Abstract
Epidemiological studies demonstrate an association between breast cancer (BC) and systemic dysregulation of glucose metabolism. However, how BC influences glucose homeostasis remains unknown. We show that BC-derived extracellular vesicles (EVs) suppress pancreatic insulin secretion to impair glucose homeostasis. EV-encapsulated miR-122 targets PKM in β-cells to suppress glycolysis and ATP-dependent insulin exocytosis. Mice receiving high-miR-122 EVs or bearing BC tumours exhibit suppressed insulin secretion, enhanced endogenous glucose production, impaired glucose tolerance and fasting hyperglycaemia. These effects contribute to tumour growth and are abolished by inhibiting EV secretion or miR-122, restoring PKM in β-cells or supplementing insulin. Compared with non-cancer controls, patients with BC have higher levels of circulating EV-encapsulated miR-122 and fasting glucose concentrations but lower fasting insulin; miR-122 levels are positively associated with glucose and negatively associated with insulin. Therefore, EV-mediated impairment of whole-body glycaemic control may contribute to tumour progression and incidence of type 2 diabetes in some patients with BC.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
RNA-seq data that support the findings of this study have been deposited in the Gene Expression Omnibus (GEO) under accession codes GSE152391, GSE173408, and GSE173276. All other data supporting the findings of this study are available from the corresponding author on reasonable request. Source data are provided with this paper.
References
Ferlay, J. et al. Cancer incidence and mortality worldwide: sources, methods and major patterns in GLOBOCAN 2012. Int. J. Cancer 136, E359–E386 (2015).
Ryu, T. Y., Park, J. & Scherer, P. E. Hyperglycemia as a risk factor for cancer progression. Diabetes Metab. J. 38, 330–336 (2014).
Maskarinec, G. et al. Type II diabetes, obesity, and breast cancer risk: the multiethnic cohort. Cancer Epidemiol. Biomark. Prev. 26, 854–861 (2017).
Bronsveld, H. K. et al. Diabetes and breast cancer subtypes. PLoS ONE 12, e0170084 (2017).
Xue, F. & Michels, K. B. Diabetes, metabolic syndrome, and breast cancer: a review of the current evidence. Am. J. Clin. Nutr. 86, s823–s835 (2007).
Larsson, S. C., Mantzoros, C. S. & Wolk, A. Diabetes mellitus and risk of breast cancer: a meta-analysis. Int J. Cancer 121, 856–862 (2007).
Liao, S. et al. Association between diabetes mellitus and breast cancer risk: a meta-analysis of the literature. Asian Pac. J. Cancer Prev. 12, 1061–1065 (2011).
Boyle, P. et al. Diabetes and breast cancer risk: a meta-analysis. Br. J. Cancer 107, 1608–1617 (2012).
Luque, R. M. et al. Breast cancer is associated to impaired glucose/insulin homeostasis in premenopausal obese/overweight patients. Oncotarget 8, 81462–81474 (2017).
Villarreal-Garza, C. et al. Impact of diabetes and hyperglycemia on survival in advanced breast cancer patients. Exp. Diabetes Res 2012, 732027 (2012).
Nam, S. et al. Association between insulin resistance and luminal B subtype breast cancer in postmenopausal women. Medicine 95, e2825 (2016).
Duggan, C. et al. Associations of insulin resistance and adiponectin with mortality in women with breast cancer. J. Clin. Oncol. 29, 32–39 (2011).
Hernandez, A. V. et al. Association between insulin resistance and breast carcinoma: a systematic review and meta-analysis. PLoS ONE 9, e99317 (2014).
Goodwin, P. J. et al. Insulin- and obesity-related variables in early-stage breast cancer: correlations and time course of prognostic associations. J. Clin. Oncol. 30, 164–171 (2012).
Rose, D. P. & Vona-Davis, L. The cellular and molecular mechanisms by which insulin influences breast cancer risk and progression. Endocr. Relat. Cancer 19, R225–R241 (2012).
Bruning, P. F. et al. Insulin resistance and breast-cancer risk. Int J. Cancer 52, 511–516 (1992).
Muti, P. et al. Fasting glucose is a risk factor for breast cancer: a prospective study. Cancer Epidemiol. Biomark. Prev. 11, 1361–1368 (2002).
Vona-Davis, L. & Rose, D. P. Type 2 diabetes and obesity metabolic interactions: common factors for breast cancer risk and novel approaches to prevention and therapy. Curr. Diabetes Rev. 8, 116–130 (2012).
Gunter, M. J. et al. Insulin, insulin-like growth factor-I, and risk of breast cancer in postmenopausal women. J. Natl Cancer Inst. 101, 48–60 (2009).
Kaaks, R. et al. Prospective study of IGF-I, IGF-binding proteins, and breast cancer risk, in northern and southern Sweden. Cancer Causes Control 13, 307–316 (2002).
Keinan-Boker, L. et al. Circulating levels of insulin-like growth factor I, its binding proteins -1,-2, -3, C-peptide and risk of postmenopausal breast cancer. Int J. Cancer 106, 90–95 (2003).
Eliassen, A. H., Tworoger, S. S., Mantzoros, C. S., Pollak, M. N. & Hankinson, S. E. Circulating insulin and C-peptide levels and risk of breast cancer among predominately premenopausal women. Cancer Epidemiol. Biomark. Prev. 16, 161–164 (2007).
Verheus, M. et al. Serum C-peptide levels and breast cancer risk: results from the European Prospective Investigation into Cancer and Nutrition (EPIC). Int J. Cancer 119, 659–667 (2006).
Lipscombe, L. L., Goodwin, P. J., Zinman, B., McLaughlin, J. R. & Hux, J. E. Increased prevalence of prior breast cancer in women with newly diagnosed diabetes. Breast Cancer Res. Treat. 98, 303–309 (2006).
Lipscombe, L. L. et al. Incidence of diabetes among postmenopausal breast cancer survivors. Diabetologia 56, 476–483 (2013).
Onitilo, A. A. et al. Breast cancer incidence before and after diagnosis of type 2 diabetes mellitus in women: increased risk in the prediabetes phase. Eur. J. Cancer Prev. 23, 76–83 (2014).
Bordeleau, L. et al. Diabetes and breast cancer among women with BRCA1 and BRCA2 mutations. Cancer 117, 1812–1818 (2011).
Onitilo, A. A. et al. Breast and prostate cancer survivors in a diabetic cohort: results from the Living with Diabetes Study. Clin. Med. Res. 11, 210–218 (2013).
Tkach, M. & Thery, C. Communication by extracellular vesicles: where we are and where we need to go. Cell 164, 1226–1232 (2016).
Becker, A. et al. Extracellular vesicles in cancer: cell-to-cell mediators of metastasis. Cancer Cell 30, 836–848 (2016).
Wang, S. E. Extracellular vesicles and metastasis. Cold Spring Harbor Perspect. Med. 10, a037275 (2019).
Valadi, H. et al. Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat. Cell Biol. 9, 654–659 (2007).
Fong, M. Y. et al. Breast-cancer-secreted miR-122 reprograms glucose metabolism in premetastatic niche to promote metastasis. Nat. Cell Biol. 17, 183–194 (2015).
Redzic, J. S., Balaj, L., van der Vos, K. E. & Breakefield, X. O. Extracellular RNA mediates and marks cancer progression. Semin. Cancer Biol. 28, 14–23 (2014).
Mitchell, P. S. et al. Circulating microRNAs as stable blood-based markers for cancer detection. Proc. Natl Acad. Sci. USA 105, 10513–10518 (2008).
Wu, X. et al. De novo sequencing of circulating miRNAs identifies novel markers predicting clinical outcome of locally advanced breast cancer. J. Transl. Med 10, 42 (2012).
Zhou, W. et al. Cancer-secreted miR-105 destroys vascular endothelial barriers to promote metastasis. Cancer Cell 25, 501–515 (2014).
Ashcroft, F. M. ATP-sensitive potassium channelopathies: focus on insulin secretion. J. Clin. Invest. 115, 2047–2058 (2005).
Nakatsu, D. et al. l-cysteine reversibly inhibits glucose-induced biphasic insulin secretion and ATP production by inactivating PKM2. Proc. Natl Acad. Sci. USA 112, E1067–E1076 (2015).
Parkes, D. G., Pittner, R., Jodka, C., Smith, P. & Young, A. Insulinotropic actions of exendin-4 and glucagon-like peptide-1 in vivo and in vitro. Metab. Clin. Exp. 50, 583–589 (2001).
Benediktsson, A. M., Schachtele, S. J., Green, S. H. & Dailey, M. E. Ballistic labeling and dynamic imaging of astrocytes in organotypic hippocampal slice cultures. J. Neurosci. Methods 141, 41–53 (2005).
Tsuyada, A. et al. CCL2 mediates cross-talk between cancer cells and stromal fibroblasts that regulates breast cancer stem cells. Cancer Res. 72, 2768–2779 (2012).
Yan, W. et al. Cancer-cell-secreted exosomal miR-105 promotes tumour growth through the MYC-dependent metabolic reprogramming of stromal cells. Nat. Cell Biol. 20, 597–609 (2018).
Tsuji, K. et al. Common bile duct injection as a novel method for establishing red fluorescent protein (RFP)-expressing human pancreatic cancer in nude mice. JOP 7, 193–199 (2006).
Thackeray, J. T. et al. Insulin supplementation attenuates cancer-induced cardiomyopathy and slows tumor disease progression. JCI Insight 2, https://doi.org/10.1172/jci.insight.93098 (2017).
Stenlof, K. et al. Efficacy and safety of canagliflozin monotherapy in subjects with type 2 diabetes mellitus inadequately controlled with diet and exercise. Diabetes Obes. Metab. 15, 372–382 (2013).
Simionescu, N. et al. Hyperglycemia determines increased specific microRNAs levels in sera and HDL of acute coronary syndrome patients and stimulates microRNAs production in human macrophages. PLoS ONE 11, e0161201 (2016).
Willeit, P. et al. Circulating microRNA-122 is associated with the risk of new-onset metabolic syndrome and type 2 diabetes. Diabetes 66, 347–357 (2017).
Rupp, A. K. et al. Loss of EpCAM expression in breast cancer derived serum exosomes: role of proteolytic cleavage. Gynecol. Oncol. 122, 437–446 (2011).
Gallagher, E. J. & LeRoith, D. Minireview: IGF, insulin, and cancer. Endocrinology 152, 2546–2551 (2011).
Belardi, V., Gallagher, E. J., Novosyadlyy, R. & LeRoith, D. Insulin and IGFs in obesity-related breast cancer. J. Mammary Gland Biol. Neoplasia 18, 277–289 (2013).
Novosyadlyy, R. et al. Insulin-mediated acceleration of breast cancer development and progression in a nonobese model of type 2 diabetes. Cancer Res. 70, 741–751 (2010).
Hopkins, B. D. et al. Suppression of insulin feedback enhances the efficacy of PI3K inhibitors. Nature 560, 499–503 (2018).
Hay, N. Reprogramming glucose metabolism in cancer: can it be exploited for cancer therapy? Nat. Rev. Cancer 16, 635–649 (2016).
Javeed, N. et al. Pancreatic cancer-derived exosomes cause paraneoplastic β-cell dysfunction. Clin. Cancer Res. 21, 1722–1733 (2015).
Schultz, N. A. et al. MicroRNA biomarkers in whole blood for detection of pancreatic cancer. JAMA 311, 392–404 (2014).
Debnath, J. et al. The role of apoptosis in creating and maintaining luminal space within normal and oncogene-expressing mammary acini. Cell 111, 29–40 (2002).
Lee, Y. S. et al. The fractalkine/CX3CR1 system regulates β cell function and insulin secretion. Cell 153, 413–425 (2013).
Ran, F. A. et al. Genome engineering using the CRISPR–Cas9 system. Nat. Protoc. 8, 2281–2308 (2013).
Gray, J. T. & Zolotukhin, S. Design and construction of functional AAV vectors. Methods Mol. Biol. 807, 25–46 (2011).
Zhao, C. et al. Overcoming insulin insufficiency by forced follistatin expression in β-cells of db/db mice. Mol. Ther. 23, 866–874 (2015).
Anastasiou, D. et al. Inhibition of pyruvate kinase M2 by reactive oxygen species contributes to cellular antioxidant responses. Science 334, 1278–1283 (2011).
Shurtleff, M. J. et al. Broad role for YBX1 in defining the small noncoding RNA composition of exosomes. Proc. Natl Acad. Sci. USA 114, E8987–E8995 (2017).
van der Ree, M. H. et al. Safety, tolerability, and antiviral effect of RG-101 in patients with chronic hepatitis C: a phase 1B, double-blind, randomised controlled trial. Lancet 389, 709–717 (2017).
Jankowska, M. M. et al. Protocol for a cross sectional study of cancer risk, environmental exposures and lifestyle behaviors in a diverse community sample: the Community of Mine study. BMC Public Health 19, 186 (2019).
Acknowledgements
This work was supported by the National Institutes of Health (NIH)/National Cancer Institute (NCI) grants R01CA218140 (S.E.W.), R01CA206911 (S.E.W.) and R01CA179977 (D.D.S.). J.D.B. is supported by a grant from the Hartwell Foundation. Research reported in this publication included work performed in the core facilities supported by the NIH/NCI under grant number P30CA23100 (UCSD Moores Cancer Center) and in the City of Hope Integrative Genomics Core supported by NIH/NCI under grant number P30CA33572. The authors would like to thank the UCSD/CMM EM facility for EM sample preparation. The EM facility is supported by NIH equipment grant 1S10OD023527. We thank X. Xiao for kindly providing plasmid pEMBOL-D(+)-Ins-GFP.
Author information
Authors and Affiliations
Contributions
S.E.W. and M.C. conceived ideas, and J.M.O., D.D.S. and W. Ying contributed to project planning. M.C. and S.E.W. designed and performed most of the experiments. R.I., W. Yan, X.R., L.J., Y.W., J.W., E.W., X.L., A.R.C. and M.Y.F. assisted with mouse experiments, cell line construction and EV preparation. S.N. and D.D. provided miR-122 ONI and control oligonucleotides and helped design experiments involving oligonucleotide treatment. D.P.P. assisted with tissue processing and histological analyses. X.W. assisted with RNA-seq. C.C. and J.D.B. assisted with flow cytometry. Z.G., K.G. and W.Z. assisted with NTA analysis of EVs. D.D.S., O.F., R.B.S., Y.Y., S.E.Y. and J.M. assisted with clinical sample assembly and assessment. S.E.W. and M.C. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
S.N. and D.D. are employees of Regulus Therapeutics, Inc. The other authors declare no competing interests.
Peer review
Peer review information
Nature Cell Biology thanks David Hodson, Luis Fajas and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Additional assessments of the EV injection model in Fig. 1.
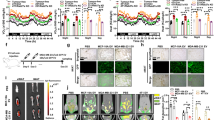
a, Food consumption was calculated by total food consumed in 48 h after the 8th EV/PBS injection (n = 3 mice per group). b, Body weight was monitored (n = 8 mice for MDA-MB-231 EV and n = 7 mice for other groups). c, Liver tissues collected at ZT10 from mice that had received 5 weeks of EV/PBS injections were subjected to RNA-seq and GSEA, showing enrichment of genes related to IRS1/2 signalling (n = 6 mice per group). d, Heatmap showing the relative levels of selected genes based on the RNA-seq data. e, Triglyceride levels measured in the liver (n = 4 mice for PBS and MCF-10A/vec EV group and n = 5 for other groups). f-h, Mouse serum glucagon (f), GLP-1 (g), and ghrelin (h) levels were determined by ELISA kits (n = 6 mice for PBS and MCF-10A/vec EV group and n = 8 for other groups). Data are shown as mean ± SEM. One-way ANOVA followed by Tukey’s multiple comparison test was used for a, e-h. Two-way ANOVA followed by Tukey’s multiple comparison test was used for b. ns: not significant. *P < 0.05. Numerical source data and statistics are available online.
Extended Data Fig. 2 Characterization of BC-cell-secreted EVs.
a, NTA of MDA-MB-231 EVs (n = 3 replicates). b, Immunoblots of indicated proteins in MDA-MB-231 whole cell lysate and EV fractions from OptiPrep gradient ultracentrifugation showing EV markers and a Golgi marker (GM130, as a negative control for EV-specific proteins). c, RT-qPCR-determined miR-122 (left) and miR-16 (right) levels in OptiPrep gradient fractions of MDA-MB-231 EVs (n = 3 replicates; normalized to an ath-miR159a spike-in control). d, miR-122 levels in EVs treated with Proteinase K (PK, 10 μg/mL) followed by RNase If (RNase, 40 U) or with PBS (as control) in the presence or absence of 1% Triton X-100 (TX-100) (n = 3 replicates). RT-qPCR data were normalized to an ath-miR159a spike-in control added after all treatments. e, RT-qPCR showing relative miR-122 levels (normalized to spike-in control) in EVs isolated from the sera of mice in Fig. 1 (n = 6 mice per group). Data are shown as mean ± SEM. One-way ANOVA followed by Tukey’s multiple comparison test was used for d and e. ns: not significant. *P < 0.05, **P < 0.01, ***P < 0.001. Numerical source data, statistics and unprocessed blots are available online.
Extended Data Fig. 3 Functional assessments of EVs and PKM2 in MIN6 β-cells.
a, Uptake of CFSE-labelled EVs by MIN6 cells (CFSE, green; DAPI, blue). Bar=50 μm. b, Immunoblots of MIN6 cells after EV/PBS treatment for 48 h. When indicated, MIN6 cells were pre-transfected with a PKM2 cDNA expression plasmid or empty vector. c, Pyruvate kinase activity in MIN6 cells (n = 3 replicates). d, Relative ATP/ADP ratio in MIN6 cells (n = 4 replicates) after low glucose (3.3 mM) or high glucose (16.7 mM) stimulation. Data were normalized to the first bar. e, Changes of the intracellular Ca2+ levels measured by Fura-2/AM in MIN6 cells after low or high glucose stimulation (n = 6 replicates). f, Insulin secretion by EV/PBS-treated MIN6 cells (n = 3 replicates) under 3.3 mM glucose, 16.7 mM glucose, or 20 mM monomethyl succinate (MMS). g, Immunoblots of MIN6 cells transfected with siRNA against Pkm2 (scrambled siRNA served as a negative control). h,i, Relative ATP/ADP ratio (h; n = 6 replicates) and insulin secretion (i; n = 4 replicates) in siRNA-transfected MIN6 cells after low glucose (3.3 mM) or high glucose (16.7 mM) stimulation. For all bar and line graphs, values are shown as mean ± SEM. One-way ANOVA followed by Tukey’s multiple comparison test was used for c and right panel of f. Two-way ANOVA with repeated measures followed by Tukey’s multiple comparison test was used for e. Two-way ANOVA followed by Tukey’s multiple comparison test was used for d and left panel of f, or by Bonferroni’s multiple comparison test used for h and i. *P < 0.05, **P < 0.01, ***P < 0.001 compared to PBS or as indicated; #P < 0.05, ##P < 0.01, ###P < 0.001 compared to MCF-10A/vec EV. In e, the upper set of signs indicate MCF-10A/miR-122 EV vs. PBS or MCF-10A/vec EV, and the lower set indicate MDA-MB-231 EV vs. PBS or MCF-10A/vec EV. Numerical source data, statistics and unprocessed blots are available online.
Extended Data Fig. 4 Characterization of MDA-MB-231-derived cells.
a, Immunoblots showing protein levels in indicated MDA-MB-231-derived cells and their EVs. EVs secreted from equal number of cells were loaded. b, NTA of EVs from indicated cells (n = 3 replicates). The inserted graph shows the numbers of EVs secreted in 24 hours per 107 cells (n = 3 replicates). c, Sanger sequencing showing genetic knockout of hsa-mir-122 gene in MDA-MB-231/miR-122 KO cells. Dashes indicate the region deleted by the CRISPR-Cas9 genome editing system. d, RT-qPCR-determined miR-122 and miR-16 levels (normalized to U6 snRNA) in indicated MDA-MB-231-derived cells (n = 6 replicates). e, Cell proliferation indicated by cell numbers counted every 24 hours (n = 4 replicates). f, Cell proliferation indicated by MTS assay. Optical density (O.D.) at 490 nm was shown (n = 8 replicates). g-j, Volcano plots showing differentially expressed miRNAs in indicated cells and EVs based on small RNA-seq data (n = 2 replicates). The position of miR-122 was noted in each plot. Data in bar and line graphs are shown as mean ± SEM. Two-way ANOVA followed by Tukey’s multiple comparison test was used for e and f. *P < 0.05, **P < 0.01. Numerical source data, statistics and unprocessed blots are available online.
Extended Data Fig. 5 Additional assessments of the MDA-MB-231-derived tumour model in Fig. 3a-n.
a, Body weight over time (n = 6 mice per group). b,c, Serum glucagon (b) and ghrelin (c) levels (n = 6 mice per group). d, Weight of gonadal white adipose tissue (WAT) (n = 7 mice per group). Values were normalized to whole body weight. e, RT-qPCR showing the relative mRNA levels (normalized to Gapdh mRNA) in gonadal WAT (n = 7 mice per group). f, Immunoblots of indicated proteins in the gastrocnemius muscle tissues from indicated groups. g, Representative images showing 2-NBDG uptake in the gastrocnemius from indicated groups (left; Bar=50μm) and overall 2-NBDG signal intensity quantified by ImageJ (right; n = 5 mice per group). Data in bar and line graphs are shown as mean ± SEM. Two-way ANOVA followed by Tukey’s multiple comparison post was used for a. One-way ANOVA followed by Tukey’s multiple comparison test was used for b-e and g. *P < 0.05, **P < 0.01, ***P < 0.001. Numerical source data, statistics and unprocessed blots are available online.
Extended Data Fig. 6 Cell proliferation and migration under different culture conditions.
a, Cell number counting every 24 hours under different glucose concentrations (n = 4 biologically independent plates of cells). Medium containing indicated glucose concentration was replenished every 24 hours. b, Wound closure migration assay under indicated glucose concentration. Bar=200μm. c, Cell number counting every 24 hours under different insulin concentrations (n = 4 biologically independent plates of cells). Medium containing indicated insulin concentration was replenished every 24 hours. Data are shown as mean ± SEM. Two-way ANOVA followed by Tukey’s multiple comparison test was used for a-c. *P < 0.05, **P < 0.01, ***P < 0.001. Numerical source data and statistics are available online.
Extended Data Fig. 7 Assessments of the pancreatic islets of mice in Fig. 7g-j.
a, Immunofluorescence of pancreas sections (left) showing PDX1 expression (green) in β-cells (insulin+; red). Bar=50μm. The overall frequency of PDX1+ among insulin+ cells was calculated and plotted in the right panel (n = 4 mice per group except n = 5 mice for MDA-MB-231/control + SGLT2i group). b, Immunofluorescence of pancreas sections (left) showing apoptosis events (indicated by cleaved caspase-3 staining; green) in β-cells (insulin+; red). Bar=50μm. The overall frequency of cleaved caspase-3+ among insulin+ cells was calculated and plotted in the right panel (n = 4 mice per group except n = 5 mice for MDA-MB-231/control + SGLT2i group). c, Representative H&E staining of pancreas sections (left) showing the regular morphology of mouse islets. Bar=50μm. Area of at least 8 islets from each mouse was calculated and plotted in the right panel (n = 4 mice per group except n = 5 mice for MDA-MB-231/control + SGLT2i group). Data are shown as mean ± SEM. One-way ANOVA followed by Tukey’s multiple comparison test was used for a-c. ns: not significant. Numerical source data and statistics are available online.
Extended Data Fig. 8 Characterization of CD24+ EVs enriched from human serum samples.
a, Immunoblots of indicated proteins in CD24+ EVs enriched from equal volume of human serum samples, including 3 non−cancer controls and 3 cases of BC. b, NTA and a representative EM image of the CD24+ EVs enriched from the serum of a BC patient. Bar=200 nm. c, Calculated ratio of miR-122 abundance in CD24+ vs. total (CD9/CD63/CD81+) circulating EVs. d, RT-qPCR showing relative miR-122 levels in CD24+ EVs (from 3 cases of BC patients) following indicated treatment. Data are shown as mean ± SEM. One-way ANOVA followed by Holm-Sidak’s multiple comparison test was used for c. *P < 0.05 and ***P < 0.001. Numerical source data, statistics and unprocessed blots are available online.
Extended Data Fig. 9 Gating strategies used for islet cell flow cytometry.
Gating strategies used to analyse uptake of CFSE-labelled EVs by insulin+ pancreatic islet β-cells presented in Fig. 2c. a-d, Islets were treated with PBS or EVs as indicated.
Supplementary information
Supplementary Table
Supplementary Tables 1–3
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 1
Unprocessed western blots.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 2
Unprocessed western blots.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 3
Unprocessed western blots.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 4
Unprocessed western blots.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 5
Unprocessed western blots.
Source Data Fig. 6
Statistical source data.
Source Data Fig. 6
Unprocessed western blots.
Source Data Fig. 7
Statistical source data.
Source Data Fig. 7
Unprocessed western blots.
Source Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 2
Unprocessed western blots.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 3
Unprocessed western blots.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 4
Unprocessed western blots.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 5
Unprocessed western blots.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 8
Unprocessed western blots.
Rights and permissions
About this article
Cite this article
Cao, M., Isaac, R., Yan, W. et al. Cancer-cell-secreted extracellular vesicles suppress insulin secretion through miR-122 to impair systemic glucose homeostasis and contribute to tumour growth. Nat Cell Biol 24, 954–967 (2022). https://doi.org/10.1038/s41556-022-00919-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-022-00919-7
This article is cited by
-
The systematic role of pancreatic cancer exosomes: distant communication, liquid biopsy and future therapy
Cancer Cell International (2024)
-
Quality control and validation of extracellular vesicles isolated from cultured human breast cancer cells
BMC Research Notes (2024)
-
The RNA binding protein IGF2BP2/IMP2 alters the cargo of cancer cell-derived extracellular vesicles supporting tumor-associated macrophages
Cell Communication and Signaling (2024)
-
Long noncoding RNA TMPO-AS1 accelerates glycolysis by regulating the miR-1270/PKM2 axis in colorectal cancer
BMC Cancer (2024)
-
Breast cancer cell-secreted miR-199b-5p hijacks neurometabolic coupling to promote brain metastasis
Nature Communications (2024)