Abstract
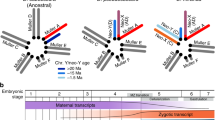
Widespread loss of genes on the Y is considered a hallmark of sex chromosome differentiation. Here we show that the initial stages of Y evolution are driven by massive amplification of distinct classes of genes. The neo-Y chromosome of Drosophila miranda initially contained about 3,000 protein-coding genes, but has gained over 3,200 genes since its formation about 1.5 million years ago primarily by tandem amplification of protein-coding genes ancestrally present on this chromosome. We show that distinct evolutionary processes may account for this drastic increase in gene number on the Y. Testis-specific and dosage-sensitive genes appear to have amplified on the Y to increase male fitness. A distinct class of meiosis-related multi-copy Y genes independently co-amplified on the X, and their expansion is probably driven by conflicts over segregation. Co-amplified X/Y genes are highly expressed in testis, enriched for meiosis and RNA interference functions and are frequently targeted by small RNAs in testis. This suggests that their amplification is driven by X versus Y antagonism for increased transmission, where sex chromosome drive suppression is probably mediated by sequence homology between the suppressor and distorter through the RNA interference mechanism. Thus, our analysis suggests that newly emerged sex chromosomes are a battleground for sexual and meiotic conflict.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All data that were used and generated for this project are given in Supplementary Table 10 and have been deposited on NCBI under BioProject ID PRJNA545539.
References
Bachtrog, D. et al. Are all sex chromosomes created equal? Trends Genet. 27, 350–357 (2011).
Charlesworth, B. Model for evolution of Y chromosomes and dosage compensation. Proc. Natl Acad. Sci. USA 75, 5618–5622 (1978).
Bachtrog, D. Y-chromosome evolution: emerging insights into processes of Y-chromosome degeneration. Nat. Rev. Genet. 14, 113–124 (2013).
Mahajan, S. & Bachtrog, D. Convergent evolution of Y chromosome gene content in flies. Nat. Commun. 8, 785 (2017).
Bellott, D. W. et al. Avian W and mammalian Y chromosomes convergently retained dosage-sensitive regulators. Nat. Genet. 49, 387–394 (2017).
Gatti, M. & Pimpinelli, S. Functional elements in Drosophila melanogaster heterochromatin. Annu. Rev. Genet. 26, 239–275 (1992).
Blackmon, H., Ross, L. & Bachtrog, D. Sex determination, sex chromosomes, and karyotype evolution in insects. J. Hered. 108, 78–93 (2017).
Hughes, J. F. et al. Conservation of Y-linked genes during human evolution revealed by comparative sequencing in chimpanzee. Nature 437, 100–103 (2005).
Soh, Y. Q. S. et al. Sequencing the mouse Y chromosome reveals convergent gene acquisition and amplification on both sex chromosomes. Cell 159, 800–813 (2014).
Bachtrog, D. & Charlesworth, B. Reduced adaptation of a non-recombining neo-Y chromosome. Nature 416, 323–326 (2002).
Muller, H. J. in The New Systematics (ed. Huxley, J.) 185–268 (Clarendon Press, 1940).
Dobzhansky, T. Drosophila miranda, a new species. Genetics 20, 377–391 (1935).
Zhou, Q. & Bachtrog, D. Sex-specific adaptation drives early sex chromosome evolution in Drosophila. Science 337, 341–345 (2012).
Mahajan, S., Wei, K. H.-C., Nalley, M. J., Gibilisco, L. & Bachtrog, D. De novo assembly of a young Drosophila Y chromosome using single-molecule sequencing and chromatin conformation capture. PLoS Biol. 16, e2006348 (2018).
Carvalho, A. B., Lazzaro, B. P. & Clark, A. G. Y chromosomal fertility factors kl-2 and kl-3 of Drosophila melanogaster encode dynein heavy chain polypeptides. Proc. Natl Acad. Sci. USA 97, 13239–13244 (2000).
Bachtrog, D., Hom, E., Wong, K. M., Maside, X. & de Jong, P. Genomic degradation of a young Y chromosome in Drosophila miranda. Genome Biol. 9, R30 (2008).
Charlesworth, B. & Charlesworth, D. The degeneration of Y chromosomes. Phil. Trans. R. Soc. Lond. B 355, 1563–1572 (2000).
Kurtz, S. et al. Versatile and open software for comparing large genomes. Genome Biol. 5, R12 (2004).
Lucotte, E. A. et al. Dynamic copy number evolution of X- and Y-linked ampliconic genes in human populations. Genetics 209, 907–920 (2018).
Bachtrog, D. Evidence that positive selection drives Y-chromosome degeneration in Drosophila miranda. Nat. Genet. 36, 518–522 (2004).
Meiklejohn, C. D. & Tao, Y. Genetic conflict and sex chromosome evolution. Trends Ecol. Evol. (Amst.) 25, 215–223 (2010).
Konkel, M. K. & Batzer, M. A. A mobile threat to genome stability: the impact of non-LTR retrotransposons upon the human genome. Semin. Cancer Biol. 20, 211–221 (2010).
Zhou, Q. et al. The epigenome of evolving Drosophila neo-sex chromosomes: dosage compensation and heterochromatin formation. PLoS Biol. 11, e1001711 (2013).
Bachtrog, D. Expression profile of a degenerating neo-Y chromosome in Drosophila. Curr. Biol. 16, 1694–1699 (2006).
Ellison, C. E. & Bachtrog, D. Dosage compensation via transposable element mediated rewiring of a regulatory network. Science 342, 846–850 (2013).
Lucchesi, J. C. & Kuroda, M. I. Dosage compensation in Drosophila. Cold Spring Harb. Perspect. Biol. 7, a019398 (2015).
Alekseyenko, A. A. et al. Conservation and de novo acquisition of dosage compensation on newly evolved sex chromosomes in Drosophila. Genes Dev. 27, 853–858 (2013).
Rice, W. R. Sex chromosomes and the evolution of sexual dimorphism. Evolution 38, 735–742 (1984).
Skaletsky, H. et al. The male-specific region of the human Y chromosome is a mosaic of discrete sequence classes. Nature 423, 825–837 (2003).
Bellott, D. W. et al. Mammalian Y chromosomes retain widely expressed dosage-sensitive regulators. Nature 508, 494–499 (2014).
Cortez, D. et al. Origins and functional evolution of Y chromosomes across mammals. Nature 508, 488–493 (2014).
Jaenike, J. Sex chromosome meiotic drive. Annu. Rev. Ecol. Syst. 32, 25–49 (2001).
Frank, S. A. Divergence of meiotic drive-suppression systems as an explanation for sex-biased hybrid sterility and inviability. Evolution 45, 262–267 (1991).
Tao, Y., Masly, J. P., Araripe, L., Ke, Y. & Hartl, D. L. A sex-ratio meiotic drive system in Drosophila simulans. I: an autosomal suppressor. PLoS Biol. 5, e292 (2007).
Tao, Y. et al. A sex-ratio meiotic drive system in Drosophila simulans. II: an X-linked distorter. PLoS Biol. 5, e293 (2007).
Lin, C.-J. et al. The hpRNA/RNAi pathway is essential to resolve intragenomic conflict in the Drosophila male germline. Dev. Cell 46, 316–326.e5 (2018).
Brashear, W. A., Raudsepp, T. & Murphy, W. J. Evolutionary conservation of Y chromosome ampliconic gene families despite extensive structural variation. Genome Res. 28, 1841–1851 (2018).
Bellott, D. W. et al. Convergent evolution of chicken Z and human X chromosomes by expansion and gene acquisition. Nature 466, 612–616 (2010).
Carvalho, A. B., Dobo, B. A., Vibranovski, M. D. & Clark, A. G. Identification of five new genes on the Y chromosome of Drosophila melanogaster. Proc. Natl Acad. Sci. USA 98, 13225–13230 (2001).
Sturgill, D., Zhang, Y., Parisi, M. & Oliver, B. Demasculinization of X chromosomes in the Drosophila genus. Nature 450, 238–241 (2007).
Assis, R., Zhou, Q. & Bachtrog, D. Sex-biased transcriptome evolution in Drosophila. Genome Biol. Evol. 4, 1189–1200 (2012).
Meiklejohn, C. D., Landeen, E. L., Cook, J. M., Kingan, S. B. & Presgraves, D. C. Sex chromosome-specific regulation in the Drosophila male germline but little evidence for chromosomal dosage compensation or meiotic inactivation. PLoS Biol. 9, e1001126 (2011).
Vibranovski, M. D., Zhang, Y. & Long, M. General gene movement off the X chromosome in the Drosophila genus. Genome Res. 19, 897–903 (2009).
Mueller, J. L. et al. The mouse X chromosome is enriched for multicopy testis genes showing postmeiotic expression. Nat. Genet. 40, 794–799 (2008).
Mueller, J. L. et al. Independent specialization of the human and mouse X chromosomes for the male germ line. Nat. Genet. 45, 1083–1087 (2013).
Helleu, Q. et al. Rapid evolution of a Y-chromosome heterochromatin protein underlies sex chromosome meiotic drive. Proc. Natl Acad. Sci. USA 113, 4110–4115 (2016).
Hurst, L. D. & Pomiankowski, A. Causes of sex ratio bias may account for unisexual sterility in hybrids: a new explanation of Haldane’s rule and related phenomena. Genetics 128, 841–858 (1991).
Larson, E. L., Keeble, S., Vanderpool, D., Dean, M. D. & Good, J. M. The composite regulatory basis of the large x-effect in mouse speciation. Mol. Biol. Evol. 34, 282–295 (2017).
Phadnis, N. & Orr, H. A. A single gene causes both male sterility and segregation distortion in Drosophila hybrids. Science 323, 376–379 (2009).
Lahn, B. T. & Page, D. C. A human sex-chromosomal gene family expressed in male germ cells and encoding variably charged proteins. Hum. Mol. Genet. 9, 311–319 (2000).
Cocquet, J. et al. A genetic basis for a postmeiotic X versus Y chromosome intragenomic conflict in the mouse. PLoS Genet. 8, e1002900 (2012).
Cocquet, J. et al. The multicopy gene sly represses the sex chromosomes in the male mouse germline after meiosis. PLoS Biol. 7, e1000244 (2009).
Larson, E. L., Kopania, E. E. K. & Good, J. M. Spermatogenesis and the evolution of mammalian sex chromosomes. Trends Genet. 34, 722–732 (2018).
Balakireva, M. D., Shevelyov, Yu. Ya, Nurminsky, D. I., Livak, K. J. & Gvozdev, V. A. Structural organization and diversification of Y-linked sequences comprising Su(Ste) genes in Drosophila melanogaster. Nucleic Acids Res. 20, 3731–3736 (1992).
Murphy, W. J. et al. Novel gene acquisition on carnivore Y chromosomes. PLoS Genet. 2, e43 (2006).
Ellison, C. & Bachtrog, D. Recurrent gene co-amplification on Drosophila X and Y chromosomes. PLoS Genet. 15, e1008251 (2019).
Smith, A., Hubley, R. & Green, P. RepeatMasker Open-4.0 (Institute for Systems Biology, accessed 30 August 2017); www.repeatmasker.org
Smith, A. & Hubley, R. RepeatModeler Open-1.0. RepeatMasker Open-4.0 (Institute for Systems Biology, accessed 30 August 2017); www.repeatmasker.org
Chu, C., Nielsen, R. & Wu, Y. REPdenovo: inferring de novo repeat motifs from short sequence reads. PLoS ONE 11, e0150719 (2016).
Bao, W., Kojima, K. K. & Kohany, O. Repbase update, a database of repetitive elements in eukaryotic genomes. Mob. DNA 6, 11 (2015).
Kim, D., Langmead, B. & Salzberg, S. L. HISAT: a fast spliced aligner with low memory requirements. Nat. Methods 12, 357–360 (2015).
Pertea, M. et al. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat. Biotechnol. 33, 290–295 (2015).
Campbell, M. S., Holt, C., Moore, B. & Yandell, M. Genome annotation and curation using MAKER and MAKER-P. Curr. Protoc. Bioinformatics 48, 4.11.1–39 (2014).
Stanke, M. & Waack, S. Gene prediction with a hidden Markov model and a new intron submodel. Bioinformatics 19 (Suppl. 2), ii215–ii225 (2003).
Korf, I. Gene finding in novel genomes. BMC Bioinformatics 5, 59 (2004).
Simão, F. A., Waterhouse, R. M., Ioannidis, P., Kriventseva, E. V. & Zdobnov, E. M. BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 31, 3210–3212 (2015).
Camacho, C. et al. BLAST+: architecture and applications. BMC Bioinformatics 10, 421 (2009).
Gel, B. & Serra, E. karyoploteR: an R/Bioconductor package to plot customizable genomes displaying arbitrary data. Bioinformatics 33, 3088–3090 (2017).
Guy, L., Kultima, J. R. & Andersson, S. G. E. genoPlotR: comparative gene and genome visualization in R. Bioinformatics 26, 2334–2335 (2010).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Li, H. et al. The sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
DePristo, M. A. et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat. Genet. 43, 491–498 (2011).
Danecek, P. et al. The variant call format and VCFtools. Bioinformatics 27, 2156–2158 (2011).
Huson, D. H. & Bryant, D. Application of phylogenetic networks in evolutionary studies. Mol. Biol. Evol. 23, 254–267 (2006).
Buchfink, B., Xie, C. & Huson, D. H. Fast and sensitive protein alignment using DIAMOND. Nat. Methods 12, 59–60 (2015).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014).
Bray, N. L., Pimentel, H., Melsted, P. & Pachter, L. Near-optimal probabilistic RNA-seq quantification. Nat. Biotechnol. 34, 525–527 (2016).
Eden, E., Navon, R., Steinfeld, I., Lipson, D. & Yakhini, Z. GOrilla: a tool for discovery and visualization of enriched GO terms in ranked gene lists. BMC Bioinformatics 10, 48 (2009).
Supek, F., Bošnjak, M., Škunca, N. & Šmuc, T. REVIGO summarizes and visualizes long lists of gene ontology terms. PLoS ONE 6, e21800 (2011).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with bowtie 2. Nat. Methods 9, 357–359 (2012).
Ramírez, F., Dündar, F., Diehl, S., Grüning, B. A. & Manke, T. deepTools: a flexible platform for exploring deep-sequencing data. Nucleic Acids Res. 42, W187–191 (2014).
Quinlan, A. R. BEDTools: the Swiss-army tool for genome feature analysis. Curr. Protoc. Bioinformatics 47, 11.12.1–34 (2014).
Acknowledgements
D.B. was funded by NIH grants (nos. R01GM076007, GM101255 and R01GM093182). We thank L. Gibilisco for generating small RNA libraries and K. Chatla and A. Tran for generating genomic libraries.
Author information
Authors and Affiliations
Contributions
D.B. conceived and oversaw the project, generated and analysed data and wrote the manuscript. S.M. analysed data. R.B. generated and analysed data.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Phylogenetic relationships of co-amplified X/Y genes in D. miranda.
Maximum-likelihood trees of D. miranda X and Y gene copies with nodes showing >70 bootstrap support highlighted with black circles. X-linked copies are shown in red, Y-linked copies shown in blue, with distinct X and Y groupings collapsed. Fasta alignments are in Data Supplement 14.
Extended Data Fig. 2 Copy number estimates for co-amplified Y genes, multi-copy Y genes, and multi-copy autosome and X genes.
For co-amplified Y genes we show all genes that were identified as co-amplified. For the multi-copy Y genes we only show genes with >3 copies on the Y. For multi-copy autosome and X genes we show only genes with >4 total copies. Multi-copy autosome and X estimates are predicted to be highly similar given that the autosome and X background in each Y-chromosome replacement line is nearly identical. Slight deviations are probably due to stochasticity in sequencing and read mapping, residual heterozygosity in the MSH22 line, or unique Y-chromosome gene amplifications.
Extended Data Fig. 3 Dosage compensation status of neo-X homologs of single-copy (left) and multi-copy (right) Y genes.
Shown are the relative numbers of neo-X genes that are bound by the MSL-complex (and are thus dosage compensated), and those not bound (and thus not dosage compensated). MSL-binding data were generated for male D. miranda larvae. Genes with multiple copies on the Y are less likely to be dosage compensated on the X. The data are presented in Data Supplement 15.
Supplementary information
Supplementary Information
Supplementary Figs. 1–8 and Tables 4 and 6–10.
Supplementary Table 1
Gene loss on the different Muller elements of D. miranda. Shown are genes in D. pseudoobscura for different chromosomes that are absent on their homologous chromosome arm in D. miranda. The table gives D. pseudoobscura gene ID (FBgn), D. melanogaster orthologue, the chromosomal location of the gene in D. pseudoobscura, the tissue of highest expression in D. pseudoobscura, the category of the gene, and whether the gene is present somewhere else in the D. miranda genome. a, Genes missing from the neo-X chromosome. b, Genes missing from the neo-Y chromosome. c, Genes missing from Muller A-AD (X chromosome). d, Genes missing from Muller B (chr4). e, Genes missing from Muller E (chr2). f, Genes that moved between chromosomes.
Supplementary Table 2
Overview of multi-copy Y genes. a, Shown are total copy numbers for multi-copy Y genes, as well as the number of full-length (>90%) and partial Y copies (50–90%; 25–50%; and less than 25% compared to the length of the orthologous gene in D. pseudoobscura). b, Expression of orthologues of multi-copy Y genes in D. pseudoobscura. c, GO analysis for orthologous genes in D. melanogaster. No significant GO enrichment terms were detected.
Supplementary Table 3
Overview of co-amplified X/Y genes. a, Shown are total copy number for co-amplified X and Y genes, as well as the numbers of full-length (>90%) and partial X and Y copies (50–90%; 25–50%; and less than 25% compared to the length of the orthologous gene in D. pseudoobscura). b, Expression of orthologues of co-amplified X/Y genes in D. pseudoobscura. c, Expression of orthologues of co-amplified X/Y genes in D. melanogaster. d, GO analysis for orthologous genes in D. melanogaster. Shown are GO terms that were significantly enriched among co-amplified X/Y genes (using either Gorilla or PantherDB).
Supplementary Table 5
Clustering of multi-copy Y and co-amplified X/Y genes. a, Genome intervals of clustered (within 100 kb of each other) multi-copy genes and their D. pseudoobscura gene ID (FBgn). b, Genome intervals of clustered (within 100 kb of each other) co-amplified genes and their D. pseudoobscura gene ID (FBgn).
Supplementary Dataset 1
Repeat library used for masking the D. miranda genome.
Supplementary Dataset 2
Repeat annotation of the D. miranda genome.
Supplementary Dataset 3
Gene annotation (all genes) of the D. miranda genome.
Supplementary Dataset 4
Gene annotation of multi-copy Y genes and their orthologues in the D. miranda genome.
Supplementary Dataset 5
Gene annotation of co-amplified X and Y genes in the D. miranda genome.
Supplementary Dataset 6
Gene expression values (TPM) for multi-copy Y genes, and co-amplified Y genes, in different D. miranda male tissues, and tissue-specificity index tau.
Supplementary Dataset 7
Gene copy numbers for co-amplified X/Y genes.
Supplementary Dataset 8
Gene expression values (TPM) for co-amplified X and Y gene families, in different D. miranda male tissues.
Supplementary Datast 9
Gene expression values (FPKM) for orthologues of co-amplified X/Y genes in different D. pseudoobscura male and female tissues.
Supplementary Dataset 10
Sense and anti-sense testis total RNA summed counts for co-amplified genes.
Supplementary Dataset 11
Sense and anti-sense testis small RNA summed counts for co-amplified genes.
Supplementary Dataset 12
Testis total RNA counts for all copies of a gene family for different categories of genes on the X/neo-X and Y/neo-Y chromosome.
Supplementary Dataset 13
Testis small RNA raw counts for different categories of genes on the X/neo-X and Y/neo-Y chromosome.
Supplementary Dataset 14
Fasta alignment of co-amplified X/Y genes and their D. pseudoobscura orthologue.
Supplementary Dataset 15
MSL ChIP and Input counts (normalized to library size) for neo-X genes whose homologue is classified as single-copy or multi-copy Y/neo-Y.
Supplementary Dataset 16
Gene expression values (TPM) for single-copy Y genes, multi-copy Y genes, and co-amplified Y genes, and their neo-X/X homologues in different D. miranda tissues. Expression values for all copies of a gene family are shown individually.
Supplementary Dataset 17
Gene expression values (TPM) for multi-copy Y genes, and co-amplified X and Y genes in different D. miranda tissues. Expression values for all copies of a gene family are summed.
Supplementary Dataset 18
Gene expression values (FPKM) for orthologues of co-amplified X/Y genes in different D. melanogaster tissues.
Rights and permissions
About this article
Cite this article
Bachtrog, D., Mahajan, S. & Bracewell, R. Massive gene amplification on a recently formed Drosophila Y chromosome. Nat Ecol Evol 3, 1587–1597 (2019). https://doi.org/10.1038/s41559-019-1009-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-019-1009-9