Abstract
Eosinophilic inflammation is a feature of allergic asthma. Despite mounting evidence showing that chromatin filaments released from neutrophils mediate various diseases, the understanding of extracellular DNA from eosinophils is limited. Here we show that eosinophil extracellular traps (EETs) in bronchoalveolar lavage fluid are associated with the severity of asthma in patients. Functionally, we find that EETs augment goblet-cell hyperplasia, mucus production, infiltration of inflammatory cells and expressions of type 2 cytokines in experimental non-infection-related asthma using both pharmaceutical and genetic approaches. Multiple clinically relevant allergens trigger EET formation at least partially via thymic stromal lymphopoietin in vivo. Mechanically, EETs activate pulmonary neuroendocrine cells via the CCDC25–ILK–PKCα–CRTC1 pathway, which is potentiated by eosinophil peroxidase. Subsequently, the pulmonary neuroendocrine cells amplify allergic immune responses via neuropeptides and neurotransmitters. Therapeutically, inhibition of CCDC25 alleviates allergic inflammation. Together, our findings demonstrate a previously unknown role of EETs in integrating immunological and neurological cues to drive asthma progression.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
Mass spectrometry data for EET-binding proteins have been deposited in ProteomeXchange with the identifier PXD027951 (https://www.ebi.ac.uk/pride/)122. Data of human CCDC25 RNA expression was derived from the FANTOM5 dataset included in the Human Protein Atlas database (https://www.proteinatlas.org/ENSG00000147419-CCDC25/tissue)37. All other data supporting the findings of this study are available from the corresponding author on reasonable request. Source data are provided with this paper.
References
Papi, A., Brightling, C., Pedersen, S. E. & Reddel, H. K. Asthma. Lancet 391, 783–800 (2018).
Global Strategy for Asthma Management and Prevention (Global Initiative for Asthma, 2020).
Holgate, S. T. et al. Asthma. Nat. Rev. Dis. Primers 1, 15025 (2015).
Lambrecht, B. N. & Hammad, H. The immunology of asthma. Nat. Immunol. 16, 45–56 (2015).
Lee, J. J. et al. Defining a link with asthma in mice congenitally deficient in eosinophils. Science 305, 1773–1776 (2004).
FitzGerald, J. M. et al. Benralizumab, an anti-interleukin-5 receptor α monoclonal antibody, as add-on treatment for patients with severe, uncontrolled, eosinophilic asthma (CALIMA): a randomised, double-blind, placebo-controlled phase 3 trial. Lancet 388, 2128–2141 (2016).
Rothenberg, M. E. Humanized anti-IL-5 antibody therapy. Cell 165, 509 (2016).
Hansel, T. T. et al. A comprehensive evaluation of nasal and bronchial cytokines and chemokines following experimental rhinovirus infection in allergic asthma: increased interferons (IFN-γ and IFN-λ) and type 2 inflammation (IL-5 and IL-13). EBioMedicine 19, 128–138 (2017).
Papayannopoulos, V. Neutrophil extracellular traps in immunity and disease. Nat. Rev. Immunol. 18, 134–147 (2018).
Lood, C. et al. Neutrophil extracellular traps enriched in oxidized mitochondrial DNA are interferogenic and contribute to lupus-like disease. Nat. Med. 22, 146–153 (2016).
Radermecker, C. et al. Locally instructed CXCR4hi neutrophils trigger environment-driven allergic asthma through the release of neutrophil extracellular traps. Nat. Immunol. 20, 1444–1455 (2019).
Albrengues, J. et al. Neutrophil extracellular traps produced during inflammation awaken dormant cancer cells in mice. Science 361, eaa04227 (2018).
Jimenez-Alcazar, M. et al. Host DNases prevent vascular occlusion by neutrophil extracellular traps. Science 358, 1202–1206 (2017).
Jorch, S. K. & Kubes, P. An emerging role for neutrophil extracellular traps in noninfectious disease. Nat. Med. 23, 279–287 (2017).
Yang, L. et al. DNA of neutrophil extracellular traps promotes cancer metastasis via CCDC25. Nature 583, 133–138 (2020).
Pavlov, V. A., Chavan, S. S. & Tracey, K. J. Molecular and functional neuroscience in immunity. Annu. Rev. Immunol. 36, 783–812 (2018).
Chavan, S. S., Pavlov, V. A. & Tracey, K. J. Mechanisms and therapeutic relevance of neuro-immune communication. Immunity 46, 927–942 (2017).
Wallrapp, A. et al. The neuropeptide NMU amplifies ILC2-driven allergic lung inflammation. Nature 549, 351–356 (2017).
Sui, P. et al. Pulmonary neuroendocrine cells amplify allergic asthma responses. Science 360, eaan8546 (2018).
Simon, D. et al. Eosinophil extracellular DNA traps in skin diseases. J. Allergy Clin. Immunol. 127, 194–199 (2011).
Ueki, S. et al. Eosinophil extracellular DNA trap cell death mediates lytic release of free secretion-competent eosinophil granules in humans. Blood 121, 2074–2083 (2013).
Yousefi, S. et al. Catapult-like release of mitochondrial DNA by eosinophils contributes to antibacterial defense. Nat. Med. 14, 949–953 (2008).
Xiao, Y. et al. Cathepsin C promotes breast cancer lung metastasis by modulating neutrophil infiltration and neutrophil extracellular trap formation. Cancer Cell 39, 423–437 (2021).
Perdomo, J. et al. Neutrophil activation and NETosis are the major drivers of thrombosis in heparin-induced thrombocytopenia. Nat. Commun. 10, 1322 (2019).
Sollberger, G. et al. Gasdermin D plays a vital role in the generation of neutrophil extracellular traps. Sci. Immunol. 3, eaar6689 (2018).
Toussaint, M. et al. Host DNA released by NETosis promotes rhinovirus-induced type-2 allergic asthma exacerbation. Nat. Med. 23, 681–691 (2017).
Thompson, E. A. et al. Role of signal transducer and activator of transcription 1 in murine allergen-induced airway remodeling and exacerbation by carbon nanotubes. Am. J. Respir. Cell Mol. Biol. 53, 625–636 (2015).
McMillan, S. J., Bishop, B., Townsend, M. J., McKenzie, A. N. & Lloyd, C. M. The absence of interleukin 9 does not affect the development of allergen-induced pulmonary inflammation nor airway hyperreactivity. J. Exp. Med. 195, 51–57 (2002).
Lambrecht, B. N., Hammad, H. & Fahy, J. V. The cytokines of asthma. Immunity 50, 975–991 (2019).
Albacker, L. A. et al. Invariant natural killer T cells recognize a fungal glycosphingolipid that can induce airway hyperreactivity. Nat. Med. 19, 1297–1304 (2013).
Corren, J. & Ziegler, S. F. TSLP: from allergy to cancer. Nat. Immunol. 20, 1603–1609 (2019).
Barnes, P. J. Targeting cytokines to treat asthma and chronic obstructive pulmonary disease. Nat. Rev. Immunol. 18, 454–466 (2018).
Urban, C. F. et al. Neutrophil extracellular traps contain calprotectin, a cytosolic protein complex involved in host defense against Candida albicans. PLoS Pathog. 5, e1000639 (2009).
Li, P. et al. PAD4 is essential for antibacterial innate immunity mediated by neutrophil extracellular traps. J. Exp. Med. 207, 1853–1862 (2010).
Mondal, S. & Thompson, P. R. Protein arginine deiminases (PADs): biochemistry and chemical biology of protein citrullination. Acc. Chem. Res. 52, 818–832 (2019).
Papayannopoulos, V., Metzler, K. D., Hakkim, A. & Zychlinsky, A. Neutrophil elastase and myeloperoxidase regulate the formation of neutrophil extracellular traps. J. Cell Biol. 191, 677–691 (2010).
Lizio, M. et al. Gateways to the FANTOM5 promoter level mammalian expression atlas. Genome Biol. 16, 22 (2015).
Veres, T. Z., Rochlitzer, S. & Braun, A. The role of neuro-immune cross-talk in the regulation of inflammation and remodelling in asthma. Pharmacol. Ther. 122, 203–214 (2009).
Yang, C., Gao, J., Du, J., Yang, X. & Jiang, J. Altered neuroendocrine immune responses, a two-sword weapon against traumatic inflammation. Int. J. Biol. Sci. 13, 1409–1419 (2017).
Branchfield, K. et al. Pulmonary neuroendocrine cells function as airway sensors to control lung immune response. Science 351, 707–710 (2016).
Costello, R. W. et al. Localization of eosinophils to airway nerves and effect on neuronal M2 muscarinic receptor function. Am. J. Physiol. 273, L93–L103 (1997).
Kemp, P. J. et al. Airway chemotransduction: from oxygen sensor to cellular effector. Am. J. Respir. Crit. Care Med. 166, S17–S24 (2002).
Gilbert, J. A., Frederick, L. M., Pobst, L. J. & Ames, M. M. Hydrogen peroxide degradation and selective carbidopa-induced cytotoxicity against human tumor lines. Biochem. Pharmacol. 69, 1159–1166 (2005).
Riera, C. E. et al. TRPV1 pain receptors regulate longevity and metabolism by neuropeptide signaling. Cell 157, 1023–1036 (2014).
Margraf, A. et al. The integrin-linked kinase is required for chemokine-triggered high-affinity conformation of the neutrophil β2-integrin LFA-1. Blood 136, 2200–2205 (2020).
Plow, E. F. & Simon, D. I. Implicating ILK in inflammation. Blood 136, 2097–2099 (2020).
Levin, R., Braiman, A. & Priel, Z. Protein kinase C induced calcium influx and sustained enhancement of ciliary beating by extracellular ATP. Cell Calcium 21, 103–113 (1997).
Shibata, K., Morita, K., Kitayama, S., Okamoto, H. & Dohi, T. Ca2+ entry induced by calcium influx factor and its regulation by protein kinase C in rabbit neutrophils. Biochem. Pharmacol. 52, 167–171 (1996).
Navedo, M. F., Amberg, G. C., Votaw, V. S. & Santana, L. F. Constitutively active L-type Ca2+ channels. Proc. Natl Acad. Sci. USA 102, 11112–11117 (2005).
Metzler, K. D., Goosmann, C., Lubojemska, A., Zychlinsky, A. & Papayannopoulos, V. A myeloperoxidase-containing complex regulates neutrophil elastase release and actin dynamics during NETosis. Cell Rep. 8, 883–896 (2014).
Apel, F., Zychlinsky, A. & Kenny, E. F. The role of neutrophil extracellular traps in rheumatic diseases. Nat. Rev. Rheumatol. 14, 467–475 (2018).
Ohnuki, L. E. et al. Differential extraction of eosinophil granule proteins. J. Immunol. Methods 307, 54–61 (2005).
Soragni, A. et al. Toxicity of eosinophil MBP is repressed by intracellular crystallization and promoted by extracellular aggregation. Mol. Cell 57, 1011–1021 (2015).
Mohanty, T. et al. Neutrophil extracellular traps in the central nervous system hinder bacterial clearance during pneumococcal meningitis. Nat. Commun. 10, 1667 (2019).
Busse, W. W. & Lemanske, R. F. Jr Asthma. N. Engl. J. Med. 344, 350–362 (2001).
Demarche, S. et al. Detailed analysis of sputum and systemic inflammation in asthma phenotypes: are paucigranulocytic asthmatics really non-inflammatory? BMC Pulm. Med. 16, 46 (2016).
Gao, W., Han, G. J., Zhu, Y. J., Mao, D. & Hu, H. Clinical characteristics and biomarkers analysis of asthma inflammatory phenotypes. Biomark. Med. 14, 211–222 (2020).
Dworski, R., Simon, H. U., Hoskins, A. & Yousefi, S. Eosinophil and neutrophil extracellular DNA traps in human allergic asthmatic airways. J. Allergy Clin. Immunol. 127, 1260–1266 (2011).
Hwang, C. S. et al. Eosinophil extracellular trap formation is closely associated with disease severity in chronic rhinosinusitis regardless of nasal polyp status. Sci. Rep. 9, 8061 (2019).
Choi, Y. et al. Eosinophil extracellular traps activate type 2 innate lymphoid cells through stimulating airway epithelium in severe asthma. Allergy 75, 95–103 (2020).
Choi, Y. et al. Biological function of eosinophil extracellular traps in patients with severe eosinophilic asthma. Exp. Mol. Med 50, 104 (2018).
Garg, A., Sui, P., Verheyden, J. M., Young, L. R. & Sun, X. Consider the lung as a sensory organ: a tip from pulmonary neuroendocrine cells. Curr. Top. Dev. Biol. 132, 67–89 (2019).
Hewitt, R. J. & Lloyd, C. M. Regulation of immune responses by the airway epithelial cell landscape. Nat. Rev. Immunol. 21, 347–362 (2021).
Sollberger, G., Tilley, D. O. & Zychlinsky, A. Neutrophil extracellular traps: the biology of chromatin externalization. Dev. Cell 44, 542–553 (2018).
Drake, M. G. et al. Eosinophils increase airway sensory nerve density in mice and in human asthma. Sci. Transl. Med. 10, eaar8477 (2018).
Gu, Q., Lim, M. E., Gleich, G. J. & Lee, L. Y. Mechanisms of eosinophil major basic protein-induced hyperexcitability of vagal pulmonary chemosensitive neurons. Am. J. Physiol. Lung Cell. Mol. Physiol. 296, L453–L461 (2009).
Jacoby, D. B., Gleich, G. J. & Fryer, A. D. Human eosinophil major basic protein is an endogenous allosteric antagonist at the inhibitory muscarinic M2 receptor. J. Clin. Invest. 91, 1314–1318 (1993).
Veiga-Fernandes, H. & Mucida, D. Neuro-immune interactions at barrier surfaces. Cell 165, 801–811 (2016).
Le, D. D. et al. Increase of mast cell-nerve association and neuropeptide receptor expression on mast cells in perennial allergic rhinitis. Neuroimmunomodulation 23, 261–270 (2016).
Hagiyama, M. et al. Enhanced nerve-mast cell interaction by a neuronal short isoform of cell adhesion molecule-1. J. Immunol. 186, 5983–5992 (2011).
Sawatzky, D. A. et al. Eosinophil adhesion to cholinergic nerves via ICAM-1 and VCAM-1 and associated eosinophil degranulation. Am. J. Physiol. Lung Cell. Mol. Physiol. 282, L1279–L1288 (2002).
Thornton, M. A. et al. Eosinophil recruitment to nasal nerves after allergen challenge in allergic rhinitis. Clin. Immunol. 147, 50–57 (2013).
Fryer, A. D. & Wills-Karp, M. Dysfunction of M2-muscarinic receptors in pulmonary parasympathetic nerves after antigen challenge. J. Appl. Physiol. 71, 2255–2261 (1991).
Ueki, S. et al. Charcot–Leyden crystal formation is closely associated with eosinophil extracellular trap cell death. Blood 132, 2183–2187 (2018).
Drake, M. G. et al. Eosinophil and airway nerve interactions in asthma. J. Leukoc. Biol. 104, 61–67 (2018).
Kenny, E. F. et al. Diverse stimuli engage different neutrophil extracellular trap pathways. eLife 6, e24437 (2017).
Yost, C. C. et al. Neonatal NET-inhibitory factor and related peptides inhibit neutrophil extracellular trap formation. J. Clin. Invest. 126, 3783–3798 (2016).
Swanney, M. P. et al. Using the lower limit of normal for the FEV1/FVC ratio reduces the misclassification of airway obstruction. Thorax 63, 1046–1051 (2008).
Bruns, S. et al. Production of extracellular traps against Aspergillus fumigatus in vitro and in infected lung tissue is dependent on invading neutrophils and influenced by hydrophobin RodA. PLoS Pathog. 6, e1000873 (2010).
Radermecker, C. et al. Locally instructed CXCR4 neutrophils trigger environment-driven allergic asthma through the release of neutrophil extracellular traps. Nat. Immunol. 20, 1444–1455 (2019).
Lu, Y. et al. Complement signals determine opposite effects of B cells in chemotherapy-induced immunity. Cell 180, 1081–1097 (2020).
Lehrman, E. K. et al. CD47 protects synapses from excess microglia-mediated pruning during development. Neuron 100, 120–134 (2018).
Diorio, C. et al. Multisystem inflammatory syndrome in children and COVID-19 are distinct presentations of SARS-CoV-2. J. Clin. Invest. 130, 5967–5975 (2020).
Sofoluwe, A., Bacchetta, M., Badaoui, M., Kwak, B. R. & Chanson, M. ATP amplifies NADPH-dependent and -independent neutrophil extracellular trap formation. Sci. Rep. 9, 16556 (2019).
Cunha, A. A. et al. Extracellular DNA traps in bronchoalveolar fluid from a murine eosinophilic pulmonary response. Allergy 69, 1696–1700 (2014).
Genschmer, K. R. et al. Activated PMN exosomes: pathogenic entities causing matrix destruction and disease in the lung. Cell 176, 113–126 (2019).
Machiels, B. et al. A gammaherpesvirus provides protection against allergic asthma by inducing the replacement of resident alveolar macrophages with regulatory monocytes. Nat. Immunol. 18, 1310–1320 (2017).
Kabata, H. et al. Thymic stromal lymphopoietin induces corticosteroid resistance in natural helper cells during airway inflammation. Nat. Commun. 4, 2675 (2013).
Cortez, M. A. et al. Infantile spasms and Down syndrome: a new animal model. Pediatr. Res. 65, 499–503 (2009).
Pinho-Ribeiro, F. A. et al. Blocking neuronal signaling to immune cells treats streptococcal invasive infection. Cell 173, 1083–1097 (2018).
Germic, N., Stojkov, D., Oberson, K., Yousefi, S. & Simon, H. U. Neither eosinophils nor neutrophils require ATG5-dependent autophagy for extracellular DNA trap formation. Immunology 152, 517–525 (2017).
Reichman, H., Rozenberg, P. & Munitz, A. Mouse eosinophils: identification, isolation, and functional analysis. Curr. Protoc. Immunol. 119, 14.43.1–14.43.22 (2017).
Dyer, K. D., Garcia-Crespo, K. E., Percopo, C. M., Sturm, E. M. & Rosenberg, H. F. Protocols for identifying, enumerating, and assessing mouse eosinophils. Methods Mol. Biol. 1032, 59–77 (2013).
Rozman, S. et al. The generation of neutrophils in the bone marrow is controlled by autophagy. Cell Death Differ. 22, 445–456 (2015).
Hasenberg, A. et al. Catchup: a mouse model for imaging-based tracking and modulation of neutrophil granulocytes. Nat. Methods 12, 445–452 (2015).
Petersen, B. C., Budelsky, A. L., Baptist, A. P., Schaller, M. A. & Lukacs, N. W. Interleukin-25 induces type 2 cytokine production in a steroid-resistant interleukin-17RB+ myeloid population that exacerbates asthmatic pathology. Nat. Med. 18, 751–758 (2012).
Islam, M. N. et al. Mitochondrial transfer from bone-marrow-derived stromal cells to pulmonary alveoli protects against acute lung injury. Nat. Med. 18, 759–765 (2012).
Wang, N. et al. Neutrophil extracellular traps induced by VP1 contribute to pulmonary edema during EV71 infection. Cell Death Discov. 5, 111 (2019).
Masuda, S. et al. Measurement of NET formation in vitro and in vivo by flow cytometry. Cytometry A 91, 822–829 (2017).
Tucker, S. L., Sarr, D. & Rada, B. Neutrophil extracellular traps are present in the airways of ENaC-overexpressing mice with cystic fibrosis-like lung disease. BMC Immunol. 22, 7 (2021).
Chevallet, M., Luche, S. & Rabilloud, T. Silver staining of proteins in polyacrylamide gels. Nat. Protoc. 1, 1852–1858 (2006).
Berndt, N., Bergmann, R., Arndt, C., Koristka, S. & Bachmann, M. Silver staining techniques of polyacrylamide gels. Methods Mol. Biol. 1853, 47–52 (2018).
Lim, C. H. et al. Thrombin and plasmin alter the proteome of neutrophil extracellular traps. Front. Immunol. 9, 1554 (2018).
Fuseya, Y. et al. The HOIL-1L ligase modulates immune signalling and cell death via monoubiquitination of LUBAC. Nat. Cell Biol. 22, 663–673 (2020).
Esposito, M. et al. TGF-β-induced DACT1 biomolecular condensates repress Wnt signalling to promote bone metastasis. Nat. Cell Biol. 23, 257–267 (2021).
Sabatel, C. et al. Exposure to bacterial CpG DNA protects from airway allergic inflammation by expanding regulatory lung interstitial macrophages. Immunity 46, 457–473 (2017).
Dong, C. C. et al. Exposure of brown Norway rats to diesel exhaust particles prior to ovalbumin (OVA) sensitization elicits IgE adjuvant activity but attenuates OVA-induced airway inflammation. Toxicol. Sci. 88, 150–160 (2005).
Noges, L. E., White, J., Cambier, J. C., Kappler, J. W. & Marrack, P. Contamination of DNase preparations confounds analysis of the role of DNA in alum-adjuvanted vaccines. J. Immunol. 197, 1221–1230 (2016).
Altarejos, J. Y. et al. The Creb1 coactivator Crtc1 is required for energy balance and fertility. Nat. Med. 14, 1112–1117 (2008).
Ch’ng, T. H. et al. Activity-dependent transport of the transcriptional coactivator CRTC1 from synapse to nucleus. Cell 150, 207–221 (2012).
Zhao, Q. et al. Targeting mitochondria-located circRNA SCAR alleviates NASH via reducing mROS Output. Cell 183, 76–93 (2020).
Huang, D. et al. NKILA lncRNA promotes tumor immune evasion by sensitizing T cells to activation-induced cell death. Nat. Immunol. 19, 1112–1125 (2018).
Chen, J., Ruan, J. W., Ye, J. X., Cheng, Z. W. & Chen, D. Z. Removal of gaseous tetrahydrofuran via a three-phase airlift bioreactor loaded with immobilized cells of GFP-tagged Pseudomonas oleovorans GDT4. Chemosphere 258, 127148 (2020).
Han, J. et al. Effects of all-trans retinoic acid on signal pathway of cyclooxygenase-2 and Smad3 in transforming growth factor-β-stimulated glomerular mesangial cells. Exp. Biol. Med. 239, 272–283 (2014).
Revelo, N. H. & Rizzoli, S. O. The membrane marker mCLING reveals the molecular composition of trafficking organelles. Curr. Protoc. Neurosci. 74, 2.25.21–22.25.21 (2016).
Slifman, N. R., Loegering, D. A., McKean, D. J. & Gleich, G. J. Ribonuclease activity associated with human eosinophil-derived neurotoxin and eosinophil cationic protein. J. Immunol. 137, 2913–2917 (1986).
Calcraft, P. J. et al. NAADP mobilizes calcium from acidic organelles through two-pore channels. Nature 459, 596–600 (2009).
Bhogaraju, S. et al. Inhibition of bacterial ubiquitin ligases by SidJ-calmodulin catalysed glutamylation. Nature 572, 382–386 (2019).
Lembrechts, R. et al. Functional expression of the multimodal extracellular calcium-sensing receptor in pulmonary neuroendocrine cells. J. Cell Sci. 126, 4490–4501 (2013).
Pintelon, I. et al. Selective visualisation of neuroepithelial bodies in vibratome slices of living lung by 4-Di-2-ASP in various animal species. Cell Tissue Res. 321, 21–33 (2005).
Chen, F. et al. Extracellular vesicle-packaged HIF-1α-stabilizing lncRNA from tumour-associated macrophages regulates aerobic glycolysis of breast cancer cells. Nat. Cell Biol. 21, 498–510 (2019).
Ma, J. et al. iProX: an integrated proteome resource. Nucleic Acids Res. 47, D1211–D1217 (2019).
Acknowledgements
This work was supported by grants from the National Key Research and Development Program of China (grant no. 2017YFA0106300 (S.S.)), Natural Science Foundation of China (grant nos 91942309 (S.S.), 92057210 (S.S.), 82071804 (S.J.) and 82002780 (Y.L.)), Science and Technology Program of Guangzhou (grant no. 202103000070 (S.S.)), Natural Science Foundation of Guangdong Province (grant no. 2020A1515010031 (Y.L.)) and Tencent Charity Foundation of China (S.J.).
Author information
Authors and Affiliations
Contributions
S.S. conceived the study, designed the experiments and wrote the manuscript. S.J. and Y.H. provided the clinical samples. Y.L., Y.H., Jiang Li, J.H. and L.Z. performed the experiments, analysed the data and wrote the manuscript. J.F., Jiaqian Li., Q.X., Q.Z. and L.H. performed the experiments and analysed the data.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Cell Biology thanks Venizelos Papayannopoulos and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 EETs are correlated with asthma progression.
a, Quantification of Fig. 1d. b, Levels of EETs were evaluated by the SYTOX Green, DAPI and ECP staining in BALF of mice immunized without or with OVA. c, Quantification of Fig. 1f. d, Quantification of SYTOX Green signals in ECP+ or MPO+ cells in BALF of mice immunized with OVA in Fig. 1e. a-d, Mean ± s.d.; n = 6 mice per group, representative of two independent experiments (a, c, d); Data are pooled from 3 independent experiments with 6-8 mice each (b); compared by two-tailed unpaired Student’s t-test.
Extended Data Fig. 2 Generation and components of EETs.
a, Venn diagram of overlapped proteins from three donors in Fig. 2g. The numbers of overlapped and unique proteins are shown in the corresponding area.
Extended Data Fig. 3 DNase treatment ameliorates asthma in mice.
a, Quantification of Fig. 3f. b, Quantification of Fig. 3h. c, Protease activity of DNase I at indicated concentrations was examined by the protease assay. Trypsin was used as a positive control (n = 3). d-i, Wild-type mice were sensitized and challenged with PBS or OVA. OVA-sensitized mice were treated intratracheally with DNase before OVA challenge. d, Representative immunofluorescence images of cytokeratin (CK), TUNEL and DAPI staining in sections of lungs harvested at baseline (day 0) and 15 (day 15) and 17 (day 17) days after the first OVA sensitization. The arrow indicates CK+ TUNEL+ cells. Scale bar, 20 μm. e, Quantification of d. f, The area (%) of ECP+ SYTOX Green+ EETs in BALF smears was quantified at baseline (day 0) and 15 (day 15) and 17 (day 17) days after the first OVA sensitization. g, Eosinophils in the BALF at baseline (day 0) and 15 (day 15) and 17 (day 17) days after the first OVA sensitization were evaluated by flow cytometric analysis. Quantification are shown. h, i, F4/80−MHC-II+CD11c+ dendritic cells (DCs) in the BALF at baseline (day 0) and 15 (day 15) and 17 (day 17) days after the first OVA sensitization were evaluated by flow cytometric analysis. Quantification (h) and representative plots (i) are shown. a, b, e-h, Mean ± s.d.; n = 6 mice per group, representative of two independent experiments; compared by one-way ANOVA with Tukey’s multiple comparisons test (a, b, f-h) and two-way ANOVA with Sidak’s multiple comparisons test (e).
Extended Data Fig. 4 EETs amplify allergic asthma responses.
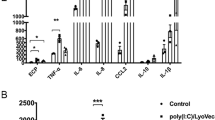
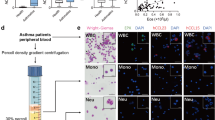
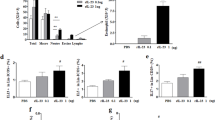
a, Representative immunoblots of PAD4 and GAPDH in human eosinophils, neutrophils, monocytes, T cell, B cells and NK cells isolated from peripheral blood of healthy donors (n = 3). b, Representative immunofluorescence images of MPO, ECP, PAD4 and DAPI staining in human neutrophils and eosinophils isolated from peripheral blood of healthy donors. Scale bar, 50 μm. c, Quantification of Fig. 4b. d, Quantification of Fig. 4c. e, f, Wild-type and PAD4−/− mice were sensitized and challenged with PBS or OVA. PAD4−/− mice were adoptively transferred with 2×105 eosinophils or neutrophils isolated from wild-type or PAD4−/− mice. e, Quantification of Fig. 4e. f, Levels of IL-25, IL-33 and TSLP in BALF were evaluated by ELISA. c-f, Mean ± s.d.; n = 5 independent experiments (c, d); n = 6 mice per group, representative of two independent experiments (e, f); compared by one-way ANOVA with Tukey’s multiple comparisons test.
Extended Data Fig. 5 EETs aggravate allergic inflammation by activating PNECs.
a, Expression profiles of CCDC25 in different tissues from the FANTOM5 dataset. b, ECP+ H3cit+DAPI+ spots (0.1-μm-diameter) in lung sections of OVA-immunized mice were defined as EETs as shown in Fig. 5a. The density of EETs (number of spots per mm2) within 50 μm of PNECs, randomly selected blood vessels and airway walls (from epithelium to serosa, excluding the cartilage and 50-μm area surrounding PNECs) was quantified. c, CCDC25 expression level of CGRP+ and CGRP− cells in mouse lung sections was quantified by immunofluorescence as shown in Fig. 5b. d, Representative immunoblots of CCDC25 of lung homogenate from C57BL/6 mice challenged with PBS or OVA (n = 3). e-g, C57BL/6 mice immunized with OVA were treated without or with BIBN4096 or CGP35348. e, The inflammatory score quantified from H&E staining of lung sections. f, Levels of indicated type 2 cytokines in BALF evaluated by ELISA. g, Percentages of PAS-stained goblet cells in total epithelial cells in lung sections. h, The relative expression level of CGRP in H146 cells with indicated treatment was measured by qRT-PCR. i, Quantification of Fig. 5e. j, The relative level of intracellular Fluo-4 immunofluorescence intensity in live lung slices with indicated treatment was quantified at indicated time point. k, Quantification of Fig. 5h. b, c, e-k, Mean ± s.d.; n = 6 mice per group, representative of two independent experiments (b, c, e-g, k); n = 4 (h, i) or 3 (j) independent experiments; compared by two-tailed unpaired Student’s t-test (c) and one-way ANOVA with Tukey’s multiple comparisons test (b, e-k).
Extended Data Fig. 6 EETs activate PNECs via CCDC25.
a, Representative immunoblots for CCDC25 in H146 cells with indicated treatment (n = 3). b, Representative immunoblots of CCDC25 in H146 cells that were transduced with sgControl or sgRNAs against CCDC25 (n = 3). c, Quantification of Fig. 6c. d-g, Wild-type and CCDC25−/− mice were immunized with OVA. d, Quantification of Fig. 6e. e, Quantification of indicated cells in immunofluorescence images of lung sections. f, The area (%) of EETs in BALF smear at baseline (day 0) and 15 (day 15) and 17 (day 17) days after the first OVA sensitization was quantified by immunofluorescence staining. g, The number of eosinophils in the BALF at indicated time point was quantified by flow cytometry. h, CRTC1 was silenced by shRNA in H146 cells. The efficiency was evaluated by western blotting (n = 3). i, Representative immunoblots of PKCα levels in membrane (Memb) and cytoplasmic (Cyto) fractions of H146 cells with indicated treatments (n = 3). j, k, ILK (j) and PKCα (k) were silenced by shRNAs in H146 cells. The efficiency was evaluated by western blotting (n = 3). l, m, H146 cells transduced with shRNAs against PKCα were treated with EETs. l, Cytosolic Ca2+ mobilization in H146 cells was examined using Fura-2/AM ratiometric fluorescence measurements. Fluorescence curves (left) and peak responses (right) are shown. Arrow indicates the time of stimulation. m, Representative immunoblots of CRTC1 in nuclear (Nuc) and cytoplasmic (Cyto) fractions of H146 cells (n = 3). c-g, l, Mean ± s.d.; n = 6 mice per group, representative of two independent experiments (d-g); n = 5 (c) or 4 (l) independent experiments; compared by one-way ANOVA with Tukey’s multiple comparisons test (d-g, l) and two-way ANOVA with Sidak’s multiple comparisons test (c).
Extended Data Fig. 7 EPX potentiates the interaction between EETs and PNECs.
a-c, Wild-type, EPX−/− and MBP−/− mice were sensitized and challenged with PBS or OVA. a, Quantification of Fig. 7d. b, Levels of indicated type 2 cytokines in BALF were evaluated by ELISA. c, The relative expression levels of CGRP in lung homogenates were examined by qRT-PCR. d, The area (%) of ECP+ SYTOX Green+ and EPX+ SYTOX Green+ signals in BALF smears as shown in Fig. 7f. e-f, PAD4−/− mice were adoptively transferred with eosinophils isolated from wild-type mice or EPX−/− mice. e, Percentages of PAS-stained cells in total epithelial cells in lung sections. f. The inflammatory score quantified from the H&E staining of lung sections. a-f, Mean ± s.d.; n = 6 (e, f) or 10 (a-d) mice per group, representative of two independent experiments; compared by one-way ANOVA with Tukey’s multiple comparisons test.
Supplementary information
Supplementary Table 1
Supplementary Tables 1–4.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 2
Unprocessed western blots and/or gels.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 4
Unprocessed western blots and/or gels.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Statistical source data.
Source Data Fig. 6
Unprocessed western blots and/or gels.
Source Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 4
Unprocessed western blots and/or gels.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 5
Unprocessed western blots and/or gels.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 6
Unprocessed western blots and/or gels.
Source Data Extended Data Fig. 7
Statistical source data.
Rights and permissions
About this article
Cite this article
Lu, Y., Huang, Y., Li, J. et al. Eosinophil extracellular traps drive asthma progression through neuro-immune signals. Nat Cell Biol 23, 1060–1072 (2021). https://doi.org/10.1038/s41556-021-00762-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-021-00762-2
This article is cited by
-
Eosinophils promote CD8+ T cell memory generation to potentiate anti-bacterial immunity
Signal Transduction and Targeted Therapy (2024)
-
Eosinophil extracellular traps in asthma: implications for pathogenesis and therapy
Respiratory Research (2023)
-
New genetic and epigenetic insights into the chemokine system: the latest discoveries aiding progression toward precision medicine
Cellular & Molecular Immunology (2023)
-
Neferine Attenuates HDM-Induced Allergic Inflammation by Inhibiting the Activation of Dendritic Cell
Inflammation (2023)
-
Chromatin-Associated Molecular Patterns (CAMPs) in sepsis
Cell Death & Disease (2022)