Abstract
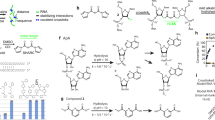
Protein–RNA interactions regulate RNA fate and function, and defects can lead to various disorders. Such interactions have mainly been studied by nucleoside-based UV crosslinking methods, which lack broad in vivo compatibility and the ability to resolve specific amino acids. In this study we genetically encoded latent bioreactive unnatural amino acids into proteins to react with bound RNA by proximity-enabled reactivity and demonstrated genetically encoded chemical crosslinking of proteins with target RNA (GECX-RNA) in vivo. Applying GECX-RNA to the RNA chaperone Hfq in Escherichia coli identified target RNAs with amino acid specificity. Combining GECX-RNA with immunoprecipitation and high-throughput sequencing of an N6-methyladenosine reader protein in mammalian cells allowed the in vivo identification of unknown N6-methyladenosine on RNA with single-nucleotide resolution throughout the transcriptome. GECX-RNA thus affords resolution at the nucleotide and amino acid level for interrogating protein–RNA interactions in vivo. It also enables the precise engineering of covalent linkages between a protein and RNA, which will inspire innovative solutions for RNA-related research and therapeutics.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All GRIP-seq data are available in the Sequence Read Archive through accession number PRJNA797913. All other data generated or analysed in this study are available within the article and its Supplementary Information. Source data are provided with this paper.
Code availability
The custom code used in this study is available at https://github.com/Shall-We-Dance/GRIP-seq.
References
Gerstberger, S., Hafner, M. & Tuschl, T. A census of human RNA-binding proteins. Nat. Rev. Genet. 15, 829–845 (2014).
Castello, A., Fischer, B., Hentze, M. W. & Preiss, T. RNA-binding proteins in Mendelian disease. Trends Genet. 29, 318–327 (2013).
Nussbacher, J. K., Batra, R., Lagier-Tourenne, C. & Yeo, G. W. RNA-binding proteins in neurodegeneration: Seq and you shall receive. Trends Neurosci. 38, 226–236 (2015).
Castello, A. et al. Comprehensive identification of RNA-binding domains in human cells. Mol. Cell 60, 696–710 (2016).
Benhalevy, D., Anastasakis, D. G. & Hafner, M. Proximity-CLIP provides a snapshot of protein-occupied RNA elements in subcellular compartments. Nat. Methods 15, 1074–1082 (2018).
Hentze, M. W., Castello, A., Schwarzl, T. & Preiss, T. A brave new world of RNA-binding proteins. Nat. Rev. Mol. Cell Biol. 19, 327–341 (2018).
Müller-McNicoll, M. & Neugebauer, K. M. How cells get the message: dynamic assembly and function of mRNA–protein complexes. Nat. Rev. Genet. 14, 275–287 (2013).
Wagenmakers, A. J. M., Reinders, R. J. & van Venrooij, W. J. Cross‐linking of mRNA to proteins by irradiation of intact cells with ultraviolet light. Eur. J. Biochem. 112, 323–330 (1980).
Saito, I. & Matsuura, T. Chemical aspects of UV-induced cross-linking of proteins to nucleic acids. Photoreactions with lysine and tryptophan. Acc. Chem. Res. 18, 134–141 (1985).
Hafner, M. et al. Transcriptome-wide identification of RNA-binding protein and microRNA target sites by PAR-CLIP. Cell 141, 129–141 (2010).
Baltz, A. G. et al. The mRNA-bound proteome and its global occupancy profile on protein-coding transcripts. Mol. Cell 46, 674–690 (2012).
Licatalosi, D. D. et al. HITS-CLIP yields genome-wide insights into brain alternative RNA processing. Nature 456, 464–469 (2008).
König, J. et al. ICLIP reveals the function of hnRNP particles in splicing at individual nucleotide resolution. Nat. Struct. Mol. Biol. 17, 909–915 (2010).
Castello, A. et al. Insights into RNA biology from an atlas of mammalian mRNA-binding proteins. Cell 149, 1393–1406 (2012).
Lee, F. C. Y. & Ule, J. Advances in CLIP technologies for studies of protein–RNA Interactions. Mol. Cell 69, 354–369 (2018).
Sugimoto, Y. et al. Analysis of CLIP and iCLIP methods for nucleotide-resolution studies of protein–RNA interactions. Genome Biol. 13, R67 (2012).
Xiang, Z. et al. Adding an unnatural covalent bond to proteins through proximity-enhanced bioreactivity. Nat. Methods 10, 885–888 (2013).
Wang, L. Genetically encoding new bioreactivity. N. Biotechnol. 38, 16–25 (2017).
Coin, I. et al. Genetically encoded chemical probes in cells reveal the binding path of urocortin-I to CRF class B GPCR. Cell 155, 1258–1269 (2013).
Yang, B. et al. Spontaneous and specific chemical cross-linking in live cells to capture and identify protein interactions. Nat. Commun. 8, 2240 (2017).
Li, Q. et al. Developing covalent protein drugs via proximity-enabled reactive therapeutics. Cell 182, 85–97 (2020).
Wang, N. et al. Genetically encoding fluorosulfate-l-tyrosine to react with lysine, histidine, and tyrosine via SuFEx in proteins in vivo. J. Am. Chem. Soc. 140, 4995–4999 (2018).
Abudayyeh, O. O. et al. C2c2 is a single-component programmable RNA-guided RNA-targeting CRISPR effector. Science 353, aaf5573 (2016).
Cox, D. B. T. et al. RNA editing with CRISPR-Cas13. Science 358, 1019–1027 (2017).
Yang, L. Z. et al. Dynamic imaging of RNA in living cells by CRISPR-Cas13 systems. Mol. Cell 76, 981–997 (2019).
Liu, L. et al. Two distant catalytic sites are responsible for C2c2 RNase activities. Cell 168, 121–134 (2017).
Smargon, A. A. et al. Cas13b is a type VI-B CRISPR-associated RNA-guided RNase differentially regulated by accessory proteins Csx27 and Csx28. Mol. Cell 65, 618–630 (2017).
Zhang, B. et al. Structural insights into Cas13b-guided CRISPR RNA maturation and recognition. Cell Res. 28, 1198–1201 (2018).
Wilusz, C. J. & Wilusz, J. Eukaryotic Lsm proteins: lessons from bacteria. Nat. Struct. Mol. Biol. 12, 1031–1036 (2005).
Bilusic, I., Popitsch, N., Rescheneder, P., Schroeder, R. & Lybecker, M. Revisiting the coding potential of the E. coli genome through Hfq co-immunoprecipitation. RNA Biol. 11, 641–654 (2014).
Holmqvist, E. et al. Global RNA recognition patterns of post‐transcriptional regulators Hfq and CsrA revealed by UV crosslinking in vivo. EMBO J. 35, 991–1011 (2016).
Chao, Y., Papenfort, K., Reinhardt, R., Sharma, C. M. & Vogel, J. An atlas of Hfq-bound transcripts reveals 3′ UTRs as a genomic reservoir of regulatory small RNAs. EMBO J. 31, 4005–4019 (2012).
Wang, W., Wang, L., Wu, J., Gong, Q. & Shi, Y. Hfq-bridged ternary complex is important for translation activation of rpoS by DsrA. Nucleic Acids Res. 41, 5938–5948 (2013).
Peng, Y., Curtis, J. E., Fang, X. & Woodson, S. A. Structural model of an mRNA in complex with the bacterial chaperone Hfq. Proc. Natl Acad. Sci. USA 111, 17134–17139 (2014).
Tree, J. J., Granneman, S., McAteer, S. P., Tollervey, D. & Gally, D. L. Identification of bacteriophage-encoded anti-sRNAs in pathogenic Escherichia coli. Mol. Cell 55, 199–213 (2014).
Schu, D. J., Zhang, A., Gottesman, S. & Storz, G. Alternative Hfq–sRNA interaction modes dictate alternative mRNA recognition. EMBO J. 34, 2557–2573 (2015).
Hoppmann, C. & Wang, L. Proximity-enabled bioreactivity to generate covalent peptide inhibitors of p53–Mdm4. Chem. Commun. 52, 5140–5143 (2016).
Liu, J. et al. Genetically encoding photocaged quinone methide to multitarget protein residues covalently in vivo. J. Am. Chem. Soc. 141, 9458–9462 (2019).
Li, S. et al. Genetically encoded chemical crosslinking of carbohydrate. Nat. Chem. https://doi.org/10.1038/s41557-022-01059-z (2022).
Nachtergaele, S. & He, C. Chemical modifications in the life of an mRNA transcript. Annu. Rev. Genet. 52, 349–372 (2018).
Meyer, K. D. et al. Comprehensive analysis of mRNA methylation reveals enrichment in 3′ UTRs and near stop codons. Cell 149, 1635–1646 (2012).
Dominissini, D. et al. Topology of the human and mouse m6A RNA methylomes revealed by m6A-seq. Nature 485, 201–206 (2012).
Linder, B. et al. Single-nucleotide-resolution mapping of m6A and m6Am throughout the transcriptome. Nat. Methods 12, 767–772 (2015).
Xu, C. et al. Structural basis for the discriminative recognition of N6-methyladenosine RNA by the human YT521-B homology domain family of proteins. J. Biol. Chem. 290, 24902–24913 (2015).
Meyer, K. D. DART-seq: an antibody-free method for global m6A detection. Nat. Methods 16, 1275–1280 (2019).
Van Nostrand, E. L. et al. Robust transcriptome-wide discovery of RNA-binding protein binding sites with enhanced CLIP (eCLIP). Nat. Methods 13, 508–514 (2016).
Lovci, M. T. et al. Rbfox proteins regulate alternative mRNA splicing through evolutionarily conserved RNA bridges. Nat. Struct. Mol. Biol. 20, 1434–1442 (2013).
Tang, Y. et al. m6A-Atlas: a comprehensive knowledgebase for unraveling the N6-methyladenosine (m6A) epitranscriptome. Nucleic Acids Res. 49, D134–D143 (2020).
Sanchez de Groot, N. et al. RNA structure drives interaction with proteins. Nat. Commun. 10, 3246 (2019).
Siegfried, N. A., Busan, S., Rice, G. M., Nelson, J. A. E. & Weeks, K. M. RNA motif discovery by SHAPE and mutational profiling (SHAPE-MaP). Nat. Methods 11, 959–965 (2014).
Gruber, A. R., Lorenz, R., Bernhart, S. H., Neuböck, R. & Hofacker, I. L. The Vienna RNA websuite. Nucleic Acids Res. 36, W70–W74 (2008).
Hwang, H.-W. et al. PAPERCLIP identifies microRNA targets and a role of CstF64/64tau in promoting non-canonical poly(A) site usage. Cell Rep. 15, 423–435 (2016).
Kini, H. K., Silverman, I. M., Ji, X., Gregory, B. D. & Liebhaber, S. A. Cytoplasmic poly(A) binding protein-1 binds to genomically encoded sequences within mammalian mRNAs. RNA 22, 61–74 (2016).
Wang, L. Engineering the genetic code in cells and animals: biological considerations and impacts. Acc. Chem. Res. 50, 2767–2776 (2017).
Mackereth, C. D. & Sattler, M. Dynamics in multi-domain protein recognition of RNA. Curr. Opin. Struct. Biol. 22, 287–296 (2012).
Lunde, B. M., Moore, C. & Varani, G. RNA-binding proteins: Modular design for efficient function. Nat. Rev. Mol. Cell Biol. 8, 479–490 (2007).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Baek, M. et al. Accurate prediction of protein structures and interactions using a three-track neural network. Science 373, 871–876 (2021).
McMahon, A. C. et al. TRIBE: hijacking an RNA-editing enzyme to identify cell-specific targets of RNA-binding proteins. Cell 165, 742–753 (2016).
Brannan, K. W. et al. Robust single-cell discovery of RNA targets of RNA-binding proteins and ribosomes. Nat. Methods 18, 507–519 (2021).
Cao, L. & Wang, L. New covalent bonding ability for proteins. Protein Sci. 31, 312–322 (2021).
Uhlén, M. et al. Tissue-based map of the human proteome. Science 347, 1260419 (2015).
Chen, S., Zhou, Y., Chen, Y. & Gu, J. Fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34, i884–i890 (2018).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Bailey, T. L. et al. MEME suite: tools for motif discovery and searching. Nucleic Acids Res. 37, W202–W208 (2009).
Olarerin-George, A. O. & Jaffrey, S. R. MetaPlotR: a Perl/R pipeline for plotting metagenes of nucleotide modifications and other transcriptomic sites. Bioinformatics 33, 1563–1564 (2017).
Lorenz, R. et al. ViennaRNA Package 2.0. Algorithms Mol. Biol. 6, 26 (2011).
Acknowledgements
L.W acknowledges the support of the NIH (R01GM118384 and R01CA258300). Y.S. acknowledges the support of the NIH (R01AG057497 and R01EY027789).
Author information
Authors and Affiliations
Contributions
W.S. designed and conducted the experiments, analysed the data and wrote the manuscript; N.W. evolved SFYRS and characterized SFY incorporation and the crosslinking of proteins; H.L. conducted the data analysis of GRIP-seq; B.Y. synthesized FSY and SFY, performed the SFY reactions with NMPs in vitro and analysed the data; L.J. helped with the dCas13b target RNA crosslinking and enrichment in mammalian cells; X.R. and Y.S. helped with the GRIP-seq experiments; L.W. conceived, directed and supported the project and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Chemistry thanks Ryan Flynn, Stephen Fried and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Identification of the positively charged residues of PsCas13b involved in pre-crRNA cleavage.
a) Multiple sequence alignment of Cas13b proteins from different species (Bzo: Bergeyella zoohelcum, Psp: Prevotella sp. P5-125, Pgu: Porphyromonas gingivalis, Pbu: Prevotella buccae, and Ran: Riemerella anatipestifer) for β-sheets 5 and 6 involved in pre-crRNA cleavage. The secondary structure of BzoCas13b is shown above the sequence28. Identical and similar residues are highlighted in red and white boxes, respectively. Positive charged catalytic residues in BzoCas13b involved in the pre-crRNA cleavage on β-sheets 5 and 6 (450 R, 452 K, 459 R) are marked with green stars on the bottom. Positive charged residues in PsCas13b located on β-sheets 5 and 6 (367 K, 370 K, 378 R, 380 R) are marked with purple squares. Multiple sequence alignment of full-length Cas13b proteins from different species is shown in Supplementary Fig. 1. b) Denaturing urea-PAGE demonstrating the pre-crRNA cleavage by dPsCas13b-WT and dPsCas13b-Ala-mutants speculatively involved in the pre-crRNA processing. dPsCas13b-WT and dPsCas13b-Ala-mutants were incubated with pre-crRNA and then separated on denaturing urea-PAGE. The Urea-gel was stained with SybrGold for fluorescent detection of RNA.
Extended Data Fig. 2 GRIP results demonstrate that site 25 of Hfq directly binds with (AAN)4 elements of rpoS RNA.
GRIP identified the binding sites of Tyr25 of Hfq on rpoS RNA in E. coli cells. Red triangles indicate cross-linking sites identified from GRIP for rpoS RNA from Hfq-25FSY expressing E. coli cells. Two examples of Sanger sequencing of clones from Hfq-25FSY sample were shown below.
Extended Data Fig. 3 Examples of GRIP-seq data for m6A identification.
Genome browser tracks of GRIP-seq data in JUN and DICER1 mRNA regions. Reverse-transcription-termination sites (RT-termination sites) from GRIP-seq were marked as yellow triangles. Known m6A sites from published datasets were marked as grey triangles.
Supplementary information
Supplementary Information
Supplementary Figs. 1–7, Tables 1–4 and source data files for the supplementary figures.
Supplementary Table 2
Sequencing statistics for GRIP-seq.
Supplementary Table 3
m6A sites identified from GRIP-seq (coordinates in hg19 human genome assembly).
Supplementary Data 1
Statistical source data for Supplementary Fig. 2b.
Supplementary Data 2
Statistical source data for Supplementary Fig. 4.
Supplementary Data 3
Statistical source data for Supplementary Fig. 5.
Supplementary Data 4
Source data for Sanger sequencing in Supplementary Fig. 2d (Ab1 files).
Supplementary Data 5
Source data for Sanger sequencing in Supplementary Fig. 2g (Ab1 files).
Supplementary Data 6
Source data for Sanger sequencing in Supplementary Fig. 7h (Ab1 files).
Source data
Source Data Fig. 1
Unprocessed gels.
Source Data Fig. 2
Unprocessed western blot and statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Uncropped images, unprocessed gels, unprocessed western blots and statistical source data.
Source Data Extended Data Fig. 1
Unprocessed gels.
Source Data Extended Data Fig. 2
Source data for Sanger sequencing (Ab1 files).
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Sun, W., Wang, N., Liu, H. et al. Genetically encoded chemical crosslinking of RNA in vivo. Nat. Chem. 15, 21–32 (2023). https://doi.org/10.1038/s41557-022-01038-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41557-022-01038-4
This article is cited by
-
Bioorthogonal masked acylating agents for proximity-dependent RNA labelling
Nature Chemistry (2024)
-
Sulfur fluoride exchange
Nature Reviews Methods Primers (2023)
-
Post-transcriptional checkpoints in autoimmunity
Nature Reviews Rheumatology (2023)
-
A (cross)link in the chains
Nature Chemistry (2023)