reaction conditions: 10 μl of FailSafe Buffer G (Epicentre), 5.8 μl of water, 0.4 μl of each primer (100 μM) (MiSeqForward, CAAGCAGAAGACGGCATACGAGATGTGACTGGAGTTCAGACGTGTGCTCTTCCGATCT; MiSeq Reverse AATGATACGGCGACCACCGAGATCTACACTCTTTCCCTACACGACGCTCTTCCGATCT), 0.4 μl of Taq and 3 μl of cleaned amplicon (50 ng to 100 ng). Amplification proceeded as follows: 95 °C for 5 min, 6 cycles of 95 °C for 30 s, 70 °C for 30 s and 72 °C for 45 s, and, finally, 72 °C for 5 min. Prior to sequencing, all PCR amplicons were quantified by gel electrophoresis and mixed in an equal molar ratio. The resulting pool was fluorometrically quantified with an HS D1000 ScreenTape (Agilent 2200 TapeStation; Agilent Technologies) and sequenced on an Illumina MiSeq instrument using Reagent Kit v3 (MS-102-3003, Illumina).
Biosynthetic profiling of NPSTs
Amplicon sequences were analysed and organized using the eSNaPD (environmental Surveyor of Natural Product Diversity) web-based tool as previously described8,9,18. The NRPS ADs from sequenced and known calcium-dependent antibiotic gene clusters (daptomycin, friulimicin, CDA, laspartomycin, A54145 and taromycin) were added to the eSNaPD reference database of domains from annotated and functionally characterized natural product gene clusters. NPSTs whose closest relatives among all reference ADs were one of these known lipopeptide domains were identified and mapped to soil locations and/or library wells. This analysis, in brief, was completed as previously described19 by debarcoding samples using a paired-end 2 × 8 bp barcode strategy. Debarcoded reads were filtered for quality and 240 bp of the forward reads, a single `N' spacer and 175 bp of the reverse-complemented reverse read were concatenated to generate a synthetic amplicon of 416 bp. The reads from each sample were clustered using UCLUST20 to generate the 95% identity centroid sequences (that is, NPST). Location information was used to map NPSTs back to soil collection locations and/or library wells. The forward read component (the first 240 bp) of each NPST was then searched using BlastN21 against a manually curated database of NRPS AD sequences. NPSTs that returned one of the known calcium-dependent antibiotics as a top match were considered hits. The resulting set of unique hits was used to generate geographic and phylogenetic distribution figures. A multiple sequence alignment of all sequences was generated using MUSCLE22, and the resulting alignment file was used to generate a maximum-likelihood tree with FastTree23.
Construction and arraying of metagenomic cosmid libraries
eDNA cosmid libraries were constructed from soil using established protocols10. Briefly, crude eDNA was isolated from ~0.5 kg of soil as outlined above, and further purified by preparative agarose gel electrophoresis to yield pure high-molecular-weight eDNA. High-molecular-weight eDNA was blunt-ended (Epicentre, End-It), ligated into pWEB-TNC (Epicentre), packaged into lambda phage and transfected into E. coli EC100 (Epicentre). Following recovery, transfected cells were inoculated into 8 ml LB with selective antibiotic (12.5 μg ml−1 chloramphenicol) in 24-well plates at a density of ~25,000 clones per well and grown overnight. Matching glycerol stocks and cosmid DNA minipreps were prepared from each well, and arrayed as 768 pools of ~25,000 unique cosmid clones. NPST data were prepared from each library pool by amplifying and sequencing ADs as described above.
Recovery of biosynthetic gene clusters from eDNA libraries
Calcium-dependent antibiotic-like NPST sequences identified within metagenomic libraries were automatically assigned to library wells by the barcode parsing functionality of the eSNaPD software package as described above. Specific primers targeting each unique sequence of interest were designed by hand. To recover single clones from library wells, a serial dilution PCR strategy18 was used as follows: library wells containing targets as 1 of ∼25,000 unique cosmids were grown overnight to confluence in LB (12.5 μg ml−1 chloramphenicol, 100 μg ml−1 carbenicillin) and diluted to a concentration of 3,000 colony-forming units (CFUs) ml−1 as judged by OD600 nm. Then, 384-well plates were inoculated with 100 μl (300 CFU) of the resulting dilution per well, grown to confluence, and screened using real-time PCR, to identify wells containing target clones as 1 of ∼300 clones. Target positive wells were then diluted to a concentration of ∼50 CFU ml−1 and the process was repeated to identify wells containing targets as 1 of ∼5 clones. Five clone pools were then plated on solid medium, and target clones were identified by colony PCR.
In silico analysis of recovered gene clusters
Recovered single cosmid clones were pooled and sequenced using ion PGM technology. Reads were assembled into contigs using an assembler program, such as Newbler24. Overlapping cosmids spanning a single pathway were initially sequenced separately and subsequently assembled into larger contigs. Assembled contigs were then annotated using an in-house pipeline consisting of open-reading-frame predictions with MetaGeneMark25, BLAST search21 and AntiSMASH predictions26. The AntiSMASH predictions employ three prediction algorithms to call the amino acid substrate specificity of an adenylation domain (NRPSPredictor2, Stachelhaus code and Minowa). These amino acid predictions were used in the initial bioinformatic characterization of clusters to predict chemical structures. Putative functions for new tailoring enzymes in eDNA pathways were assigned on the basis of the predicted function of the closest characterized relative identified by Blast in NCBI.
Assembly of DFD0097-735 pTARa BAC for heterologous expression
For assembly of the DFD0097-735 pTARa bacterial artificial chromosome (BAC), transformation-associated recombination (TAR) in yeast was employed11,27. Initially, the three overlapping cosmids (DFD0097-644, DFD0097-735 and DFD0097-388) containing the full biosynthetic pathway were digested and linearized with DraI. A custom E. coli:yeast:Streptomyces shuttle capture vector, pTARa, containing two 500 bp homology arms to the terminal overlapping cosmid clones was constructed as previously described11,27. This vector was subsequently linearized with PmeI and gel-purified. The linearized cosmids and capture vector were then co-transformed into Saccharomyces cerevisiae (BY4727 Δdnl4) using a standard LiAc/ss carrier DNA/PEG yeast transformation protocol28. Briefly, yeast were grown overnight in 50 ml of YPD medium containing G418 (200 μg ml−1) at 30 °C. In the morning, 2 ml of the overnight culture was reinoculated into 50 ml of fresh YPD medium containing G418 (200 μg ml−1) and grown for ∼4 h (OD600 nm = 2.0). This culture was harvested by centrifugation (10 min, 3,200g), washed twice with sterile 4 °C water and resuspended in 1 ml of sterile 4 °C water. For each transformation 100 μl of washed cells was transferred to a Microfuge tube. The cells were collected by centrifugation (30 s, 18,000g) and resuspended in a transformation mix containing 36 μl of 1 M LiAc solution, 50 μl of 2 mg ml−1 carrier DNA (salmon sperm DNA) solution, 240 μl of 50% (wt/vol) PEG 3350 solution, and 34 μl of Tris-EDTA containing 2 μg of each cosmid and 1 μg of vector. This transformation mix was incubated at 42 °C for 40 min. Cells were then collected by centrifugation (30 s, 18,000g), resuspended in 100 μl of water and plated on appropriate synthetic composite dropout medium agar plates. Agar plates were incubated at 30 °C until colonies appeared. Colonies were checked by PCR. DNA was isolated from PCR-positive yeast clones, transferred into E. coli ET12567/pUZ8002 cells and then moved into Streptomyces spp. by intergeneric conjugation for heterologous expression.
Heterologous expression
The assembled BAC, DFD0097-735 pTARa and an empty pTARa vector control were separately integrated into the chromosome of Streptomyces albus J1074. Spore suspensions of these recombinant strains were used to seed starter cultures in 50 ml trypticase soy broth (Oxoid). These cultures were grown for 48 h (30 °C/200 rpm) and 0.4 ml of the resulting confluent culture was used to inoculate 50 ml flasks of production medium, R5a medium: 100 g l−1 sucrose, 0.25 g l−1 K2SO4, 10.12 g l−1 MgCl2, 10.0 g l−1 d-glucose, 0.1 g l−1 casamino acids, 21 g l−1 MOPS, 2 g l−1 NaOH, 40 μg l−1 ZnCl2, 20 μg l−1 FeCl3 6H2O, 10 μg l−1 MnCl2, 10 μg l−1 (NH4)6Mo7O24 4H2O. Fifty-millilitre liquid cultures were grown in 125 ml baffled flasks (22 °C, 220 rpm) for 14 days.
Isolation of malacidin A and B
After 14 days, 4 l of cultures were combined, and mycelia were removed by centrifugation at 4,000g for 20 min. The mycelium-free medium supernatant was applied to 150 g of pre-equilibrated Diaion HP-20 resin packed in a column (40 × 220 mm). The HP-20 column was subsequently washed with 2 l of H2O, and then eluted with 2 l of 100% methanol. The methanolic elution was concentrated by rotary evaporation, and then combined with 2 g octadecyl-functionalized silica resin (Sigma-Aldrich) per 10 ml concentrate. This resin/concentrate mixture was dried overnight on a Savant Speedvac Concentrator (Thermo-Fisher). The dried loaded resin was used to dry-load a 100 g Gold HP C18 column for medium-pressure reversed-phase chromatography (Teledyne Isco Combiflash Rf150). This chromatography was performed using a linear gradient of 0.1% acetic acid–acetonitrile from 10% to 100% over 20 min at 60 ml min−1. To identify column fractions containing active compound, aliquots of 10 ml fractions were analysed by ultra-performance liquid chromatography–mass spectrometry (UPLC–MS). Fractions containing the malacidins were pooled, re-dried on resin, and subjected to a second-round medium-pressure, narrow-range reversed-phase chromatography. Using a 100 g Gold HP C18 column, chromatography was performed with a linear gradient of 0.1% acetic acid–acetonitrile from 30% to 60% over 20 min at 60 ml min−1. This enabled the initial separation of malacidin-A-containing and malacidin-B-containing fractions. Combined fractions containing either malacidin A or B were subsequently cleaned up individually using preparative high-performance liquid chromatography (HPLC; XBridge Prep C18, 10 × 150 mM, 5 μM, Agilent HPLC System) using a linear gradient of 0.1% trifloroacetic acid–acetonitrile from 30% to 50% over 30 min at a flow rate of 4 ml min−1. Malacidin A and B had a retention time of 12 min and 16 min, respectively. For analysis of purity and detection of fractions by UPLC–MS throughout the isolation process, 5 μl was injected onto a UPLC–MS system (Waters Corporation) and analysed by a linear gradient of 0.1% formic acid–acetonitrile from 30% to 50% over 3.4 min.
Structural determination by NMR and ESI–MS/MS
See the Supplementary Discussion for details on structural determination. 1H and 13C NMR spectra were obtained at 600 and 150 MHz, respectively, on a Bruker Avance DMX600 NMR. Spectra were taken at 298 K using either 11.21 mM malacidin A or 7.92 mM malacidin B in 3.59 mM triethylamine in D2O, unless otherwise noted. The chemical shifts were referenced to the methyl group of triethylamine in D2O (δC 8.189, δH 1.292). For electrospray ionization with tandem mass spectrometry (ESI–MS/MS), samples in methanol were diluted 1:50 with 50% methanol/0.1% formic acid and infused (5 μl min−1) for analysis by high-resolution (60,000 at m/z 200)/high-mass-accuracy MS, MS2 and MS3 (Fusion Lumos, ThermoFischer Scientific). In addition, each sample was diluted 1:1 with 0.1 M ammonium bicarbonate for propionylation of primary amines followed by a 1:25 dilution in methanol/0.1% formic acid and analysis by ESI–MS, MS2 and MS3. Positive-ion ESI conditions: 3.9 kV, heated capillary set at 300 °C and sheath gas setting of ‘1’. Both ion-trap-based collision-induced dissociation (CID) and beam-type fragmentation (HCD) were used. For fragmentation experiments, ions were isolated using a window of 2.0 m/z.
Determination of absolute configuration of amino acids of malacidins
Malacidin A and B (0.5 mg) were dissolved in 6 N HCl (500 µl) separately and heated at 115 °C for 10 h. For each antibiotic, four separate reactions were set up. After hydrolysis, the reaction mixtures were cooled in ice water for 5 min. The reaction solvent was evaporated in vacuo. The dried reaction was resuspended in 500 µl of water and the water was evaporated in vacuo. This process was repeated three times. The hydrolysates, containing free amino acids, were dissolved in 100 µl of 1 N NaHCO3. Either 100 µl of l-FDAA (1-fluro-2,4-dinitrophenyl-5-L-alanine amide) or 100 µl of d-FDAA in acetone (10 mg ml−1) was added to each of the four vials and they were incubated at 42 °C for 1 h. To neutralize the reaction, 100 µl of 2 N HCl was added to each reaction mixture. Reactions were then diluted with 300 µl of 50% acetonitrile/water. Five microlitres of each reaction mixture was analysed by liquid chromatography–high-resolution mass spectrometry with a gradient solvent system (20%–60% acetonitrile/water with 0.1% formic acid over 40 min; flow rate 0.2 ml min−1) on the RP column (Thermo Acclaim 120, C18 2.1 × 150 mm).
Microbial susceptibility assays
The malacidins were screened against a panel of assay strains and pathogenic bacteria as indicated in Supplementary Table 3. MIC assays were performed in duplicate in 96-well microtiter plates on the basis of the protocol recommended by the Clinical and Laboratory Standards Institute29. All presented data are the average of at least three independent assays. Stock solutions of malacidin or daptomycin (2 mg ml−1 in H2O) were added to the first well in a row and serially diluted (twofold per transfer) across the microtiter plate. CaCl2 was supplemented to media at a final concentration of 15 mM and fetal bovine serum (ATCC) was added to media (1:10) to test the effect of serum. Overnight cultures of bacteria were diluted 5,000-fold, and 50 μl was used as an inoculum in each well. MIC values were determined by visual inspection after 18 h incubation (30 °C, static growth). For the enhanced calcium titration experiments, standard MIC assays were performed with methicillin-resistant Staphylococcus aureus (MRSA) PFGE strain type USA300 in media supplemented with CaCl2 at: 25.0, 18.8, 14.1, 7.03, 3.52, 2.50, 1.76, 0.880, 0.440, 0.250 and 0 mM. To assess the effects of monovalent and divalent cations on malacidin activity, standard MIC assays were performed with MRSA USA300 in media supplemented with 15 mM CaCl2, MgCl2, MnCl2, ZnCl2, SrCl2, NaCl and KCl. To evaluate the effects of pulmonary surfactants on activity against a community-acquired pneumonia-causing pathogen, standard MIC assays were performed with Streptococcus pneumoniae TCH8431 in media supplemented with both 15 mM CaCl2 and Survanta (beractant, Abbvie) at a final concentration of 5, 1, 0.5, 0.25 or 0% (volume/volume (v/v) per cent).
Mammalian cytotoxicity assays
Cytotoxicity against human cell lines was tested using an ATP release assay, CellTiter-Glo (Promega), according to the manufacturer’s instructions. HEK293 cells (293FT, Thermo Fisher Scientific, no. R700-07) and MRC5 cells (ATCC CCL-171) were grown in complete DMEM media supplemented with 10% FBS, and were inspected visually for authentification and tested for mycoplasma contamination using a MycoAlert detection kit (Lonza). Cells were grown to confluence, trypsinized, counted and plated in 384-well cell culture plates at an appropriate density (2,500 cells per well for HEK293 and 1000 cells per well for MRC5). The test compound was added 24 h later in the presence of calcium, and viability was determined after 4.5 h of incubation. Experiments were performed with biological replicates. Haemolytic activity was evaluated by a red blood cell disc diffusion assay. Twenty-microlitre stocks of malacidin A and Triton X-100 were infused on filter discs, dried completely and then overlaid on 5% sheep blood agar plates (Hardy Diagnostics). The plates were incubated for 24 h at 20 °C, and then checked for lysis.
Rat cutaneous wound infection model
Methicillin-resistant Staphylococcus aureus strain MW2 was grown in Mueller Hinton broth at 37°C with shaking overnight. The culture was centrifuged, supernatant aspirated and the bacteria were gently washed once in sterile saline. The optical density was determined at 600 nm. The bacterial suspension was diluted to provide a challenge inoculum of approximately 500 CFU per wound in a volume of 0.05 ml in sterile 0.9% NaCl. The inoculum count was verified by viable counts on Mannitol Salt Agar plates spread with proper dilutions of the inoculum and incubated at 37 °C for 24–48 h. For the wound infection model, 8-week-old male (~200 g) Sprague Dawley rats were given two wounds each. Two rats were used at each time point (day 1 and day 3) for each treatment group for a total of 4 rats (8 wounds) per drug. This sample size was statistically calculated on the basis of previous in-house wound burden studies comparing vehicle-treated groups with antibiotic controls30. Rats were randomly selected into the treatment groups. To generate wounds, the rats were anaesthetized by intraperitoneal injection of 100 mg kg−1 ketamine + 10 mg kg−1 xylazine and the dorsal side of the rats was shaved with electrical clippers and then depilated with Nair. The exposed skin was wiped with betadine. Two symmetrical wounds were made on the dorsum of each rat using a 0.8-cm-diameter disposable biopsy punch. Sterile polyurethane rings serving as wound chambers were placed over the fresh wounds and attached by surgical adhesive. After the wound creation, rats from each group were infected with 0.05 ml of the bacterial suspension for a final infection dose of 500 CFU per wound. Wounds were covered with Tegaderm visible adhesive dressing, and the rats were rehydrated with physiological saline administered via intraperitoneal injection. The analgesic buprenorphine (0.05 mg kg−1) was administered to minimize pain during surgical recovery. At 30 min post infection, rats were given single daily topical treatments of vehicle (25 mM CaCl2 in sterile water), or 0.5 mg malacidin A or daptomycin suspended in 25 mM CaCl2, and the wounds were covered in fresh Tegaderm dressing. At 1 day and 3 days post infection, the rats were humanely euthanized and wounds were excised and assessed for bacterial burdens by plating on MSA. Rats were observed twice daily for morbidity and possible signs of acute toxicity. Abnormal clinical signs were recorded if observed. Rats were housed in the Public Health Research Institute’s Animal Biosafety Level-2 Research Animal Facility (ICPH RAF), a centre of the New Jersey Medical School, Rutgers University (NJMS-Rutgers). The animal facility follows the Public Health Service and National Institute of Health Policy of Humane Care and Use of Laboratory Animals. All experimental protocols were approved by the Rutgers Institutional Animal Care and Use Committee (IACUC).
Selection for malacidin-resistant mutants
To select for resistant mutants, a single MRSA USA300 colony from a freshly struck plate was inoculated into LB media and grown overnight at 37 °C. The saturated overnight culture was diluted 100-fold, supplemented with a sub-lethal dose (0.5× MIC) of malacidin A, vancomycin, daptomycin or rifamycin and 15 mM calcium. Two-hundred-microlitre aliquots were then distributed into microtiter plate wells. The next day, 3 μl of culture from each well was used to inoculate 200 μl of calcium-supplemented media (15 mM calcium) with fresh antibiotic at 0.5× and 4× MIC. This process was repeated for 20 days. In the cases where bacterial growth was observed in the 4× MIC overnight cultures, the resistant culture was plated in successively higher concentrations of antibiotic the following day. This was repeated over the course of the experiment to assess fold change in MIC at day 0 to day 20.
Membrane leakage and depolarization assays
The effects of malacidin on membrane integrity was assessed using SYTOX green. In brief, single colonies of MRSA USA300 were grown in LB media with and without 15 mM CaCl2 to an OD600 nm of 0.35. Nine hundred microlitres of cells were mixed with 100 μl of 17 μM SYTOX green dye (Thermo Fisher). The resulting mixture was incubated for 5 min at 22 °C, and then distributed to a microtiter plate at 50 μl per well. An initial reading of fluorescence pre-antibiotic addition was measured at excitation and emission wavelengths of 488 nm and 523 nm, respectively. Fifty microlitres of antibiotics was added to respective wells at a final concentration of 20 μg ml−1. Measurements were immediately collected for 10 min. To assess the effects of malacidin on membrane depolarization, similar assays were set up using the membrane potential probe, DiBAC4 (bis-(1,3-dibutylbarbituric acid)trimethine oxonol). Single colonies of MRSA USA300 were grown in LB media with and without 15 mM CaCl2 to an OD600 nm of 0.35. Nine hundred microlitres of cells were mixed with 100 μl of 20 μg ml−1 DiBAC4 dye (Thermo Fisher). The resulting mixture was incubated for 5 min at 22 °C, and then distributed to a microtiter plate at 50 μl per well. An initial reading of fluorescence pre-antibiotic addition was measured at excitation and emission wavelengths of 492 nm and 515 nm, respectively. Fifty microlitres of antibiotics was added to respective wells at a final concentration of 20 μg ml−1. Measurements were immediately collected for 10 min. Representative examples from three technical replicates are shown.
UDP-MurNAc-pentapeptide accumulation assay
The intracellular accumulation of the cell wall precursor UDP-MurNAc-pentapeptide after treatment of MRSA USA300 with malacidin was assessed as previously described16. In brief, single colonies of MRSA USA300 were grown in LB media with and without 15 mM CaCl2 to an OD600 nm of 0.6. One microlitre of cells and medium were incubated with a final concentration of 130 μg ml−1 chloramphenicol for 15 min at 37 °C. Antibiotics to be assayed were added at 10 μg ml−1 and incubated for 60 min at 37 °C. Vancomycin, known to form a complex with lipid II, was used as a positive control. Cells were collected by centrifugation, and resuspended in 30 μl dH2O and incubated in boiling water for 15 min. The cell extract was then centrifuged at 14,000g. Supernatant was analysed for UDP-linked cell wall precursors by a UPLC–MS system (Waters Corporation). Experiments were performed with biological replicates.
Complex formation with cell wall precursors
Binding of malacidin to C55-P and lipid II was evaluated by incubating 1 nmol of each purified precursor with 0.5 nmol of malacidin or daptomycin in 100 mM Tris-HCl, pH 7.5, 0.1% Triton X-100, 13 mM MgCl2, and with or without 25 mM CaCl2, for 60 min at 37 °C. Subsequently, the mixture was extracted twice with n-BuOH/6 M pyridinium acetate buffer, pH 4.2 (3:1, v/v). The butanol fraction was evaporated and the residue was dissolved in CHCl3/methanol (1:1, v/v). The resuspension was analysed for the loss of unbound malacidin or daptomycin to a complex by TLC analysis using chloroform/methanol/water/ammonia (88:48:10:1, v/v/v/v) as the solvent and detection by 254/366 nm visualization. Experiments were performed with biological replicates.
Life Sciences Reporting Summary
Further information on experimental design is available in the Life Sciences Reporting Summary.
Data availability
The DNA sequence to the malacidin gene cluster has been deposited as GenBank Accession KY654519. The eSNaDP2.0 software is available at http://esnapd2.rockefeller.edu/. The remaining data that support the findings of this study are available from the corresponding author upon request.
References
O’Neill, J. Tackling Drug-Resistant Infections Globally: Final Report and Recommendations (The Review on Antimicrobial Resistance, Wellcome Trust & UK Government, 2016).
Tringe, S. G. et al. Comparative metagenomics of microbial communities. Science 308, 554–557 (2005).
Reddy, B. V. et al. Natural product biosynthetic gene diversity in geographically distinct soil microbiomes. Appl. Environ. Microbiol. 78, 3744–3752 (2012).
Strieker, M. & Marahiel, M. A. The structural diversity of acidic lipopeptide antibiotics. ChemBioChem 10, 607–616 (2009).
Jung, D., Rozek, A., Okon, M. & Hancock, R. E. Structural transitions as determinants of the action of the calcium-dependent antibiotic daptomycin. Chem. Biol. 11, 949–957 (2004).
Bunkoczi, G., Vertesy, L. & Sheldrick, G. M. Structure of the lipopeptide antibiotic tsushimycin. Acta Crystallogr. Sect. D. 61, 1160–1164 (2005).
Katz, M., Hover, B. M. & Brady, S. F. Culture-independent discovery of natural products from soil metagenomes. J. Ind. Microbiol. Biotechnol. 43, 129–141 (2016).
Owen, J. G. et al. Mapping gene clusters within arrayed metagenomic libraries to expand the structural diversity of biomedically relevant natural products. Proc. Natl Acad. Sci. USA 110, 11797–11802 (2013).
Reddy, B. V., Milshteyn, A., Charlop-Powers, Z. & Brady, S. F. eSNaPD: a versatile, web-based bioinformatics platform for surveying and mining natural product biosynthetic diversity from metagenomes. Chem. Biol. 21, 1023–1033 (2014).
Brady, S. F. Construction of soil environmental DNA cosmid libraries and screening for clones that produce biologically active small molecules. Nat. Protoc. 2, 1297–1305 (2007).
Kim, J. H. et al. Cloning large natural product gene clusters from the environment: piecing environmental DNA gene clusters back together with TAR. Biopolymers 93, 833–844 (2010).
Silverman, J. A., Mortin, L. I., Vanpraagh, A. D., Li, T. & Alder, J. Inhibition of daptomycin by pulmonary surfactant: in vitro modeling and clinical impact. J. Infect. Dis. 191, 2149–2152 (2005).
Zhang, T., Taylor, S. D., Palmer, M. & Duhamel, J. Membrane binding and oligomerization of the lipopeptide A54145 studied by pyrene fluorescence. Biophys. J. 111, 1267–1277 (2016).
Straus, S. K. & Hancock, R. E. Mode of action of the new antibiotic for Gram-positive pathogens daptomycin: comparison with cationic antimicrobial peptides and lipopeptides. Biochim. Biophys. Acta 1758, 1215–1223 (2006).
Muller, A. et al. Daptomycin inhibits cell envelope synthesis by interfering with fluid membrane microdomains. Proc. Natl Acad. Sci. USA 113, E7077–E7086 (2016).
Schneider, T. et al. The lipopeptide antibiotic Friulimicin B inhibits cell wall biosynthesis through complex formation with bactoprenol phosphate. Antimicrob. Agents Ch. 53, 1610–1618 (2009).
Kleijn, L. H. et al. Total synthesis of Laspartomycin C and characterization of its antibacterial mechanism of action. J. Med. Chem. 59, 3569–3574 (2016).
Owen, J. G. et al. Multiplexed metagenome mining using short DNA sequence tags facilitates targeted discovery of epoxyketone proteasome inhibitors. Proc. Natl Acad. Sci. USA 112, 4221–4226 (2015).
Charlop-Powers, Z. et al. Urban park soil microbiomes are a rich reservoir of natural product biosynthetic diversity. Proc. Natl Acad. Sci. USA 113, 14811–14816 (2016).
Edgar, R. C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26, 2460–2461 (2010).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Price, M. N., Dehal, P. S. & Arkin, A. P. FastTree: computing large minimum evolution trees with profiles instead of a distance matrix. Mol. Biol. Evol. 26, 1641–1650 (2009).
Zhang, T., Luo, Y., Chen, Y., Li, X. & Yu, J. BIGrat: a repeat resolver for pyrosequencing-based re-sequencing with Newbler. BMC Res. Notes 5, 567 (2012).
Zhu, W., Lomsadze, A. & Borodovsky, M. Ab initio gene identification in metagenomic sequences. Nucleic Acids Res. 38, e132 (2010).
Medema, M. H. et al. antiSMASH: rapid identification, annotation and analysis of secondary metabolite biosynthesis gene clusters in bacterial and fungal genome sequences. Nucleic Acids Res. 39, W339–W346 (2011).
Kallifidas, D. & Brady, S. F. Reassembly of functionally intact environmental DNA-derived biosynthetic gene clusters. Methods Enzymol. 517, 225–239 (2012).
Gietz, R. D. & Schiestl, R. H. Large-scale high-efficiency yeast transformation using the LiAc/SS carrier DNA/PEG method. Nat. Protoc. 2, 38–41 (2007).
Cockerill, F. R. Methods for Dilution Antimicrobial Susceptibility Tests for Bacteria that Grow Aerobically; Approved Standard—Ninth edition (Clinical and Laboratory Standards Institute, Wayne, PA, 2012).
Zhao, Y. et al. Carbohydrate-derived fulvic acid is a highly promising topical agent to enhance healing of wounds infected with drug-resistant pathogens. J. Trauma Acute Care Surg. 79, S121–S129 (2015).
Acknowledgements
We thank F. Rubino and D. Kahne for discussion and an aliquot of lipid II. HEK293 cells and MRC5 cells were kindly provided by the High-throughput Screening Resource Center at the Rockefeller University. This work was supported in part by a grant from the Gates Foundation and NIH U19AI109713. B.M.H. was supported by NIH Grant F32 AI124479. Z.C.-P. was supported by NIH Grant F32 AI1100029.
Author information
Authors and Affiliations
Contributions
B.M.H. and S.F.B. designed research; B.M.H., S.-H.K., M.K., J.G.O., M.A.T., J.M., A.E. and H.M. performed research; B.M.H., S.-H.K. and Z.C.-P. analysed data; and B.M.H. and S.F.B. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The following authors, S.F.B., B.M.H., M.K., Z.C.-P., declare competing financial interests as they are employees or consultants of Lodo Therapeutics.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Discussion, Supplementary Tables 1–4, Supplementary Figures 1–29 and Supplementary References.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hover, B.M., Kim, SH., Katz, M. et al. Culture-independent discovery of the malacidins as calcium-dependent antibiotics with activity against multidrug-resistant Gram-positive pathogens. Nat Microbiol 3, 415–422 (2018). https://doi.org/10.1038/s41564-018-0110-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-018-0110-1
This article is cited by
-
Geministatins: new depside antibiotics from the fungus Austroacremonium gemini
The Journal of Antibiotics (2024)
-
Metagenomic domain substitution for the high-throughput modification of nonribosomal peptides
Nature Chemical Biology (2024)
-
Metagenomics- Paving the Path for Sustainable Circular Bioeconomy
Circular Economy and Sustainability (2024)
-
The overlooked bacterial pandemic
Seminars in Immunopathology (2024)
-
Formation of zinc oxide composites of doxycycline with high antibacterial activity based on DC-magnetron deposition of ZnO nanoscale particles on the drug surface
Applied Physics A (2024)