Abstract
In animals, maternal diet and environment can influence the health of offspring. Whether and how maternal dietary choice impacts the nervous system across multiple generations is not well understood. Here we show that feeding Caenorhabditis elegans with ursolic acid, a natural plant product, improves axon transport and reduces adult-onset axon fragility intergenerationally. Ursolic acid provides neuroprotection by enhancing maternal provisioning of sphingosine-1-phosphate, a bioactive sphingolipid. Intestine-to-oocyte sphingosine-1-phosphate transfer is required for intergenerational neuroprotection and is dependent on the RME-2 lipoprotein yolk receptor. Sphingosine-1-phosphate acts intergenerationally by upregulating the transcription of the acid ceramidase-1 (asah-1) gene in the intestine. Spatial regulation of sphingolipid metabolism is critical, as inappropriate asah-1 expression in neurons causes developmental axon outgrowth defects. Our results show that sphingolipid homeostasis impacts the development and intergenerational health of the nervous system. The ability of specific lipid metabolites to act as messengers between generations may have broad implications for dietary choice during reproduction.
Similar content being viewed by others
Main
Animal exposure to environmental and dietary changes can modify the physiology and development of offspring1,2. Certain parentally acquired traits may also be inherited over multiple generations3,4. Epigenetic mechanisms of inheritance exploiting small RNAs, DNA methylation and chromatin modifications have previously been defined2,3,4,5,6,7. However, the role of maternal provisioning to offspring as a means of multigenerational inheritance has not been well explored. Maternal provisioning potentially enables the inheritance of information beyond nucleic acids—affording the transmission of lipids, proteins and metabolites8. Such information flow could directly report ancestral experience to alter the physiology and development of offspring across generations, and potentially shape the evolutionary trajectory.
Whether altered maternal metabolism and provisioning impacts the neuronal health of offspring across generations is an open question and challenging to resolve. The Caenorhabditis elegans model is used to dissect epigenetic mechanisms due to its short generation time and straightforward genetics3,4,7. C. elegans is also suitable for analysing the temporal effects of dietary and environmental changes, as such conditions can be precisely controlled. Furthermore, defined C. elegans neurodevelopmental and disease models allow potential multigenerational effects to be examined9,10. We therefore used C. elegans to identify molecules that may regulate neuronal health across generations.
Axons are long cytoplasmic projections that transmit information between neurons. Neuronal health maintenance requires the transport of cargo (organelles, RNA, proteins and lipids) along the axonal cytoskeleton11,12. Microtubules are major cytoskeletal components comprised of cylindrical structures assembled from α- and β-tubulin heterodimers that mediate intracellular transport11,13. Defective microtubule structure disrupts the supply of essential materials and is associated with multiple neurodegenerative disorders14. Therefore, identification of molecular mechanisms supporting axonal health under conditions of suboptimal microtubule-associated intracellular transport is important. Posterior lateral mechanosensory (PLM) neurons in C. elegans extend axons along the length of the animal to coordinate touch responses15. A previous study showed that loss of the MEC-17 (a homologue of αTAT1) α-tubulin acetyltransferase causes progressive adult-onset PLM axon degeneration9. MEC-17 loss causes microtubule instability and aberrant axonal transport, resulting in disrupted distribution of mitochondria and synaptic proteins9. Degeneration of PLM axons is caused by axon fragility, as the phenotype is suppressed by paralyzing mec-17-mutant animals9. Furthermore, PLM axon fragility in mec-17 mutants is exacerbated in animals with increased body length (for example, lon-2 mutants), probably due to the added demand on axonal transport that is required to maintain a longer axon9. We used the well-defined MEC-17 and LON-2-deficient axon fragility model to identify mechanisms that maintain axon integrity across generations.
In this paper we show that supplementation with the natural product ursolic acid (UA), specifically during oocyte production, promotes axon transport and reduces axon fragility intergenerationally in C. elegans. Ursolic acid protects axons by upregulating expression of the sphingolipid biosynthetic enzyme acid ceramidase-1 (asah-1) gene in the intestine. Intestine-to-oocyte transport of sphingosine-1-phosphate (S1P), a downstream bioactive sphingolipid metabolite, in maternal yolk is required for intergenerational neuroprotection through upregulation of asah-1 expression in subsequent generations. Furthermore, we show that transcriptional regulation of asah-1 is dependent on the intestinal transcription factors PQM-1 (GATA zinc finger) and CEH-60 (three-amino-acid loop extension (TALE) class). Together, our study reveals that a short-term dietary supplement during the maternal reproductive period can be neuroprotective over multiple generations—providing a paradigm for intergenerational inheritance of health-promoting traits. We propose that intergenerational regulation of metabolism and metabolic gene expression is a universal principle underlying intergenerational inheritance across evolution.
UA reduces axon fragility intergenerationally
To identify axon health-promoting molecules, we examined the morphology of PLM neurons in wild-type and mec-17(ok2109); lon-2(e678) animals (Fig. 1). As shown previously, the mec-17(ok2109); lon-2(e678) animals exhibited approximately 50% penetrant PLM axon breaks in day 3 adults (Fig. 1b,d)9. In a screen of natural products, we identified UA as a suppressor of axon fragility in mec-17(ok2109); lon-2(e678) animals (Fig. 1 and Extended Data Fig. 1). Ursolic acid is a lipophilic pentacyclic triterpenoid acid found in plants that has broad biological functions, acting as an anti-inflammatory, antioxidant and neuroprotective molecule16,17 (Fig. 1c). To assess the potency of UA-induced neuroprotection, we fed mec-17(ok2109); lon-2(e678) mothers with different concentrations of UA and examined the fragility of the PLM axons (Fig. 1a–d and Extended Data Fig. 2a). We found that the mec-17(ok2109); lon-2(e678) F1 progeny of animals incubated with 50 μM UA from larval stage 4 (L4) had reduced axon degeneration (Fig. 1d and Extended Data Fig. 2a). To determine whether the UA-induced suppression of axon fragility was due to reduced body length or motility, we used WormLab tracking. We found no change in the motility or body length of mec-17(ok2109); lon-2(e678) animals exposed to UA compared with the controls, which suggested a molecular rather than physical effect (Extended Data Fig. 2b–e).
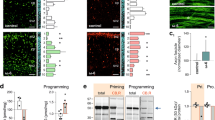
a,b, Schematics (top) and fluorescence micrographs (bottom) of posterior lateral mechanosensory left (PLML) anatomy in wild-type (a) and mec-17(ok2109); lon-2(e678) (b) animals expressing the Pmec-4::gfp transgene (zdIs5). Left lateral view, anterior to the left. Scale bars, 25 μm. b, A typical PLM axon break (red line) observed in adult (3-d-old) mec-17(ok2109); lon-2(e678) animals is indicated. c, Chemical structure of UA. d, Adult mec-17(ok2109); lon-2(e678) F1 animals exposed to UA (50 μM) from P0 L4 to day 3 of F1 had reduced PLM axon breaks. e, Timeline of UA exposure. The stages of C. elegans development relevant to UA exposure in the P0 and F1 generations are shown (top). Vertical dashed line, demarcation between the P0 and F1 generations. f, Continuous UA exposure (P0 L4 larva to F1 adult) reduces PLM axon breaks in mec-17(ok2109); lon-2(e678) animals. Specifically, UA exposure of P0 animals from the L4 larval stage to adult (ii), but not earlier (i) or later ((iii) and (iv)) stages results in a reduction in PLM axon breaks in mec-17(ok2109); lon-2(e678) animals. g, Experimental scheme for the intergenerational inheritance experiment. P0 animals (L4 larvae) were treated with DMSO (control) or UA for 16 h. Animals of each generation were allowed to lay eggs on untreated plates for 3 h and the adults (3-d-old) were assessed for axon breaks. h, The progeny of mec-17(ok2109); lon-2(e678) P0 mothers exposed to UA for 16 h (L4 to young adult) have reduced PLM axon breaks for two generations (F1 and F2). d,f,h, n = 101 and 103 (d); f, n = 249, 199, 173, 151, 166 and 107 (f); and n = 126, 128, 103, 128, 77 and 97 (h) hermaphrodite animals per condition (left to right). P values were determined using a one-way analysis of variance (ANOVA; f) or unpaired Student’s t-test (d,h). ****P ≤ 0.0001; ***P ≤ 0.001; **P ≤ 0.01; and NS, not significant. Error bars indicate the s.e.m. Source data are provided.
We investigated whether there was a critical developmental period for UA to reduce PLM axon fragility. For this, we exposed mec-17(ok2109); lon-2(e678) animals to UA at the following stages of C. elegans development: P0 L3 to L4 (during sperm generation and before oocyte production), P0 L4 to adult (oocytes and sperm present), F1 embryo to L1 (embryogenesis) and F1 L1 to adult (larval stages and adult; Fig. 1e). We found that axon breaks in the F1 adult progeny were only reduced when P0 hermaphrodites containing oocytes and sperm (P0 L4 to adult) were exposed to UA (Fig. 1e,f). Furthermore, a reduction in PLM axon breaks was not observed in P0 animals or their F1 progeny when the P0 embryos were exposed to UA, suggesting that UA does not penetrate the eggshell (Extended Data Fig. 2f). These data suggest that the UA health-promoting effect is deposited in gametes to maintain axonal health through to adulthood. We thus examined whether UA can protect the nervous system in subsequent generations. We incubated P0 L4 hermaphrodites with dimethylsulfoxide (DMSO; control) or UA for 16 h and then transferred the now adults to untreated plates to lay eggs for 3 h (Fig. 1g). These F1 eggs therefore underwent oocyte maturation during UA exposure. When the F1 animals reached the L4 stage, a cohort were transferred to lay eggs for analysis of the next generation and the remainder matured until day 3 of adulthood to examine PLM axon health (Fig. 1g,h). We repeated this process for the subsequent generations. We found that incubation with UA during P0 oocyte maturation reduced PLM axon breaks in the F1 and F2 generations but not the F3 generation (Fig. 1g,h), revealing an intergenerational neuroprotective effect of UA.
Inherited neuroprotection requires intestine–oocyte transport
How does UA protect the nervous system intergenerationally? As the functional period is during oocyte production and maturation, we hypothesized that UA may affect maternal yolk provisioning to oocytes. Yolk synthesized in the intestine of C. elegans hermaphrodites contains lipids and lipoproteins that provide oocytes, and thus embryos, with nutrients for development8. Oocyte yolk import occurs through endocytosis and requires receptor-mediated endocytosis-2 (RME-2), a low-density lipoprotein receptor18. We found that RME-2 is required for UA to reduce PLM neuron fragility, providing support for a role for intestine–oocyte transport (Fig. 2a). Animals lacking RME-2 are also defective in the transport of RNAs that are major transmitters of epigenetic inheritance19. The HRDE-1 argonaute and ZNFX-1 helicase are critical for small-RNA inheritance; however, hrde-1 and znfx-1 are dispensable for UA to reduce PLM breaks intergenerationally (Fig. 2b,c)20,21,22. These data suggest that UA stimulates alternative factors in the maternal yolk and that this information optimizes the oocyte/embryonic environment to promote axon health.
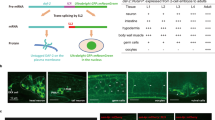
a, RNAi-mediated knockdown of rme-2 suppresses UA-induced reduction of PLM axon breaks in mec-17(ok2109); lon-2(e678) animals. b,c, The progeny of P0 mothers (mec-17(ok2109); lon-2(e678), hrde-1(tm1200); mec-17(ok2109); lon-2(e678) (b) and znfx-1(gg561); mec-17(ok2109); lon-2(e678) (c) animals) exposed to UA for 16 h (L4 to young adult) have reduced PLM axon breaks for two generations (F1 and F2). Experimental timing as in Fig. 1g. d, Schematic of transcriptome analysis following UA exposure (top). Total RNA was isolated from L4 larvae exposed to UA for 12 h (compared with the DMSO control). asah-1 was upregulated following UA exposure (Supplementary Table 1). The upregulation of asah-1 messenger RNA in wild-type L4 larvae after 12 h of UA exposure was independently confirmed using qPCR (bottom; n = 3 independent experiments). The housekeeping gene cdc-42 was used as the control. e,f, Loss of asah-1 by RNAi (e) or gene deletion (f) suppresses the ability of UA to reduce PLM axon breaks in mec-17(ok2109); lon-2(e678) animals. g, Representative fluorescence micrographs (top) and calculated levels (bottom) of the Pasah-1::nls::gfp transcriptional reporter. Expression was detected in the intestine from late embryogenesis through to adult (see Extended Data Fig. 4). Pasah-1::nls::gfp expression increased in animals exposed to UA for 12 h compared with the controls (DMSO); a.u. arbitrary units. Lateral views, anterior to the left, of L4 larvae are shown. Pharynx marked by a white asterisk. Scale bar, 25 μm. a–c,e–g, n = 186, 186, 162 and 154 (a); n = 90, 69, 61, 65, 71, 71, 160, 156, 145, 158, 73 and 68 (b); n = 98, 101, 99, 100, 76, 71, 115, 115, 115, 114, 116 and 121 (c); n = 126, 101, 148 and 94 (e); n = 223, 220, 202 and 194 (f); and n = 37 and 43 (g) hermaphrodite animals per condition (left to right). a–g, P values were determined using an ANOVA (a–c,e) or unpaired Student’s t-test (d,f,g). ****P ≤ 0.0001; ***P ≤ 0.001; **P ≤ 0.01; *P ≤ 0.05; and NS, not significant. Error bars indicate the s.e.m. Source data are provided.
UA induces intestinal acid ceramidase expression
To identify the factor(s) regulated by UA, we examined the transcriptomes of synchronized L4 larvae that had been exposed to UA (or DMSO) for 12 h (Fig. 2d, Extended Data Fig. 3 and Supplementary Table 1). We identified 49 dysregulated genes (false discovery rate of 0.02) and surveyed this dataset for genes expressed in the intestine (the source of yolk) that potentially control lipid metabolism. We detected increased asah-1 transcript levels in the animals exposed to UA, which we confirmed by quantitative PCR (qPCR) analysis (Fig. 2d). The asah-1 gene encodes an acid ceramidase that hydrolyses ceramide into fatty acid and sphingosine23. To assess the requirement of asah-1 for UA to reduce PLM axon fragility, we performed RNA-mediated interference (RNAi) to knockdown asah-1 in mec-17(ok2109); lon-2(e678) animals incubated with UA (Fig. 2e). Treatment with UA did not reduce the PLM axon fragility of asah-1-knockdown animals (Fig. 2e). This result was independently confirmed in animals with a genetic asah-1 deletion (asah-1(tm495) deletion allele) and showed that asah-1 loss caused an increase in PLM axon breaks in untreated animals (Fig. 2f). To identify the tissue in which asah-1 is expressed and potentially regulated by UA, we monitored the spatiotemporal expression of asah-1 using a 2,000-bp sequence upstream of asah-1 to drive nuclear-localized green fluorescent protein (GFP; Fig. 2g and Extended Data Fig. 4). We detected GFP exclusively in the intestinal cells of Pasah-1::gfp animals from late embryos through to adult (Fig. 2g and Extended Data Fig. 4). At all stages, we observed an anterior bias in intestinal GFP expression (Fig. 2g and Extended Data Fig. 4). Increased fluorescence was observed in the intestinal nuclei of Pasah-1::gfp L4 animals exposed to UA for 12 h (Fig. 2g). Thus, UA induces asah-1 transcription in the intestine.
Intestinal ASAH-1 expression protects axons
As asah-1 was upregulated in animals exposed to UA, we investigated whether asah-1 overexpression in mec-17(ok2109); lon-2(e678) animals could mimic UA-induced neuroprotection. Single-cell sequencing corroborates asah-1 intestinal expression; however, low level expression can be detected in other cells/tissues, including the PLM neurons24,25. We therefore overexpressed asah-1 in mec-17(ok2109); lon-2(e678) animals using the following heterologous promoters: intestine (ges-1), hypodermis (dpy-7), muscle (myo-3) and mechanosensory neurons (mec-4; Fig. 3a and Extended Data Fig. 5)26,27,28,29. Overexpression of asah-1 in the intestine, but not hypodermis or muscle, reduced PLM axon breaks in the mec-17(ok2109); lon-2(e678) animals (Fig. 3a). We further found that presence of the Pges-1::asah-1 transgene in progeny is not required for PLM neuroprotection (Fig. 3b), providing support for the idea that ASAH-1 activity in the hermaphrodite intestine protects the PLM neurons in the next generation.
a, Expression of asah-1 cDNA driven by the heterologous intestinal promoter (ges-1), but not the hypodermal (dpy-7) or muscle (myo-3) promoters, resulted in reduced PLM axon breaks in mec-17(ok2109); lon-2(e678) animals. b, PLM axon breaks were reduced in mec-17(ok2109); lon-2(e678) animals derived from Pges-1::asah-1 transgenic animals independently of inheritance of the transgene. The mec-17(ok2109); lon-2(e678) animals were either injected with Pges-1::asah-1 (transgenic lines nos. 1 and 2) or uninjected (−). c,d, Overexpression of asah-1 in the nervous system (rab-3 promoter) caused developmental axon outgrowth defects in the mechanosensory neurons of wild-type animals expressing the Pmec-4::gfp transgene (zdIs5). c, Proportion of PLM axon outgrowth defects in Pmec-4::gfp- and Pmec-4::gfp; Prab-3::asah-1-expressing L1 larvae. d, Schematic (top) and fluorescence micrographs (bottom) of ALM (left/right) and PLM (left/right) axons in wild-type (left) and Prab-3::asah-1-expressing animals (right). The typical axon outgrowth defect observed is marked in red. Left lateral view, anterior to the left. Scale bars, 25 μm. a–c, n = 305, 72, 80, 75, 72, 98 and 97 (a); n = 126, 147, 149, 123 and 124 (b); and n = 62 and 95 (c) hermaphrodite animals per condition (left to right). P values were determined using an ANOVA (a) or unpaired Student’s t-test (b,c). Error bars indicate the s.e.m. ****P ≤ 0.0001; ***P ≤ 0.001; **P ≤ 0.01; *P ≤ 0.05; and NS, not significant. Source data are provided.
Overexpression of asah-1 in the mechanosensory neurons caused PLM axon outgrowth defects in mec-17(ok2109); lon-2(e678) animals (Extended Data Fig. 5a,c), precluding analysis of PLM fragility. Overexpression of asah-1 in the neurons of wild-type animals—either in the mechanosensory neurons (mec-4 promoter) or pan-neuronally (rab-3 promoter)—also caused extensive axon outgrowth defects in the anterior lateral mechanosensory (ALM) and PLM neurons, which develop embryonically, but not the post-embryonic posterior ventral microtubule neurons (Fig. 3c,d and Extended Data Fig. 5a–c). This potential embryonic effect was supported by the detection of PVQ axon guidance defects when asah-1 was neuronally overexpressed (Extended Data Fig. 5f,g). However, neuronal asah-1 overexpression did not cause overt motility defects, suggesting intact global nervous system architecture. These data reveal that intestinal asah-1 reduces PLM axon fragility in animals with defective microtubule stability, and inappropriate neuronal asah-1 expression—which is likely to be associated with disrupted sphingolipid homeostasis—causes cell-autonomous neurodevelopmental defects.
Sphingolipids are amphipathic bioactive molecules with multiple cellular functions, including cell adhesion and migration, cell death and cell proliferation30. Sphingolipid homeostasis is maintained through the de novo or salvage pathways (Fig. 4a)31. The serine palmitoyltransferase (SPT) protein complex is the rate-limiting enzyme in the de novo pathway that generates ceramide, the ASAH-1 substrate (Fig. 4a)32. SPTL-1 knockout causes C. elegans embryonic lethality/larval arrest, probably due to the loss of multiple sphingolipids. Therefore, to assess SPTL-1 regulation of PLM axon fragility, we overexpressed sptl-1 complementary DNA in the intestine. Overexpression of sptl-1 reduced PLM axon breaks in mec-17(ok2109); lon-2(e678) animals, confirming that ceramide or its derivatives are important for axon health (Fig. 4b). Indeed, when mec-17(ok2109); lon-2(e678) animals were incubated with two distinct ceramides containing different fatty-acid-chain lengths (Cer20 (d18:1/20:0) and Cer22 (d18:1/22:0)), there was a reduction in PLM axon breaks (Fig. 4c). In the salvage pathway, lysosomal membrane ceramide is hydrolysed to sphingosine by acid ceramidases, and sphingosine phosphorylation by sphingosine kinases (SphK) generates S1P (Fig. 4a)31. We found that intestinal sphk-1 expression reduced PLM axon breaks in mec-17(ok2109); lon-2(e678) animals, as also shown by overexpression of sptl-1 or asah-1 (Fig. 4b,d,e). Furthermore, sphk-1 loss increased PLM axon fragility and prevented UA-induced reduction of PLM axon breaks in mec-17(ok2109); lon-2(e678) animals (Extended Data Fig. 6a,b). Overexpression of asah-1 in the intestine reduced PLM axon breaks (Fig. 4d). To determine whether sphk-1, and thus potentially S1P generation, is required for ASAH-1 to perform this neuroprotective function, we knocked down sphk-1 in animals overexpressing intestinal asah-1 and evaluated the PLM axon fragility of mec-17(ok2109); lon-2(e678) animals (Fig. 4f). We found that sphk-1 RNAi suppressed the beneficial effect of asah-1 overexpression on PLM axon fragility (Fig. 4f). These data suggest that UA neuroprotection is S1P-dependent.
a, C. elegans orthologues of de novo- and salvage sphingolipid-pathway components. Sphingolipid metabolic enzymes (nematode and mammalian orthologue) and sphingolipid intermediates are shown. CerS, ceramide synthase; CDase, ceramidase; SPP, S1P phosphatase; and SPL, S1P lyase. b,d,e, The function of SPTL-1, ASAH-1 and SPHK-1 (coloured lettering in a) in PLM axon fragility was examined. Overexpression of sptl-1 (b), asah-1 (d) or sphk-1 (e) cDNA in the intestine (ges-1 promoter) reduces PLM axon breaks in mec-17(ok2109); lon-2(e678) animals. Two independent transgenic lines (nos. 1 and 2, as indicated in the coloured bars) were analysed. c, Exposure of P0 mothers to ceramide for 16 h (L4 to young adult) results in a reduction of PLM axon breaks in mec-17(ok2109); lon-2(e678) animals. f, RNAi-induced knockdown of sphk-1 suppresses the ability of intestinal asah-1 expression to reduce PLM axon breaks in mec-17(ok2109); lon-2(e678) animals. Transgenic line no. 2 from d was used. b–f, n = 69, 73, 75 and 73 (b); n = 92, 81 and 88 (c); n = 76, 72, 79 and 80 (d); n = 94, 102, 99 and 98 (e); and n = 99, 97 and 116 (f) hermaphrodite animals per condition (left to right). P values were determined using an ANOVA (c,f) or unpaired Student’s t-test (b,d,e). ****P ≤ 0.0001; ***P ≤ 0.001; **P ≤ 0.01; *P ≤ 0.05; and NS, not significant. Error bars indicate the s.e.m. Source data are provided.
S1P prevents axon fragility intergenerationally
We examined the role of S1P in PLM axonal health directly by incubating mec-17(ok2109); lon-2(e678) animals with different S1P concentrations for one generation (P0 L4 to 3-d-old F1 adult). A concentration of 20 μM S1P reduced PLM axon fragility (Fig. 5a,b). To determine the functional period of S1P, we exposed mec-17(ok2109); lon-2(e678) animals to S1P for two time periods: P0 L4 larvae to adult and L1 larvae to adult. S1P only reduced axon fragility in the F1 progeny when provided to hermaphrodites containing oocytes (P0 L4 to adult)—the same functional period as UA (compare Figs. 5c and 1f). Furthermore, two generations of the progeny of P0 animals incubated with S1P from L4 larvae to adult (16 h) had to reduced axonal fragility, revealing an intergenerational effect (Fig. 5d,e). These data suggest that intestinal S1P is transported within the yolk to oocytes to promote PLM axonal health in subsequent generations. To examine whether S1P can undergo intestine–oocyte transport, we fed wild-type L4 larvae with S1P–fluorescein, a fluorescently labelled S1P analogue (Fig. 5f). Fluorescence was observed in the intestinal tract within 1 h of feeding, suggesting that S1P–fluorescein is not immediately metabolized (Extended Data Fig. 6c). After 16 h of S1P–fluorescein feeding, we detected fluorescence in proximal oocytes, suggesting yolk-dependent intestine–oocyte transport (Fig. 5f). To determine the importance of intestine–oocyte S1P transport to prevent axon fragility, we knocked down rme-2 in mec-17(ok2109); lon-2(e678) animals incubated with S1P. We found that S1P neuroprotection requires rme-2 (Fig. 5g), revealing that S1P intergenerational neuroprotection requires intestine–oocyte transport.
a, Chemical structure of S1P. b, Continuous exposure (P0 L4 larva to F1 adult) to 20 µM S1P reduces PLM axon breaks in mec-17(ok2109); lon-2(e678) animals. c, PLM axon breaks were reduced in mec-17(ok2109); lon-2(e678) animals when P0 mothers (L4 to young adult), but not F1 larvae, were exposed to S1P for 16 h. d, Experimental scheme for the intergenerational inheritance experiment. P0 animals (L4 larvae) were treated with methanol (control) or S1P for 16 h. Animals of each generation were allowed to lay eggs on untreated plates for 3 h and 3-d-old adults were assessed for axon breaks. e, The progeny of P0 mothers (mec-17(ok2109); lon-2(e678) animals) exposed to S1P for 16 h (L4 to young adult) had reduced PLM axon breaks for two generations (F1 and F2). f, S1P–fluorescein was detected in the oocytes of wild-type adult hermaphrodites after 16 h of feeding. A vehicle-treated control (left) and an animal following S1P–fluorescein treatment (right) are shown. Nomarski micrographs (top) and fluorescence images (bottom) of the same animals are provided. The dashed white lines outline oocytes. Lateral view, anterior to the left. Scale bar, 25 μm. g, RNAi-mediated knockdown of rme-2 suppresses S1P-induced reduction of PLM axon breaks in mec-17(ok2109); lon-2(e678) animals. b,c,e,g, n = 73, 73, 78, 74, 76 and 79 (b); n = 105, 104, 135 and 107 (c); n = 158, 163, 153, 157, 157 and 155 (e); and n = 104, 105, 107 and 104 (g) hermaphrodite animals per condition (left to right). P values were determined using an ANOVA (c,g) or unpaired Student’s t-test (b,e). ***P ≤ 0.001; **P ≤ 0.01; *P ≤ 0.05; and NS, not significant. Error bars indicate the s.e.m. Source data are provided.
UA and S1P enhance PLM axon transport
A previous study showed that MEC-17 controls the efficient transport of protein cargos throughout the PLM axon to maintain neuronal health9. We therefore investigated whether reduced PLM axon fragility afforded by UA and S1P to animals lacking MEC-17 is due to improved axon transport. We examined the localization of fluorescent reporters for UNC-104 (a kinesin-3 motor protein) and RAB-3 (a synaptic vesicle-associated small guanosine triphosphatase), which exhibited inappropriate posterior axonal pooling in mec-17(ok2109) animals (Fig. 6a,b). We found that mec-17(ok2109) animals exposed to UA or S1P had reduced axon transport defects (Fig. 6a,b), suggesting that UA and S1P promote the transport of protein cargo along the PLM axon. In a parallel approach to examine the potential effect of UA and S1P on microtubule health, we treated animals with colchicine—a microtubule destabilizing agent33. Previous work showed that colchicine increases PLM axon breaks in animals lacking MEC-17 (ref. 9). We found that UA and S1P reduce PLM axon breaks in mec-17(ok2109); lon-2(e678) animals exposed to colchicine (Fig. 6c). Thus, in the presence of two microtubule destabilizing conditions (MEC-17 loss and colchicine) UA and S1P reduce PLM axon fragility. Critically, we found that UA and S1P protect wild-type PLM axons exposed to high levels of colchicine, revealing that the neuroprotective effect is not directly associated with MEC-17 loss (Fig. 6d).
a,b, Continuous exposure to UA or S1P reduces inappropriate posterior accumulation of UNC-104::GFP (kinesin; a) and mCherry::RAB-3 (pre-synaptic guanosine triphosphatase; b) in the PLM axons of mec-17(ok2109) animals. Scale bars, 25 μm. c,d, Continuous exposure to UA or S1P reduces colchicine-induced PLM axon breaks in mec-17(ok2109); lon-2(e678) (c) and wild-type (d) animals. Wild-type and mec-17(ok2109); lon-2(e678) animals were exposed to 200 µM and 100 µM colchicine, respectively. Two-day-old adult animals were scored. e, Continuous UA or S1P exposure reduces PLM axon breaks in lin-14(n355n679) lon-2(e678) animals. f, Continuous exposure to S1P, but not UA, reduces D-type motor neuron commissure defects in lin-14(n355n679) animals. g, Continuous exposure to UA or S1P reduces PVQ axon defects in ced-10(rp100) animals (g). a–g, Continuous exposure: P0 to L4 larva to F1 to adult; n = 245, 247, 103 and 99 (a); n = 107, 119, 96 and 94 (b); n = 112, 152, 146, 99, 101 and 102 (c); n = 76, 74, 96, 76, 75 and 67 (d); n = 149, 175, 138 and 144 (e); n = 109, 106, 97 and 107 (f); and n = 102, 117, 156 and 150 (g) hermaphrodite animals per condition (left to right). P values were determined using an ANOVA (c,d) or unpaired Student’s t-test (a,b,e–g). ****P ≤ 0.0001; ***P ≤ 0.001; **P ≤ 0.01; *P ≤ 0.05; and NS, not significant. Error bars indicate the s.e.m. Source data are provided.
UA and S1P provide general neuroprotection
To assess the potential broad applicability of UA and S1P neuroprotection, we examined multiple distinct genetic and neuronal models (Fig. 6e–g). A previous study showed that increased microtubule stability can reduce PLM axon breaks caused by mutations to the LIN-14 transcription factor34. We exposed lin-14(n355n679) lon-2(e678) animals to UA or S1P and found that PLM breaks were reduced, suggesting that UA and S1P can generally promote PLM axonal health (Fig. 6e). Next, we examined two other neuron classes—the D-type GABAergic motor neurons and the PVQ interneurons (Fig. 6f,g). LIN-14 loss causes axon breaks of GABAergic D-type motor neuron commissures34. Interestingly, we found that S1P, but not UA, suppresses D-type motor axon breaks in lin-14(n355n679) animals (Fig. 6f), suggesting distinct mechanisms of action or route of entry for S1P in this context. Finally, we explored the potential function of UA and S1P in a neurodevelopmental model. We previously identified a point mutation in the CED-10 Rac GTPase that causes PVQ axon defects35. We found that both UA and S1P reduced PVQ axon defects in ced-10(rp100) animals (Fig. 6g). Together, these data reveal that UA and S1P can promote axonal health in distinct genetic and neuronal contexts.
PQM-1 and CEH-60 control ASAH-1 expression
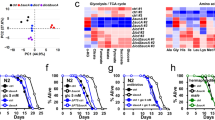
How does UA induce asah-1 intestinal expression to provide intergenerational neuroprotection? As UA induces asah-1 transcription (Fig. 2d,g), we reasoned that transcription factors related to intestinal stress or metabolic control may regulate asah-1. We examined the asah-1 promoter for conserved transcription factor-binding motifs and surveyed publicly available chromatin immunoprecipitation–sequencing (ChIP–seq) data for transcription factors exhibiting peaks upstream of the asah-1 locus (Fig. 7a and Extended Data Fig. 7a). We identified ChIP–seq peaks for two transcription factors in the asah-1 promoter: PQM-1 (a GATA zinc-finger transcription factor) and CEH-60 (an orthologue of mammalian PBX; a TALE class transcription factor; Fig. 7a and Extended Data Fig. 7a). These ChIP–seq peaks coincide with putative binding sites we identified in silico (Fig. 7a and Extended Data Fig. 7a). PQM-1 and CEH-60 function in the intestine to balance transcriptional networks governing stress responses and nutrient supply to progeny36,37. We therefore investigated their potential role in controlling PLM axon fragility and asah-1 expression.
a, PQM-1 and CEH-60 transcription factor ChIP–seq peaks at the asah-1 gene locus (top) aligned with a schematic of the asah-1 endogenous reporter (bottom). The asah-1 upstream region contains binding motifs for CEH-60 and PQM-1 (red line). The location of each binding motif (arrows) is indicated as the distance from the ATG (+1). b, Expression of asah-1::f2a::gfp::h2b in the intestine of an L4 larva (see Extended Data Fig. 7b for other developmental stages). Schematic (top) and overlay of Nomarski and fluorescence image (bottom). Filled circles indicate that expression was detected and open circles that weak/no expression was detected. Lateral view, anterior to the left. Pharynx is marked by a white asterisk; red line indicates bright asah-1::f2a::gfp::h2b expression. Scale bar, 25 μm. c, The expression of asah-1::f2a::gfp::h2b increases following exposure to UA (see Extended Data Fig. 8a for individual intestinal cell measurements). d, Loss of pqm-1 or ceh-60 prevents UA-induced induction of asah-1::f2a::gfp::h2b expression. e–g, Loss of pqm-1 (via RNAi (e) or gene deletion (f)) or ceh-60 (via RNAi (g)) suppresses UA-induced PLM neuroprotection in mec-17(ok2109); lon-2(e678) animals. h, Mutation of the putative PQM-1 binding motif in the asah-1 promoter induces asah-1::f2a::gfp::h2b expression. Int cell pair 1–6, second–seventh pair, respectively, of intestinal cells from the pharynx. i, Mutation of the putative PQM-1 binding motif in the asah-1 promoter reduces PLM axon breaks in mec-17(ok2109); lon-2(e678) animals. j, The progeny of P0 mothers exposed to S1P for 16 h (L4 to young adult) increases asah-1::f2a::gfp::h2b expression for two generations (F1 and F2). c–j, n = 28 and 30 (c); n = 18, 19, 21, 26, 22 and 24 (d); n = 74, 77, 99 and 103 (e); n = 73, 75, 122 and147 (f); n = 65, 71, 150 and 151 (g); n = 32, 30, 32, 30, 32, 30, 32, 30, 32, 30, 32 and 30 (h); n = 100 and 104 (i); and n = 27, 28, 26, 26, 23 and 25 (j) hermaphrodite animals per condition (left to right). P values were determined using an ANOVA (d–g,j) or unpaired Student’s t-test (c,h,i). ****P ≤ 0.0001; ***P ≤ 0.001; **P ≤ 0.01; *P ≤ 0.05; and NS, not significant. Error bars indicate the s.e.m.; a.u., arbitrary units. Source data are provided.
To directly assess whether PQM-1 and CEH-60 transcriptionally regulate endogenous asah-1, we generated a fluorescent reporter by inserting a f2A-gfp-h2B cassette immediately downstream of the asah-1 coding sequence (Fig. 7a,b). As ribosomal skipping occurs at the f2A sequence, independently translated GFP–H2B protein is visualized in nuclei (Fig. 7b and Extended Data Fig. 7b)38. We detected GFP–H2B expression in anterior intestinal nuclei (except the most anterior nuclei pair), with weak/undetectable expression in the posterior (Fig. 7b and Extended Data Fig. 7b). This anteriorly biased expression mirrors the asah-1 transcriptional reporter (compare Fig. 7b with Extended Data Fig. 4), suggesting transcriptional regulation of spatial expression. GFP–H2B expression by the asah-1 endogenous reporter was induced by exposure to UA (Fig. 7c and Extended Data Fig. 8a), providing support for our earlier analysis (Fig. 2g). We next crossed pqm-1 and ceh-60 loss-of-function mutants into asah-1::f2A-gfp-h2B animals and measured GFP–H2B fluorescence in the intestinal nuclei. Loss of PQM-1 elevated GFP–H2B expression and loss of either PQM-1 or CEH-60 prevented GFP–H2B induction by UA (Fig. 7d). As PQM-1 and CEH-60 are also required for yolk delivery to oocytes due to their control of intestinal vitellogenin expression36,39, we examined their role in UA-induced neuroprotection. We found that loss of PQM-1 or CEH-60 inhibited UA neuroprotection in mec-17(ok2109); lon-2(e678) animals (Fig. 7e–g). Therefore, PQM-1 and CEH-60 are required for induction of asah-1 by UA and for neuroprotection in response to UA exposure.
As PQM-1 and CEH-60 control metabolic homeostasis and yolk production36,37, asah-1 dysregulation in these transcription factor mutants may be a secondary consequence of disrupted lipid homeostasis. To directly examine the importance of PQM-1 and CEH-60 binding for asah-1 expression, we mutated the putative binding motifs that coincide with the PQM-1 and CEH-60 ChIP peaks in the asah-1 promoter (Fig. 7a and Extended Data Fig. 7a). We found that GFP–H2B intestinal expression was induced by mutations to the PQM-1 binding motif; mutations to the CEH-60 site had a weaker effect (Fig. 7h and Extended Data Fig. 8b). These data support a direct role for these transcription factors in asah-1 transcriptional repression and suggest that PQM-1 and CEH-60 occupancy at the asah-1 promoter may control sphingolipid homeostasis. We also noticed that PQM-1 motif mutations caused a posterior shift in GFP–H2B expression, revealing spatial regulation of intestinal expression (Fig. 7h). Given that PQM-1 motif mutations increased asah-1 expression, we investigated whether this is neuroprotective. When the asah-1::f2A-gfp-h2B (PQM-1 motif mutant) strain was crossed into mec-17(ok2109); lon-2(e678) animals, we observed robust reduction in PLM axon fragility (Fig. 7i). This revealed that asah-1 derepression has a functionally relevant outcome for axonal health.
S1P levels regulate ASAH-1 expression intergenerationally
Ceramide hydrolysis is the only catabolic source of sphingosine and therefore ceramidase (for example, ASAH-1) activity is a rate-limiting step in governing intracellular sphingosine and S1P levels40. We hypothesized that S1P intergenerational inheritance is triggered by an imbalance of sphingolipid homeostasis, such as increased S1P. Thus, detection and recycling of S1P to sphingosine and then ceramide may feedback to induce asah-1 expression in progeny. To explore this possibility, we exposed asah-1::f2A-gfp-h2B P0 animals to S1P for 16 h (L4 to young adult) and measured the intestinal expression of GFP–H2B in the subsequent generations. Remarkably, the levels of GFP–H2B were increased in the F1 and F2 progeny following P0 S1P exposure (Fig. 7j), establishing a regulatory mechanism for S1P-induced intergenerational inheritance.
Discussion
Here we show that short-term maternal dietary supplements can prevent axon fragility intergenerationally in C. elegans (Extended Data Fig. 8c). Supplementation with UA induces intestinal expression of asah-1, which encodes a rate-limiting enzyme in the sphingolipid salvage pathway. Our genetic evidence reveals that intestinal overexpression of either the de novo or salvage sphingolipid pathway prevents axon fragility and that SphK expression is required to promote axon health. The metabolite generated by SphK is S1P, and we found S1P supplementation also prevents axon fragility intergenerationally. Neuroprotection is dependent on S1P transport in intestinally derived yolk to oocytes18. Axon stability requires the correct transport of protein cargo along microtubules41. We found that UA and S1P improve axon transport and protect against axon fragility caused by a microtubule-destabilizing drug. Thus, S1P probably promotes microtubule stability, as has previously been shown for ceramide in a cell culture model42. We discovered that the transcription factors PQM-1 and CEH-60, known regulators of intestinal lipid homeostasis36,37, are required for asah-1 regulation and UA neuroprotection. Remarkably, mutation of a PQM-1-repressive element in the asah-1 promoter is sufficient to reduce axon fragility, corroborating the critical role for asah-1 in the regulation of neuronal health.
Sphingolipid locale impacts neuron development and health
We showed that neuronal development and health require correct spatial regulation of sphingolipid biosynthetic enzymes. Elevated intestinal asah-1 expression reduced PLM axon fragility in an SphK-dependant manner. In contrast, overexpression of asah-1 in the nervous system caused developmental axon outgrowth defects in the ALM and PLM neurons. However, the absence of overt motility defects in animals overexpressing asah-1 in neurons suggests that axon architecture is largely intact and disrupted sphingolipid homeostasis has context-specific effects. asah-1 is expressed at low levels in embryonic ALM and PLM neurons, as are other members of the sphingolipid salvage pathway such as sphk-1 (ref. 24). In contrast, expression of sptl-1, which encodes the rate-limiting enzyme of the sphingolipid de novo pathway, is undetected in these neurons. Perhaps asah-1 overexpression in embryonic ALM and PLM neurons causes depletion of ceramide and/or other ceramide derivates that are not replenished by the de novo pathway. Such disrupted ceramide homeostasis in neurons could affect multiple signalling pathways controlling axon development. Future genetic and biochemical studies are required to dissect the roles of ASAH-1 and the other C. elegans ceramidase enzymes (ASAH-2 and W02F12.2) in the nervous system and more broadly.
We showed that maternal S1P transport to oocytes reduces the fragility of axons with impaired cytoskeletal function. We propose that elevated S1P in the oocyte/embryo influences the axonal environment to limit axon fragility. The important roles of sphingolipid homeostasis in plasma-membrane fluidity, lipid-raft integrity and molecular trafficking may be central to the protective role S1P plays in this context43,44. Disrupted sphingolipid metabolism has been implicated in neurodegenerative disorders including Alzheimer’s disease, Parkinson’s disease, Huntington’s disease and multiple sclerosis45. The distinct autonomous and non-autonomous effects of sphingolipid homeostasis we revealed in C. elegans may also be relevant in these disease contexts.
Sphingolipid regulation of intergenerational inheritance
The neuroprotective effect of intestinal asah-1 is dependent on the expression of SphK-1, an enzyme that phosphorylates sphingosine to generate S1P. In support of this, maternally supplied S1P is transferred from the intestine to oocytes to protect axons. We showed that maternal S1P is probably transported within low-density yolk lipoproteins to promote neuroprotection. Essential roles for S1P and SphK in protecting vertebrate oocytes from radiation-induced apoptosis and the development of embryonic cardiovascular tissue have also been revealed46,47,48,49. In vertebrates, plasma lipoproteins also carry S1P, highlighting a conserved mode of carriage50.
How does S1P act intergenerationally? Intestinal asah-1 expression is elevated for two generations following exposure of reproductive stage hermaphrodites to S1P. This suggests that mechanisms in intestinal cells detect changes in sphingolipid levels, including S1P, that alter sphingolipid-pathway gene expression (for example, asah-1) in response. Examples of such sphingolipid autoregulatory networks have previously been identified in cellular and zebrafish models where sphingolipid metabolic enzymes are induced by accumulation of their sphingolipid substrate51,52. Increased asah-1 in the intestines of F1 animals could generate high S1P that would be transferred to the F2 generation to maintain the intergenerational effect. We suggest that a threshold of intestinal S1P must be maintained to initiate asah-1 upregulation and this threshold may be lost over time—hence, a reason for the intergenerational rather than transgenerational effect. At the level of asah-1 gene regulation, subtle changes in the subcellular localization or promoter occupancy of asah-1 regulators, including the transcription factors PQM-1 and CEH-60, could be responsible. PQM-1 nuclear localization is indeed responsive to certain stressors and perturbed insulin growth factor signalling37.
Periods of prenatal and postnatal interactions enable mammalian mothers to influence offspring53. Alterations in the reproductive behaviour of offspring permits the transmission of these effects to succeeding generations53. We show that the transmission of a lipid metabolite in the yolk can intergenerationally protect specific neurons in C. elegans. We suggest that our findings point to a broader underlying role for metabolism and metabolic gene expression in controlling intergenerational effects across species.
Methods
C. elegans strains and culture
C. elegans hermaphrodites were maintained according to standard protocols at 20 °C on nematode growth medium (NGM) plates with Escherichia coli (OP50) bacteria as a food source. The wild-type strain used was Bristol, N2. Mutant strains were backcrossed to N2 a minimum of three times and the animals were well-fed for at least two generations before performing experiments. A list of the strains and transgenes used in this study is provided in Supplementary Table 2.
Natural compound screen
All compounds were solubilized in DMSO and diluted to 50 μM by mixing with NGM agar. For the control plates, animals were grown with 1% DMSO. None of the compounds had an overt effect on OP50 growth. P0 animals were grown from L4 stage on compound plates and their F1 progeny were scored as 3-d-old adults. All plates were prepared fresh before each experiment and used within 2 d. To reduce metabolic breakdown of the compound by bacteria, NGM plates (containing compound or DMSO) were seeded with 150 µl heat-killed OP50 (65 °C for 30 min) and dried overnight at 20 °C. A 5-µl volume of live OP50 bacteria (1/10 dilution with Luria broth (LB)) was added to the dead bacteria to enable worm growth.
C. elegans expression constructs and transgenic strain generation
Reporter gene constructs were generated by PCR amplifying DNA elements and cloning these into Fire vectors. The constructs were injected into young adult hermaphrodites using standard methods.
Pasah-1::nls::gfp reporter construct
A 2,044 bp sequence upstream of the asah-1 start codon was amplified from C. elegans genomic DNA using forward (5′-GAAATGAAATAAGCTTAAAGAGAGAATAATAATCGAGTGAG-3′) and reverse (5′-GCAGGCATGCAAGCTTCTTTCTTGACTAGCTCTGAATAGTG-3′) oligonucleotides incorporating an HindIII restriction site. The HindIII-digested promoter fragment was ligated into HindIII-digested pPD95.67 vector, resulting in Pasah-1::NLS::gfp. Pasah-1::NLS::gfp was injected at 25 ng µl−1, with 5 ng µl−1 Pmyo-2::mCherry vector and 120 ng µl−1 bacterial DNA. The resultant Pasah-1::NLS::gfp extrachromosomal array line was integrated using ultraviolet light irradiation. The integrated Pasah-1::NLS::gfp (rp165) was backcrossed to N2 six times before analysis.
Pges-1::asah-1 overexpression construct
Complementary DNA to asah-1 (1,183 bp) was amplified from an N2 cDNA library using forward (5′-AGGACCCTTGGCTAGCATGCTCCGAGAATTGTCGG-3′) and reverse (5′-GATATCAATACCATGGCTACCATGGATAGCATTCTCCCGG-3′) oligonucleotides, and then cloned into a Pges-1::egl-8 vector (digested with NheI and NcoI to remove the egl-8 sequence) using an In-Fusion HD cloning kit (Takara Bio).
Pdpy-7::asah-1 overexpression construct
Complementary DNA to asah-1 (1,183 bp) was amplified from an N2 cDNA library using forward (5′-CGACTCTAGAGGATCCATGCTCCGAGAATTGTCGG-3′) and reverse (5′-CGCTCAGTTGGAATTCCTACCATGGATAGCATTCTCCCGG-3′) oligonucleotides, and then cloned into a Pdpy-7::nls-dsRed2 vector (digested with BamHI and EcoRI to remove the nls-dsRed2 sequence) using an In-Fusion HD cloning kit (Takara Bio).
Pmyo-3::asah-1 overexpression construct
Complementary DNA to asah-1 (1,183 bp) was amplified from an N2 cDNA library using forward (5′-AGGACCCTTGGCTAGCATGCTCCGAGAATTGTCGG-3′) and reverse (5′-GATATCAATACCATGGCTACCATGGATAGCATTCTCCCGG-3′) oligonucleotides, and then cloned into the pPD95.86 vector using an In-Fusion HD cloning kit (Takara Bio) after linearizing the vector with NheI and NcoI.
Prab-3::asah-1 overexpression construct
Complementary DNA to asah-1 (1,183 bp) was amplified from an N2 cDNA library using forward (5′-TGGCTAGCGTCGACGGTACCATGCTCCGAGAATTGTCGG-3′) and reverse (5′-GATATCAATACCATGGCTACCATGGATAGCATTCTCCCGG-3′) oligonucleotides, and then cloned into a Prab-3::nfya-1 vector (digested with KpnI and NcoI to remove the nfya-1 sequence) using an In-Fusion HD cloning kit (Takara Bio).
Pmec-4::asah-1 overexpression construct
Complementary DNA to asah-1 (1,183 bp) was amplified from an N2 cDNA library using forward (5′-AGGACCCTTGGCTAGATGCTCCGAGAATTGTCGG-3′) and reverse (5′-TACCGTCGACGCTAGCTACCATGGATAGCATTCTCCCGG-3′) oligonucleotides, and then cloned into the pSM::Pmec-4::unc-54 3′UTR vector using an In-Fusion HD cloning kit (Takara Bio) after linearizing the vector with NheI.
Pges-1::sptl-1 overexpression construct
Complementary DNA to sptl-1 (1,378 bp) was amplified from an N2 cDNA library using forward (5′-AGGACCCTTGGCTAGCATGGGATTTCTACCAGATTCGTGG-3′) and reverse (5′-GATATCAATACCATGTTAGAATTTATGAGCAACAACTCGG-3′) oligonucleotides, and then cloned into a Pges-1::egl-8 vector (digested with NheI and NcoI to remove the egl-8 sequence) using an In-Fusion HD cloning kit (Takara Bio).
Pges-1::sphk-1 overexpression construct
Complementary DNA to sphk-1 (1,423 bp) was amplified from an N2 cDNA library using forward (5′-AGGACCCTTGGCTAGCATGTTCATAGTAGTGGTAAC-3′) and reverse (5′-GATATCAATACCATGGCTAGGCAGTTGATGAGAAAA-3′) oligonucleotides, and then cloned into a Pges-1::egl-8 vector (digested with NheI and NcoI to remove the egl-8 sequence) using an In-Fusion HD cloning kit (Takara Bio).
UA treatment
A 10 mg ml−1 stock solution of UA (Sigma-Aldrich, 89797) dissolved in DMSO was prepared. UA (or DMSO for the control plates) was added to a 3.5-cm petri dish before the addition of NGM agar. All plates were freshy prepared before each experiment and used within 2 d. To reduce metabolic breakdown of UA by bacteria, 150 µl of heat-killed OP50 (65 °C for 30 min) was seeded on NGM plates (containing UA or DMSO) and dried overnight at 20 °C. A 5-µl volume of live OP50 (1/10 dilution with LB broth) was added to the dead bacteria to enable worm growth.
S1P treatment
A 2 mM stock solution of S1P (Sigma-Aldrich, 73914) dissolved in methanol was prepared. NGM agar was poured into a 3.5 cm petri dish, to which S1P (or methanol for the control plates) was added and mixed. All plates were freshly prepared before each experiment and used within 2 d. To reduce metabolic breakdown of S1P by bacteria, 150 µl of heat-killed OP50 (65 °C for 30 min) was seeded and spread on NGM plates (containing S1P or methanol) and dried overnight at 20 °C. A 5-µl volume of live OP50 (1/10 dilution with LB broth) was added to the dead bacteria to enable worm growth.
Ceramide treatment
Stock solutions of 0.5 mg ml−1 C20 (d18:1/20:0; Sigma-Aldrich, 860520P) and C22 (d18:1/22:0; Sigma-Aldrich, 860501P) dissolved in ethanol were prepared. Before seeding with bacteria, 60 µg ceramide was added to the surface of NGM plates. To reduce metabolic breakdown of ceramides by bacteria, 150 µl of heat-killed OP50 (65 °C for 30 min) was seeded and spread on NGM plates (containing ceramide or ethanol as the control) and dried overnight at 20 °C. A 5-µl volume of live OP50 (1/10 dilution with LB broth) was added to the dead bacteria to enable worm growth.
S1P–fluorescein application
A stock solution of 1 mM S1P–fluorescein (Echelon, S-200F) dissolved in methanol was prepared. To reduce metabolic breakdown of S1P–fluorescein by bacteria, 150 µl of heat-killed (65 °C for 30 min) OP50 was mixed with S1P–fluorescein or methanol (as the control), seeded on NGM plates and used immediately once it dried.
Microtubule drug treatment
A stock solution of 20 mM colchicine (Sigma-Aldrich, C9754) dissolved in DMSO was prepared. Colchicine (or DMSO for the control plates) was added to a 3.5 cm petri dish before the addition of NGM agar. All plates were freshly prepared before each experiment and used within 2 d. To reduce metabolic breakdown of colchicine by bacteria, 150 µl of heat-killed OP50 (65 °C for 30 min) was seeded on NGM plates (containing colchicine or DMSO) and dried overnight at 20 °C. A 5-µl volume of live OP50 (1/10 dilution with LB broth) was added to the dead bacteria to enable worm growth.
Intergenerational experiments
P0 animals were treated from mid-L4 to young adult (16 h) with UA or S1P (DMSO and methanol were used the respective controls) and the worms were then transferred to untreated plates for 3 h to lay semi-synchronized eggs. The mothers were removed after laying and the progeny (F1 generation) were cultured on untreated plates. When the F1 animals reached the 1-d-old adult stage, they were transferred to new untreated plates for 3 h to lay eggs (F2 generation) and this was repeated for the progeny of these animals (F3 generation). For each generation, PLM fragility defects were scored in 3-d-old adults.
CRISPR–Cas9 genome editing
Endogenous asah-1::f2a::gfp::h2b reporter
Cas9 protein, trans-activating CRISPR RNA and a crispr RNA (crRNA; TGGTAGATTGGATTCAACAT) from IDT were used to generate asah-1(rp176[asah-1::f2a::gfp::h2b]). A double-stranded DNA hybrid donor sequence of f2a::gfp::h2b was amplified by PCR, as described previously54. To facilitate the scoring of asah-1 expression, a nuclear-localized gfp was inserted downstream of the asah-1 coding region, separated by an F2A sequence that splits the endogenous ASAH-1 and GFP–H2B proteins.
Mutagenesis of the asah-1 promoter
For the mutagenesis of asah-1 promoter motifs, crRNAs and single-stranded oligonucleotide (ssODN) donors were used as follows.
CEH-60 motif: crRNA, 5′-AAAACCGGTGTGATTGATGA-3′; and ssODN, (5′-TTGTAAATTATGCTCCATAAACCAAAAACCGGTGTGATTGATGACATTTTCATGAGGGAAATAAATTAAA-3′). Mutation of the motif introduced a NotI restriction site (GCGGCCGC) to facilitate genotyping.
PQM-1 motif site: crRNA, 5′-AAGTGATAAGGAGTAAAGTG-3′; and ssODN, 5′-ACATTTTCATGAGGGAAATAAATTAAAAAGTGATAAGGAGTAAAGTGTGGCCACGTGTTTTCCGCAAAAA-3′. Mutation of the motif introduced a BamHI restriction site (GGATCC) to facilitate genotyping.
RNAi experiments
HT115(DE3) E. coli bacteria expressing RNAi plasmids for specific genes or empty vector (L4440) were cultured at 37 °C for 16 h in LB broth containing 50 µg ml−1 ampicillin. Saturated RNAi cultures were plated on RNAi plates and dried for 24 h. L4 hermaphrodites were plated on RNAi plates and incubated for one generation at 20 °C, unless otherwise stated. Due to the potency of rme-2 RNAi in preventing progeny survival, we diluted the rme-2 RNAi bacteria to 10% with 90% L4440 RNAi bacteria. Dilution of rme-2 RNAi bacteria to 10% was shown previously to reduce embryonic yolk levels55.
Microscopy
Worms were anaesthetized using 20 mM sodium azide for neuroanatomical scoring and levamisole (0.1 ng μl−1) for fluorescence measurements. The worms were mounted on 5% agarose pads on glass slides. Images were acquired using a Zeiss Axio Imager M2 and Zen software tools. Figures were prepared using ImageJ and Adobe Photoshop and Illustrator.
Phenotypic analysis
Phenotypic analysis of PLM neurons was performed on L1 larvae and 3-d-old adults where indicated. Axonal breaks were identified only when the distal fragment was visible. PVQ and D-type motor neuron analyses were performed on L1 and L4 larvae, respectively. All phenotypic scoring was performed at least in triplicate on independent days.
Axon transport analysis
UNC-104::GFP and mCherry::RAB-3 pooling at the distal end of PLM neurites was scored in 3-d-old adults. All phenotypic scoring was performed at least in triplicate on independent days.
Analysis of asah-1 expression
Expression of Pasah-1::gfp and asah-1::f2a::gfp::h2b was imaged and measured in six pairs of intestinal nuclei from the second anterior pair backwards (the expression in the first pair of anterior nuclei was weak and inconsistent). Texas red fluorescence in each nucleus was used as background.
Sequence alignment
Sequences of the asah-1 promoter in nematode species were extracted from WormBase (http://www.wormbase.org) and aligned using Clustal Omega56.
RNA sequencing
N2 gravid hermaphrodites were bleached with hypochlorite solution and eggs were allowed to hatch overnight in M9 buffer. The resultant synchronized L1 larvae were fed with OP50 and incubated for 34 h until they reached the L3 stage. These worms were washed from the plates with M9 buffer and exposed to UA (50 µM), DMSO (vehicle) or no treatment for 12 h. After treatment, the worms (now L4 larvae) were harvested and washed three times with M9 buffer to remove bacteria before being resuspended in TRIzol and flash-frozen in liquid nitrogen. Total RNA was extracted by phase separation with chloroform, followed by the RNeasy mini kit (Qiagen) protocol. RNase-free DNase was applied to the RNA samples to remove DNA contamination. Library preparation and sequencing on Illumina HiSeq instruments were performed by the Monash Micromon Sequencing Facility and data analysis was performed by the Monash Bioinformatics Platform. The transcriptome of five independent replicates for each group was analysed. The RNA-sequencing data were converted to a per-gene read-count matrix using RNAsik pipeline v1.5.0. Briefly, single-end reads were mapped to the C. elegans genome (WBcel235) using STAR57 in splice-aware mode, duplicates were marked using Picard Markduplicates and reads were assigned to gene models (exonic only) from the WBcel235 annotation using featureCounts v1.5.2 (ref. 58). Genes were only considered ‘testable’ if they had at least ten read counts in a sample and at least 1.0 counts-per-million in at least three samples. After applying this filter for testable genes, 14,037 protein-coding genes were retained. Genes that were differentially expressed between the treatment and control groups were identified using the EdgeR-quasi workflow59. The RNA-sequencing data are available at the Gene Expression Omnibus under the accession number GSE214425.
cDNA preparation and qPCR
Total RNA was isolated as described for RNA sequencing but with independent samples. An iScript kit (BioRad) was used for cDNA synthesis. PowerUp SYBR Green (Thermo Fisher) was used for the qPCR. The cDNA samples from three biological replicates were run in triplicate. The relative transcript levels were normalized to the housekeeping gene cdc-42. The following oligonucleotide pairs were used: asah-1 F, 5′-CTACTGTTCCTTGTGTCGGA-3′ and asah-1 R, 5′-GTTGGAGGCATTTGTTGCA-3′; and cdc-42 F, 5′-GACAATTACGCCGTCACAG-3′ and cdc-42 R, 5′-CGTAATCTTCCTGTCCAGCA-3′.
ChIP–seq data analysis
Published ChIP–seq datasets for CEH-60 (ref. 36) and PQM-1 (refs. 37,60) were used to identify binding peaks in the asah-1 promoter using IGV61.
Locomotion analysis
Worm locomotion was analysed at room temperature on day 1 and 3 adults using WormLab imaging (MBF Bioscience). NGM plates used for tracking were freshly seeded with 20 µl OP50 10 min before each experiment. Animals treated with UA or DMSO were randomly picked for tracking. Six animals were placed on each plate and their tracks were recorded for 6 min. To allow worms to adapt to the new conditions, the initial minute of tracking was not included for analysis. The average speed was calculated from the total distance moved (forwards and backwards) during the next 5 min.
Body-length analysis
The body length of worms was analysed at room temperature on day 1 and 3 adults using WormLab imaging (MBF Bioscience). NGM plates used for body-length measurements were freshly seeded with 20 µl OP50 10 min before each experiment. Animals treated with UA or DMSO were randomly picked for measurement. Six animals were placed on each plate for imaging and their body length was automatically measured by the WormLab software.
Statistics and reproducibility
Parameters such as the n value, mean ± s.e.m. and significant P values are reported in the figures, figure legends and Source Data. Significance was defined as P < 0.05.
Statistical analysis was performed in GraphPad Prism (v9.5) using an ANOVA, followed by Dunnett’s or Tukey’s multiple comparison tests for three or more conditions. An unpaired Student’s t-test was performed if the comparison was for two conditions. All experiments were performed with at least three independent biological replicates, with similar results (unless specified otherwise in the figure legends); the investigator was blinded to the genotype. No statistical method was used to pre-determine sample size. No data were excluded from analysis. The experiments were not randomized.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The RNA-sequencing data obtained in this work have been deposited at NCBI Gene Expression Omnibus under the accession number GSE214425. Previously published ChIP–seq datasets for the transcription factors CEH-60 and PQM-1 that were re-analysed here and used to identify binding peaks in the asah-1 promoter using IGV are available under the accession codes GSE112981 and GSE25811. All data are available in the main text or the extended data. Source data are provided with this paper.
References
Burton, N. O. et al. Insulin-like signalling to the maternal germline controls progeny response to osmotic stress. Nat. Cell Biol. 19, 252–257 (2017).
Ost, A. et al. Paternal diet defines offspring chromatin state and intergenerational obesity. Cell 159, 1352–1364 (2014).
Rechavi, O. et al. Starvation-induced transgenerational inheritance of small RNAs in C. elegans. Cell 158, 277–287 (2014).
Klosin, A., Casas, E., Hidalgo-Carcedo, C., Vavouri, T. & Lehner, B. Transgenerational transmission of environmental information in C. elegans. Science 356, 320–323 (2017).
Minkina, O. & Hunter, C. P. Intergenerational transmission of gene regulatory information in Caenorhabditis elegans. Trends Genet. 34, 54–64 (2018).
Perez, M. F. & Lehner, B. Intergenerational and transgenerational epigenetic inheritance in animals. Nat. Cell Biol. 21, 143–151 (2019).
Kaletsky, R. et al. C. elegans interprets bacterial non-coding RNAs to learn pathogenic avoidance. Nature 586, 445–451 (2020).
Perez, M. F. & Lehner, B. Vitellogenins—yolk gene function and regulation in Caenorhabditis elegans. Front. Physiol. 10, 1067 (2019).
Neumann, B. & Hilliard, M. A. Loss of MEC-17 leads to microtubule instability and axonal degeneration. Cell Rep. 6, 93–103 (2014).
Rapti, G. A perspective on C. elegans neurodevelopment: from early visionaries to a booming neuroscience research. J. Neurogenet. 34, 259–272 (2020).
Westermann, S. & Weber, K. Post-translational modifications regulate microtubule function. Nat. Rev. Mol. Cell Biol. 4, 938–947 (2003).
Dalla Costa, I. et al. The functional organization of axonal mRNA transport and translation. Nat. Rev. Neurosci. 22, 77–91 (2021).
Desai, A. & Mitchison, T. J. Microtubule polymerization dynamics. Annu. Rev. Cell Dev. Biol. 13, 83–117 (1997).
Sleigh, J. N., Rossor, A. M., Fellows, A. D., Tosolini, A. P. & Schiavo, G. Axonal transport and neurological disease. Nat. Rev. Neurol. 15, 691–703 (2019).
Chalfie, M. & Sulston, J. Developmental genetics of the mechanosensory neurons of Caenorhabditis elegans. Dev. Biol. 82, 358–370 (1981).
Kashyap, D., Tuli, H. S. & Sharma, A. K. Ursolic acid (UA): a metabolite with promising therapeutic potential. Life Sci. 146, 201–213 (2016).
Ramos-Hryb, A. B., Pazini, F. L., Kaster, M. P. & Rodrigues, A. L. S. Therapeutic potential of ursolic acid to manage neurodegenerative and psychiatric diseases. CNS Drugs 31, 1029–1041 (2017).
Grant, B. & Hirsh, D. Receptor-mediated endocytosis in the Caenorhabditis elegans oocyte. Mol. Biol. Cell 10, 4311–4326 (1999).
Marre, J., Traver, E. C. & Jose, A. M. Extracellular RNA is transported from one generation to the next in Caenorhabditis elegans. Proc. Natl Acad. Sci. USA 113, 12496–12501 (2016).
Buckley, B. A. et al. A nuclear Argonaute promotes multigenerational epigenetic inheritance and germline immortality. Nature 489, 447–451 (2012).
Wan, G. et al. Spatiotemporal regulation of liquid-like condensates in epigenetic inheritance. Nature 557, 679–683 (2018).
Ouyang, J. P. T., Zhang, W. L. & Seydoux, G. The conserved helicase ZNFX-1 memorializes silenced RNAs in perinuclear condensates. Nat. Cell Biol. 24, 1129–1140 (2022).
Gebai, A., Gorelik, A., Li, Z., Illes, K. & Nagar, B. Structural basis for the activation of acid ceramidase. Nat. Commun. 9, 1621 (2018).
Packer, J. S. et al. A lineage-resolved molecular atlas of C. elegans embryogenesis at single-cell resolution. Science 365, eaax1971 (2019).
Cao, J. et al. Comprehensive single-cell transcriptional profiling of a multicellular organism. Science 357, 661–667 (2017).
Okkema, P. G., Harrison, S. W., Plunger, V., Aryana, A. & Fire, A. Sequence requirements for myosin gene expression and regulation in Caenorhabditis elegans. Genetics 135, 385–404 (1993).
Gilleard, J. S., Barry, J. D. & Johnstone, I. L. Cis regulatory requirements for hypodermal cell-specific expression of the Caenorhabditis elegans cuticle collagen gene dpy-7. Mol. Cell. Biol. 17, 2301–2311 (1997).
Mitani, S., Du, H., Hall, D. H., Driscoll, M. & Chalfie, M. Combinatorial control of touch receptor neuron expression in Caenorhabditis elegans. Development 119, 773–783 (1993).
Aamodt, E. J., Chung, M. A. & McGhee, J. D. Spatial control of gut-specific gene expression during Caenorhabditis elegans development. Science 252, 579–582 (1991).
Maceyka, M. & Spiegel, S. Sphingolipid metabolites in inflammatory disease. Nature 510, 58–67 (2014).
Hannun, Y. A. & Obeid, L. M. Principles of bioactive lipid signalling: lessons from sphingolipids. Nat. Rev. Mol. Cell Biol. 9, 139–150 (2008).
Hanada, K. Serine palmitoyltransferase, a key enzyme of sphingolipid metabolism. Biochim. Biophys. Acta 1632, 16–30 (2003).
The Role of Microtubules in Cell Biology, Neurobiology, and Oncology (ed Fojo, A. T.) (Humana Press, 2008).
Ritchie, F. K. et al. The heterochronic gene lin-14 controls axonal degeneration in C. elegans neurons. Cell Rep. 20, 2955–2965 (2017).
Norgaard, S., Deng, S., Cao, W. & Pocock, R. Distinct CED-10/Rac1 domains confer context-specific functions in development. PLoS Genet. 14, e1007670 (2018).
Dowen, R. H. CEH-60/PBX and UNC-62/MEIS coordinate a metabolic switch that supports reproduction in C. elegans. Dev. Cell 49, 235–250 (2019).
Tepper, R. G. et al. PQM-1 complements DAF-16 as a key transcriptional regulator of DAF-2-mediated development and longevity. Cell 154, 676–690 (2013).
Ahier, A. & Jarriault, S. Simultaneous expression of multiple proteins under a single promoter in Caenorhabditis elegans via a versatile 2A-based toolkit. Genetics 196, 605–613 (2014).
Dowen, R. H., Breen, P. C., Tullius, T., Conery, A. L. & Ruvkun, G. A microRNA program in the C. elegans hypodermis couples to intestinal mTORC2/PQM-1 signaling to modulate fat transport. Genes Dev. 30, 1515–1528 (2016).
Okino, N. et al. The reverse activity of human acid ceramidase. J. Biol. Chem. 278, 29948–29953 (2003).
Coleman, M. Axon degeneration mechanisms: commonality amid diversity. Nat. Rev. Neurosci. 6, 889–898 (2005).
Tripathi, P. et al. Palmitoylation of acetylated tubulin and association with ceramide-rich platforms is critical for ciliogenesis. J. Lipid Res. 62, 100021 (2021).
Head, B. P., Patel, H. H. & Insel, P. A. Interaction of membrane/lipid rafts with the cytoskeleton: impact on signaling and function: membrane/lipid rafts, mediators of cytoskeletal arrangement and cell signaling. Biochim. Biophys. Acta 1838, 532–545 (2014).
Mencarelli, C. & Martinez-Martinez, P. Ceramide function in the brain: when a slight tilt is enough. Cell. Mol. Life Sci. 70, 181–203 (2013).
Di Pardo, A. & Maglione, V. Sphingolipid metabolism: a new therapeutic opportunity for brain degenerative disorders. Front. Neurosci. 12, 249 (2018).
Mendelson, K., Zygmunt, T., Torres-Vazquez, J., Evans, T. & Hla, T. Sphingosine 1-phosphate receptor signaling regulates proper embryonic vascular patterning. J. Biol. Chem. 288, 2143–2156 (2013).
Mizugishi, K. et al. Essential role for sphingosine kinases in neural and vascular development. Mol. Cell. Biol. 25, 11113–11121 (2005).
Kono, M. et al. The sphingosine-1-phosphate receptors S1P1, S1P2, and S1P3 function coordinately during embryonic angiogenesis. J. Biol. Chem. 279, 29367–29373 (2004).
Morita, Y. et al. Oocyte apoptosis is suppressed by disruption of the acid sphingomyelinase gene or by sphingosine-1-phosphate therapy. Nat. Med. 6, 1109–1114 (2000).
Cartier, A. & Hla, T. Sphingosine 1-phosphate: lipid signaling in pathology and therapy. Science 366, eaar5551 (2019).
Xiong, Y. et al. Sphingosine kinases are not required for inflammatory responses in macrophages. J. Biol. Chem. 288, 32563–32573 (2013).
Mendelson, K. et al. The ceramide synthase 2b gene mediates genomic sensing and regulation of sphingosine levels during zebrafish embryogenesis. eLife 6, e21992 (2017).
Champagne, F. A. Epigenetic mechanisms and the transgenerational effects of maternal care. Front. Neuroendocrinol. 29, 386–397 (2008).
Dokshin, G. A., Ghanta, K. S., Piscopo, K. M. & Mello, C. C. Robust genome editing with short single-stranded and long, partially single-stranded DNA donors in Caenorhabditis elegans. Genetics 210, 781–787 (2018).
Perez, M. F., Francesconi, M., Hidalgo-Carcedo, C. & Lehner, B. Maternal age generates phenotypic variation in Caenorhabditis elegans. Nature 552, 106–109 (2017).
Sievers, F. et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 7, 539 (2011).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Chen, Y., Lun, A. T. & Smyth, G. K. From reads to genes to pathways: differential expression analysis of RNA-Seq experiments using Rsubread and the edgeR quasi-likelihood pipeline. F1000Research 5, 1438 (2016).
Niu, W. et al. Diverse transcription factor binding features revealed by genome-wide ChIP–seq in C. elegans. Genome Res. 21, 245–254 (2011).
Robinson, J. T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
Acknowledgements
We thank B. Neumann and the members of the Pocock laboratory for their comments on the manuscript. We thank the Monash Micromon Sequencing Facility and Monash Bioinformatics Platform for the RNA sequencing and data analysis. Some strains used in this study were provided by the Caenorhabditis Genetics Center, which is funded by NIH Office of Research and Infrastructure Programs (P40 OD10440) and the National BioResource Project (NBRP) Japan. This work was supported by the following funding: National Health and Medical Research Council grant nos. GNT1105374, GNT1137645 and GNT2000766 (R.P.); and Apex Biotech Research Pty Ltd donation 281589068 (Z.X.).
Author information
Authors and Affiliations
Contributions
Conceptualization: W.W., Z.X. and R.P. Methodology: W.W., J.L. and R.P. Investigation: W.W., T.S., X.C., Q.F., R.C. and R.P. Visualization: W.W., X.C., R.C. and R.P. Funding acquisition: Z.X. and R.P. Project administration: W.W. and R.P. Supervision: J.L., Z.X. and R.P. Writing of the original draft: R.P. Writing, review and editing: W.W., T.S., X.C., Q.F., R.C., J.L., Z.X. and R.P.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Cell Biology thanks Timothy Hla, Nick Burton and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Natural-product screen for regulators of axon fragility.
a, Natural-product exposure timeline. Upper schematic shows the stages of C. elegans development relevant to continuous natural-product exposure in the P0 and F1 generations. b, Quantification of PLM axon breaks in mec-17(ok2109); lon-2(e678) animals following continuous exposure (P0 to L4 larva to F1 to adult) of a natural-product library at 50 μM. Orange bars. reduced PLM breaks; red bars, increased PLM breaks; grey bars, not significant. Quantification of PLM axon breaks shown in b, n = 122, 102, 100, 85, 101, 100, 98, 94, 97, 100, 97, 94, 99, 95, 98, 98, 100, 102, 100, 91, 99 and 100 hermaphrodite animals per condition (left to right). P values were determined using an unpaired t-test. ***P ≤ 0.001; *P ≤ 0.05. Error bars indicate the s.e.m. Source data are provided.
Extended Data Fig. 2 Effect of UA on axon fragility and worm growth/behaviour.
a, Continuous exposure (P0 to L4 larva to F1 to adult) of UA at 50 or 100 μM, but not 25 μM, rescues PLM axon breaks in mec-17(ok2109); lon-2(e678) animals. b,c, Quantification of locomotion speed of zdIs5 and zdIs5; mec-17(ok2109); lon-2(e678) F1 animals on day 1 (b) and 3 (c) of adulthood treated with DMSO (control) or UA (P0 mothers treated for 16 h, L4 to young adult). Data presented as speed in μm s−1 (points). d,e, Quantification of body length of zdIs5 and zdIs5; mec-17(ok2109); lon-2(e678) F1 animals at day 1 (d) and 3 (e) of adulthood treated with DMSO (control) or UA (P0 mothers treated for 16 h, L4 to young adult). Data presented as individual lengths in μm (points). f, Exposure of UA (50 μM) from P0 embryo to F1 3-d-old adults (F1 to A3, continuous) reduces PLM axon breaks in mec-17(ok2109); lon-2(e678) F1 animals. In contrast, UA exposure (50 μM) from P0 embryo to P0 3-d-old adults (P0 to A3, continuous), or only during embryogenesis (P0 to A3 and F1 to A3, time-specific) does not reduce PLM axon breaks in mec-17(ok2109); lon-2(e678) F1 animals. Quantification of PLM axon breaks shown in a, n = 101, 102, 101, 103, 101 and 100; f, n = 99, 98, 96, 101, 99, 99, 96 and 97 hermaphrodite animals per condition (left to right). Quantification of locomotion speed shown in b, n = 65 and 40; c, n = 49 and 38 hermaphrodite animals per condition (left to right). Quantification of body length shown in d, n = 66 and 46; e, n = 50 and 50 hermaphrodite animals per condition (left to right). P values were calculated using a one-way analysis of variance (ANOVA; b–e) or unpaired t-test (a,f). ***P ≤ 0.001; *P ≤ 0.05; and NS, not significant. Error bars indicate the s.e.m. Source data are provided.
Extended Data Fig. 3 UA RNA-sequencing analysis.
a, Schematic of transcriptome analysis following UA exposure. Total RNA isolated from L4 larvae exposed to DMSO or UA for 12 h; 49 genes were regulated by UA exposure (false discovery rate, 0.02). b, Multidimensional scale plot (MDS) of untreated, DMSO-treated (control) and UA-treated RNA-sequencing samples. Source numerical data are provided in Supplementary Table 1.
Extended Data Fig. 4 asah-1 transcriptional reporter expression analysis.
The Pasah-1::NLS::gfp transcriptional reporter (rpIs165) is expressed in the intestine from the threefold stage of embryogenesis throughout larval development and into adulthood. A schematic of a young adult is shown below the fluorescence micrograph. Note that Pasah-1::NLS::gfp expression is brighter in the anterior intestine (red line). Anterior to the left in larval image. Pharynx marked by a white asterisk. Scale bars, 20 μm.
Extended Data Fig. 5 asah-1 overexpression analysis.
a, Schematic and fluorescence micrographs of ALM and PLM anatomy in L1 larval wild-type (top micrograph) and Pmec-4::asah-1-overexpressing (bottom micrographs) animals expressing the Pmec-4::gfp transgene (zdIs5). Outgrowth defects are marked with red lines and arrows. Scale bar, 20 μm. b,c, Quantification of ALM (b) and PLM (c) outgrowth defects in wild-type, mec-17(ok2109); lon-2(e678); Pmec-4::asah-1, Pmec-4::asah-1 and Prab-3::asah-1 animals. The ALM and PLM outgrowth defects correspond to the micrographs in a. d,e, Schematics and fluorescence micrographs (d) as well as quantification (e) of PVM axon guidance in L4 wild-type and Prab-3::asah-1-overexpressing animals expressing the Pmec-4::gfp transgene (zdIs5). Scale bar, 25 μm. f,g, Schematics (f) and quantification (g) of PVQ axon guidance in L1 wild-type and Prab-3::asah-1-overexpressing animals expressing the Psra-6::gfp transgene (oyIs14). Quantification of ALM outgrowth defects shown in b, n = 60, 42, 39, 46, 48 and 54 hermaphrodite animals per condition (left to right). Quantification of PLM outgrowth defects shown in c, n = 60, 42, 39, 46, 48 and 54 hermaphrodite animals per condition (left to right). Quantification of PVM guidance defects shown in e, n = 45 and 75 hermaphrodite animals per condition (left to right). Quantification of PVQ axon defects shown in g, n = 65 and 73 hermaphrodite animals per condition (left to right). P values were determined using a two-way analysis of variance (ANOVA; b,c) or unpaired t-test (e,g). ****P ≤ 0.0001; *P ≤ 0.05; and NS, not significant. Error bars indicate the s.e.m. Source data are provided.
Extended Data Fig. 6 Analysis of sphk-1 function and S1P–fluorescein detection.
a,b, Loss of sphk-1 via RNAi (a) and gene deletion (sphk-1-(ok1097) deletion mutant; b) enhances PLM axon breaks in mec-17(ok2109); lon-2(e678) animals and prevents the protective effect of UA. c, Brightfield and fluorescence image overlay of L4 hermaphrodites exposed to control (methanol) or S1P–fluorescein for 1 h. S1P–fluorescein was detected in the intestinal lumen. Scale bar, 20 μm. Quantification of PLM axon breaks shown in a, n = 99, 97, 128 and 138; b, n = 100, 105, 100 and 100 hermaphrodite animals per condition (left to right). P values were determined using a one-way analysis of variance (ANOVA). **P ≤ 0.01; *P ≤ 0.05; and NS, not significant. Error bars indicate the s.e.m. Source data are provided.
Extended Data Fig. 7 Analysis of the asah-1 endogenous reporter.
a, Nucleotide sequence 200 bp upstream of the asah-1 start codon. Upper sequence shows the wild-type sequence highlighting putative transcription factor-binding motifs for CEH-60 (red) and PQM-1 (blue, motif). The bottom sequence shows the independent mutations introduced to delete each putative transcription factor-binding motif (yellow highlights) that introduced the following restriction enzyme recognition sequences: CEH-60 motif (NotI) and PQM-1 motif (BamHI). b, GFP–H2B expression from the asah-1::f2a::gfp::h2b endogenous reporter (rpIs176) was detected in the intestine from late embryogenesis (threefold stage) throughout larval development and into adulthood. A schematic of each animal is shown next to the fluorescence micrographs. Note that Pasah-1::NLS::gfp expression is consistently brighter in the anterior intestine (red line). Anterior to the left in the worm image. Pharynx marked by a white asterisk. Scale bar, 20 μm.
Extended Data Fig. 8 Regulation of endogenous asah-1 expression.
a, GFP–H2B expression from the asah-1::f2a::gfp::h2b endogenous reporter increases when animals are exposed to UA. Quantification of nuclear fluorescence from each pair of intestinal cells per worm is shown (Int cell pair 1 = second pair of intestinal cells from the pharynx). b, Mutation of putative CEH-60 binding motif in the asah-1 promoter induces GFP–H2B expression from the asah-1 endogenous reporter. Quantification of nuclear fluorescence from each pair of intestinal cells per worm is shown (Int cell pair 1, second pair of intestinal cells from the pharynx). c, Proposed model of UA/S1P intergenerational neuroprotection. UA exposure of P0 animals (L4 to young adult) elevates intestinal asah-1 expression (probably through inhibition of PQM-1/CEH-60-directed transcriptional repression) and increases S1P levels. S1P is transferred within intestinally derived yolk to oocytes via the RME-2 receptor and promotes axon health in these animals (top). In F1 progeny, elevated S1P is detected in the intestine, which causes asah-1 expression to increase and generate additional S1P that is transmitted, via the yolk, to F2 oocytes and promotes axon health in these animals (bottom). UA/S1P neuroprotection is not observed in F3 progeny, probably due to reduced S1P levels that are below the threshold required for neuroprotection. Quantification of asah-1::f2a::gfp::h2b fluorescence shown in a, n = 28, 29, 28, 29, 28, 29, 28, 29, 28, 29, 28 and 29; b, n = 29 animals per condition (left to right). P values were determined using and unpaired t-test. ****P ≤ 0.0001; ***P ≤ 0.001; **P ≤ 0.01; *P ≤ 0.05. Error bars indicate the s.e.m. Source data are provided.
Supplementary information
Supplementary Tables 1 and 2
Supplementary Table 1. RNA-sequencing data. Table 2. C. elegans strains used in this study.
Source data
Source Data Figs. 1–7 and Extended Data Figs. 1, 2, 5, 6 and 8.
Statistical source data for Figs. 1–7 and Extended Data Figs. 1,2,5,6,8.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wang, W., Sherry, T., Cheng, X. et al. An intestinal sphingolipid confers intergenerational neuroprotection. Nat Cell Biol 25, 1196–1207 (2023). https://doi.org/10.1038/s41556-023-01195-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-023-01195-9
This article is cited by
-
A mother to offspring metabolic link
Nature Cell Biology (2023)