Abstract
Sex chromosomes are susceptible to the evolution of selfish meiotic drive elements that bias transmission and distort progeny sex ratios. Conflict between such sex-ratio drivers and the rest of the genome can trigger evolutionary arms races resulting in genetically suppressed ‘cryptic’ drive systems. The Winters cryptic sex-ratio drive system of Drosophila simulans comprises a driver, Distorter on the X (Dox) and an autosomal suppressor, Not much yang, a retroduplicate of Dox that suppresses via production of endogenous small interfering RNAs (esiRNAs). Here we report that over 22 Dox-like (Dxl) sequences originated, amplified and diversified over the ~250,000-year history of the three closely related species, D. simulans, D. mauritiana and D. sechellia. The Dxl sequences encode a rapidly evolving family of protamines. Dxl copy numbers amplified by ectopic exchange among euchromatic islands of satellite DNAs on the X chromosome and separately spawned four esiRNA-producing suppressors on the autosomes. Our results reveal the genomic consequences of evolutionary arms races and highlight complex interactions among different classes of selfish DNAs.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Sandler, L. & Novitski, E. Meiotic drive as an evolutionary force. Am. Nat. 91, 105–110 (1957).
Lindholm, A. K. et al. The ecology and evolutionary dynamics of meiotic drive. Trends Ecol. Evol. 31, 315–326 (2016).
Lyttle, T. W. Segregation distorters. Annu. Rev. Genet. 25, 511–557 (1991).
Lyttle, T. W. Cheaters sometimes prosper: distortion of Mendelian segregation by meiotic drive. Trends Genet. 9, 205–210 (1993).
Presgraves, D. C. in Sperm Biology: An Evolutionary Perspective (eds Birkhead, T. R. et al.) Ch. 12, 472–506 (Elsevier Press, 2008).
Hartl, D. L. Genetic dissection of segregation distortion. I. Suicide combinations of SD genes. Genetics 76, 477–486 (1974).
Charlesworth, B. & Hartl, D. L. Population dynamics of the segregation distorter polymorphism of Drosophila melanogaster. Genetics 89, 171–192 (1978).
Hamilton, W. D. Extraordinary sex ratios. Science 156, 477–488 (1967).
Hurst, L. D. & Pomiankowski, A. Causes of sex ratio bias may account for unisexual sterility in hybrids: a new explanation of Haldane’s rule and related phenomena. Genetics 128, 841–858 (1991).
Frank, S. H. Divergence of meiotic drive-suppressors as an explanation for sex-biased hybrid sterility and inviability. Evolution 45, 262–267 (1991).
Gershenson, S. A new sex ratio abnormality in Drosophila obscura. Genetics 13, 488–507 (1928).
Fisher, R. A. The Genetical Theory of Natural Selection (Oxford Univ. Press, 1930).
Jaenike, J. Sex chromosome meiotic drive. Annu. Rev. Ecol. Syst. 32, 25–49 (2001).
Vaz, S. C. & Carvalho, A. B. Evolution of autosomal suppression of the sex-ratio trait in Drosophila. Genetics 166, 265–277 (2004).
Hall, D. W. Meiotic drive and sex chromosome cycling. Evolution 58, 925–931 (2004).
Hartl, D. L. Modifier theory and meiotic drive. Theor. Popul. Biol. 7, 168–174 (1975).
Thomson, G. J. & Feldman, M. W. Population genetics of modifiers of meiotic drive. II. Linkage modification in the Segregation Distorter system. Theor. Popul. Biol. 5, 155–162 (1974).
Bastide, H., Gerard, P. R., Ogereau, D., Cazemajor, M. & Montchamp-Moreau, C. Local dynamics of a fast-evolving sex-ratio system in Drosophila simulans. Mol. Ecol. 22, 5352–5367 (2013).
Haig, D. & Grafen, A. Genetic scrambling as a defence against meiotic drive. J. Theor. Biol. 153, 531–558 (1991).
Burt, A. & Trivers, R. A. Genes in Conflict (Harvard Univ. Press, 2006).
Meiklejohn, C. D. & Tao, Y. Genetic conflict and sex chromosome evolution. Trends Ecol. Evol. 25, 215–223 (2010).
Bachtrog, D. The Y chromosome as a battleground for intragenomic conflict. Trends Genet. 36, 510–522 (2020).
Tao, Y. et al. A sex-ratio meiotic drive system in Drosophila simulans. II: An X-linked distorter. Public Libr. Sci. Biol. 5, e293 (2007).
Tao, Y., Masly, J. P., Araripe, L., Ke, Y. & Hartl, D. L. A sex-ratio meiotic drive system in Drosophila simulans. I: An autosomal suppressor. Public Libr. Sci. Biol. 5, e292 (2007).
Lin, C. J. et al. The hpRNA/RNAi pathway is essential to resolve intragenomic conflict in the Drosophila male germline. Dev. Cell 46, 316–326 (2018).
Kingan, S. B., Garrigan, D. & Hartl, D. L. Recurrent selection on the Winters sex-ratio genes in Drosophila simulans. Genetics 184, 253–265 (2010).
Meiklejohn, C. D. et al. Gene flow mediates the role of sex chromosome meiotic drive during complex speciation. eLife 7, e35468 (2018).
Nolte, V., Pandey, R. V., Kofler, R. & Schlotterer, C. Genome-wide patterns of natural variation reveal strong selective sweeps and ongoing genomic conflict in Drosophila mauritiana. Genome Res. 23, 99–110 (2013).
Garrigan, D., Kingan, S. B., Geneva, A. J., Vedanayagam, J. P. & Presgraves, D. C. Genome diversity and divergence in Drosophila mauritiana: multiple signatures of faster X evolution. Genome Biol. Evol. 6, 2444–2458 (2014).
Tao, Y., Hartl, D. L. & Laurie, C. C. Sex-ratio segregation distortion associated with reproductive isolation in Drosophila. Proc. Natl Acad. Sci. USA 98, 13183–13188 (2001).
Chakraborty, M. et al. Evolution of genome structure in the Drosophila simulans species complex. Genome Res. 31, 380–396 (2021).
Sproul, J. S. et al. Dynamic evolution of euchromatic satellites on the X chromosome in Drosophila melanogaster and the simulans clade. Mol. Biol. Evol. 37, 2241–2256 (2020).
Joshi, S. S. & Meller, V. H. Satellite repeats identify X chromatin for dosage compensation in Drosophila melanogaster males. Curr. Biol. 27, 1393–1402 (2017).
Garrigan, D. et al. Genome sequencing reveals complex speciation in the Drosophila simulans clade. Genome Res. 22, 1499–1511 (2012).
Kliman, R. M. et al. The population genetics of the origin and divergence of the Drosophila simulans complex species. Genetics 156, 1913–1931 (2000).
Miller, D., Brinkworth, M. & Iles, D. Paternal DNA packaging in spermatozoa: more than the sum of its parts? DNA, histones, protamines and epigenetics. Reproduction 139, 287–301 (2010).
Gingell, L. F. & McLean, J. R. A protamine knockdown mimics the function of Sd in Drosophila melanogaster. G3 10, 2111–2115 (2020).
Larracuente, A. M. & Presgraves, D. C. The selfish Segregation Distorter complex of Drosophila melanogaster. Genetics 192, 33–53 (2012).
Wu, C.-I., Lyttle, T. W., Wu, M.-L. & Lin, G. F. Association between DNA satellite sequences and the responder of Segregation Distorter in D. melanogaster. Cell 54, 179–189 (1988).
Thomas, J., Phillips, C. D., Baker, R. J. & Pritham, E. J. Rolling-circle transposons catalyze genomic innovation in a mammalian lineage. Genome Biol. Evol. 6, 2595–2610 (2014).
Hurst, L. D. Is Stellate a relict meiotic driver? Genetics 130, 229–230 (1992).
Cocquet, J. et al. The multicopy gene Sly represses the sex chromosomes in the male mouse germline after meiosis. PLoS Genet. 7, e1000244 (2009).
Kruger, A. N. et al. A neofunctionalized X-linked ampliconic gene family is essential for male fertility and equal sex ratio in mice. Curr. Biol. 29, 3699–3706 (2019).
Hu, W. et al. A large gene family in fission yeast encodes spore killers that subvert Mendel’s law. eLife 6, e26057 (2017).
Eickbush, M. T., Young, J. M. & Zanders, S. E. Killer meiotic drive and dynamic evolution of the wtf gene family. Mol. Biol. Evol. 36, 1201–1214 (2019).
Vogan, A. A. et al. Combinations of Spok genes create multiple meiotic drivers in Podospora. eLife 8, e46454 (2019).
Derome, N., Metayer, K., Montchamp-Moreau, C. & Veuille, M. Signature of selective sweep associated with the evolution of sex-ratio drive in Drosophila simulans. Genetics 1166, 1357–1366 (2004).
Presgraves, D. C., Gerard, P. R., Cherukuri, A. & Lyttle, T. W. Large-scale selective sweep among Segregation Distorter chromosomes in African populations of Drosophila melanogaster. PLoS Genet. 5, e1000463 (2009).
Nam, K. et al. Extreme selective sweeps independently targeted the X chromosomes of the great apes. Proc. Natl Acad. Sci. USA 112, 6413–6418 (2015).
Aravin, A. A. et al. Double-stranded RNA-mediated silencing of genomic tandem repeats and transposable elements in the D. melanogaster germline. Curr. Biol. 11, 1017–1027 (2001).
Daugherty, M. D. & Zanders, S. E. Gene conversion generates evolutionary novelty that fuels genetic conflicts. Curr. Opin. Genet. Dev. 58–59, 49–54 (2019).
Beckmann, J. F., Sharma, G. D., Mendez, L., Chen, H. & Hochstrasser, M. The Wolbachia cytoplasmic incompatibility enzyme CidB targets nuclear import and protamine-histone exchange factors. eLife 8, e50026 (2019).
Acknowledgements
This work was supported by funds from NIH grant no. R01 GM123194 and the University of Rochester to D.C.P. We thank J. J. Emerson, A. Larracuente, C. Meiklejohn, K. Montooth and J. Vedanayagam for early access to the PacBio-based genome assemblies. And we thank B. Navarro Dominguez and C. Meiklejohn for valuable feedback on the manuscript.
Author information
Authors and Affiliations
Contributions
C.A.M. and D.C.P. conceived and designed the study. C.A.M. performed all analyses. C.A.M. and D.C.P. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Ecology & Evolution thanks Aaron Vogan and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
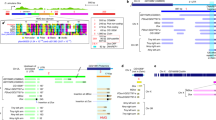
Extended Data Fig. 1 Schematic alignment of inserted sat359 islands in X:9.4-10.4 Mb, colour-coded by putative sequence homology.
Segments of the same colour aligned vertically are high-confidence nucleotide alignments, whereas segments of different colour do not share sequence homology regardless of vertical alignment in the figure. One insertion into a sat359 island, the transposase-like sequence inserted into the D. sechellia Dxl-2 location, does not share sequence homology with any of the other loci, and has been omitted from this figure.
Extended Data Fig. 2 Alignment of the putative source material for Dxl genes found at approximate coordinates X:17.1 and X:17.2 Mb.
Segments of the same colour aligned vertically are high-confidence nucleotide alignments. At X:17.2, the three D. simulans clade species all have remnants of an insertion (total span = ~3 kb) that interrupts the CG8664 gene. This sequence, along with some of CG8664 and additional material, is alignable with insertion at the second position, X:17.1 Mb. Putative homology of the inserted sequence at both locations, along with CG8664, are colour-coded and the span of present-day Dxl homology is indicated.
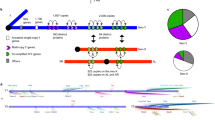
Extended Data Fig. 3 Chromosomal distribution of sat359 islands in X:9.4-10.4 Mb region with all identified inserted sequence, including Dxl genes, in D. simulans, D. mauritiana, and D. sechellia.
Tick marks indicate locations of conserved sat359 islands, blue dots indicate protein-coding genes of interest in the region, and the different coloured squares represent the homologies of sequences inserted into sat359 islands (green=Dxl; purple=Ptpmeg2 fragment; brown=mkg-p retrotransposition; red=transposase-like sequence; yellow=cubn fragment; pink=CARPB intron fragments).
Extended Data Fig. 4 Evidence for transfer of Dxl material via a circular DNA intermediate molecule.
Dxl-12mau (green) is flanked by sat359 repeats (blue arrows) and by CARPB intronic sequence (boxes labelled A and B). These flanking sequence match sequences present in Dxl-10mau and Dxl-14mau, but the order of the two CARPB segments A and B differs from Dxl-12mau. The re-ordering of homologous sequences, as well as their intervening sequence, is consistent with a transfer of material via circular DNA intermediate.
Supplementary information
Supplementary Information
Supplementary text and Figs. 1–4.
Supplementary Tables
Supplementary Tables 1–4.
Supplementary Data 1
Alignment of inserted sat359 clusters.
Supplementary Data 2
Dxl protamines—positions 1–66, aligned with known protamines.
Supplementary Data 3
D. simulans r2.02, curated transcript set (‘base’ sequence set).
Supplementary Data 4
Consensus hairpin sequences of autosomal suppressors.
Supplementary Data 5
Species-specific consensus sequences, all Dxl-related sequence.
Rights and permissions
About this article
Cite this article
Muirhead, C.A., Presgraves, D.C. Satellite DNA-mediated diversification of a sex-ratio meiotic drive gene family in Drosophila. Nat Ecol Evol 5, 1604–1612 (2021). https://doi.org/10.1038/s41559-021-01543-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-021-01543-8
This article is cited by
-
A meiotic driver alters sperm form and function in house mice: a possible example of spite
Chromosome Research (2022)
-
Non-Mendelian segregation and transmission drive of B chromosomes
Chromosome Research (2022)
-
A flurry of sex-ratio distorters
Nature Ecology & Evolution (2021)