Abstract
Epigenetic inheritance describes the transmission of gene regulatory information across generations without altering DNA sequences, enabling offspring to adapt to environmental conditions. Small RNAs have been implicated in this, through both the oocyte and the sperm. However, as much of the cellular content is extruded during spermatogenesis, it is unclear whether cytoplasmic small RNAs can contribute to epigenetic inheritance through sperm. Here we identify a sperm-specific germ granule, termed the paternal epigenetic inheritance (PEI) granule, that mediates paternal epigenetic inheritance by retaining the cytoplasmic Argonaute protein WAGO-3 during spermatogenesis in Caenorhabditis elegans. We identify the PEI granule proteins PEI-1 and PEI-2, which have distinct functions in this process: granule formation, Argonaute selectivity and subcellular localization. We show that PEI granule segregation is coupled to the transport of sperm-specific secretory vesicles through PEI-2 in an S-palmitoylation-dependent manner. PEI-like proteins are found in humans, suggesting that the identified mechanism may be conserved.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
The accession number for the small-RNA-seq data generated in this study is PRJNA629991. The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE95 partner repository under dataset identifier PXD019099. The BigDataViewer supporting the CLEM analyses is available at Mendeley Data (V1; https://doi.org/10.17632/dgb8d7h2hz.1; https://data.mendeley.com/datasets/dgb8d7h2hz/1). All other data supporting the findings of this study are available from the corresponding author on reasonable request. Source data are provided with this paper.
Code availability
All code developed for this analysis is available via https://github.com/Tunphie/SequencingTools/blob/main/smRNA_TypeCounter.py and https://github.com/Tunphie/SequencingTools/blob/main/CoverageOnProteinCodingGenes.py.
References
Bošković, A. & Rando, O. J. Transgenerational epigenetic inheritance. Annu. Rev. Genet. 52, 21–41 (2018).
Perez, M. F. & Lehner, B. Intergenerational and transgenerational epigenetic inheritance in animals. Nat. Cell Biol. 21, 143–151 (2019).
Hutvagner, G. & Simard, M. J. Argonaute proteins: key players in RNA silencing. Nat. Rev. Mol. Cell Biol. 9, 22–32 (2008).
Peters, L. & Meister, G. Argonaute proteins: mediators of RNA silencing. Mol. Cell 26, 611–623 (2007).
Castel, S. E. & Martienssen, R. A. RNA interference in the nucleus: Roles for small RNAs in transcription, epigenetics and beyond. Nat. Rev. Genet. 14, 100–112 (2013).
Xu, F., Guang, S. & Feng, X. Distinct nuclear and cytoplasmic machineries cooperatively promote the inheritance of RNAi in Caenorhabditis elegans. Biol. Cell 110, 217–224 (2018).
Wan, G. et al. Spatiotemporal regulation of liquid-like condensates in epigenetic inheritance. Nature 557, 679–683 (2018).
de Albuquerque, B. F. M., Placentino, M. & Ketting, R. F. Maternal piRNAs are essential for germline development following de novo establishment of endo-siRNAs in Caenorhabditis elegans. Dev. Cell 34, 448–456 (2015).
Phillips, C. M., Brown, K. C., Montgomery, B. E., Ruvkun, G. & Montgomery, T. A. PiRNAs and piRNA-dependent siRNAs protect conserved and essential C. elegans genes from misrouting into the RNAi pathway. Dev. Cell 34, 457–465 (2015).
Buckley, B. A. et al. A nuclear Argonaute promotes multigenerational epigenetic inheritance and germline immortality. Nature 489, 447–451 (2012).
Mao, H. et al. The Nrde pathway mediates small-RNA-directed histone H3 lysine 27 trimethylation in Caenorhabditis elegans. Curr. Biol. 25, 2398–2403 (2015).
Gent, J. I. et al. Distinct phases of siRNA synthesis in an endogenous RNAi pathway in C. elegans soma. Mol. Cell 37, 679–689 (2010).
Ketting, R. F., Haverkamp, T. H. A., Van Luenen, H. G. A. M. & Plasterk, R. H. A. mut-7 of C. elegans, required for transposon silencing and RNA interference, is a homolog of Werner syndrome helicase and RNaseD. Cell 99, 133–141 (1999).
Zhang, C. et al. mut-16 and other mutator class genes modulate 22G and 26G siRNA pathways in Caenorhabditis elegans. Proc. Natl Acad. Sci. USA 108, 1201–1208 (2011).
Phillips, C. M., Montgomery, T. A., Breen, P. C. & Ruvkun, G. MUT-16 promotes formation of perinuclear Mutator foci required for RNA silencing in the C. elegans germline. Genes Dev. 26, 1433–1444 (2012).
Gu, W. et al. Distinct Argonaute-mediated 22G-RNA pathways direct genome surveillance in the C. elegans Germline. Mol. Cell 36, 231–244 (2009).
Yigit, E. et al. Analysis of the C. elegans argonaute family reveals that distinct Argonautes act sequentially during RNAi. Cell 127, 747–757 (2006).
Xu, F. et al. A cytoplasmic Argonaute protein promotes the inheritance of RNAi. Cell Rep. 23, 2482–2494 (2018).
Ashe, A. et al. PiRNAs can trigger a multigenerational epigenetic memory in the germline of C. elegans. Cell 150, 88–99 (2012).
Shirayama, M. et al. PiRNAs initiate an epigenetic memory of nonself RNA in the C. elegans germline. Cell 150, 65–77 (2012).
Luteijn, M. J. et al. Extremely stable Piwi-induced gene silencing in Caenorhabditis elegans. EMBO J. 31, 3422–3430 (2012).
Claycomb, J. M. et al. The Argonaute CSR-1 and Its 22G-RNA cofactors are required for holocentric chromosome segregation. Cell 139, 123–134 (2009).
Updike, D. & Strome, S. P granule assembly and function in Caenorhabditis elegans germ cells. J. Androl. 31, 53–60 (2010).
Banani, S. F., Lee, H. O., Hyman, A. A. & Rosen, M. K. Biomolecular condensates: organizers of cellular biochemistry. Nat. Rev. Mol. Cell Biol. 18, 285–298 (2017).
Voronina, E., Seydoux, G., Sassone-Corsi, P. & Nagamori, I. RNA granules in germ cells. Cold Spring Harb. Perspect. Biol. 3, a002774 (2011).
Ellis, R. E. & Stanfield, G. M. The regulation of spermatogenesis and sperm function in nematodes. Semin. Cell Dev. Biol. 29, 17–30 (2014).
Conine, C. C. et al. Argonautes ALG-3 and ALG-4 are required for spermatogenesis-specific 26G-RNAs and thermotolerant sperm in Caenorhabditis elegans. Proc. Natl Acad. Sci. USA 107, 3588–3593 (2010).
Grishok, A., Tabara, H. & Mello, C. C. Genetic requirements for inheritance of RNAi in C. elegans. Science 287, 2494–2497 (2000).
Alcazar, R. M., Lin, R. & Fire, A. Z. Transmission dynamics of heritable silencing induced by double-stranded RNA in Caenorhabditis elegans. Genetics 180, 1275–1288 (2008).
Lev, I. et al. Germ granules govern small RNA inheritance. Curr. Biol. 29, 2880–2891.e4 (2019).
Ozata, D. M., Gainetdinov, I., Zoch, A., O’Carroll, D. & Zamore, P. D. PIWI-interacting RNAs: small RNAs with big functions. Nat. Rev. Genet. 20, 89–108 (2019).
Bagijn, M. P. et al. Function, targets, and evolution of Caenorhabditis elegans piRNAs. Science 337, 574–578 (2012).
Lee, H. C. et al. C. elegans piRNAs mediate the genome-wide surveillance of germline transcripts. Cell 150, 78–87 (2012).
Gudipati, R. K. et al. Protease-mediated processing of Argonaute proteins controls small RNA association. Mol. Cell https://doi.org/10.1016/j.molcel.2021.03.029 (2021).
Robert, V. P. V., Sijen, T., van Wolfswinkel, J. & Plasterk, R. H. A. Chromatin and RNAi factors protect the C. elegans germline against repetitive sequences. Genes Dev. 19, 782–787 (2005).
Vastenhouw, N. L. et al. A genome-wide screen identifies 27 genes involved in transposon silencing in C. elegans. Curr. Biol. 13, 1311–1316 (2003).
Stoeckius, M., Grün, D. & Rajewsky, N. Paternal RNA contributions in the Caenorhabditis elegans zygote. EMBO J. 33, 1740–1750 (2014).
Barucci, G. et al. Small RNA-mediated transgenerational silencing of histone genes impairs fertility in piRNA mutants. Nat. Cell Biol. 22, 235–245 (2020).
Spike, C. A., Bader, J., Reinke, V. & Strome, S. DEPS-1 promotes P-granule assembly and RNA interference in C. elegans germ cells. Development 135, 983–993 (2008).
Collins, T., Stone, J. R. & Williams, A. J. All in the family: the BTB/POZ, KRAB, and SCAN domains. Mol. Cell. Biol. 21, 3609–3615 (2001).
Stogios, P. J. & Privé, G. G. The BACK domain in BTB-kelch proteins. Trends Biochem. Sci. 29, 634–637 (2004).
Kroschwald, S., Maharana, S. & Simon, A. Hexanediol: a chemical probe to investigate the material properties of membrane-less compartments. Matters https://doi.org/10.19185/matters.201702000010 (2017).
Marzahn, M. R. et al. Higher‐order oligomerization promotes localization of SPOP to liquid nuclear speckles. EMBO J. 35, 1254–1275 (2016).
Brangwynne, C. P. et al. Germline P granules are liquid droplets that localize by controlled dissolution/condensation. Science 324, 1729–1732 (2009).
Putnam, A., Cassani, M., Smith, J. & Seydoux, G. A gel phase promotes condensation of liquid P granules in Caenorhabditis elegans embryos. Nat. Struct. Mol. Biol. 26, 220–226 (2019).
Wang, J. et al. A molecular grammar governing the driving forces for phase separation of prion-like RNA binding proteins. Cell 174, 688–699 (2018).
Hanazawa, M., Yonetani, M. & Sugimoto, A. PGL proteins self associate and bind RNPs to mediate germ granule assembly in C. elegans. J. Cell Biol. 192, 929–937 (2011).
Kawasaki, I. et al. The PGL family proteins associate with germ granules and function redundantly in Caenorhabditis elegans germline development. Genetics 167, 645–661 (2004).
Kawasaki, I. et al. PGL-1, a predicted RNA-binding component of germ granules, is essential for fertility in C. elegans. Cell 94, 635–645 (1998).
Kelleher, J. F. et al. Myosin VI is required for asymmetric segregation of cellular components during C. elegans spermatogenesis. Curr. Biol. 10, 1489–1496 (2000).
Tabaczar, S., Czogalla, A., Podkalicka, J., Biernatowska, A. & Sikorski, A. F. Protein palmitoylation: palmitoyltransferases and their specificity. Exp. Biol. Med. 242, 1150–1157 (2017).
Gleason, E. J., Lindsey, W. C., Kroft, T. L., Singson, A. W. & L’Hernault, S. W. Spe-10 encodes a DHHC-CRD zinc-finger membrane protein required for endoplasmic reticulum/golgi membrane morphogenesis during Caenorhabditis elegans spermatogenesis. Genetics 172, 145–158 (2006).
Gonzalo, S. & Linder, M. E. SNAP-25 palmitoylation and plasma membrane targeting require a functional secretory pathway. Mol. Biol. Cell 9, 585–597 (1998).
Fukata, Y., Bredt, D. S. & Fukata, M. in The Dynamic Synapse: Molecular Methods in Ionotropic Receptor Biology (eds Kittler, J. T. & Moss, S. J.) Ch. 5 (CRC Press/Taylor & Francis, 2006).
Yao, H. et al. Inhibiting PD-L1 palmitoylation enhances T-cell immune responses against tumours. Nat. Biomed. Eng. 3, 306–317 (2019).
Wu, H. & Fuxreiter, M. The structure and dynamics of higher-order assemblies: amyloids, signalosomes, and granules. Cell 165, 1055–1066 (2016).
Conine, C. C. et al. Argonautes promote male fertility and provide a paternal memory of germline gene expression in C. elegans. Cell 155, 1532–1544 (2013).
Zhou, L. et al. BTBD18 regulates a subset of piRNA-generating loci through transcription elongation in mice. Dev. Cell 40, 453–466 (2017).
Kleiman, S. E. et al. Reduced human germ cell-less (HGCL) expression in azoospermic men with severe germinal cell impairment. J. Androl. 24, 670–675 (2003).
Gjerstorff, M. F. et al. GAGE cancer-germline antigens are recruited to the nuclear envelope by germ cell-less (GCL). PLoS ONE 7, e45819 (2012).
Brenner, S. The genetics of Caenorhabditis elegans. Genetics 77, 71–94 (1974).
Schweinsberg, P. J. & Grant, B. D. C. elegans gene transformation by microparticle bombardment. WormBook https://doi.org/10.1895/wormbook.1.166.1 (2013).
Haeussler, M. et al. Evaluation of off-target and on-target scoring algorithms and integration into the guide RNA selection tool CRISPOR. Genome Biol. 17, 148 (2016).
Chen, B. et al. Dynamic imaging of genomic loci in living human cells by an optimized CRISPR/Cas system. Cell 155, 1479–1491 (2013).
Chiu, J., March, P. E., Lee, R. & Tillett, D. Site-directed, ligase-independent mutagenesis (SLIM): a single-tube methodology approaching 100% efficiency in 4h. Nucleic Acids Res. 32, e174 (2004).
Chiu, J., Tillett, D., Dawes, I. W. & March, P. E. Site-directed, ligase-independent mutagenesis (SLIM) for highly efficient mutagenesis of plasmids greater than 8kb. J. Microbiol. Methods 73, 195–198 (2008).
Dickinson, D. J., Ward, J. D., Reiner, D. J. & Goldstein, B. Engineering the Caenorhabditis elegans genome using Cas9-triggered homologous recombination. Nat. Methods 10, 1028–1034 (2013).
Dickinson, D. J., Pani, A. M., Heppert, J. K., Higgins, C. D. & Goldstein, B. Streamlined genome engineering with a self-excising drug selection cassette. Genetics 200, 1035–1049 (2015).
Ward, J. D. Rapid and precise engineering of the caenorhabditis elegans genome with lethal mutation co-conversion and inactivation of NHEJ repair. Genetics 199, 363–377 (2014).
Frøkjær-Jensen, C. et al. Single-copy insertion of transgenes in Caenorhabditis elegans. Nat. Genet. 40, 1375–1383 (2008).
Paix, A. et al. Scalable and versatile genome editing using linear DNAs with microhomology to Cas9 sites in Caenorhabditis elegans. Genetics 198, 1347–1356 (2014).
Paix, A., Schmidt, H. & Seydoux, G. Cas9-assisted recombineering in C. elegans: genome editing using in vivo assembly of linear DNAs. Nucleic Acids Res. 44, e128 (2016).
Arribere, J. A. et al. Efficient marker-free recovery of custom genetic modifications with CRISPR/Cas9 in Caenorhabditis elegans. Genetics 198, 837–846 (2014).
El Mouridi, S. et al. Reliable CRISPR/Cas9 genome engineering in Caenorhabditis elegans using a single efficient sgRNA and an easily recognizable phenotype. G3 7, 1429–1437 (2017).
Klass, M. R. & Hirsh, D. Sperm isolation and biochemical analysis of the major sperm protein from Caenorhabditis elegans. Dev. Biol. 84, 299–312 (1981).
Shevchenko, A., Tomas, H., Havliš, J., Olsen, J. V. & Mann, M. In-gel digestion for mass spectrometric characterization of proteins and proteomes. Nat. Protoc. 1, 2856–2860 (2007).
Kappei, D. et al. HOT1 is a mammalian direct telomere repeat-binding protein contributing to telomerase recruitment. EMBO J. 32, 1681–1701 (2013).
Rappsilber, J., Mann, M. & Ishihama, Y. Protocol for micro-purification, enrichment, pre-fractionation and storage of peptides for proteomics using StageTips. Nat. Protoc. 2, 1896–1906 (2007).
Bluhm, A., Casas-Vila, N., Scheibe, M. & Butter, F. Reader interactome of epigenetic histone marks in birds. Proteomics 16, 427–436 (2016).
Cox, J. & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 26, 1367–1372 (2008).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 17, 10–12 (2011).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Anders, S., Pyl, P. T. & Huber, W. HTSeq-A Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Phillips, C. M. et al. MUT-14 and SMUT-1 DEAD box RNA helicases have overlapping roles in germline RNAi and endogenous siRNA formation. Curr. Biol. 24, 839–844 (2014).
Ortiz, M. A., Noble, D., Sorokin, E. P. & Kimble, J. A new dataset of spermatogenic vs. oogenic transcriptomes in the nematode Caenorhabditis elegans. G3 4, 1765–1772 (2014).
Ramírez, F., Dündar, F., Diehl, S., Grüning, B. A. & Manke, T. DeepTools: a flexible platform for exploring deep-sequencing data. Nucleic Acids Res. 42, 187–191 (2014).
Koulouras, G. et al. EasyFRAP-web: a web-based tool for the analysis of fluorescence recovery after photobleaching data. Nucleic Acids Res. 46, W467–W472 (2018).
Gleason, E. J. et al. Developmental genetics of secretory vesicle acidification during Caenorhabditis elegans spermatogenesis. Genetics 191, 477–491 (2012).
Kukulski, W. et al. Correlated fluorescence and 3D electron microscopy with high sensitivity and spatial precision. J. Cell Biol. 192, 111–119 (2011).
Kremer, J. R., Mastronarde, D. N. & McIntosh, J. R. Computer visualization of three-dimensional image data using IMOD. J. Struct. Biol. 116, 71–76 (1996).
Paul-Gilloteaux, P. et al. eC-CLEM: flexible multidimensional registration software for correlative microscopies. Nat. Methods 14, 102–103 (2017).
de Chaumont, F. et al. Icy: an open bioimage informatics platform for extended reproducible research. Nat. Methods 9, 690–696 (2012).
Perez-Riverol, Y. et al. The PRIDE database and related tools and resources in 2019: Improving support for quantification data. Nucleic Acids Res. 47, D442–D450 (2019).
Acknowledgements
We thank the members of the Ketting laboratory for discussions; H. Grosshans for reading the manuscript; M. Dörr and S. Hellmann for technical support; C. Werner and A. Dold of the IMB Genomics Core Facility for small RNA library preparation; the staff at the IMB Media Laboratory, Microscopy, Proteomic and Genomic Core Facilities for consumables, equipment and experimental support; and S. Köhler and M. Schorb for their help in CLEM sample preparation and visualization. Some of the strains were provided by the Caenorhabditis Genetics Center (CGC), funded by NIH Office of Research Infrastructure Programs (P40 OD010440). We acknowledge the GenEvo RTG funded by the Deutsche Forschungsgemeinschaft (DFG; 407023052/GRK2526/1), which enabled the conception of this project. E.J.G., S.P. and S.W.L. were supported for this work by Emory College of Arts and Sciences. This work was supported by grants of the DFG KE 1888/1-1, KE1888/1-2 and KE 1888/6-1 (to R.F.K.) and the National Institute of Health R35 GM119656 (to C.M.P.), and T32 GM118289 (to D.A.H.N.).
Author information
Authors and Affiliations
Contributions
J.S. and R.F.K. conceived the study and designed experiments. J.S. executed experiments and performed data analysis. S.D. and F.B. performed MS analysis. A.M.d.J.D. and A.-S.S. performed small-RNA-seq analysis. M.B. and V.O. performed the CLEM experiments. A.W.B. performed PEI-1/2 studies in cell culture. D.A.H.N. and C.M.P. provided unpublished strains. E.J.G., S.P. and S.W.L. shared unpublished data on MO counts. R.F.K. supervised the project. J.S. and R.F.K. wrote the manuscript with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Cell Biology thanks Ben Lehner, Oded Rechavi and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 WAGO-3 is required for germline immortality and transgenerational maintenance of RNAe.
a, Mortal germline assay representing loss of fertility of strains with indicated genotype at 25 °C. Statistical significance was tested with a log-rank-test (n = 15 populations per strain assayed in a single experiment). b, Diagram displaying mCherry::H2B(RNAe) reactivation in prg-1(n4357), prg-1(n4357);mut-7(xf125) and prg-1(n4357);wago-3(pk1673) mutant generations. F2-F5: second-fifth homozygous generation. For each generation, reactivation in 10 populations of 50 animals each was scored. Each plotted point represents the fraction of 50 animals that express the mCherry::H2B transgene. Since no prg-1(n4357) single mutant animal was found to reactivate mCherry::H2B expression, the value of this group is deterministically zero due to lack of variability/statistical noise. Thus, any positive number of animals that expresses the mCherry::H2B transgene in either the prg-1(n4357);mut-7(xf125) or prg-1(n4357);wago-3(pk1673) group causes a significant difference from the prg-1(n4357) group. c, Micrographs of three example animals with the mCherry::H2B transgene in either RNAe (prg-1(n4357)) or activated (prg-1(n4357);wago-3(pk1673) and prg-1(n4357);mut-7(xf125)) status. Top panel shows schematic representation of an adult hermaphroditic gonad. Activity status of the transgene was homogeneous in F2 homozygous prg-1(n4357) and prg-1(n4357);mut-7(xf125) mutants. Images represent two biologically independent experiments. Scale bars: 30 μm. Source data are provided.
Extended Data Fig. 2 WAGO-3 is associated with 22G RNAs.
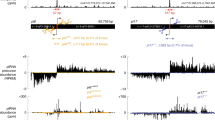
a, Read length distribution and first nucleotide bias of indicated small RNA libraries. Both hermaphrodite and male libraries were prepared and sequenced in biological triplicates. Each panel represents a replicate, top and bottom bars represent sense and anti-sense small RNAs mapping to annotated loci. b, Heat map showing the enrichment or depletion of small RNA species of the libraries shown in a. >27 nt are non-specific RNA fragments, also including fragments of structural RNAs, such as rRNA and tRNA. 21U, 22G, 26 G and miRNA are known Argonaute-associated small RNA species. Statistically significant differences between small RNA types were determined by Fisher’s exact tests (***: p ≤ 0.001, ns: p > 0.05). The exact P values are provided as source data. c, Box plots displaying the distribution of WAGO-3 associated 22G RNAs mapping to protein-coding genes across the indicated gene segments. Each box plot represents average data from three biological replicates. Boxplot centre and box edges indicate median and 25th or 75th percentiles, respectively, while whiskers indicate the median ± 1.5 x interquartile range. d, Metagene analysis of 22G RNA reads mapping to protein-coding WAGO-3 target genes. TSS – transcription start site, TES – transcription end site. Shading around each line represents the standard error of the median of each bin. Source data are provided.
Extended Data Fig. 3 WAGO-3 targets overlap with Mutator targets and sperm-derived 22 G RNAs.
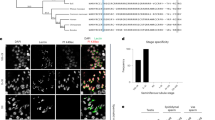
a, Scatter plots displaying the RPKM of 22 G RNAs mapping to individual genes in immunoprecipitation (IP) (Y-axis) versus input (X-axis) samples for all six sequenced libraries. Transposons are not included in these plots. Light blue: significant enrichment of genes in at least two replicates. Dark blue: significant enrichment of genes in only one replicate. b, Germline expression status of protein-coding WAGO-3 target genes. Left column shows the distribution of expression profiles of all annotated protein-coding genes. Columns 2 and 3 show the same, but for hermaphrodite and male-specific WAGO-3 targets, respectively. Column 4 shows the same for targets found in both sexes. Statistically significant differences with respect to complete gene annotations were determined by Chi-square tests. c, Scatter plots displaying the RPKM of 22G RNAs mapping to transposons in the six individual experiments in input (X-axes) and IP samples (Y-axes). Red line represents the diagonal. d, Venn diagrams displaying the overlaps of WAGO-3 targets (protein-coding) called in hermaphrodites and males with previously determined CSR-1 and MUT-16 targets. e, Stacked bar plot showing number and types of WAGO-3 targets. f, Heat map showing the overlap of WAGO-3 targets (protein-coding) with previously determined sperm-derived 22G RNA targets (protein-coding), which were binned into indicated groups. Values inside the boxes indicate fraction of overlap. Source data are provided.
Extended Data Fig. 4 PEI granules specifically recruit WAGO-3.
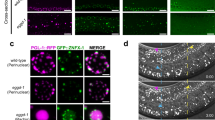
a,b, Confocal maximum intensity projections of an adult hermaphrodite (a) and gastrula-staged embryo (b) expressing indicated proteins. Zooms show perinuclear co-localization of GFP::3xFLAG::WAGO-3 and PGL-1::mTagRFP-T in meiotic (a) and primordial (b) germ cells. Z2/Z3 are the primordial germ cells of C. elegans. Scale bars: 20 μm (a, adult), 20 μm (b, embryo), 10 μm (a, zoom), 4 μm (b, zoom). c, Quantification of GFP::3xFLAG::WAGO-3 foci number within indicated, male-derived germ cells (n = 10 cells pooled from two independent experiments, for each condition). Statistically significant differences were determined by one-way ANOVA (p ≤ 0.001) followed by Tukey’s honestly significant difference post hoc test (p ≤ 0.05). Different letters represent significant differences. Note that the values for primary spermatocytes and budding spermatids (c) are the same as those shown in Fig. 3j (FL) and Extended Data Fig. 10f (wild-type), and Fig. 4n (FL) and Fig. 7j (wild-type), respectively. Secondary spermatocyte and budding spermatid stages are separated into ‘c’ and ‘s’. c: coupled; due to incomplete cytokinesis, leaving the two sister cells coupled and both cells were analyzed; s: separate, each of the coupled cells in ‘c’ was analyzed individually. Boxplot centre and box edges indicate median and 25th or 75th percentiles, respectively, while whiskers indicate the median ± 1.5 x interquartile range. d, Co-immunoprecipitation experiments using whole-worm extracts of late-L4 stage hermaphrodites analyzed by Western blotting. Sample processing control was run on a different gel. Data represent two biologically independent experiments. e-j, Confocal micrographs of an adult male (f) and late-L4 stage hermaphrodites (e,g-j) expressing indicated proteins. sc – spermatocyte, st – spermatid, rb – residual body. Scale bars: 10 μm (e-j). a,b,e-j, Images represent two biologically independent experiments. k-n, Co-localization analyses between PEI-1::mTagRFP-T and DEPS-1::GFP (k), GFP::3xFLAG::WAGO-1 (l), GFP::3xFLAG::CSR-1 (m) and GFP::3xFLAG::ALG-3 (n) based on the images shown in g-j, respectively (n = 10 worms for each condition). X and Y axes indicate fluorescence intensity. PC: Pearson’s correlation coefficient. Exact P values (c), unprocessed original scans of blots and the source data for all graphical representations are provided.
Extended Data Fig. 5 Presence of WAGO-3 in spermatozoa is dependent on the IDR of PEI-1.
a, Confocal micrographs showing spermatogenesis of late-L4 stage hermaphrodites expressing PEI 1::mTagRFP-T in indicated mutants. sc – spermatocyte, rb – residual body, st – spermatid. Images represent two biologically independent experiments. Scale bars: 10 μm. b-h, Confocal maximum intensity projections of hermaphrodite-derived spermatozoa within the spermatheca expressing indicated proteins. In all panels, except c, a piece of a gonad arm expressing GFP::3xFLAG::WAGO-3 is visible in the top part of the image. Dashed lines indicate spermatheca. Images represent two biologically independent experiments. Scale bars: 10 μm.
Extended Data Fig. 6 PEI granules are static condensates with liquid-like properties.
a,b, Confocal micrographs of isolated, male-derived spermatocytes (a) and budding spermatids (b) expressing GFP::3xFLAG::WAGO-3 and PEI-1::mTagRFP-T. Images were taken after a 30 minute treatment with 1,6-hexanediol. Hoechst33342 was used to stain DNA. Residual bodies are marked by a dashed circle. Images represent two biologically independent experiments. Scale bars: 4 μm. c, FRAP recovery curve of GFP::3xFLAG::WAGO-3 localizing to either P granules in L4 gonads or PEI granules in male-derived spermatids. Normalized data is presented as mean+/− SD and was fitted to a double exponential curve (n = 4 granules pooled from one independent experiment). d, Time sequence showing fluorescence recovery after photobleaching (FRAP) of GFP::3xFLAG::WAGO-3 localizing to either P granules in L4 gonads or PEI granules in male-derived spermatids. Residual bodies are marked by a dashed circle. Images represent two biologically independent experiments. Scale bars: 4 μm. e, Time sequence of GFP::3xFLAG::WAGO-3, taken from Extended Data Movie 1. Images are confocal maximum intensity projections of an isolated, male-derived spermatocyte. Images represent two biologically independent experiments. Scale bar: 4 μm. Source data are provided.
Extended Data Fig. 7 The amino acid composition of the PEI-1 IDR affects exchange dynamics of WAGO-3.
a,b, Occurrence of glycine and serine residues in PEI-1 (a) and PGL-1 (b) was counted in amino acid 50-mers, starting at position one, shifting 5 residues at a time, and displayed as stacked columns. Indicated residue positions in the diagrams are the mid-point of the 50-mer. Y-axes display number of relevant residues in amino acid 50-mers. X-axes indicate the position along the respective proteins. The various domains are indicated by vertical, dashed lines. NtDD and CDD indicate the N-terminal and C-terminal dimerization domains of PGL-1, respectively. The exact positions of glycine and serine residues for each protein are indicated above the stacked bar diagrams. IDR – intrinsically disordered region. c-h, Amino acid composition profiles of the intrinsically disordered region of the indicated proteins. Bars representing serine and glycine residues are highlighted in blue and orange, respectively. Panel d reflects a fusion between the PEI-1 IDR and the very C-terminal end of PGL-1(A711-F771). The profiles were generated using Composition profiler. Sequences were analyzed against the SwissProt database using 10,000 bootstrap iterations. Statistical significance was tested using the two sample t test (***: p ≤ 0.001, **: p ≤ 0.01, *: p ≤ 0.05, ns: p > 0.05). The exact P values are provided as source data. i-j, Time lapse images showing fluorescence recovery after photobleaching (FRAP) of GFP::3xFLAG::WAGO-3 localizing to PEI granules via PEI-1::mTagRFP-T (i) or via PEI-1::PGL-1(A711-F771)::mTagRFP-T (j) in isolated, male-derived budding spermatids. Residual bodies are marked by a dashed circle. Images represent two biologically independent experiments. Scale bars: 4 μm. k, FRAP recovery curves of GFP::3xFLAG::WAGO-3 localizing to PEI granules, containing indicated PEI-1 proteins, in male-derived budding spermatids. Normalized data is presented as mean + /− SD and was fitted to a double exponential curve (n = 5 granules pooled from one independent experiment). Source data are provided.
Extended Data Fig. 8 PEI-1 peptides at the N- and C-termini localize to asymmetrically segregated structures of defined shape.
a, Confocal micrograph of an L4 hermaphrodite expressing free mTagRFP-T from the pei-1 locus. The dashed line indicates the outline of the gonad. Scale bar: 20 μm. b, Confocal maximum intensity projection of spermatozoa within the spermatheca of an adult hermaphrodite expressing free mTagRFP-T from the pei-1 locus. The dashed line indicates the outline of the spermatheca. Scale bar: 10 μm. c, Confocal Z-stack of an isolated, male-derived spermatocyte expressing PEI-1_ΔH15-Q558::mTagRFP-T (see Fig. 4a, middle construct). Z-size: 125.9 nm. Scale bar: 4 μm. d, Confocal micrograph of two isolated, male-derived secondary spermatocytes expressing GFP::3xFLAG::WAGO-3 and PEI-1_ΔH15-Q558::mTagRFP-T. The pei-1 locus was heterozygous: pei-1_ΔH15-Q558::mTagRfp-t/pei-1(+). This allowed the formation and visualization of both PEI granules and PEI-1_ΔH15-Q558-specific structures within the same animal. Scale bar: 4 μm. e, Line profiles displaying relative fluorescence intensity for PEI-1_ΔH15-Q558::mTagRFP-T signals versus GFP::3xFLAG::WAGO-3 signals over the indicated, dashed line shown in panel d. Vertical lines and colored circles indicate fluorescence peaks. a.u. – arbitrary unit. f, Confocal micrograph of an isolated, male-derived spermatocyte (left) and budding spermatids (right) showing the subcellular distribution of mitochondria and PEI-1_ΔH15-Q558-specific structures. MitoTracker Green FM was used to stain mitochondria. Residual bodies are marked by dashed circles. Scale bar: 4 μm. g, Line profiles displaying relative fluorescence intensity for PEI-1_ΔH15-Q558::mTagRFP-T signals versus mitochondria signals over the indicated, dashed line shown in panel f. Vertical lines and colored circles indicate fluorescence peaks. a.u. – arbitrary unit. All images represent two biologically independent experiments. Source data are provided.
Extended Data Fig. 9 PEI granules are associated with membranous organelles throughout spermatogenesis.
a-c, Overview GFP::3xFLAG::WAGO-3 CLEM montages acquired from three high-pressure frozen adult males expressing GFP::3xFLAG::WAGO-3 and PEI-1::mTagRFP-T. The depicted animals were used to collect the high-resolution CLEM images shown in Fig. 6a-e. In all three panels, germ cell development progresses from left to right. The GFP::3xFLAG::WAGO-3 signal is fitted locally, implying that fluorescence signal was fitted using landmarks (tetraspecks and Hoechst staining) spread over the entire region of interest, spanning a larger field of view. The panels a’-a”’, b’ and c’ depict zoom-ins of the indicated areas in panels a-c. The indicated ‘Tomo’ regions within these zoomed-in panels are shown in greater detail in Fig. 6a-e. Precise CLEM overlays at specific ROIs (Tomos 1 to 5) were done using landmarks more locally and close to each ROI, covering a smaller field of view. The EM grids shown in panels a-c can be navigated using Fiji software and data deposited to Mendeley Data (https://data.mendeley.com/datasets/dgb8d7h2hz/1), following the instruction listed in Supplementary Information. All images represent two biologically independent experiments.
Extended Data Fig. 10 PEI-2 interacts with PEI-1.
a, Co-immunoprecipitation experiments using whole-worm extracts of late-L4 stage hermaphrodites analyzed by Western blotting. Labels above the blots indicate the presence (+) or absence (-) of the respective tags. Asterisks indicate non-specific signals. b, Pull-down experiments on extracts of transfected BmN4 cells expressing the indicated PEI-1 and PEI-2 variants. Full-length (FL) PEI-1-mCherry was pulled down, followed by detection of the various PEI-2 fragments. Expression of free 3xFLAG-eGFP served as negative control. c, Fraction of total mTagRFP-T signal within the residual body of male-derived budding spermatids expressing PEI-1_ΔH15-Q558::mTagRFP-T in wild-type or pei-2(xf270) mutant background (n = 10 cells pooled from two independent experiments, for each condition). Note that the wild-type data is the same as shown in Fig. 5c (ΔH15-Q558). d, Confocal maximum intensity projection of isolated, male-derived budding spermatids expressing PEI-1_ΔH15-Q558::mTagRFP-T in pei-2(xf270) mutant background. The residual body is marked by a dashed circle. Scale bar: 4 μm. e, Confocal maximum intensity projection of an isolated, male-derived primary spermatocyte expressing GFP::3xFLAG::WAGO-3 and PEI-1::mTagRFP-T in pei-2(xf270) mutant background. Scale bar: 4μm. d,e, Images represent two biologically independent experiments. f-g, Co-localization analysis between GFP::3xFLAG::WAGO-3 and PEI-1::mTagRFP-T (g), and quantification of GFP::3xFLAG::WAGO-3 foci number (f) in wild-type and pei-2(xf270) mutant, male-derived primary spermatocytes (n = 10 cells pooled from two independent experiments, for each condition). c,f,g, Statistically significant differences were determined by one-way ANOVA (p ≤ 0.001). Boxplot centre and box edges indicate median and 25th or 75th percentiles, respectively, while whiskers indicate the median ± 1.5 x interquartile range. Note that the wild-type data in f and g are the same as those shown in Fig. 3j (FL) and Extended Data Fig. 4c (primary spermatocyte), and Fig. 3k (FL), respectively. h-i, Transfected BmN4 cells were treated with the palmitoylation inhibitor 2-BP at indicated concentrations, followed by Western Blot detection of PEI-1-mCherry (h) and PEI-2-HA-eGFP (i). α-tubulin served as loading control. a,b,h,i, The experiment has been performed once. Unprocessed original scans of blots are provided in source data.
Supplementary information
Supplementary Information
Information about the BigDataViewer supporting the CLEM analyses.
Supplementary Video 1
Time sequence of an isolated, male-derived spermatocyte expressing GFP::3xFLAG::WAGO-3 as shown in Extended Data Fig. 6e. Images are confocal maximum intensity projections. Scale bar, 4 μm.
Supplementary Video 2
Time sequence of an isolated, male-derived spermatocyte expressing GFP::3xFLAG::WAGO-3 in absence of SPE-10. Images are confocal maximum intensity projections. Scale bar, 4 μm.
Supplementary Tables 1–3
Supplementary Table 1: list of C. elegans strains used in this study. Supplementary Table 2: list of sgRNA sequences used in this study. Supplementary Table 3: list of DNA donor templates used in this study.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 7
Statistical source data.
Source Data Fig. 8
Unprocessed western blots.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 4
Unprocessed western blots.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 10
Statistical source data.
Source Data Extended Data Fig. 10
Unprocessed western blots.
Rights and permissions
About this article
Cite this article
Schreier, J., Dietz, S., Boermel, M. et al. Membrane-associated cytoplasmic granules carrying the Argonaute protein WAGO-3 enable paternal epigenetic inheritance in Caenorhabditis elegans. Nat Cell Biol 24, 217–229 (2022). https://doi.org/10.1038/s41556-021-00827-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-021-00827-2
This article is cited by
-
A paternal protein facilitates sperm RNA delivery to regulate zygotic development
Science China Life Sciences (2023)
-
Systematic characterization of small RNAs associated with C. elegans Argonautes
Science China Life Sciences (2023)
-
Sperm granules mediate epigenetic inheritance
Nature Cell Biology (2022)