Abstract
The ability of populations to expand their geographical ranges, whether as invaders, agricultural strains or climate migrants, is currently one of the most serious global problems. However, fundamental mechanisms remain poorly understood regarding factors that enable certain populations, such as biological invaders, to rapidly transition to novel habitats. According to one hypothesis, environmental fluctuations in the native range could promote successful invasions by imposing balancing selection on key traits and maintaining the genetic variation that enables rapid adaptation in novel habitats. Here we test the genomic predictions of this hypothesis by performing whole-genome sequencing of multiple independent invasive freshwater and native saline populations of the copepod Eurytemora affinis complex. We found that invasive populations have repeatedly responded to selection through the parallel use of the same single-nucleotide polymorphisms and genomic loci, to a much greater degree than expected. These same loci were enriched for signatures of long-term balancing selection in the native ranges, with 15–47% of loci exhibiting significant signatures of balancing selection. The strong association between parallel evolution in the invaded range and balancing selection in the native range supports the hypothesis that fluctuating habitats can promote invasive success and that balancing selection might serve as a widespread and important mechanism that enables rapid adaptation in nature.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Raw sequence data and aligned reads have been deposited in the NCBI Sequence Read Archive under BioProject ID PRJNA610547.
Code availablility
Custom scripts used throughout this analysis are available online at https://github.com/TheDBStern/NEE2020.
References
Paini, D. R. et al. Global threat to agriculture from invasive species. Proc. Natl Acad. Sci. USA 113, 7575–7579 (2016).
Bradshaw, C. J. A. et al. Massive yet grossly underestimated global costs of invasive insects. Nat. Commun. 7, 12986 (2016).
Bellard, C. et al. Will climate change promote future invasions? Glob. Change Biol. 19, 3740–3748 (2013).
Hellmann, J. J., Byers, J. E., Bierwagen, B. G. & Dukes, J. S. Five potential consequences of climate change for invasive species. Conserv. Biol. 22, 534–543 (2008).
Chown, S. L. et al. Biological invasions, climate change and genomics. Evol. Appl. 8, 23–46 (2015).
Rahel, F. J. & Olden, J. D. Assessing the effects of climate change on aquatic invasive species. Conserv. Biol. 22, 521–533 (2008).
Chapman, D. S. et al. Modelling the introduction and spread of non-native species: international trade and climate change drive ragweed invasion. Glob. Change Biol. 22, 3067–3079 (2016).
Dam, H. G. Evolutionary adaptation of marine zooplankton to global change. Annu. Rev. Mar. Sci. 5, 349–370 (2013).
Blackburn, T. M. et al. A proposed unified framework for biological invasions. Trends Ecol. Evol. 26, 333–339 (2011).
Lee, C. E. Evolutionary genetics of invasive species. Trends Ecol. Evol. 17, 386–391 (2002).
Lee, C. E. in Encyclopedia of Biological Invasions (eds Simberloff, D. & Rejmanek, M.) 215–222 (Univ. California Press 2010).
Colautti, R. I., Alexander, J. M., Dlugosch, K. M., Keller, S. R. & Sultan, S. E. Invasions and extinctions through the looking glass of evolutionary ecology. Phil. Trans. R. Soc. B 372, 20160031 (2017).
Williamson, M. & Fitter, A. The varying success of invaders. Ecology 77, 1661–1666 (1996).
Hayes, K. R. & Barry, S. C. Are there any consistent predictors of invasion success? Biol. Invasions 10, 483–506 (2008).
Bock, D. G. et al. What we still don’t know about invasion genetics. Mol. Ecol. 24, 2277–2297 (2015).
Kolar, C. S. & Lodge, D. M. Progress in invasion biology: predicting invaders. Trends Ecol. Evol. 16, 199–204 (2001).
Lee, C. E. & Bell, M. A. Causes and consequences of recent freshwater invasions by saltwater animals. Trends Ecol. Evol. 14, 284–288 (1999).
Casties, I., Seebens, H. & Briski, E. Importance of geographic origin for invasion success: a case study of the North and Baltic seas versus the Great Lakes–St. Lawrence River region. Ecol. Evol. 6, 8318–8329 (2016).
Lee, C. E. & Gelembiuk, G. W. Evolutionary origins of invasive populations. Evol. Appl. 1, 427–448 (2008).
Winkler, G., Dodson, J. J. & Lee, C. E. Heterogeneity within the native range: population genetic analyses of sympatric invasive and noninvasive clades of the freshwater invading copepod Eurytemora affinis. Mol. Ecol. 17, 415–430 (2008).
Barrett, R. D. H. & Schluter, D. Adaptation from standing genetic variation. Trends Ecol. Evol. 23, 38–44 (2008).
Lee, C. E. Evolutionary mechanisms of habitat invasions, using the copepod Eurytemora affinis as a model system. Evol. Appl. 9, 248–270 (2016).
Peischl, S. & Excoffier, L. Expansion load: recessive mutations and the role of standing genetic variation. Mol. Ecol. 24, 2084–2094 (2015).
Messer, P. W., Ellner, S. P. & Hairston, N. G. Can population genetics adapt to rapid evolution? Trends Genet. 32, 408–418 (2016).
Lewontin, R. The Genetic Basis of Evolutionary Change (Columbia Univ. Press, 1974).
Crow, J. F. Muller, Dobzhansky, and overdominance. J. Hist. Biol. 20, 351–380 (1987).
Beatty, J. Weighing the risks: stalemate in the classical/balance controversy. J. Hist. Biol. 20, 289–319 (1987).
Asthana, S., Schmidt, S. & Sunyaev, S. A limited role for balancing selection. Trends Genet. 21, 30–32 (2005).
Muller, H. J. Our load of mutations. Am. J. Hum. Genet. 2, 111–176 (1950).
Dobzhansky, T. A review of some fundamental concepts and problems of population genetics. Cold Spring Harb. Symp. Quant. Biol. 20, 1–15 (1955).
Hedrick, P. W. Genetic variation in a heterogeneous environment. I. Temporal heterogeneity and absolute dominance model. Genetics 78, 757–770 (1974).
Dempster, E. R. Maintenance of genetic heterogeneity. Cold Spring Harb. Symp. Quant. Biol. 20, 25–32 (1955).
Haldane, J. B. S. & Jayakar, S. D. Polymorphism due to selection of varying direction. J. Genet. 58, 237–242 (1963).
Gillespie, J. Polymorphism in random environments. Theor. Popul. Biol. 4, 193–195 (1973).
Felsenstein, J. Theoretical population genetics of variable selection and migration. Annu. Rev. Genet. 10, 253–280 (1976).
Holt, R. D., Barfield, M. & Gomulkiewicz, R. Temporal variation can facilitate niche evolution in harsh sink environments. Am. Nat. 164, 187–200 (2004).
Huang, Y., Tran, I. & Agrawal, A. F. Does genetic variation maintained by environmental heterogeneity facilitate adaptation to novel selection? Am. Nat. 188, 27–37 (2016).
de Filippo, C. et al. Recent selection changes in human genes under long-term balancing selection. Mol. Biol. Evol. 33, 1435–1447 (2016).
Hermisson, J. & Pennings, P. S. Soft sweeps: molecular population genetics of adaptation from standing genetic variation. Genetics 169, 2335–2352 (2005).
Hermisson, J. & Pennings, P. S. Soft sweeps and beyond: understanding the patterns and probabilities of selection footprints under rapid adaptation. Methods Ecol. Evol. 8, 700–716 (2017).
Jensen, J. D. On the unfounded enthusiasm for soft selective sweeps. Nat. Commun. 5, 6281 (2014).
Ralph, P. L. & Coop, G. The role of standing variation in geographic convergent adaptation. Am. Nat. 186, S5–S23 (2015).
Brennan, R. S., Garrett, A. D., Huber, K. E., Hargarten, H. & Pespeni, M. H. Rare genetic variation and balanced polymorphisms are important for survival in global change conditions. Proc. R. Soc. B 286, 20190943 (2019).
Mallard, F., Nolte, V., Tobler, R., Kapun, M. & Schlotterer, C. A simple genetic basis of adaptation to a novel thermal environment results in complex metabolic rewiring in Drosophila. Genome Biol. 19, 119 (2018).
Kelly, J. K. & Hughes, K. A. Pervasive linked selection and intermediate-frequency alleles are implicated in an evolve-and-resequencing experiment of Drosophila simulans. Genetics 211, 943–961 (2019).
Stern, D. L. The genetic causes of convergent evolution. Nat. Rev. Genet. 14, 751–764 (2013).
Lee, K. M. & Coop, G. Population genomics perspectives on convergent adaptation. Phil. Trans. R. Soc. B 374, 20180236 (2019).
Waldvogel, A. M. et al. Evolutionary genomics can improve prediction of species’ responses to climate change. Evol. Lett. 4, 4–18 (2019).
Lee, C. E. Rapid and repeated invasions of fresh water by the copepod Eurytemora affinis. Evolution 53, 1423–1434 (1999).
Katajisto, T. Copepod eggs survive a decade in the sediments of the Baltic Sea. Hydrobiologia 320, 153–159 (1996).
Ban, S. & Minoda, T. Hatching of diapause eggs of Eurytemora affinis (Copepoda: Calanoida) collected from lake-bottom sediments. J. Crustac. Biol. 12, 51–56 (1992).
Glippa, O., Denis, L., Lesourd, S. & Souissi, S. Seasonal fluctuations of the copepod resting egg bank in the middle Seine estuary, France: impact on the nauplii recruitment. Estuar. Coast. Mar. Sci. 142, 60–67 (2014).
Posavi, M., Gelembiuk, G. W., Larget, B. & Lee, C. E. Testing for beneficial reversal of dominance during salinity shifts in the invasive copepod Eurytemora affinis, and implications for the maintenance of genetic variation. Evolution 68, 3166–3183 (2014).
Lee, C. E. Global phylogeography of a cryptic copepod species complex and reproductive isolation between genetically proximate “populations”. Evolution 54, 2014–2027 (2000).
Pickrell, J. K. & Pritchard, J. K. Inference of population splits and mixtures from genome-wide allele frequency data. PLoS Genet. 8, e1002967 (2012).
Foll, M., Gaggiotti, O. E., Daub, J. T., Vatsiou, A. & Excoffier, L. Widespread signals of convergent adaptation to high altitude in Asia and America. Am. J. Hum. Genet. 95, 394–407 (2014).
Foll, M. & Gaggiotti, O. A genome-scan method to identify selected loci appropriate for both dominant and codominant markers: a Bayesian perspective. Genetics 180, 977–993 (2008).
Gautier, M. Genome-wide scan for adaptive divergence and association with population-specific covariates. Genetics 201, 1555–1579 (2015).
Jones, F. C. et al. The genomic basis of adaptive evolution in threespine sticklebacks. Nature 484, 55–61 (2012).
Alberto, F. J. et al. Convergent genomic signatures of domestication in sheep and goats. Nat. Commun. 9, 813 (2018).
Gerber, L. et al. The legs have it: in situ expression of ion transporters V-Type H+ ATPase and Na+/K+-ATPase in osmoregulatory leg organs of the invading copepod Eurytemora affinis. Physiol. Biochem. Zool. 89, 233–250 (2016).
Johnson, K. E., Perreau, L., Charmantier, G., Charmantier-Daures, M. & Lee, C. E. Without gills: localization of osmoregulatory function in the copepod Eurytemora affinis. Physiol. Biochem. Zool. 87, 310–324 (2014).
Lee, C. E., Kiergaard, M., Gelembiuk, G. W., Eads, B. D. & Posavi, M. Pumping ions: rapid parallel evolution of ionic regulation following habitat invasions. Evolution 65, 2229–2244 (2011).
Siewert, K. M. & Voight, B. F. Detecting long-term balancing selection using allele frequency correlation. Mol. Biol. Evol. 34, 2996–3005 (2017).
Bitarello, B. D. et al. Signatures of long-term balancing selection in human genomes. Genome Biol. Evol. 10, 939–955 (2018).
Siewert, K. M. & Voight, B. F. BetaScan2: standardized statistics to detect balancing selection utilizing substitution data. Genome Biol. Evol. 12, 3873–3877 (2020).
Tajima, F. Statistical method for testing the neutral mutation hypothesis by DNA polymorphism. Genetics 123, 585–595 (1989).
Bolnick, D. I., Barrett, R. D. H., Oke, K. B., Rennison, D. J. & Stuart, Y. E. (Non)parallel evolution. Ann. Rev. Ecol. Evol. Syst. 49, 303–330 (2018).
Hedrick, P. W., Ginevan, M. E. & Ewing, E. P. Genetic polymorphism in heterogeneous environments. Ann. Rev. Ecol. Syst. 7, 1–32 (1976).
Wittmann, M. J., Bergland, A. O., Feldman, M. W., Schmidt, P. S. & Petrov, D. A. Seasonally fluctuating selection can maintain polymorphism at many loci via segregation lift. Proc. Natl Acad. Sci. USA 114, E9932–E9941 (2017).
Bergland, A. O., Behrman, E. L., O’Brien, K. R., Schmidt, P. S. & Petrov, D. A. Genomic evidence of rapid and stable adaptive oscillations over seasonal time scales in Drosophila. PLoS Genet. 10, e1004775 (2014).
Troth, A., Puzey, J. R., Kim, R. S., Willis, J. H. & Kelly, J. K. Selective trade-offs maintain alleles underpinning complex trait variation in plants. Science 361, 475–478 (2018).
Siepielski, A. M., DiBattista, J. D. & Carlson, S. M. It’s about time: the temporal dynamics of phenotypic selection in the wild. Ecol. Lett. 12, 1261–1276 (2009).
Thurman, T. J. & Barrett, R. D. H. The genetic consequences of selection in natural populations. Mol. Ecol. 25, 1429–1448 (2016).
Gelembiuk, G. W., May, G. E. & Lee, C. E. Phylogeography and systematics of zebra mussels and related species. Mol. Ecol. 15, 1033–1050 (2006).
May, G. E., Gelembiuk, G. W., Panov, V. E., Orlova, M. I. & Lee, C. E. Molecular ecology of zebra mussel invasions. Mol. Ecol. 15, 1021–1031 (2006).
Bertram, J. & Masel, J. Different mechanisms drive the maintenance of polymorphism at loci subject to strong versus weak fluctuating selection. Evolution 73, 883–896 (2019).
Chen, J., Nolte, V. & Schlotterer, C. Temperature stress mediates decanalization and dominance of gene expression in Drosophila melanogaster. PLoS Genet. 11, e1004883 (2015).
Panov, V. E., Krylov, P. I. & Riccardi, N. Role of diapause in dispersal and invasion success by aquatic invertebrates. J. Limnol. 63, 56–69 (2004).
Lee, C. E. & Frost, B. W. Morphological stasis in the Eurytemora affinis species complex (Copepoda: Temoridae). Hydrobiologia 480, 111–128 (2002).
Alekseev, V. R. & Souissi, A. A new species within the Eurytemora affinis complex (Copepoda: Calanoida) from the Atlantic Coast of USA, with observations on eight morphologically different European populations. Zootaxa 2767, 41–56 (2011).
Smit, A. F. A., Hubley, R. & Green, P. RepeatMasker Open-4.0 (2013–2015); http://www.repeatmasker.org
Eyun, S. I. et al. Evolutionary history of chemosensory-related gene families across the Arthropoda. Mol. Biol. Evol. 34, 1838–1862 (2017).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Sedlazeck, F. J., Rescheneder, P. & von Haeseler, A. NextGenMap: fast and accurate read mapping in highly polymorphic genomes. Bioinformatics 29, 2790–2791 (2013).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Kofler, R., Pandey, R. V. & Schlotterer, C. PoPoolation2: identifying differentiation between populations using sequencing of pooled DNA samples (Pool-Seq). Bioinformatics 27, 3435–3436 (2011).
Hivert, V., Leblois, R., Petit, E. J., Gautier, M. & Vitalis, R. Measuring genetic differentiation from Pool-Seq data. Genetics 210, 315–330 (2018).
Kofler, R. et al. PoPoolation: a toolbox for population genetic analysis of next generation sequencing data from pooled individuals. PLoS ONE 6, e0015925 (2011).
Revell, L. J. phytools: an R package for phylogenetic comparative biology (and other things). Methods Ecol. Evol. 3, 217–223 (2012).
Ives, A. R., Midford, P. E. & Garland, T. Within-species variation and measurement error in phylogenetic comparative methods. Syst. Biol. 56, 252–270 (2007).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Neph, S. et al. BEDOPS: high-performance genomic feature operations. Bioinformatics 28, 1919–1920 (2012).
Lee, E. et al. Web Apollo: a web-based genomic annotation editing platform. Genome Biol. 14, R93 (2013).
Kofler, R. & Schlotterer, C. Gowinda: unbiased analysis of gene set enrichment for genome-wide association studies. Bioinformatics 28, 2084–2085 (2012).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate—a practical and powerful approach to multiple testing. J. R. Stat. Soc. B 57, 289–300 (1995).
Foley, B. R.et al. A gene-based SNP resource and linkage map for the copepod Tigriopus californicus. BMC Genom. 12, 568 (2011).
Dymowska, A. K., Hwang, P. P. & Goss, G. G. Structure and function of ionocytes in the freshwater fish gill. Respir. Physiol. Neurobiol. 184, 282–292 (2012).
Lee, C. E. & Petersen, C. H. Effects of developmental acclimation on adult salinity tolerance in the freshwater-invading copepod Eurytemora affinis. Physiol. Biochem. Zool. 76, 296–301 (2003).
Lee, C. E., Remfert, J. L. & Chang, Y. M. Response to selection and evolvability of invasive populations. Genetica 129, 179–192 (2007).
Lee, C. E., Remfert, J. L. & Gelembiuk, G. W. Evolution of physiological tolerance and performance during freshwater invasions. Integr. Comp. Biol. 43, 439–449 (2003).
Ellner, S. & Sasaki, A. Patterns of genetic polymorphism maintained by fluctuating selection with overlapping generations. Theor. Popul. Biol. 50, 31–65 (1996).
Turelli, M. & Barton, N. H. Polygenic variation maintained by balancing selection: pleiotropy, sex-dependent allelic effects and GxE interactions. Genetics 166, 1053–1079 (2004).
Turelli, M., Schemske, D. W. & Bierzychudek, P. Stable two-allele polymorphisms maintained by fluctuating fitnesses and seed banks: protecting the blues in Linanthus parryae. Evolution 55, 1283–1298 (2001).
Ellner, S. & Hairston, N. G. Role of overlapping generations in maintaining genetic variation in a fluctuating environment. Am. Nat. 143, 403–417 (1994).
Wright, S. Physiological and evolutionary theories of dominance. Am. Nat. 68, 24–53 (1934).
Curtsinger, J. W., Service, P. M. & Prout, T. Antagonistic pleiotropy, reversal of dominance, and genetic polymorphism. Am. Nat. 144, 210–228 (1994).
Gulisija, D., Kim, Y. & Plotkin, J. B. Phenotypic plasticity promotes balanced polymorphism in periodic environments by a genomic storage effect. Genetics 202, 1437–1448 (2016).
Gulisija, D. & Plotkin, J. B. Phenotypic plasticity promotes recombination and gene clustering in periodic environments. Nat. Commun. 8, 2041 (2017).
Acknowledgements
This research was funded by National Science Foundation grants OCE-1046372 and OCE-1658517 to C.E.L. and the Michael Guyer Postdoctoral Fellowship to D.B.S. We thank J. Pool, S. Schoville and N. Sharpe for comments on the manuscript. The samples used in this project were collected by M. Bontrager. Computation was performed using the computational resources and assistance of the UW-Madison Center for High Throughput Computing (CHTC) in the Department of Computer Sciences.
Author information
Authors and Affiliations
Contributions
C.E.L. contributed to the design of the study and data collection. D.B.S. contributed to the conception and execution of the analyses. Both authors contributed to the writing and approval of the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
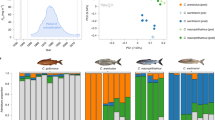
Extended Data Fig. 1 β(2) and NCD2 score density plot for the native, saline Baie de L’Isle Verte population (St. Lawrence drainage, Atlantic clade).
Higher β(2) scores and lower NCD2 scores signify stronger signatures of long-term balancing selection. β(2) scores are higher and NCD2 scores are lower on average for ‘parallel’ candidate SNPs (blue) relative to the rest of the genome (grey).
Extended Data Fig. 2 β(2) and NCD2 score density plot for the native, saline Taylor Bayou (Gulf of Mexico, Gulf clade).
Higher β(2) scores and lower NCD2 scores signify stronger signatures of long-term balancing selection. β(2) scores are higher and NCD2 scores are lower on average for ‘parallel’ candidate SNPs (blue) relative to the rest of the genome (grey).
Supplementary information
Supplementary Information
Supplementary Box 1, Materials and Methods.
Supplementary Tables
Supplementary Tables 1–12.
Rights and permissions
About this article
Cite this article
Stern, D.B., Lee, C.E. Evolutionary origins of genomic adaptations in an invasive copepod. Nat Ecol Evol 4, 1084–1094 (2020). https://doi.org/10.1038/s41559-020-1201-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-020-1201-y
This article is cited by
-
Balancing selection on an MYB transcription factor maintains the twig trichome color variation in Melastoma normale
BMC Biology (2023)
-
Rapid evolution of phenotypic plasticity in patchy habitats
Scientific Reports (2023)
-
Epigenetic plasticity enables copepods to cope with ocean acidification
Nature Climate Change (2022)
-
Genome-wide signatures of synergistic epistasis during parallel adaptation in a Baltic Sea copepod
Nature Communications (2022)
-
Rapid genetic adaptation to recently colonized environments is driven by genes underlying life history traits
BMC Genomics (2021)