Abstract
Nuclear localization of HIPPO-YAP fusion proteins has been implicated in supratentorial ependymoma development. Here, unexpectedly, we find that liquid–liquid phase separation, rather than nuclear localization, of recurrent patient-derived YAP fusions, YAP-MAMLD1 and C11ORF95-YAP, underlies ependymoma tumourigenesis from neural progenitor cells. Mutagenesis and chimaera assays demonstrate that an intrinsically disordered region promotes oligomerization of the YAP fusions into nuclear, puncta-like, membrane-less condensates. Oligomerization and nuclear condensates induced by YAP fusion with a coiled-coil domain of transcriptional activator GCN4 also promote ependymoma formation. YAP-MAMLD1 concentrates transcription factors and co-activators, including BRD4, MED1 and TEAD, in condensates while excluding transcriptional repressive PRC2, and induces long-range enhancer–promoter interactions that promote transcription and oncogenic programmes. Blocking condensate-mediated transcriptional co-activator activity inhibits tumourigenesis, indicating a critical role of liquid phase separation for YAP fusion oncogenic activity in ependymoma. YAP fusions containing the intrinsically disordered region features are common in human tumours, suggesting that nuclear condensates could be targeted to treat YAP-fusion-induced cancers.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
Sequencing data that support the findings of this study have been deposited in the NCBI Gene Expression Omnibus (GEO) under accession codes GSE207293 including RNA-seq and ChIP–seq data with the accession number GSE181867, and methyl-seq with accession number GSE206926 and HiChIP with accession number GSE207801. Molecular Signatures Database v5.1 was obtained from the following website: https://www.gsea-msigdb.org/gsea/msigdb. Reference genome mm10 was obtained from the following website: https://hgdownload.soe.ucsc.edu/downloads.html. Source data are provided with this paper. All other data supporting the findings of this study are available from the corresponding author on reasonable request
Code availability
No custom code was used in this study. Open-source algorithms were used as detailed in analysis methods including FastQC v0.11.6 (https://www.bioinformatics.babraham.ac.uk/projects/fastqc/), STAR v2.7.8a (https://github.com/STAR-Fusion/STAR-Fusion/wiki/STAR-Fusion-release-and-CTAT-Genome-Lib-Compatibility-Matrix), Bowtie2 v2.4.0 (https://bowtie-bio.sourceforge.net/bowtie2/index.shtml), DEseq2 1.36.0 (https://bioconductor.org/packages/release/bioc/html/DESeq2.html), MACS v1.4.2 (http://liulab.dfci.harvard.edu/MACS), HOMER (http://homer.salk.edu/homer/index.html), GSEA 4.2.3 (http://www.broadinstitute.org/gsea/index.jsp), Cytoscape 3.9.1 (https://cytoscape.org/), Toppcluster (https://toppcluster.cchmc.org/), bwameth v0.20 (https://github.com/brentp/bwa-meth), MarkDuplicates v1.122.0 (http://broadinstitute.github.io/picard), MethylDackel v0.5.2 (https://github.com/dpryan79/MethylDackel.git), Metilene v0.2-8 (http://www.bioinf.uni-leipzig.de/Software/metilene) and MAPS pipeline for HiChIP–seq (https://github.com/ijuric/MAPS).
References
Wang, Q. et al. Supratentorial pediatric cortical ependymomas: a comprehensive retrospective study. Neurosurg. Rev. 44, 1543–1551 (2021).
Hubner, J. M., Kool, M., Pfister, S. M. & Pajtler, K. W. Epidemiology, molecular classification and WHO grading of ependymoma. J. Neurosurg. Sci. 62, 46–50 (2018).
Zacharoulis, S. et al. Treatment and outcome of children with relapsed ependymoma: a multi-institutional retrospective analysis. Childs Nerv. Syst. 26, 905–911 (2010).
Taylor, M. D. et al. Radial glia cells are candidate stem cells of ependymoma. Cancer Cell 8, 323–335 (2005).
Pajtler, K. W. et al. Molecular classification of ependymal tumors across all CNS compartments, histopathological grades, and age groups. Cancer Cell 27, 728–743 (2015).
Johnson, R. A. et al. Cross-species genomics matches driver mutations and cell compartments to model ependymoma. Nature 466, 632–636 (2010).
Gojo, J. et al. Single-cell RNA-seq reveals cellular hierarchies and impaired developmental trajectories in pediatric ependymoma. Cancer Cell 38, 44–59.e49 (2020).
Zanconato, F., Battilana, G., Cordenonsi, M. & Piccolo, S. YAP/TAZ as therapeutic targets in cancer. Curr. Opin. Pharmacol. 29, 26–33 (2016).
Harvey, K. F., Zhang, X. & Thomas, D. M. The Hippo pathway and human cancer. Nat. Rev. Cancer 13, 246–257 (2013).
Hong, W. & Guan, K. L. The YAP and TAZ transcription co-activators: key downstream effectors of the mammalian Hippo pathway. Semin. Cell Dev. Biol. 23, 785–793 (2012).
Varelas, X. The Hippo pathway effectors TAZ and YAP in development, homeostasis and disease. Development 141, 1614–1626 (2014).
Pajtler, K. W. et al. YAP1 subgroup supratentorial ependymoma requires TEAD and nuclear factor I-mediated transcriptional programmes for tumorigenesis. Nat. Commun. 10, 3914 (2019).
Parker, M. et al. C11orf95-RELA fusions drive oncogenic NF-κB signalling in ependymoma. Nature 506, 451–455 (2014).
Szulzewsky, F. et al. Comparison of tumor-associated YAP1 fusions identifies a recurrent set of functions critical for oncogenesis. Genes Dev. 34, 1051–1064 (2020).
Meng, Z., Moroishi, T. & Guan, K. L. Mechanisms of Hippo pathway regulation. Genes Dev. 30, 1–17 (2016).
Wu, L. M. N. et al. Programming of Schwann cells by Lats1/2-TAZ/YAP signaling drives malignant peripheral nerve sheath tumorigenesis. Cancer Cell 33, 292–308 (2018).
Zhao, S. et al. PiggyBac transposon vectors: the tools of the human gene encoding. Transl. Lung Cancer Res. 5, 120–125 (2016).
Dong, J. et al. Elucidation of a universal size-control mechanism in Drosophila and mammals. Cell 130, 1120–1133 (2007).
Zhao, B. et al. Inactivation of YAP oncoprotein by the Hippo pathway is involved in cell contact inhibition and tissue growth control. Genes Dev. 21, 2747–2761 (2007).
Chen, Q. Z. et al. Homeostatic control of Hippo signaling activity revealed by an endogenous activating mutation in YAP. Genes Dev. 29, 1285–1297 (2015).
Andreiuolo, F. et al. Childhood supratentorial ependymomas with YAP1-MAMLD1 fusion: an entity with characteristic clinical, radiological, cytogenetic and histopathological features. Brain Pathol. 29, 205–216 (2019).
Takadera, M. et al. Phenotypic characterization with somatic genome editing and gene transfer reveals the diverse oncogenicity of ependymoma fusion genes. Acta Neuropathol. Commun. 8, 203 (2020).
Eder, N. et al. YAP1/TAZ drives ependymoma-like tumour formation in mice. Nat. Commun. 11, 2380 (2020).
Ligon, K. L. et al. The oligodendroglial lineage marker OLIG2 is universally expressed in diffuse gliomas. J. Neuropathol. Exp. Neurol. 63, 499–509 (2004).
Lu, F. et al. Olig2-dependent reciprocal shift in PDGF and EGF receptor signaling regulates tumor phenotype and mitotic growth in malignant glioma. Cancer Cell 29, 669–683 (2016).
Venugopal, C., McFarlane, N. M., Nolte, S., Manoranjan, B. & Singh, S. K. Processing of primary brain tumor tissue for stem cell assays and flow sorting. J. Vis. Exp. 67, 4111 (2012).
Jiang, S., Fagman, J. B., Chen, C., Alberti, S. & Liu, B. Protein phase separation and its role in tumorigenesis. eLife 9, e60264 (2020).
Alberti, S., Gladfelter, A. & Mittag, T. Considerations and challenges in studying liquid–liquid phase separation and biomolecular condensates. Cell 176, 419–434 (2019).
Bolognesi, A., Sliwa-Gonzalez, A., Prasad, R. & Barral, Y. Fluorescence recovery after photo-bleaching (FRAP) and fluorescence loss in photo-bleaching (FLIP) experiments to study protein dynamics during budding yeast cell division. Methods Mol. Biol. 1369, 25–44 (2016).
Chong, S. et al. Imaging dynamic and selective low-complexity domain interactions that control gene transcription. Science 361, eaar2555 (2018).
Sabari, B. R. et al. Coactivator condensation at super-enhancers links phase separation and gene control. Science 361, eaar3958 (2018).
Zhao, B. et al. TEAD mediates YAP-dependent gene induction and growth control. Genes Dev. 22, 1962–1971 (2008).
Liu-Chittenden, Y. et al. Genetic and pharmacological disruption of the TEAD-YAP complex suppresses the oncogenic activity of YAP. Genes Dev. 26, 1300–1305 (2012).
Na Ayutthaya, P. P., Lundberg, D., Weigel, D. & Li, L. Blue native polyacrylamide gel electrophoresis (BN-PAGE) for the analysis of protein oligomers in plants. Curr. Protoc. Plant Biol. 5, e20107 (2020).
Ciani, B. et al. Molecular basis of coiled-coil oligomerization-state specificity. Proc. Natl Acad. Sci. USA 107, 19850–19855 (2010).
Banani, S. F., Lee, H. O., Hyman, A. A. & Rosen, M. K. Biomolecular condensates: organizers of cellular biochemistry. Nat. Rev. Mol. Cell Biol. 18, 285–298 (2017).
Forsova, O. S. & Zakharov, V. V. High-order oligomers of intrinsically disordered brain proteins BASP1 and GAP-43 preserve the structural disorder. FEBS J. 283, 1550–1569 (2016).
Cao, R. et al. Role of histone H3 lysine 27 methylation in Polycomb-group silencing. Science 298, 1039–1043 (2002).
Laugesen, A., Hojfeldt, J. W. & Helin, K. Molecular mechanisms directing PRC2 recruitment and H3K27 methylation. Mol. Cell 74, 8–18 (2019).
Yu, J. R., Lee, C. H., Oksuz, O., Stafford, J. M. & Reinberg, D. PRC2 is high maintenance. Genes Dev. 33, 903–935 (2019).
Jain, S. U. et al. PFA ependymoma-associated protein EZHIP inhibits PRC2 activity through a H3 K27M-like mechanism. Nat. Commun. 10, 2146 (2019).
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: the next generation. Cell 144, 646–674 (2011).
Creyghton, M. P. et al. Histone H3K27ac separates active from poised enhancers and predicts developmental state. Proc. Natl Acad. Sci. USA 107, 21931–21936 (2010).
Hnisz, D. et al. Convergence of developmental and oncogenic signaling pathways at transcriptional super-enhancers. Mol. Cell 58, 362–370 (2015).
Tang, F., Yang, Z., Tan, Y. & Li, Y. Super-enhancer function and its application in cancer targeted therapy. NPJ Precis. Oncol. 4, 2 (2020).
Wang, X., Cairns, M. J. & Yan, J. Super-enhancers in transcriptional regulation and genome organization. Nucleic Acids Res. 47, 11481–11496 (2019).
Mumbach, M. R. et al. HiChIP: efficient and sensitive analysis of protein-directed genome architecture. Nat. Methods 13, 919–922 (2016).
McKeown, M. R. & Bradner, J. E. Therapeutic strategies to inhibit MYC. Cold Spring Harb. Perspect. Med. 4, a014266 (2014).
Pearson, A. D. et al. Bromodomain and extra-terminal inhibitors—a consensus prioritisation after the Paediatric Strategy Forum for medicinal product development of epigenetic modifiers in children—ACCELERATE. Eur. J. Cancer 146, 115–124 (2021).
Lu, Y. et al. Phase separation of TAZ compartmentalizes the transcription machinery to promote gene expression. Nat. Cell Biol. 22, 453–464 (2020).
Cai, D. et al. Phase separation of YAP reorganizes genome topology for long-term YAP target gene expression. Nat. Cell Biol. 21, 1578–1589 (2019).
Hnisz, D., Shrinivas, K., Young, R. A., Chakraborty, A. K. & Sharp, P. A. A phase separation model for transcriptional control. Cell 169, 13–23 (2017).
Li, H. et al. SUZ12 promotes human epithelial ovarian cancer by suppressing apoptosis via silencing HRK. Mol. Cancer Res. 10, 1462–1472 (2012).
Varambally, S. et al. The polycomb group protein EZH2 is involved in progression of prostate cancer. Nature 419, 624–629 (2002).
Bender, S. et al. Reduced H3K27me3 and DNA hypomethylation are major drivers of gene expression in K27M mutant pediatric high-grade gliomas. Cancer Cell 24, 660–672 (2013).
Lewis, P. W. et al. Inhibition of PRC2 activity by a gain-of-function H3 mutation found in pediatric glioblastoma. Science 340, 857–861 (2013).
Bayliss, J. et al. Lowered H3K27me3 and DNA hypomethylation define poorly prognostic pediatric posterior fossa ependymomas. Sci. Transl. Med. 8, 366ra161 (2016).
Li, Y. et al. Genome-wide analyses reveal a role of Polycomb in promoting hypomethylation of DNA methylation valleys. Genome Biol. 19, 18 (2018).
Takeshima, H., Yamashita, S., Shimazu, T., Niwa, T. & Ushijima, T. The presence of RNA polymerase II, active or stalled, predicts epigenetic fate of promoter CpG islands. Genome Res. 19, 1974–1982 (2009).
Antonescu, C. R. et al. Novel YAP1-TFE3 fusion defines a distinct subset of epithelioid hemangioendothelioma. Genes Chromosomes Cancer 52, 775–784 (2013).
Veeraraghavan, J., Ma, J., Hu, Y. & Wang, X. S. Recurrent and pathological gene fusions in breast cancer: current advances in genomic discovery and clinical implications. Breast Cancer Res. Treat. 158, 219–232 (2016).
Szulzewsky, F., Holland, E. C. & Vasioukhin, V. YAP1 and its fusion proteins in cancer initiation, progression and therapeutic resistance. Dev. Biol. 475, 205–221 (2021).
Sekine, S. et al. Recurrent YAP1-MAML2 and YAP1-NUTM1 fusions in poroma and porocarcinoma. J. Clin. Invest. 129, 3827–3832 (2019).
Heallen, T. et al. Hippo pathway inhibits Wnt signaling to restrain cardiomyocyte proliferation and heart size. Science 332, 458–461 (2011).
Liao, Y. et al. OLIG2 maintenance is not essential for diffuse intrinsic pontine glioma cell line growth but regulates tumor phenotypes. Neuro Oncol. 23, 1183–1196 (2021).
Lastowska, M. et al. Transcriptional profiling of paediatric ependymomas identifies prognostically significant groups. J. Pathol. Clin. Res. 7, 565–576 (2021).
Wansink, D. G. et al. Fluorescent labeling of nascent RNA reveals transcription by RNA polymerase II in domains scattered throughout the nucleus. J. Cell Biol. 122, 283–293 (1993).
Gao, Y. et al. Single Cas9 nickase induced generation of NRAMP1 knockin cattle with reduced off-target effects. Genome Biol. 18, 13 (2017).
Zhang, L. et al. Single-cell transcriptomics in medulloblastoma reveals tumor-initiating progenitors and oncogenic cascades during tumorigenesis and relapse. Cancer Cell 36, 302–318 (2019).
Skene, P. J. & Henikoff, S. An efficient targeted nuclease strategy for high-resolution mapping of DNA binding sites. eLife 16, 21856 (2017).
Juhling, F. et al. metilene: fast and sensitive calling of differentially methylated regions from bisulfite sequencing data. Genome Res. 26, 256–262 (2016).
Juric, I. et al. MAPS: model-based analysis of long-range chromatin interactions from PLAC-seq and HiChIP experiments. PLoS Comput. Biol. 15, e1006982 (2019).
Acknowledgements
We thank R. J. Gilbertson for C11ORF95-YAP1 construct and R. Tjian for LacO-LacI and RPB1 constructs, R. L Johnson for Lats1/2 floxed mice, X. Zhang for MED1 antibody, X. Zhou for cartoon drawing and Y. Wang for technical assistance and plasmids of CRISPR/Cas9-mediated knock-in. X. Hu and X. Wu performed experiments at the Cincinnati Children’s Hospital, and current affiliation is Fudan University. This study was in part funded by grants from the CancerFree KIDS, Pray-Hope-Believe Foundation and TeamConnor Childhood Cancer Foundation, and CureStartsNow Foundation to Q.R.L.
Author information
Authors and Affiliations
Contributions
X.H. and Q.R.L. designed the project and analysed data. X.H. and X.W. performed cell, molecular and animal experiments. X.H. and K.B. performed IUE animal experiments. C.Z. performed HiChIP–seq and constructed YAP-NLS. D.X. performed native gel study and CUT&RUN–seq experiments. L.Z. performed ChIP–seq. S.O. performed allografts and drug treatment. X.L. and M.S. performed immunohistochemistry. Z.L. performed methyl-seq and short hairpin RNA knockdown. J.T. and M.L. provided EPN samples. F.S. and E.C.H. provided YAP-TFE3, YAP-SS18 and YAP-FAM118B. L.L. and M.H. analysed HiChIP. M.X. provided resources. X.H. and Q.R.L. wrote and revised the manuscript. Q.R.L. supervised the study.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Cell Biology thanks Howard Fine, Peter St George-Hyslop and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 The activation of HIPPO-YAP/TAZ in human ependymoma.
a, Enrichment of HIPPO signaling pathway in ependymoma (EPN) relative to high grade glioma (HGG), low grade glioma (LGG), diffuse intrinsic pontine glioma (DIPG), and medulloblastoma (MB). Kolmogorov-Smirnov test. p and q (FDR-corrected p) values were indicated here. b-c, YAP and TAZ (b) YAP target genes CTGF and CYR61 (c) expression in human brain tumors. Black lines indicate mean. EPN (n = 98) stands for ependymoma, HGG (n = 139) for high-grade glioma, LGG (n = 261) for low-grade glioma, DIPG (n = 32) for diffuse intrinsic pontine glioma, and MB (n = 133) for medulloblastoma. Mann-Whitney test. d, Mutation rates of the core components of the HIPPO signaling pathway in human brain tumor. Chi-squared analysis (df=4, n = 2302), p = 1.4e-4. All ‘n’ here referred to number of individual cases.
Extended Data Fig. 2 Activation of HIPPO-YAP/TAZ leads to ependymoma in mice.
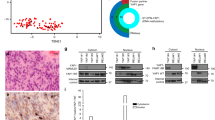
a, Diagram of Lats1/2 knockout by transduction of Cre in Lats1fl/f;Lats2fl/fl mice via in utero electroporation (IUE). b, Immunostaining of YAP in the brain of Lats1fl/f;Lats2fl/fl E17.5 embryos electroporated with pCAG-Cre or control constructs. Arrows indicate nuclear puncta formation. Scale bars, 10 μm. c, Hematoxylin and eosin (H&E) staining showing control and Lat1/2 double conditional knockout (cKO) tumors from 4-month-old mice. Scale bars, 1 mm (left) and 20 μm (right). d, Quantification of tumorigenic phenotypes of indicated mice. i, e,g, Immunostaining of ependymoma-related markers (e) and neural stemness-related markers (g) in Lats1/2 cKO tumors. Scale bars, 20 μm. f, h, Quantification of cells positive for ependymoma markers (f) and neural stemness-related markers (h) in Lats1/2 cKO tumors and control tissues. Data are presented as mean ± s.e.m (unpaired two-tailed Student’s t-test). n = 3 independent experiments. i, Sections of cortical tissues from mice transduced with YAP-M or YAP-C and mouse glioma tissue immunostained for OLIG2. Scale bars, 10 μm.
Extended Data Fig. 3 Sphere, allograft formation and methylation profiling of YAP-M fusion-driven mouse tumors.
a, Representative images of neurospheres formed by NSCs isolated from E17.5 embryos of wild-type (NSC) and tumor cells that express YAP fusion proteins YAP-M and YAP-C. Scale bars, 500 μm. b, Percentage of wells containing NSCs, YAP-M-expressing cells, and YAP-C-expressing cells that formed spheres in continuous passages. Data are presented as mean ±s.e.m (unpaired two-tailed Student’s t-test). n = 9 independent experiments. c, Sphere diameters from the fourth passage of NSCs and YAP-M- and YAP-C-expressing cells. Data are presented as mean ± s.e.m (unpaired two-tailed Student’s t-test). n = 30 independent spheres. d, Representative bioluminescence images of the NSG mice engrafted with 1.8×105 and 6×104 YAP-M-expressing tumor cells at 50 day after injection. e, Allograft tumor formation at different doses of YAP-M-expressing tumor cells. f, Heat-map representation of an unsupervised clustering of DNA methylation profiles of human ST-EPN-YAP tumors (n = 13), mouse tumors induced by YAP-MAMLD1 (n = 3) and mouse neural stem cells (NSC) (n = 3) across promoter regions of around top 600 variable methylated gene orthologs among human EPN tumors. The signature genes specific for ST-EPN-YAP are shown in the right side. Each row represents a gene; each column represents a sample.
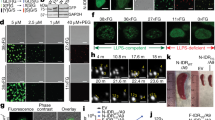
Extended Data Fig. 4 YAP-M and YAP-C fusion proteins form liquid-like nuclear condensates.
a, Sections of cortical tumor tissues from mice transduced with YAP-M or YAP-C immunostained for YAP. Scale bar, 10 μm. b, 293 T cells transfected with empty vector or vectors expressing YAP, YAP-S112A, YAP-NLS, YAP-S112A-NLS, YAP-M, and YAP-C fusions immunostained for YAP (red). DAPI (blue) was used to stain nuclei. Scale bar, 5 μm. c, C2C12 cells transfected with YAP-M construct. Scale bar, 20 μm (left) and 5 μm (Right). d, Quantification of nuclear fluorescence intensity of the cells transfected with YAP-derivatives. Data shown are means; n = 30 cells for each group, pooled from 2 independent experiments. one-tailed unpaired t-test. e,f, YAP western blot analysis (e) and quantifications (f) showing the nuclear levels of YAP-NLS, YAP-M or YAP-C. Histone H1, loading control. n = 2 independent experiments with similar results. g, Live-cell imaging of C2C12 cells expressing YAP-C-GFP. The arrows indicate representative YAP-C nuclear puncta that fused over time. This assay was performed twice (two independent transfections) with similar results. Scale bar, 5 μm. h, Time-lapse images of YAP-M-GFP or YAP-C-GFP after 5% 1,6-hexanediol treatment.
Extended Data Fig. 5 Endogenous YAP-MAMLD1 proteins undergo phase separation.
a, Tissue sections of a human ST-EPN tumor carrying YAP-MAMLD1 fusion and a low-grade glioma (LGG) were immunostained for YAP (green). DAPI (blue), nuclei. Scale bar, 10 μm. b, Scheme for site-specific knock-in (KI) of YAP-M fusion at the YAP1 locus via CRISPR/cas9n. c, HeLa cells with YAP-M knock-in were immunostained for YAP (red) and HA-tagged YAP-M (green). DAPI (blue), nuclei. Scale bar, 5 μm. d, Western blot analysis of nuclear and total protein levels of YAP and YAP-M in HeLa cells without or with knocked-in YAP-M. GAPDH and histone H1 as loading controls. e, Quantification of nuclear and total protein levels of YAP and YAP-M in HeLa cells without (Ctrl) or with YAP-M knock-in. n = 3 independent experiments. Data are presented as mean ± s.e.m (unpaired two-tailed Student’s t-test).
Extended Data Fig. 6 Disruption of YAP-TEAD interaction does not abolish phase-separation of YAP-M proteins.
a, YAPS94A-MAMLD1, which lost the ability of YAP to interact with TEAD, undergoes phase separation in C2C12 cells. Scale bar, 5 μm. b, YAP-dependent luciferase activity in NIH 3T3 cells expressing 8×GTIIC and YAP-MAMLD1 or YAPS94A-MAMLD1 constructs. Data are means ± s.e.m.; p values were determined using unpaired two-tailed Student’s t-tests; n = 3 biologically independent samples. c, C2C12 cells transfected with YAP-M, treated with 5 μM of verteporfin for 24hrs and stained for YAP (red). DAPI (blue) was used to stain nuclei. Scale bar, 5 μm.
Extended Data Fig. 7 YAP fusions contain IDRs and form oligomers in the nucleus to promote tumorigenesis.
a, Plot of disordered regions in YAP-C fusion identified using PONDR. VSL2 scores are shown on the y axis and positions are shown on the x axis. The blue bar designates the IDR investigated. b, Amino acid compositions of YAP-M (upper) and YAP-C (lower). c, Immunostaining of YAP in the C2C12 cells transfected with YAP-IDR construct. Arrows indicate condensate-like structures. Scale bar, 5 μm. d, C2C12 cells transfected with CC-YAP or CC-YAP-NLS stained for YAP (red). DAPI (blue) was used to stain nuclei. Scale bar, 5 μm. e, Representative bioluminescence images of the mice electroporated with CC-YAP or CC-YAP-NLS during tumor progression. f, H&E staining of brain tumor tissues of the mice that express CC-YAP-NLS. Arrow indicates tumor lesion. Scale bar, 1 mm.
Extended Data Fig. 8 Expression of YAP-M promotes oncogenic programs in NSCs.
a, Volcano plot of transcriptome profiles of YAP-M-expressing NSCs versus YAP-expressing NSCs. Red and green dots represent genes significantly upregulated and downregulated upon YAP-M expression (P < 0.01). Differential expression was determined by FDR-corrected p-value from Wald test. b, GSEA analysis of genes differentially expressed in YAP-M-expressing NSCs (n = 4) versus SVZ of wild-type mice (n = 3). n = biologically independent replicates. c, Pathway enrichment analysis of genes differentially expressed in YAP-M-expressing NSCs versus YAP-expressing NSCs (red) or SVZ of wild-type mice (orange). Distinct pathways and biological processes between the molecular subgroups of ependymal tumors are shown (FDR corrected p < 0.01). Nodes represent enriched gene sets, which are grouped by their similarity. Node size is proportional to the total number of genes within each gene set. The illustrated network map was simplified by manual curation to remove general and uninformative sub-networks. Fisher’s exact test was applied to determine the significance level.
Extended Data Fig. 9 YAP-M fusion induces long-range super-enhancer interactions to promote oncogenic signaling.
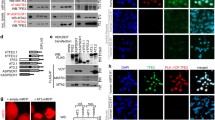
a, Circos map of combined omic profiles of YAP-M tumors. The innermost layer represents oncogenic genes near super-enhancers (SEs). The second layer includes SEs (n = 627). The third layer includes all enhancers (n = 16,381). The fourth layer represents the top 10,000 binding loci of YAP. The fifth layer includes differentially expressed genes in tumors from mice transfected with the YAP-M expression construct relative to NSCs that express YAP. The sixth layer represents the mouse genome (mm10). b, Distribution of H3K27ac binding genomic regions in the indicated cells. c, YAP-M fusion induces long-range enhancer/promoter interactions at the gene loci associated oncogenic signaling. Genomic tracks depicting long-range enhancer/promoter interactions at the indicated loci of oncogenesis-promoting genes in the control and YAP-M transduced NSCs. Pink arcs, DNA contacts; histograms, YAP1 and H3K27ac ChIP-seq peaks. d,e, Co-staining of HA-tagged YAP-M or YAP-MΔIDR (green) and H3K27ac (red) and immunoblot analysis in C2C12 cells transfected with YAP-M or YAP-MΔIDR constructs. Scale bars, 10 μm.
Extended Data Fig. 10 The effect of pharmacological and genetic inhibition of Brd4 on nuclear puncta formation and tumor growth, and IDR-containing YAP fusions in different cancers.
a, C2C12 cells transfected with YAP-M, treated with 0.1 μM of JQ1 or vehicle and stained for YAP (red). Scale bars, 10 μm. b, Western blot showing the effect of shRNAs to repress BRD4 or MED1 expression. c, Growth of murine NPCs expressing YAP-M fusion was inhibited by knock-down of BRD4 or MED1. 3 independent transfections for each group. Data are presented as mean ± s.e.m. Unpaired student’s two-sided t-test was applied to compare cell numbers at the terminal time point. d, Representative bioluminescence images of the mice electroporated with YAP-M construct after treated with vehicle or BMS-986158 (a chemical inhibitor of Brd4) during tumor progression. e, Survival analysis of mice transfected with the YAP-M expression construct treated with BMS-986158 or vehicle. N = 6 animals/group. Log rank test. f, IDR regions in YAP fusion proteins are predicted in multiple cancer types. g, YAP immunostaining in the HEK293T cells transfected with vectors expressing YAP fusions (YAP1-TFE3, YAP1-SS18 and YAP1-FAM118B) shows nuclear puncta-like structures. Scale bars: 10 μm.
Supplementary information
Supplementary Video 1
Dynamic fusion of nuclear puncta of YAP-MAMLD1-GFP.
Source data
Source Data Fig. 4
Unprocessed western blots.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 5
Unprocessed western blots.
Source Data Fig. 6
Statistical source data.
Source Data Fig. 7
Statistical source data.
Source Data Fig. 7
Unprocessed western blots.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 4
Unprocessed western blots.
Source Data Extended Data Fig. 5
XH_unmodified_gels_ED_Fig4e.
Source Data Extended Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 9
Statistical source data.
Source Data Extended Data Fig. 9
Unprocessed western blots.
Source Data Extended Data Fig. 10
Unprocessed western blots.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Hu, X., Wu, X., Berry, K. et al. Nuclear condensates of YAP fusion proteins alter transcription to drive ependymoma tumourigenesis. Nat Cell Biol 25, 323–336 (2023). https://doi.org/10.1038/s41556-022-01069-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-022-01069-6