Abstract
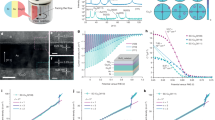
Microbe–semiconductor biohybrids, which integrate microbial enzymatic synthesis with the light-harvesting capabilities of inorganic semiconductors, have emerged as promising solar-to-chemical conversion systems. Improving the electron transport at the nano–bio interface and inside cells is important for boosting conversion efficiencies, yet the underlying mechanism is challenging to study by bulk measurements owing to the heterogeneities of both constituents. Here we develop a generalizable, quantitative multimodal microscopy platform that combines multi-channel optical imaging and photocurrent mapping to probe such biohybrids down to single- to sub-cell/particle levels. We uncover and differentiate the critical roles of different hydrogenases in the lithoautotrophic bacterium Ralstonia eutropha for bioplastic formation, discover this bacterium’s surprisingly large nanoampere-level electron-uptake capability, and dissect the cross-membrane electron-transport pathways. This imaging platform, and the associated analytical framework, can uncover electron-transport mechanisms in various types of biohybrid, and potentially offers a means to use and engineer R. eutropha for efficient chemical production coupled with photocatalytic materials.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All data are available in the main text or the Supplementary Information. Raw data supporting the findings of this study are available upon reasonable request. Source data are provided with this paper.
Code availability
MATLAB codes for data analysis and simulations supporting the findings of this study are provided with this paper.
References
Cestellos-Blanco, S., Zhang, H., Kim, J. M., Shen, Y. X. & Yang, P. Photosynthetic semiconductor biohybrids for solar-driven biocatalysis. Nat. Catal. 3, 245–255 (2020).
Rabaey, K. & Rozendal, R. A. Microbial electrosynthesis—revisiting the electrical route for microbial production. Nat. Rev. Microbiol. 8, 706–716 (2010).
Kracke, F., Vassilev, I. & Krömer, J. O. Microbial electron transport and energy conservation—the foundation for optimizing bioelectrochemical systems. Front. Microbiol. 6, 575 (2015).
Kornienko, N., Zhang, J. Z., Sakimoto, K. K., Yang, P. & Reisner, E. Interfacing nature’s catalytic machinery with synthetic materials for semi-artificial photosynthesis. Nat. Nanotechnol. 13, 890–899 (2018).
Dogutan, D. K. & Nocera, D. G. Artificial photosynthesis at efficiencies greatly exceeding that of natural photosynthesis. Acc. Chem. Res. 52, 3143–3148 (2019).
Nangle, S. N., Sakimoto, K. K., Silver, P. A. & Nocera, D. G. Biological-inorganic hybrid systems as a generalized platform for chemical production. Curr. Opin. Chem. Biol. 41, 107–113 (2017).
Liu, C., Colón, B. C., Ziesack, M., Silver, P. A. & Nocera, D. G. Water splitting-biosynthetic system with CO2 reduction efficiencies exceeding photosynthesis. Science 352, 1210–1213 (2016).
Nichols, E. M. et al. Hybrid bioinorganic approach to solar-to-chemical conversion. Proc. Natl Acad. Sci. USA 112, 11461–11466 (2015).
Liu, C. et al. Nanowire-bacteria hybrids for unassisted solar carbon dioxide fixation to value-added chemicals. Nano Lett. 15, 3634–3639 (2015).
Sakimoto, K. K., Wong Andrew, B. & Yang, P. Self-photosensitization of nonphotosynthetic bacteria for solar-to-chemical production. Science 351, 74–77 (2016).
Wang, B., Jiang, Z., Yu, J. C., Wang, J. & Wong, P. K. Enhanced CO2 reduction and valuable C2+ chemical production by a CdS-photosynthetic hybrid system. Nanoscale 11, 9296–9301 (2019).
Guo, J. et al. Light-driven fine chemical production in yeast biohybrids. Science 362, 813–816 (2018).
Kornienko, N. et al. Spectroscopic elucidation of energy transfer in hybrid inorganic-biological organisms for solar-to-chemical production. Proc. Natl Acad. Sci. USA 113, 11750–11755 (2016).
Brown Katherine, A. et al. Light-driven dinitrogen reduction catalyzed by a CdS:nitrogenase MoFe protein biohybrid. Science 352, 448–450 (2016).
Dong, F., Lee, Y. S., Gaffney, E. M., Liou, W. & Minteer, S. D. Engineering cyanobacterium with transmembrane electron transfer ability for bioelectrochemical nitrogen fixation. ACS Catal. 11, 13169–13179 (2021).
Knoche, K. L., Aoyama, E., Hasan, K. & Minteer, S. D. Role of nitrogenase and ferredoxin in the mechanism of bioelectrocatalytic nitrogen fixation by the cyanobacteria Anabaena variabilis SA-1 mutant immobilized on indium tin oxide (ITO) electrodes. Electrochim. Acta 232, 396–403 (2017).
Taniguchi, Y. et al. Quantifying E. coli proteome and transcriptome with single-molecule sensitivity in single cells. Science 329, 533–538 (2010).
Debroye, E. et al. Facet-dependent photoreduction on single ZnO crystals. J. Phys. Chem. Lett. 8, 340–346 (2017).
Mao, X. & Chen, P. Inter-facet junction effects on particulate photoelectrodes. Nat. Mater. 21, 331–337 (2021).
Reinecke, F. & Steinbüchel, A. Ralstonia eutropha strain H16 as model organism for PHA metabolism and for biotechnological production of technically interesting biopolymers. J. Mol. Microbiol. Biotechnol. 16, 91–108 (2009).
Burgdorf, T. et al. [NiFe]-hydrogenases of Ralstonia eutropha H16: modular enzymes for oxygen-tolerant biological hydrogen oxidation. J. Mol. Microbiol. Biotechnol. 10, 181–196 (2005).
Pohlmann, A. et al. Genome sequence of the bioplastic-producing ‘Knallgas’ bacterium Ralstonia eutropha H16. Nat. Biotechnol. 24, 1257–1262 (2006).
Cramm, R. Genomic view of energy metabolism in Ralstonia eutropha H16. J. Mol. Microbiol. Biotechnol. 16, 38–52 (2009).
Pötter, M. et al. The complex structure of polyhydroxybutyrate (PHB) granules: four orthologous and paralogous phasins occur in Ralstonia eutropha. Microbiology 150, 2301–2311 (2004).
Peoples, O. P. & Sinskey, A. J. Poly-P-hydroxybutyrate (PHB) biosynthesis in Alcaligenes eutrophus H16. J. Biol. Chem. 264, 15298–15303 (1989).
Bresan, S. et al. Polyhydroxyalkanoate (PHA) granules have no phospholipids. Sci. Rep. 6, 26612 (2016).
York, G. M., Junker, B. H., Stubbe, J. & Sinskey, A. J. Accumulation of the PhaP phasin of Ralstonia eutropha is dependent on production of polyhydroxybutyrate in cells. J. Bacteriol. 183, 4217–4226 (2001).
Fu, B. et al. Metal-induced sensor mobilization turns on affinity to activate regulator for metal detoxification in live bacteria. Proc. Natl Acad. Sci. USA 117, 13248–13255 (2020).
Mullineaux, C. W., Nenninger, A., Ray, N. & Robinson, C. Diffusion of green fluorescent protein in three cell environments in Escherichia coli. J. Bacteriol. 188, 3442–3448 (2006).
Deutzmann, J. S., Sahin, M., Spormann, A. M. & Harwood, C. S. Extracellular enzymes facilitate electron uptake in biocorrosion and bioelectrosynthesis. mBio 6, e00496–00415 (2015).
Kohlmann, Y. et al. Analyses of soluble and membrane proteomes of Ralstonia eutropha H16 reveal major changes in the protein complement in adaptation to lithoautotrophy. J. Proteome Res. 10, 2767–2776 (2011).
Friedrich, B., Buhrke, T., Burgdorf, T. & Lenz, O. A hydrogen-sensing multiprotein complex controls aerobic hydrogen metabolism in Ralstonia eutropha. Biochem. Soc. Trans. 33, 97–101 (2005).
Aslan, E. et al. Photocatalytic H2 evolution with a Cu2WS4 catalyst on a metal free D-π-A organic dye-sensitized TiO2. Appl. Catal. B 210, 320–327 (2017).
Sivula, K. & van de Krol, R. Semiconducting materials for photoelectrochemical energy conversion. Nat. Rev. Mater. 1, 15010 (2016).
Nasir, J. A. et al. Recent developments and perspectives in CdS-based photocatalysts for water splitting. J. Mater. Chem. A 8, 20752–20780 (2020).
Lanni, E. J. et al. Correlated imaging with C60-SIMS and confocal Raman microscopy: visualization of cell-scale molecular distributions in bacterial biofilms. Anal. Chem. 86, 10885–10891 (2014).
Chadwick, G. L., Jiménez, O. F., Gralnick, J. A., Bond, D. R. & Orphan, V. J. NanoSIMS imaging reveals metabolic stratification within current-producing biofilms. Proc. Natl Acad. Sci. USA 116, 20716–20724 (2019).
Saggu, M. et al. Spectroscopic insights into the oxygen-tolerant membrane-associated [NiFe] hydrogenase of Ralstonia eutropha H16. J. Biol. Chem. 284, 16264–16276 (2009).
Edwards, R. A., Keller, L. H. & Schifferli, D. M. Improved allelic exchange vectors and their use to analyze 987P fimbria gene expression. Gene 207, 149–157 (1998).
Torella, J. P. et al. Efficient solar-to-fuels production from a hybrid microbial-water-splitting catalyst system. Proc. Natl Acad. Sci. USA 112, 2337–2342 (2015).
Chen, T.-Y. et al. Concentration- and chromosome-organization-dependent regulator unbinding from DNA for transcription regulation in living cells. Nat. Commun. 6, 7445 (2015).
Acknowledgements
This research is supported by the US Department of Energy (Office of Science, Office of Biological and Environmental Research, Biological Systems Science Division, under award no. DE-SC0020179). It uses Cornell Center for Materials Research Shared Facilities supported through the NSF MRSEC programme (DMR-1719875). The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript. We thank C. Brigham at Wentworth Institute of Technology for providing the R. eutropha H16 strain, W. Metcalf at the University of Illinois at Urbana Champaign for E. coli WM3064, O. Lenz at Technische Universität Berlin for R. eutropha ΔhoxH, L. Krzeminski formerly at Cornell University for constructing tagged E. coli strains, and S. Murphy, T. Doerr and A. Schmitz at Cornell University for discussions on genetics.
Author information
Authors and Affiliations
Contributions
B.F. constructed strains. X.M. synthesized semiconductor materials. B.F. and X.M. designed experiments, performed measurements, wrote codes and analysed data. Y.P., Z.Z. and T.Y. contributed to strain construction, materials synthesis and/or imaging experiments, where Y.P. and Z.Z. contributed equally. W.J. and D.H.F. contributed to cell culturing. W.L. helped with instrument set-up. B.P., F.S. and B.B. contributed to genetic engineering. M.S. and T.H. contributed to discussions on materials synthesis. P.C. conceived and directed the research. B.F., X.M. and P.C. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Chemistry thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–28, sections 1–13, Tables 1–6 and references 42–74.
Supplementary Code 1
MATLAB codes for data analysis and simulations supporting the findings of this study.
Supplementary Table 1
List of primers used in manuscript.
Source data
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 5
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Fu, B., Mao, X., Park, Y. et al. Single-cell multimodal imaging uncovers energy conversion pathways in biohybrids. Nat. Chem. 15, 1400–1407 (2023). https://doi.org/10.1038/s41557-023-01285-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41557-023-01285-z