Abstract
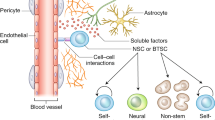
Brain metastases represent an important clinical problem for patients with small-cell lung cancer (SCLC). However, the mechanisms underlying SCLC growth in the brain remain poorly understood. Here, using intracranial injections in mice and assembloids between SCLC aggregates and human cortical organoids in culture, we found that SCLC cells recruit reactive astrocytes to the tumour microenvironment. This crosstalk between SCLC cells and astrocytes drives the induction of gene expression programmes that are similar to those found during early brain development in neurons and astrocytes. Mechanistically, the brain development factor Reelin, secreted by SCLC cells, recruits astrocytes to brain metastases. These astrocytes in turn promote SCLC growth by secreting neuronal pro-survival factors such as SERPINE1. Thus, SCLC brain metastases grow by co-opting mechanisms involved in reciprocal neuron–astrocyte interactions during brain development. Targeting such developmental programmes activated in this cancer ecosystem may help prevent and treat brain metastases.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
ATAC–seq and RNA-seq data have been deposited in the Gene Expression Omnibus (GSE179032 and GSE178743). All other data supporting the findings of this study are available from the corresponding author on reasonable request. The ENSEMBL mm10 genome (GRCm38) can be accessed at https://www.ncbi.nlm.nih.gov/datasets/genome/GCF_000001635.20/Source data are provided with this paper.
Code availability
The code for the RNA sequencing analyses is available at Zenodo, https://doi.org/10.5281/zenodo.5068366.
References
Chaffer, C. L. & Weinberg, R. A. A perspective on cancer cell metastasis. Science 331, 1559–1564 (2011).
Valiente, M. et al. The evolving landscape of brain metastasis. Trends Cancer 4, 176–196 (2018).
Suh, J. H. et al. Current approaches to the management of brain metastases. Nat. Rev. Clin. Oncol. 17, 279–299 (2020).
Bos, P. D. et al. Genes that mediate breast-cancer metastasis to the brain. Nature 459, 1005–1009 (2009).
Lorger, M. & Felding-Habermann, B. Capturing changes in the brain microenvironment during initial steps of breast-cancer brain metastasis. Am. J. Pathol. 176, 2958–2971 (2010).
Zeng, Q. et al. Synaptic proximity enables NMDAR signalling to promote brain metastasis. Nature 573, 526–531 (2019).
Ebright, R. Y. et al. HIF1A signaling selectively supports proliferation of breast cancer in the brain. Nat. Commun. 11, 6311 (2020).
Adler, O. et al. Reciprocal interactions between innate immune cells and astrocytes facilitate neuroinflammation and brain metastasis via lipocalin-2. Nat. Cancer 4, 401–418 (2023).
Biermann, J. et al. Dissecting the treatment-naïve ecosystem of human melanoma brain metastasis. Cell 185, 2591–2608.e30 (2022).
Achrol, A. S. et al. Brain metastases. Nat. Rev. Dis. Prim. 5, 5 (2019).
Hebert, J. D., Neal, J. W. & Winslow, M. M. Dissecting metastasis using preclinical models and methods. Nat. Rev. Cancer 23, 391–407 (2023).
Rudin, C. M., Brambilla, E., Faivre-Finn, C. & Sage, J. Small-cell lung cancer. Nat. Rev. Dis. Prim. 7, 3 (2021).
Boire, A., Brastianos, P. K., Garzia, L. & Valiente, M. Brain metastasis. Nat. Rev. Cancer 20, 4–11 (2020).
Ko, J., Winslow, M. M. & Sage, J. Mechanisms of small-cell lung cancer metastasis. EMBO Mol. Med. 13, e13122 (2021).
Megyesfalvi, Z. et al. The landscape of small-cell lung cancer metastases: organ specificity and timing. Thorac. Cancer 12, 914–923 (2021).
Rusthoven, C. G., Camidge, D. R., Robin, T. P. & Brown, P. D. Radiosurgery for small-cell brain metastases: challenging the last bastion of preferential whole-brain radiotherapy delivery. J. Clin. Oncol. 38, 3587–3591 (2020).
Neal, J. W., Gubens, M. A. & Wakelee, H. A. Current management of small-cell lung cancer. Clin. Chest Med. 32, 853–863 (2011).
Takahashi, T. et al. Prophylactic cranial irradiation versus observation in patients with extensive-disease small-cell lung cancer: a multicentre, randomised, open-label, phase 3 trial. Lancet Oncol. 18, 663–671 (2017).
Lukas, R. V., Gondi, V., Kamson, D. O., Kumthekar, P. & Salgia, R. State-of-the-art considerations in small-cell lung cancer brain metastases. Oncotarget 8, 71223–71233 (2017).
Drapkin, B. J. & Rudin, C. M. Advances in small-cell lung cancer (SCLC) translational research. Cold Spring Harb. Perspect. Med. 11, a038240 (2020).
Ilhan-Mutlu, A. et al. Expression profiling of angiogenesis-related genes in brain metastases of lung cancer and melanoma. Tumour Biol. 37, 1173–1182 (2016).
Thompson, E. M., Pishko, G. L., Muldoon, L. L. & Neuwelt, E. A. Inhibition of SUR1 decreases the vascular permeability of cerebral metastases. Neoplasia 15, 535–543 (2013).
Li, B. et al. Elevated PLGF contributes to small-cell lung cancer brain metastasis. Oncogene 32, 2952–2962 (2013).
Xu, Z.-H. et al. Brain microvascular endothelial cell exosome-mediated S100A16 up-regulation confers small-cell lung cancer cell survival in brain. FASEB J. 33, 1742–1757 (2019).
Wang, A. & Komiya, T. Brain metastasis as exclusion criteria in clinical trials involving extensive-stage small-cell lung cancer. J. Cancer Res. Clin. Oncol. 145, 3099–3104 (2019).
Stewart, C. A. et al. Single-cell analyses reveal increased intratumoral heterogeneity after the onset of therapy resistance in small-cell lung cancer. Nat. Cancer 1, 423–436 (2020).
Meuwissen, R. et al. Induction of small-cell lung cancer by somatic inactivation of both Trp53 and Rb1 in a conditional mouse model. Cancer Cell. 4, 181–189 (2003).
Guo, W.-L. et al. Clinical features of brain metastases in small-cell lung cancer: an implication for hippocampal sparing whole brain radiation therapy. Transl. Oncol. 10, 54–58 (2017).
Seute, T., Leffers, P., ten Velde, G. P. M. & Twijnstra, A. Leptomeningeal metastases from small-cell lung carcinoma. Cancer 104, 1700–1705 (2005).
Uña, E. Intraventricular metastases from small-cell carcinoma of the lung. BMJ Case Rep. 2012, bcr1220115440 (2012).
Denny, S. K. et al. Nfib promotes metastasis through a widespread increase in chromatin accessibility. Cell 166, 328–342 (2016).
Yang, D. et al. Axon-like protrusions promote small-cell lung cancer migration and metastasis. eLife 8, e50616 (2019).
Wasilewski, D., Priego, N., Fustero-Torre, C. & Valiente, M. Reactive astrocytes in brain metastasis. Front Oncol. 7, 298 (2017).
Escartin, C. et al. Reactive astrocyte nomenclature, definitions, and future directions. Nat. Neurosci. 24, 312–325 (2021).
Henrik Heiland, D. et al. Tumor-associated reactive astrocytes aid the evolution of immunosuppressive environment in glioblastoma. Nat. Commun. 10, 2541 (2019).
Zhang, Y. et al. Purification and characterization of progenitor and mature human astrocytes reveals transcriptional and functional differences with mouse. Neuron 89, 37–53 (2016).
Sirkisoon, S. R. et al. TGLI1 transcription factor mediates breast-cancer brain metastasis via activating metastasis-initiating cancer stem cells and astrocytes in the tumor microenvironment. Oncogene 39, 64–78 (2020).
Priego, N. et al. STAT3 labels a subpopulation of reactive astrocytes required for brain metastasis. Nat. Med. 24, 1024–1035 (2018).
Williamson, S. C. et al. Vasculogenic mimicry in small-cell lung cancer. Nat. Commun. 7, 13322 (2016).
Simpson, K. L. et al. A biobank of small-cell lung cancer CDX models elucidates inter- and intratumoral phenotypic heterogeneity. Nat. Cancer 1, 437–451 (2020).
Jahchan, N. S. et al. Identification and targeting of long-term tumor-propagating cells in small-cell lung cancer. Cell Rep. 16, 644–656 (2016).
Guo, D. et al. Tumor cells generate astrocyte-like cells that contribute to SHH-driven medulloblastoma relapse. J. Exp. Med. 218, e20202350 (2021).
Yao, M. et al. Astrocytic trans-differentiation completes a multicellular paracrine feedback loop required for medulloblastoma tumor growth. Cell 180, 502–520.e19 (2020).
Voskuhl, R. R. et al. Reactive astrocytes form scar-like perivascular barriers to leukocytes during adaptive immune inflammation of the CNS. J. Neurosci. 29, 11511–11522 (2009).
Taylor, X. et al. Activated endothelial cells induce a distinct type of astrocytic reactivity. Commun. Biol. 5, 282 (2022).
Liddelow, S. A. & Barres, B. A. Reactive astrocytes: production, function, and therapeutic potential. Immunity 46, 957–967 (2017).
Weimer, J. M. et al. MARCKS modulates radial progenitor placement, proliferation and organization in the developing cerebral cortex. Development 136, 2965–2975 (2009).
Luo, J. Dysregulation of polarity proteins in astrocyte reactivity. Ageing Res. Rev. 86, 101869 (2023).
Paşca, A. M. et al. Functional cortical neurons and astrocytes from human pluripotent stem cells in 3D culture. Nat. Methods 12, 671–678 (2015).
Yoon, S.-J. et al. Reliability of human cortical organoid generation. Nat. Methods 16, 75–78 (2019).
Sloan, S. A. et al. Human astrocyte maturation captured in 3D cerebral cortical spheroids derived from pluripotent stem cells. Neuron 95, 779–790.e6 (2017).
Tsai, H.-H. et al. Regional astrocyte allocation regulates CNS synaptogenesis and repair. Science 337, 358–362 (2012).
Allen, N. J. & Lyons, D. A. Glia as architects of central nervous system formation and function. Science 362, 181–185 (2018).
Jacobsen, C. T. & Miller, R. H. Control of astrocyte migration in the developing cerebral cortex. Dev. Neurosci. 25, 207–216 (2003).
Yang, D. et al. Intertumoral heterogeneity in SCLC is influenced by the cell type of origin. Cancer Discov. 8, 1316–1331 (2018).
Carney, D. N. et al. Serum neuron-specific enolase: a marker for disease extent and response to therapy of small-cell lung cancer. Lancet 1, 583–585 (1982).
van Zandwijk, N., Jassem, E., Bonfrer, J. M., Mooi, W. J. & van Tinteren, H. Serum neuron-specific enolase and lactate dehydrogenase as predictors of response to chemotherapy and survival in non-small-cell lung cancer. Semin. Oncol. 19, 37–43 (1992).
Semenova, E. A. et al. Transcription factor NFIB is a driver of small-cell lung cancer progression in mice and marks metastatic disease in patients. Cell Rep. 16, 631–643 (2016).
Wu, N. et al. NFIB overexpression cooperates with Rb/p53 deletion to promote small-cell lung cancer. Oncotarget 7, 57514–57524 (2016).
Dooley, A. L. et al. Nuclear factor I/B is an oncogene in small-cell lung cancer. Genes Dev. 25, 1470–1475 (2011).
Zhan, J. S. et al. Astrocytes in migration. Neurochem. Res. 42, 272–282 (2017).
Brunkhorst, R., Bock, H. & Derouiche, A. Reelin induces process growth in cultured astrocytes: implication for glia-synaptic plasticity. Arch. Ital. Biol. 153, 249–254 (2015).
Courtès, S. et al. Reelin controls progenitor cell migration in the healthy and pathological adult mouse brain. PLoS ONE 6, e20430 (2011).
Niu, S., Renfro, A., Quattrocchi, C. C., Sheldon, M. & D’Arcangelo, G. Reelin promotes hippocampal dendrite development through the VLDLR/ApoER2-Dab1 pathway. Neuron 41, 71–84 (2004).
Khialeeva, E. et al. Reelin deficiency delays mammary tumor growth and metastatic progression. J. Mammary Gland Biol. Neoplasia. 22, 59–69 (2017).
Wang, Y., Ye, F., Liang, Y. & Yang, Q. Breast-cancer brain metastasis: insight into molecular mechanisms and therapeutic strategies. Br. J. Cancer 125, 1056–1067 (2021).
Yasui, N. et al. Structure of a receptor-binding fragment of reelin and mutational analysis reveal a recognition mechanism similar to endocytic receptors. Proc. Natl Acad. Sci. USA 104, 9988–9993 (2007).
Davies, A. M. Regulation of neuronal survival and death by extracellular signals during development. EMBO J. 22, 2537–2545 (2003).
Gabriel, C. et al. Transforming growth factor α-induced expression of type-1 plasminogen activator inhibitor in astrocytes rescues neurons from excitotoxicity. FASEB J. 17, 277–279 (2003).
Pavón, M. et al. uPA/uPAR and SERPINE1 in head and neck cancer: role in tumor resistance, metastasis, prognosis and therapy. Oncotarget 7, 57351–57366 (2016).
Che, Y. et al. Cisplatin-activated PAI-1 secretion in the cancer-associated fibroblasts with paracrine effects promoting esophageal squamous cell carcinoma progression and causing chemoresistance. Cell Death Dis. 9, 759 (2018).
Schneider, D. J., Chen, Y. & Sobel, B. E. The effect of plasminogen activator inhibitor type 1 on apoptosis. Thromb. Haemost. 100, 1037–1040 (2008).
Valiente, M. et al. Serpins promote cancer cell survival and vascular Co-option in brain metastasis. Cell 156, 1002–1016 (2014).
Elokdah, H. et al. Tiplaxtinin, a novel, orally efficacious inhibitor of plasminogen activator inhibitor-1: design, synthesis, and preclinical characterization. J. Med. Chem. 47, 3491–3494 (2004).
Bydoun, M., Sterea, A., Weaver, I. C. G., Bharadwaj, A. G. & Waisman, D. M. A novel mechanism of plasminogen activation in epithelial and mesenchymal cells. Sci. Rep. 8, 14091 (2018).
Caino, M. C. et al. A neuronal network of mitochondrial dynamics regulates metastasis. Nat. Commun. 7, 13730 (2016).
Neman, J. et al. Human breast-cancer metastases to the brain display GABAergic properties in the neural niche. Proc. Natl Acad. Sci. USA 111, 984–989 (2014).
Tan, R. et al. Neural functions play different roles in triple negative breast cancer (TNBC) and non-TNBC. Sci. Rep. 10, 3065 (2020).
Wingrove, E. et al. Transcriptomic hallmarks of tumor plasticity and stromal interactions in brain metastasis. Cell Rep. 27, 1277–1292.e7 (2019).
Quail, D. & Joyce, J. Microenvironmental regulation of tumor progression and metastasis. Nat. Med. 19, 1423–1437 (2013).
Sevenich, L. et al. Analysis of tumour- and stroma-supplied proteolytic networks reveals a brain-metastasis-promoting role for cathepsin S. Nat. Cell Biol. 16, 876–888 (2014).
Schmitt, A. et al. Functional synapses between small-cell lung cancer and glutamatergic neurons. Preprint at bioRxiv https://doi.org/10.1101/2023.01.19.524045 (2023).
Savchuk S., et al. Neuronal-activity dependent mechanisms of small-cell lung cancer progression. Preprint at bioRxiv https://doi.org/10.1101/2023.01.19.524430 (2023).
Lim, J. S. et al. Intratumoral heterogeneity generated by Notch signaling promotes small-cell lung cancer. Nature 545, 360–364 (2017).
Pan, Y. et al. NF1 mutation drives neuronal activity-dependent initiation of optic glioma. Nature 594, 277–282 (2021).
Hodgkinson, C. L. et al. Tumorigenicity and genetic profiling of circulating tumor cells in small-cell lung cancer. Nat. Med. 20, 897–903 (2014).
Calbo, J. et al. A functional role for tumor cell heterogeneity in a mouse model of small-cell lung cancer. Cancer Cell 19, 244–256 (2011).
Seike, T. et al. Interaction between lung-cancer cells and astrocytes via specific inflammatory cytokines in the microenvironment of brain metastasis. Clin. Exp. Metastasis 28, 13–25 (2011).
Chen, Q. et al. Carcinoma-astrocyte gap junctions promote brain metastasis by cGAMP transfer. Nature 533, 493–498 (2016).
Kim, S.-J. et al. Astrocytes upregulate survival genes in tumor cells and induce protection from chemotherapy. Neoplasia 13, 286–298 (2011).
Berghoff, A. S. et al. Tumor infiltrating lymphocytes and PD-L1 expression in brain metastases of small-cell lung cancer (SCLC). J. Neurooncol. 130, 19–29 (2016).
Hohensee, I. et al. PTEN mediates the cross talk between breast and glial cells in brain metastases leading to rapid disease progression. Oncotarget 8, 6155–6168 (2017).
Strickland, M. R., Alvarez-Breckenridge, C., Gainor, J. F. & Brastianos, P. K. Tumor immune microenvironment of brain metastases: toward unlocking antitumor immunity. Cancer Discov. 12, 1199–1216 (2022).
Placencio, V. R. & DeClerck, Y. A. Plasminogen activator inhibitor-1 in cancer: rationale and insight for future therapeutic testing. Cancer Res. 75, 2969–2974 (2015).
Lee, S. H., Eren, M., Vaughan, D. E., Schleimer, R. P. & Cho, S. H. A plasminogen activator inhibitor-1 inhibitor reduces airway remodeling in a murine model of chronic asthma. Am. J. Respir. Cell Mol. Biol. 46, 842–846 (2012).
Brown, N. J. Therapeutic potential of plasminogen activator inhibitor-1 inhibitors. Ther. Adv. Cardiovasc Dis. 4, 315–324 (2010).
Kutz, S. M., Higgins, C. E. & Higgins, P. J. Novel combinatorial therapeutic targeting of PAI-1 (SERPINE1) gene expression in Alzheimer’s disease. Mol. Med Ther. 1, 106 (2012).
Jacobsen, J. S. et al. Enhanced clearance of Aβ in brain by sustaining the plasmin proteolysis cascade. Proc. Natl Acad. Sci. USA 105, 8754–8759 (2008).
Lin, L. et al. Reelin promotes the adhesion and drug resistance of multiple myeloma cells via integrin β1 signaling and STAT3. Oncotarget 7, 9844–9858 (2016).
Qin, X. et al. Extracellular matrix protein Reelin promotes myeloma progression by facilitating tumor cell proliferation and glycolysis. Sci. Rep. 7, 45305 (2017).
Jandial, R., Choy, C., Levy, D. M., Chen, M. Y. & Ansari, K. I. Astrocyte-induced Reelin expression drives proliferation of Her2+-breast-cancer metastases. Clin. Exp. Metastasis 34, 185–196 (2017).
Deshpande, K. et al. Neuronal exposure induces neurotransmitter signaling and synaptic mediators in tumors early in brain metastasis. Neurooncol 24, 914–924 (2022).
Park, K.-S. et al. A crucial requirement for Hedgehog signaling in small-cell lung cancer. Nat. Med. 17, 1504–1508 (2011).
Jahchan, N. S. et al. A drug repositioning approach identifies tricyclic antidepressants as inhibitors of small-cell lung cancer and other neuroendocrine tumors. Cancer Discov. 3, 1364–1377 (2013).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies and species. Nat. Biotechnol. 36, 411–420 (2018).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902.e21 (2019).
Corces, M. R. et al. An improved ATAC–seq protocol reduces background and enables interrogation of frozen tissues. Nat. Methods 14, 959–962 (2017).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinforma. Oxf. Engl. 25, 2078–2079 (2009).
Zhang, Y. et al. Model-based analysis of ChIP–seq (MACS). Genome Biol. 9, R137 (2008).
Stark R., Brown G. DiffBind: differential binding analysis of ChIP–seq peak data. v. 3.17 (Bioconductor, 2023).
Patro, R., Duggal, G., Love, M. I., Irizarry, R. A. & Kingsford, C. Salmon provides fast and bias-aware quantification of transcript expression. Nat. Methods 14, 417–419 (2017).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Yu, G., Wang, L.-G., Han, Y. & He, Q.-Y. clusterProfiler: an R Package for comparing biological themes among gene clusters. OMICS 16, 284–287 (2012).
Ge, S. X., Jung, D. & Yao, R. ShinyGO: a graphical gene-set enrichment tool for animals and plants. Bioinformatics 36, 2628–2629 (2020).
Zamanian, J. L. et al. Genomic analysis of reactive astrogliosis. J. Neurosci. 32, 6391–6410 (2012).
Durinck, S., Spellman, P. T., Birney, E. & Huber, W. Mapping identifiers for the integration of genomic datasets with the R/Bioconductor package biomaRt. Nat. Protoc. 4, 1184–1191 (2009).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Acknowledgements
We thank members of the Sage laboratory for their help throughout this study (including A. He for his help with SUBr1 cells), P. Chu for her help with tissue sections and K. Guttenplan for his help with astrocyte cultures. We thank C. Paiato and M. Kleinberger for their technical assistance with immunostaining of patient brain metastasis samples. This work was supported by the NIH (J.S., CA231997 and CA217450; C.K., CA231997), a Damon Runyon Cancer Research Foundation fellowship (F.Q., DRG-2322-18), Stanford Graduate Fellowships (D.Y. and M.C.L.) and a Tobacco-Related Disease Research Program (TRDRP) Predoctoral Fellowship (M.C.L., T32DT4747). J.S. is the Elaine and John Chambers Professor in Pediatric Cancer. All illustrations were created with BioRender.com.
Author information
Authors and Affiliations
Contributions
F.Q., M.M.W. and J.S. conceived the study based on observations from D.Y. F.Q. designed, performed and analysed most of the experiments with cells and mice, with help from D.Y., J.K., M.C.L. and G.G.H. S.C.B., C.J.M., A.P.D. and D. B. performed computational analyses. F.Q., W.M. and A.P. developed and analysed the assembloid assays under the supervision of A.M.P. M.P.-P., K.L.S. and C.D. provided the CDX14P model. A.T. and C.K. provided human samples and helped with histopathological analyses. M.D. helped generate the SuBr1 model. E.T. and M.P. provided human SCLC brain metastasis samples and helped with histology. F.Q. and J.S. wrote the paper and prepared the figures with inputs from all authors.
Corresponding author
Ethics declarations
Competing interests
J.S. licensed a patent to Forty Seven/Gilead on the use of CD47-blocking strategies in SCLC and has equity in, and is an advisor for, DISCO Pharmaceuticals. M.M.W. has equity in, and is an advisor for, D2G Oncology. M.D. has received recent research support from Novartis, Abbvie, United Therapeutics, Verily and Varian, and has consulted with Beigene, Astra Zeneca and Jazz Pharmaceuticals. M.P. has received honoraria for lectures, consultation or advisory board participation from the following for-profit companies: Bayer, Bristol-Myers Squibb, Novartis, Gerson Lehrman Group (GLG), CMC Contrast, GlaxoSmithKline, Mundipharma, Roche, BMJ Journals, MedMedia, Astra Zeneca, AbbVie, Lilly, Medahead, Daiichi Sankyo, Sanofi, Merck Sharp & Dome, Tocagen, Adastra, Gan & Lee Pharmaceuticals and Servier. The other authors declare no competing interests.
Peer review
Peer review information
Nature Cell Biology thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Modeling SCLC brain metastasis in mice following injection of mouse SCLC cells.
a. Survival data of NSG mice with N2N1G and 16 T mouse SCLC brain allografts (n = 6). b. Representative images of hematoxylin and eosin (H&E) counterstained-brain sections for the N2N1G mouse brain allografts and for human SCLC brain metastases. Similar to human tumors, mouse tumors display stippled chromatin, nuclear molding, scant cytoplasm, and frequent mitotic figures and apoptosis. Dark purple: tumor. Scale, 200 μm. Similar results were observed from 3 biologically independent samples. c. Representative images of luciferase (luc) bioluminescence activity from growing tumors after injection of N2N1G cells stably expressing luciferase (N2N1G-luc). d. Quantification of luciferase bioluminescence signal from 5 mice as in (c). e. Representative image of luciferase bioluminescence following intracarotid injection of N2N1G-luc cells (28 days). White arrow: subcranial tumor; asterisk: liver metastases. f. Representative images of hematoxylin and eosin (H&E) counterstained-brain and liver sections with parenchymal (PA), leptomeningeal (LP), ventricular (VC) brain metastases and liver metastases following injections of N2N1G-luc cells. Scale bar, 1 mm. g. Quantification of (f) for intracranial, intracardiac, and intracarotid injections. n = 9 mice per approach. h. Quantification of N2N1G tumors at different sites (n = 3 each); these tumors were used for analyses in Fig. 1b-d. Data show mean with SD. P value calculated by one-way ANOVA.
Extended Data Fig. 2 Reactivation of astrocytes in mouse SCLC brain metastases analyzed by RNA sequencing.
a. Representative fluorescent image of an N2N1G mouse brain allograft (cancer cells are GFP+) 18 days post-injection. Similar results were observed from 6 biologically independent samples from 2 experiments. Scale bar, 1 mm. b. Representative flow cytometry data of all GFP+ cells isolated from an N2N1G brain tumor core, its edge, and the sham control side. c. Uniform Manifold Approximation and Projection (UMAP) analysis of scRNA-seq of GFPneg stromal cells isolated from an N2N1G brain tumor core, its edge, and the sham control side. d. Percentage of each cell type in GFPneg stromal cell populations as in (c). e. Uniform Manifold Approximation and Projection (UMAP) of astrocyte populations from scRNA-seq data from (c). More than 3 populations of astrocytes as in (d) were found when the analysis was focused on astrocytes. f. Expression plot of reactive astrocyte signature genes in astrocyte populations within tumors, at the tumor edge, and in sham injection control regions. g. Schematic representation of the immunopanning protocol to isolate SCLC-associated and injury-associated (surgery and sham injection) control astrocytes before bulk RNA sequencing and analysis. Astrocytes were isolated from the two sides of the brain of the same mice (N2N1G allograft model). Created with BioRender.com h. Gene ontology (GO) enrichment (top 10) for the genes that are upregulated in tumor-associated astrocytes (TA) compared to control astrocytes (CA) as in (g). FDR, false discovery rate. i. Gene expression heatmap of TA and CA grouped by signature genes of alternative activation, inflammatory genes, mature, and progenitor astrocytes (RNA-seq). j. Gene Set Enrichment Analysis (GSEA) for an MCAO (middle cerebral artery occlusion)-induced activation signature (Heiland et al., 2019) for genes upregulated in N2N1G brain TA compared to CA.
Extended Data Fig. 3 Reactive astrocytes infiltrate SCLC brain metastases in mice and humans.
a. Representative images showing immunofluorescent staining of GFAP (red, astrocytes) and human mitochondria (hMito, green, SCLC CDX3 xenograft model) on sections from a leptomeningeal metastasis (LP). Scale, 100 μm. Similar results were observed from 3 biologically independent samples. b. Representative images showing immunofluorescent staining of GFAP (red) and GFP (green for SCLC cells) on sections from an immunocompetent mouse SCLC allograft model (KP1-GFP cells). Similar results were observed from 4 biologically independent samples. Blue, DAPI DNA stain. Scale, 50 μm. c, d. Representative images (c) showing immunofluorescent staining of GFAP (red) and GFP+ N2N1G SCLC cells (green) and quantification (d) of colocalization (Pearson’s coefficient for the red and green channels). n = 3 biologically independent samples. Blue, DAPI DNA stain. Scale, 50 μm. e, f. Representative immunofluorescent staining images (e) and quantification (f) of cleaved-caspase 3 (c-cas3) expression on a section from a GFP+ N2N1G SCLC brain allograft. n = 3 biologically independent samples. P values calculated by one-way ANOVA for (d) and (f). g-h. Representative immunofluorescent staining images (g) and quantification (h) of perivascular and non-perivascular astrocytes (GFAP+). CD31 stains endothelial cells in the vasculature. n = 3 biologically independent samples. Data show mean with SD. P values calculated by two-sided t-test when comparing two groups and one-way ANOVA when comparing three and more groups.
Extended Data Fig. 4 Astrocytes activated by SCLC cells promote the growth and survival of mouse and human SCLC cells.
a. Expression of upregulated cell migration genes in astrocyte populations (as in Extended Data Fig. 2e) from tumor or tumor edge compared to a sham injection control. n = 3 animals. b. Representative high magnification images of immunofluorescent staining of GFAP (red) and GFP (green for human cancer cells) at an SCLC-hCO assembloid fusion interface showing the cell body of astrocytes within the SCLC aggregate. Showing separate channel images from Fig. 3d. Scale, 20 μm. c. Quantification of human astrocyte chemotaxis in control medium and human cancer cell-conditioned medium. d. Schematic description of the astrocyte-SCLC co-culture assay. Created with BioRender.com e, f. Cell viability (AlamarBlue assay, RFU: relative fluorescent unit) (d) and apoptosis (caspase3/7 activity, RLU: relative luminescence unit) (e) measured in N2N1G cells cultured with (red) or without (black) mouse astrocytes (mA). n = 3 independent experiments. g. Representative images showing immunofluorescent staining of human astrocytes (hA) cultured alone or co-cultured with human SCLC cell lines (SUBr1, NCI-H69, and NCI-H82). GFAP (green), Vimentin (VIM, red). Blue, DAPI DNA stain. Scale, 20 μm. h. Quantification of GFAP fluorescent intensity relative to Vimentin (n = 6 from 3 independent experiments). i. Quantification of hA chemotaxis toward human SCLC-conditioned medium. N = 3 independent experiments. Data show mean with SD. P values calculated by two-sided t-test for (c)(f)(h)(i), and two-way ANOVA for (e).
Extended Data Fig. 5 Paracrine mechanisms in the promotion of the growth and survival of SCLC cells by reactive astrocytes.
a, b. Cell viability (AlamarBlue assay, RFU: relative fluorescent unit) (a) and apoptosis (caspase3/7 activity, RLU: relative luminescence unit) (b) measured in NCI-H69 cells cultured with (red) or without (black) human astrocytes (hA), as in Extended Data Fig. 3c. c, d. As in (a-b) with NCI-H82 cells. e, f. As in (a-b) with SUBr1 cells. g, h. Relative cell viability (luciferase activity) of NCI-H69 (g) or NCI-H82 (h) cells co-cultured with (blue) or without (red) direct contact with hA during co-culture. i. Relative cell viability of NCI-H69 cells cultured in astrocyte-conditioned medium (hA Con-AM), SCLC-astrocyte co-culture-conditioned medium (hA + SCLC Con-AM), or SCLC-conditioned medium (SCLC Con-AM). n = 3 independent experiments. P values calculated by two-sided t-test for (b)(d)(f)(i) and two-way ANOVA for (a)(c)(e)(g)(h).
Extended Data Fig. 6 Correlation between NFIB levels and the growth of SCLC brain metastases.
a. Immunoassay (by WES capillary transfer) for NFIB expression in NFIB-high 16 T and N2N1G mouse SCLC cells and in NFIB-low KP22 and 5PFBl cells. HSP90 serves as a loading control. Similar results were observed from 2 independent experiments. b. Representative image of immunofluorescent staining for NFIB (red) and N2N1G NFIB-high SCLC cells (green) on sections from a brain allograft (right). Similar results were observed from 3 biologically independent samples. Scale bar, 20 μm. c. Summary of the ability of various mouse and human SCLC tumors to grow in the brain following intra-cranial injection and subcutaneously. d. Representative images of subcutaneous xenografts and allografts from models in (c) after 14 days in NSG mice. Scale bar, 5 mm. e. Representative images of brains sections from models in (c) after 21 days. f. Quantification of tumor growth of shCtrl (control) and shNfib N2N1G-luc cells measured by bioluminescence imaging (n = 5 tumors from 5 mice each genotype). g. Cell viability measured by AlamarBlue assay of shCtrl and shNfib N2N1G cells at 48 hours in culture. n = 3 independent experiments. h. Immunoassay for NFIB expression in 16 T cells with control and shNfib shRNAs (n = 1 experiment). HSP90: loading control. i-j. Quantification of tumor size of control and shNfib 16 T brain allografts (i) and subcutaneous allografts (j). n = 5 (control and shNfib-2) and 6 (shNfib-1) mice from 2 experiments. k. Representative immunohistochemistry (IHC) for NFIB and GFAP expression (brown signal) in brain metastases sections from SCLC patients. Scale bar, 50 μm. l. Quantification of (k) (n = 20 regions from 10 patient metastases) for GFAP staining area and NFIB intensity (brown signal). Data show mean with SD. P-values calculated by two-sided t-test when comparing two groups and by one-way ANOVA when comparing three groups.
Extended Data Fig. 7 NFIB levels and activity do not significantly change in the brain microenvironment compared to other tumor sites.
a. Immunoassay (by WES capillary transfer) for NFIB expression in N2N1G cells isolated from brain, liver, and subcutaneous (SubQ) tumors. HSP90 serves as a loading control. b. Quantification of NFIB protein levels from (a) (n = 3 tumors from 3 mice). Data show mean with SD. P values calculated by two-sided t-test. c. Schematic description for the ATAC-seq analysis of N2N1G cells isolated from brain allograft, liver metastases, SubQ tumors, and cells in culture. Created with BioRender.com d. Clustering of ATAC-seq data of 3 brains, 3 livers, 3 SubQ, and 2 N2N1G in culture. e. Box plot showing accessibility at NFIB peaks across the different samples, as indicated (ATAC-seq datasets). For each box, the line represents the median of the spread. Box, 75% of all peaks; whiskers, 25% outlier peaks. n = 3 biologically independent samples. P values calculated by two-sided Wilcox’s test for non-parametric data. f. Number and percentage of site-specific and shared peaks that are more accessible in N2N1G cells from brain and liver tumors compared to subcutaneous tumors (cut-off: 1.5x more accessible).
Extended Data Fig. 8 Reelin-Vldlr signaling regulates SCLC-induced astrocyte migration.
a. Chromatin accessibility at two sites in the Reln locus assessed by ATAC-seq in shCtrl (control) and shNfib 16 T and KP1 cells, and KP22 cells overexpressing empty vector (oe empty) or mouse NFIB (oe Nfib). Chromatin immunoprecipitation (ChIP) for NFIB in 16 T, KP1, and KP22 cells. b. Immunoassay for NFIB in shCtrl and shReln N2N1G cells. HSP90 serves as a loading control. c. Quantification of (b) (n = 3 independent experiments). d. Representative images showing immunofluorescence staining of Reelin (red) and GFP (green, cancer cells), or Vldlr (gray) and GFAP (red) on an N2N1G brain allograft section. Blue: DAPI DNA stain. Scale bar, 50 μm. Similar results were observed from 3 biologically independent samples. e. Immunoassay for Reelin in shCtrl and shRELN NCI-H69 cells. HSP90 serves as a loading control (n = 1). f. Representative images showing immunofluorescent staining of human astrocytes (hA, marked by Vimentin) for the reactivation marker GFAP (green) when co-cultured (CC) with shCtrl or shRELN NCI-H69 cells. Blue: DAPI to stain DNA. Scale bar, 20 μm. g. Quantification of fluorescent intensity for GFAP normalized to Vimentin (n = 3 independent experiments) as in (f). h. Representative images showing rare immunofluorescent staining of GFAP (green) in Vimentin+ (red) hA treated with or without recombinant human Reelin. Blue: DAPI DNA stain. Scale bar, 20 μm. Similar results were observed from 2 independent experiments. i. Cell viability (AlamarBlue assay) measured in shCtrl and shRELN NCI-H69 cells cultured with hA compared to without hA (n = 3 independent experiments). j. Chemotaxis of mouse astrocytes (mA) in control medium or medium with recombinant mouse Reelin protein and the mouse Reelin-blocking antibody CR-50. k. Relative viability of mA in shCtrl and shReln N2N1G-conditioned medium with or without the addition of recombinant mouse Reelin compared to control medium (n = 3 independent experiments). l. Same as (i) for hA and human Reelin. m. Immunoassay for VLDLR in shCtrl and shVLDLR hA. HSP90 serves as a loading control (n = 1 experiment). n. Chemotaxis assay for shCtrl and shVLDLR hA with or without hSCLC-conditioned medium (n = 3 independent experiments). o. Viability of shDAB1 hA cells at 48 hours. Data show mean with SD. n = 3 independent experiments. P values calculated via two-sided t-test when comparing two groups and via one-way ANOVA when comparing 3-4 groups.
Extended Data Fig. 9 Reelin expression is critical for the recruitment of GFAP+ astrocytes and the growth of SCLC brain metastases.
a. Cell viability of shCtrl (control) and shReln N2N1G cells measured by AlamarBlue assay (n = 3). b, c. Representative image and tumor size of shCtrl and shReln N2N1G subcutaneous allografts 21 days post-transplant. Scale bar, 5 mm. N = 9 from 2 experiments. d-f. Tumor size of shCtrl and shReln 16 T brain allografts (d), subcutaneous allografts (e), and liver metastases (f) 21 days post-transplant. N = 7 (shCtrl) and 8 (shReln) from 2 experiments in (d). N = 9 mice from 3 experiments in (e) and (f). g. Representative images of immunohistochemistry staining of Reelin (brown, with blue hematoxylin counterstain) on SCLC brain tumors generated by shCtrl and shReln N2N1G cells. Scale bar, 100 μm. Similar results were observed from 3 independent experiments. h-i. Representative images (h) and quantification (i, n = 3 independent experiments) of shCtrl and shReln SCLC cell migration from 3D cancer spheres in Matrigel. j. Representative images of immunofluorescent staining of GFAP (white) in SCLC brain tumors grown from a 1:1 mix of shCtrl and shReln2 16 T cells expressing mCherry (red) or GFP (green), as indicated. Blue: DAPI DNA stain. Scale bar, 20 μm. Yellow asterisk shows regions where astrocytes are absent. k. Quantification of tumor size in (j). l-m. Quantification of the ratio between mCherry+ and GFP+ of 16T cells growing in the brain from (Fig. 6h and Extended Data Fig. 9j). N = 6 mice from 2 independent experiments. n. Representative images of immunofluorescent staining of CD31 and GFAP in shCtrl and shReln N2N1G brain allografts. GFP labels N2N1G cells, DAPI stain for nucleus. Scale bar, 20 μm. o-p. Quantification of CD31 level and perivascular GFAP-positive astrocytes in shCtrl and shReln N2N1G brain allografts. N = 4 tumors. q. Chemotaxis of mouse astrocytes in conditioned media from 4T1 cells, 4T1 cells expressing control vector, and 4T1 cells expressing mouse Reelin central domain. n = 3 independent experiments. r. Quantification of liver metastasis size by luciferase imaging of the abdomen of mice 14 days after intracardiac injections of 4T1 cells expressing control vector or mouse Reelin central domain (Leu1221-Ile 2661). N = 6 mice from 2 experiments. Data show mean with SD. P values calculated by two-sided t-test when comparing two groups and one-way ANOVA when comparing three or more groups.
Extended Data Fig. 10 Astrocytes activated by SCLC cells secrete factors promoting the survival of SCLC cells.
a. Volcano plot showing differentially expressed genes in hA co-cultured with NCI-H82 compared to hA alone. b. Gene ontology (GO) enrichment (top 9) for the genes that are upregulated in hA co-cultured with NCI-H82 compared to hA alone. c. Immunoassay (by WES capillary transfer) for SERPINE1 expression in naive hA and hA co-cultured with the NCI-H69 and SUBr1 human SCLC cell lines. HSP90 serves as a loading control. d. Quantification of (c) (n = 2 independent experiments). e. The expression level of SERPINE1 in astrocyte populations (as in Extended Data Fig. 2f) from N2N1G brain allograft tumor, tumor edge, and sham control brain. n = 3 biologically independent samples. f-g. Representative images and quantification of immunofluorescent staining of SERPINE1 in patient SCLC brain metastases. NCAM stains SCLC cells. GFAP stains astrocytes. DAPI stains nucleus. Scale bar, 20 μm. n = 3 samples. h. Ratio of active SERPINE1 measured by ELISA in culture medium from mouse astrocytes co-cultured with N2N1G cells for 2 days and then treated with DMSO (control) and Tiplaxtinin (5 μM). i. Cell viability (AlamarBlue assay) measured in N2N1G cells treated with recombinant mSERPINE1 and Tip (n = 3 experiments). j, k. Cell viability (AlamarBlue assay) measured in NCI-H69 cells (j) and SUBr1 cells (k) treated with recombinant hSERPINE1 and the SERPINE1 inhibitor Tiplaxtinin (Tip) (n = 3 experiments). l-m. Relative cell viability (AlamarBlue assay) measured in NCI-H69 cells (l) and SUBr1 cells (m) cultured with hA compared to without hA, with or without Tiplaxtinin treatment (n = 3 experiments). n-o. Relative apoptosis (caspase3/7 activity) measured in NCI-H69 cells (n) and SUBr1 cells (o) cultured with hA compared to without hA, with or without Tip treatment (n = 3 experiments). p. Overlap between upregulated genes in NCI-H82 cultured with hA and N2N1G cells growing in the brain compared to subcutaneously. q. Apoptosis measured by cleaved caspase3/7 in shCtrl (control) and shReln N2N1G and 16T cells in culture. r. Immunoassay for SERPINE1 expression in N2N1G cells in Fig. 7m, n. Data show mean with SD. P values calculated via two-sided t-test when comparing two groups and one-way ANOVA when comparing three groups.
Supplementary information
Supplementary Data 1
FACS gates for GFP+ and GFP− cells.
Supplementary Table
Supplementary Tables
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Statistical source data.
Source Data Fig. 7
Statistical source data.
Source Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 9
Statistical source data.
Source Data Extended Data Fig. 10
Statistical source data.
Source Data Fig. 4
Unprocessed immunoassay.
Source Data Fig. 5
Unprocessed immunoassay.
Source Data Extended Data Fig. 6
Unprocessed mmunoassay.
Source Data Extended Data Fig. 7
Unprocessed immunoassay.
Source Data Extended Data Fig. 8
Unprocessed immunoassay.
Source Data Extended Data Fig. 10
Unprocessed immunoassay.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Qu, F., Brough, S.C., Michno, W. et al. Crosstalk between small-cell lung cancer cells and astrocytes mimics brain development to promote brain metastasis. Nat Cell Biol 25, 1506–1519 (2023). https://doi.org/10.1038/s41556-023-01241-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-023-01241-6