Abstract
Cancers adapt to increasingly potent targeted therapies by reprogramming their phenotype. Here we investigated such a phenomenon in prostate cancer, in which tumours can escape epithelial lineage confinement and transition to a high-plasticity state as an adaptive response to potent androgen receptor (AR) antagonism. We found that AR activity can be maintained as tumours adopt alternative lineage identities, with changes in chromatin architecture guiding AR transcriptional rerouting. The epigenetic regulator enhancer of zeste homologue 2 (EZH2) co-occupies the reprogrammed AR cistrome to transcriptionally modulate stem cell and neuronal gene networks—granting privileges associated with both fates. This function of EZH2 was associated with T350 phosphorylation and establishment of a non-canonical polycomb subcomplex. Our study provides mechanistic insights into the plasticity of the lineage-infidelity state governed by AR reprogramming that enabled us to redirect cell fate by modulating EZH2 and AR, highlighting the clinical potential of reversing resistance phenotypes.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
RNA-seq, ChIP-seq and ATAC-seq data have been deposited in the GEO under the accession GSE138460. Previously published sequencing data were re-analysed here: publicly available data from the SU2C/PCF-West Coast Dream Team cohort was obtained from ref. 5, the SU2C cohort from GitHub (https://github.com/cBioPortal/datahub/tree/master/public/prad_su2c_2019)37, the University of Washington Rapid Autopsy cohort6 from GEO accession GSE126078, the Beltran 2016 cohort4, and the CALGB 90203 cohort60. The SKO/DKO/TKO prostate cancer GEMM dataset7 was downloaded from GEO accession GSE90891 and the NPp53 GEMM13 was downloaded from GEO accession GSE92721. Source data are provided with this paper.
References
Davies, A. H., Beltran, H. & Zoubeidi, A. Cellular plasticity and the neuroendocrine phenotype in prostate cancer. Nat. Rev. Urol. 15, 271–286 (2018).
Le Magnen, C., Shen, M. M. & Abate-Shen, C. Lineage plasticity in cancer progression and treatment. Annu. Rev. Cancer Biol. 2, 271–289 (2018).
Boumahdi, S. & de Sauvage, F. J. The great escape: tumour cell plasticity in resistance to targeted therapy. Nat. Rev. Drug Discov. 19, 39–56 (2020).
Beltran, H. et al. Divergent clonal evolution of castration-resistant neuroendocrine prostate cancer. Nat. Med. 22, 298–305 (2016).
Aggarwal, R. et al. Clinical and genomic characterization of treatment-emergent small-cell neuroendocrine prostate cancer: a multi-institutional prospective study. J. Clin. Oncol. 36, 2492–2503 (2018).
Labrecque, M. P. et al. Molecular profiling stratifies diverse phenotypes of treatment-refractory metastatic castration-resistant prostate cancer. J. Clin. Invest. 129, 4492–4505 (2019).
Ku, S. Y. et al. Rb1 and Trp53 cooperate to suppress prostate cancer lineage plasticity, metastasis, and antiandrogen resistance. Science 355, 78–83 (2017).
Mu, P. et al. SOX2 promotes lineage plasticity and antiandrogen resistance in TP53- and RB1-deficient prostate cancer. Science 355, 84–88 (2017).
Beltran, H. et al. Molecular characterization of neuroendocrine prostate cancer and identification of new drug targets. Cancer Discov. 1, 487–495 (2011).
Aparicio, A. M. et al. Platinum-based chemotherapy for variant castrate-resistant prostate cancer. Clin. Cancer Res. 19, 3621–3630 (2013).
Tzelepi, V. et al. Modeling a lethal prostate cancer variant with small-cell carcinoma features. Clin. Cancer Res. 18, 666–677 (2012).
Aparicio, A. M. et al. Combined tumor suppressor defects characterize clinically defined aggressive variant prostate cancers. Clin. Cancer Res. 22, 1520–1530 (2016).
Zou, M. et al. Transdifferentiation as a mechanism of treatment resistance in a mouse model of castration-resistant prostate cancer. Cancer Discov. 7, 736–749 (2017).
Smith, B. A. et al. A basal stem cell signature identifies aggressive prostate cancer phenotypes. Proc. Natl Acad. Sci. USA 112, E6544–E6552 (2015).
Smith, B. A. et al. A human adult stem cell signature marks aggressive variants across epithelial cancers. Cell Rep. 24, 3353–3366 e3355 (2018).
Williams, S. G. et al. Immune molecular profiling of a multiresistant primary prostate cancer with a neuroendocrine-like phenotype: a case report. BMC Urol. 20, 171 (2020).
Alumkal, J. J. et al. Transcriptional profiling identifies an androgen receptor activity-low, stemness program associated with enzalutamide resistance. Proc. Natl Acad. Sci. USA 117, 12315–12323 (2020).
Ge, R. et al. Epigenetic modulations and lineage plasticity in advanced prostate cancer. Ann. Oncol. 31, 470–479 (2020).
Zhang, Z. et al. Loss of CHD1 promotes heterogeneous mechanisms of resistance to AR-targeted therapy via chromatin dysregulation. Cancer Cell 37, 584–598 e511 (2020).
Clermont, P. L. et al. Polycomb-mediated silencing in neuroendocrine prostate cancer. Clin. Epigenetics 7, 40 (2015).
Kim, J. et al. Polycomb- and methylation-independent roles of EZH2 as a transcription activator. Cell Rep. 25, 2808–2820 e2804 (2018).
Xu, K. et al. EZH2 oncogenic activity in castration-resistant prostate cancer cells is Polycomb-independent. Science 338, 1465–1469 (2012).
Bluemn, E. G. et al. Androgen receptor pathway-independent prostate cancer is sustained through FGF signaling. Cancer Cell 32, 474–489(2017).
Bishop, J. L. et al. The master neural transcription factor BRN2 is an androgen receptor-suppressed driver of neuroendocrine differentiation in prostate cancer. Cancer Discov. 7, 54–71 (2017).
Korpal, M. et al. An F876L mutation in androgen receptor confers genetic and phenotypic resistance to MDV3100 (enzalutamide). Cancer Discov. 3, 1030–1043 (2013).
Nyquist, M. D. et al. Combined TP53 and RB1 loss promotes prostate cancer resistance to a spectrum of therapeutics and confers vulnerability to replication stress. Cell Rep. 31, 107669 (2020).
Burger, P. E. et al. High aldehyde dehydrogenase activity: a novel functional marker of murine prostate stem/progenitor cells. Stem Cells 27, 2220–2228 (2009).
Pomerantz, M. M. et al. Prostate cancer reactivates developmental epigenomic programs during metastatic progression. Nat. Genet. 52, 790–799 (2020).
Park, J. W. et al. Reprogramming normal human epithelial tissues to a common, lethal neuroendocrine cancer lineage. Science 362, 91–95 (2018).
Xu, J. et al. Developmental control of polycomb subunit composition by GATA factors mediates a switch to non-canonical functions. Mol. Cell 57, 304–316 (2015).
Liu, P., Chen, M., Liu, Y., Qi, L. S. & Ding, S. CRISPR-based chromatin remodeling of the endogenous Oct4 or Sox2 locus enables reprogramming to pluripotency. Cell Stem Cell 22, 252–261 (2018).
Park, N. I. et al. ASCL1 reorganizes chromatin to direct neuronal fate and suppress tumorigenicity of glioblastoma stem cells. Cell Stem Cell 21, 209–224 e207 (2017).
Wan, L. et al. Phosphorylation of EZH2 by AMPK suppresses PRC2 methyltransferase activity and oncogenic function. Mol. Cell 69, 279–291 (2018).
Wei, Y. et al. CDK1-dependent phosphorylation of EZH2 suppresses methylation of H3K27 and promotes osteogenic differentiation of human mesenchymal stem cells. Nat. Cell Biol. 13, 87–94 (2011).
Kaneko, S. et al. Phosphorylation of the PRC2 component Ezh2 is cell cycle-regulated and up-regulates its binding to ncRNA. Genes Dev. 24, 2615–2620 (2010).
Chen, S. et al. Cyclin-dependent kinases regulate epigenetic gene silencing through phosphorylation of EZH2. Nat. Cell Biol. 12, 1108–1114 (2010).
Abida, W. et al. Genomic correlates of clinical outcome in advanced prostate cancer. Proc. Natl Acad. Sci. USA 116, 11428–11436 (2019).
Kim, K. H. et al. SWI/SNF-mutant cancers depend on catalytic and non-catalytic activity of EZH2. Nat. Med. 21, 1491–1496 (2015).
Wong, D. J. et al. Module map of stem cell genes guides creation of epithelial cancer stem cells. Cell Stem Cell 2, 333–344 (2008).
Ben-Porath, I. et al. An embryonic stem cell-like gene expression signature in poorly differentiated aggressive human tumors. Nat. Genet. 40, 499–507 (2008).
Chen, S., Xu, Y., Yuan, X., Bubley, G. J. & Balk, S. P. Androgen receptor phosphorylation and stabilization in prostate cancer by cyclin-dependent kinase 1. Proc. Natl Acad. Sci. USA 103, 15969–15974 (2006).
Wang, X. Q. et al. CDK1–PDK1–PI3K/Akt signaling pathway regulates embryonic and induced pluripotency. Cell Death Differ. 24, 38–48 (2017).
Zeng, X., Chen, S. & Huang, H. Phosphorylation of EZH2 by CDK1 and CDK2: a possible regulatory mechanism of transmission of the H3K27me3 epigenetic mark through cell divisions. Cell Cycle 10, 579–583 (2011).
McKay, R. R. et al. Evaluation of intense androgen deprivation before prostatectomy: A randomized phase II trial of enzalutamide and leuprolide with or without abiraterone. J. Clin. Oncol. 37, 923–931 (2019).
Hussain, M. et al. Enzalutamide in men with nonmetastatic, castration-resistant prostate cancer. N. Engl. J. Med. 378, 2465–2474 (2018).
Bai, Y. et al. Inhibition of enhancer of zeste homolog 2 (EZH2) overcomes enzalutamide resistance in castration-resistant prostate cancer. J. Biol. Chem. 294, 9911–9923 (2019).
Xiao, L. et al. Epigenetic reprogramming with antisense oligonucleotides enhances the effectiveness of androgen receptor inhibition in castration-resistant prostate cancer. Cancer Res. 78, 5731–5740 (2018).
Puca, L. et al. Patient derived organoids to model rare prostate cancer phenotypes. Nat. Commun. 9, 2404 (2018).
Kuruma, H. et al. A novel antiandrogen, compound 30, suppresses castration-resistant and MDV3100-resistant prostate cancer growth in vitro and in vivo. Mol. Cancer Ther. 12, 567–576 (2013).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Trapnell, C. et al. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat. Protoc. 7, 562–578 (2012).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Buenrostro, J. D. et al. ATAC-seq: a method for assaying chromatin accessibility genome-wide. Curr. Protoc. Mol. Biol. 109, 21.29.1–21.29.9 (2015).
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics 26, 589–595 (2010).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Yu, G., Wang, L. G. & He, Q. Y. ChIPseeker: an R/Bioconductor package for ChIP peak annotation, comparison and visualization. Bioinformatics 31, 2382–2383 (2015).
Ramirez, F., Dundar, F., Diehl, S., Gruning, B. A. & Manke, T. deepTools: a flexible platform for exploring deep-sequencing data. Nucleic Acids Res. 42, W187–W191 (2014).
Kumar, V. et al. Uniform, optimal signal processing of mapped deep-sequencing data. Nat. Biotechnol. 31, 615–622 (2013).
Beltran, H. et al. Impact of therapy on genomics and transcriptomics in high-risk prostate cancer treated with neoadjuvant docetaxel and androgen deprivation therapy. Clin. Cancer Res. 23, 6802–6811 (2017).
Ertel, A. et al. RB-pathway disruption in breast cancer: differential association with disease subtypes, disease-specific prognosis and therapeutic response. Cell Cycle 9, 4153–4163 (2010).
Acknowledgements
This research was supported by funding from the Terry Fox Research Institute New Frontiers Program (F15-05505 to A.Z.), the Prostate Cancer Foundation (to A.Z. and H.B.), the Canadian Institutes of Health Research (399802 to A.D. and A.Z.), Dutch Cancer Society/Alpe d’HuZes (10084 to W.Z.), and a Prostate Cancer Foundation Young Investigator Award (to A.D.). We thank members of the Zoubeidi laboratory for valuable inputs in designing this research, the Molecular Pathology Core (Vancouver Prostate Centre) for assisting with tissue processing and immunohistochemistry, the Flow Cytometry Core Facility (BC Cancer Research Centre) for cell sorting, the Genomic Analysis Core (Vancouver Prostate Centre) for ChIP-seq and ATAC-seq processing, the Genomics Core Facility (Netherlands Cancer Institute) for ChIP-seq analyses on prostate tumour specimens, the Core Facility for Molecular Pathology and Biobanking (Netherlands Cancer Institute) for tissue processing and haematoxylin and eosin analyses using Slide Score, the Biomedical Research Centre Sequencing Core (University of British Columbia) for RNA-seq, and O. Witte and J. Lee (UCLA) for providing LASCPC-1 cells. L.S. is supported by a Principal Cancer Research Fellowships awarded by Cancer Council’s Beat Cancer project on behalf of its donors, the State Government through the Department of Health and the Australian Government through the Medical Research Future Fund.
Author information
Authors and Affiliations
Contributions
A.D. and A.Z. conceptualized and designed the study. S.N., T.N. and A.T generated cell lines and performed functionalization experiments. D. Ganguli, F.K., M.A., F.H. and H.H.H. performed peak calling, quality control and visualization for ChIP-seq and/or ATAC-seq. T.N., D.T. and M.K performed RIME and ChIP experiments. S. Kim and C.B. characterized cell lines. S. Kumar performed PLA. O.S. and J.B. generated xenografts. M.E.O. analysed RIME. S.L., M.H., S.S., H.v.d.P., A.M.B. and W.Z. procured and/or analysed clinical specimens from the DARANA trial. L.F. reviewed pathology and scored all immunohistochemistry staining. H.H. provided the pEZH2-T350 antibody and EZH2 T350 mutant constructs for initial exploratory studies. D. Goodrich shared GEMMs. M.G. and H.B. provided access to clinical specimens. D. Goodrich, W.T. and F.Y.F. reviewed the manuscript. L.S. and A.Z. edited the manuscript. A.D. performed all other experiments, generated figures, and wrote the manuscript. All authors provided intellectual input and vetted and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Cell Biology thanks Li Xin and the other, anonymous, reviewers for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Characterization of the enzalutamide resistance model.
(a) Schematic depicting generation of the ENZ-driven resistance model. (b) Tumour volume and serum PSA of PSA+ (T49) and PSA- (T42) ENZ-resistant tumours at time following ENZ treatment. (c) Immunoblot of AR and PSA in cell lines derived from CRPC (16DCRPC) and ENZ-resistant AR-driven (49FENZR) and lineage plastic (42DENZR, 42FENZR) tumours. (d) Frequency of activating AR F876L mutation in CRPC and ENZ-resistant cell lines. (e) PCA of the global transcriptome in the indicated cell lines. (f) Significantly enriched (p < 0.05) gene ontology pathways in CRPC and ENZ-resistant cell lines ranked by normalized enrichment score. The following keywords were used to define functional categories: Plasticity (morphogenesis, plasticity, differentiation, mesenchymal); Neuronal (cerebral, axon, synap, neuro); Migration (chemotaxis, migration); Hormone (androgen, hormone); Translation. (g) Expression of ‘Core 9’ embryonic stem cell genes and neuronal lineage markers in the indicated cell lines, reported relative to LNCaP. (h) ASC scores in the indicated prostate cancer cell lines (n = 3) and AR+/NE+ and AR-/NE+ patient tumours from Aggarwal et al. Statistical analysis was performed using a two-tailed unpaired t-test. Error bars represent mean ± SD. (i) Transcript expression of RB1 and TP53 in cell lines and SU2C patient samples with wild-type RB1/TP53 (SU2CWT) or biallelic RB1 and TP53 deletion (SU2CRB1/TP53). An RB1/TP53 signature score was applied to cell lines and tumours (higher score indicative of functional RB1/TP53 loss). (j) Immunoblot of pRB1-S780 in the indicated cell lines. (k) Partial least squares discriminant analysis (PLS-DA) of global transcriptome separates AR+/NE+ and AR-/NE+ patient tumours from the Labrecque et al cohort (AR+/NE+, n = 11; AR-/NE+, n = 11; GEO: GSE126078). RNA-seq data from cell lines were projected on the PLS-DA plot. Probability ellipse=95% confidence. (l) Spheroid formation quantified at 8 days following seeding of single cells from the indicated cell lines (mean ± SD; two-tailed unpaired t-test, n = 3). Phase contrast images are shown. Scale bar, 100 μm. (m) Flow cytometry plots of CD44 and NCAM1 cell surface expression (top) and ALDH activity (bottom) in the indicated cell lines (mean ± SD). Diethylaminobenzaldehyde (DEAB) is as a control for background fluorescence.
Extended Data Fig. 2 ENZ-resistant lineage plastic tumours exhibit a distinct AR cistrome.
(a) Principal component analysis (PCA) reveals distinct AR binding patterns across prostate states. Each dot represents the genome-wide AR cistrome in an individual clinical specimen (6 normal prostate epithelial, 18 primary prostate cancer tumours, 15 PDX tumours derived from patient mCRPC; GEO:GSE130408) or indicated cell line. (b) Gene ontology (GO) pathways enriched surrounding AR binding sites in ENZ-resistant AR-driven (49FENZR) and lineage plastic (42DENZR) cells. The closest 2000 peaks in proximity to a transcriptional start site were used for pathway analysis. Statistical significance was determined using a hypergeometric test. Representative AR ChIP-seq tracks surrounding the KLK3/PSA locus are shown. (c) Heatmap indicating AR ChIP-seq signal intensity in 42DENZR cells and DHT-stimulated LNCaP cells from Jin et al and Zhang et al. The shade of green reflects binding intensity. Each horizontal line represents a 6-kb locus.
Extended Data Fig. 3 Lineage plastic NE-like tumours exhibit a unique chromatin accessibility profile.
(a) Heatmap of ATAC-seq signal intensity in a GEMM of prostate adenocarcinoma (SKO, Ptenf/f) evolution to a plastic, NE-like state (DKO, Ptenf/f/Rb1f/f; TKO, Ptenf/f/Rb1f/f/Tp53f/f). Each horizontal line represents a 6-kb locus. (b) Motif analysis surrounding ATAC-seq peaks (250-bp) in NE-like DKO and TKO GEMMs compared to SKO. Transcription factor motifs identified by HOMER were plotted by ranks generated from their associated differential p values. (c) Significantly enriched pathways (gene set enrichment analysis) in accessible chromatin regions specific to NE-like DKO and TKO GEMMs compared to SKO.
Extended Data Fig. 4 The EZH2 cistrome is expanded in the lineage plastic state.
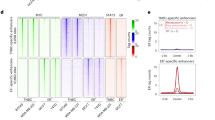
(a) Heatmap of EZH2 ChIP-seq signal intensity in CRPC 16DCRPC and 42DENZR cell lines (left), with overlaid H3K27Ac and H3K27Me3 histone mark ChIP-seq (right). Each horizontal line represents a 6-kb locus. (b) Representative ChIP-seq tracks surrounding the WNT5A locus in 16DCRPC and 42ENZR cells. Regions of EZH2 co-occupancy with the active H3K27Ac histone mark are highlighted. (c) Relative expression of genes bound by EZH2 alone (EZH2-none) or co-operatively with H3K27Me3 (EZH2-me) and H3K27Ac (EZH2-ac) histone marks in 42DENZR and 42FENZR cell lines. Box plot shows mean and interquartile range. (d) Heatmap of H3K27Me3 and K3K27Ac ChIP-seq signal intensity surrounding AR:EZH2 co-occupied regions in 42DENZR cells. (e) Heatmap indicating AR and EZH2 ChIP-seq signal intensity at AR:EZH2 co-occupied sites (n = 2155) in 42DENZR cells, and EZH2 signal intensity at the corresponding sites in AR-negative cell lines: NCI-H660, DU145 (GEO: GSE135623), and PC-3 (GEO: GSE123204). The shade of green (AR) or blue (EZH2) reflects binding intensity. Each horizontal line represents a 6-kb locus.
Extended Data Fig. 5 Characterization of EZH2 phosphorylation.
(a) EZH2 was immunoprecipitated in 42ENZR cells, trypsin digested, and analyzed by mass spectrometry. Peptides covering 36% of EZH2 were recovered and analyzed for post-translational modifications. (n = 4 independent replicates). (b) Expression of total and phosphorylated (T350, S21, and T311 residues) EZH2 in the indicated cell lines. Protein abundance was assessed by densitometry and is reported relative to total EZH2. (c) IHC staining of pEZH2-S21 and pEZH2-T350 in serial sections from representative CRPC (n = 39) and NEPC (n = 26) patient tumours (Scale bar, 100 μm). Staining area and intensity was quantified and reported (mean ± SD; two-tailed unpaired t-test). (d) Expression of genes positively regulated by EZH2 when phosphorylated at S21 [defined by Xu et al.] in the indicated cell lines and patient tumours from the Beltran 2016 cohort. Statistical analysis was performed using a two-tailed unpaired t-test. Box plots show mean and interquartile range. ns, not significant. (e) qRT-PCR of NE lineage markers in CRPCcrEZH2 cells expressing myc-tagged EZH2S21A or EZH2S21D mutants, reported relative to empty vector transfected cells. (mean ± SD; two-tailed unpaired t-test, n = 3). Immunoblotting confirmed transgene expression. (f) Proliferation of parental 16DCRPC (control) and CRPCcrEZH2 cells stably expressing EZH2T350A and EZH2T350D phospho-mutants assessed by IncuCyte (mean ± SD, n = 3 replicates). Immunoblotting confirmed transgene expression. (g) qRT-PCR of plasticity and NE markers in VCaP and C4-2 cell lines co-transfected with EZH2 siRNA and siRNA-resistant myc-tagged EZH2WT, EZH2T350A, or EZH2T350D plasmid following treatment with ENZ (10 μM) for 7 days (mean ± SD; two-tailed unpaired t-test, n = 3).
Extended Data Fig. 6 pEZH2-T350 is associated with lineage plasticity.
(a) Frequency of patients with low and high EMT and ASC signature scores in the SU2C37 and Labrecque et al clinical cohorts that exhibit a high pEZH2-T350 score (indicative of pEZH2-T350 phosphorylation). Patients were defined as “lo” or “hi” for each signature based on +/- 1 standard deviation from the mean signature score in the respective cohort. (b) Frequency of patients with high pEZH2-T350 score (defined as ≥1 standard deviation from cohort mean) in adenocarcinoma, adenocarcinoma with genomic RB1/TP53 loss, and NEPC patient tumours from the SU2C clinical cohort. (c) CDK1 transcript abundance in a GEMM model of prostate adenocarcinoma to NE-like tumour evolution (GEO: GSE90891). DKO and TKO tumours mimic NE-like tumours. SKO, PBCre4:Ptenf/f; DKO, PBCre4:Ptenf/f;Rb1f/f; TKO, PBCre4:Ptenf/f;Rb1f/f;Trp53f/f. Statistical analysis was performed using a two-tailed unpaired t-test. Box plots show mean and interquartile range. (d) CDK1 transcript abundance in benign prostate, adenocarcinoma (AdPC), and NEPC patient specimens from the 2011 Beltran cohort and 2016 Beltran cohort. Statistical analysis was performed using a two-tailed unpaired t-test. Box plots show mean and interquartile range. (e) CDK1 transcript abundance in a patient-derived xenograft (PDX) model of adenocarcinoma (LTL331) to NEPC (LTL331R) lineage conversion following androgen deprivation (castration) (ENA: PRJEB9660). Statistical analysis was performed using a two-tailed unpaired t-test. Mean with min/max range is reported. (f) Expression (qRT-PCR) of genes in 42DENZR cells following treatment with CDK1 inhibitor (5 µM RO-3306) for 24 hours. Data are reported relative to vehicle treated cells (mean ± SD; two-tailed unpaired t-test, n = 2).
Extended Data Fig. 7 Assessment of neuroendocrine differentiation following AR silencing.
(a) Immunoblot AR, EZH2, and H3K27Me3 (a surrogate marker of EZH2 activity) in 42DENZR cells following CRISPR-mediated AR deletion (crAR) or EZH2 inhibition (10 µM GSK126, 96 hrs). (b) Relative expression (qRT-PCR) of neuroendocrine lineage markers in 16DCRPC and C4-2 cell lines following siRNA-mediated AR silencing for 96 hours. Data are reported relative to cells transfected with a non-silencing scrambled control (mean ± SD, n = 3). A fold change >2 is considered significant. Immunoblotting confirmed AR knockdown.
Extended Data Fig. 8 Silencing EZH2 expression and/or activity reverts the lineage-infidelity phenotype.
(a-b) qRT-PCR in 42DENZR (a) and 42FENZR (b) cells following siRNA-mediated EZH2 silencing (siEZH2) for the indicated time, reported relative to non-transfected control cells at day 0 (mean ± SD; two-tailed unpaired t-test, n = 3). NTC, non-targeting control. (c-d) Spheroid formation and ALDH activity in 42DENZR (c) and 42FENZR (d) cells following siRNA-mediated EZH2 silencing (siEZH2; left) or treatment with increasing dose of EZH2 inhibitor (GSK126; right) for 8 days (mean ± SD; two-tailed unpaired t-test, n = 2). (e) qRT-PCR in 42DENZR cells treated with EZH2 inhibitor (10 μM GSK126) for 7 days, followed by removal (washout) for 14 days. Expression is reported relative to cells at day 0 (mean ± SD; two-tailed unpaired t-test, n = 3). Immunoblotting confirmed on-target effect.
Supplementary information
Supplementary Information
Supplementary Figs. 1–3 and CRISPR vector sequences.
Supplementary Video 1
Live-cell imaging of CRPCreporter cells treated with ENZ.
Supplementary Tables
Supplementary Tables 1–5.
Source data
Source Data Fig. 1
Statistical Source Data
Source Data Fig. 2
Statistical Source Data
Source Data Fig. 2
Unprocessed gel
Source Data Fig. 3
Statistical Source Data
Source Data Fig. 3
Unprocessed gel
Source Data Fig. 4
Statistical Source Data
Source Data Fig. 4
Unprocessed gel
Source Data Fig. 5
Statistical Source Data
Source Data Fig. 5
Unprocessed gel
Source Data Fig. 6
Statistical Source Data
Source Data Fig. 7
Statistical Source Data
Source Data Fig. 7
Unprocessed gel
Source Data Extended Data Fig. 1
Statistical Source Data
Source Data Extended Data Fig. 1
Unprocessed gel
Source Data Extended Data Fig. 5
Statistical Source Data
Source Data Extended Data Fig. 5
Unprocessed gel
Source Data Extended Data Fig. 6
Statistical Source Data
Source Data Extended Data Fig. 7
Statistical Source Data
Source Data Extended Data Fig. 7
Unprocessed gel
Source Data Extended Data Fig. 8
Statistical Source Data
Source Data Extended Data Fig. 8
Unprocessed gel
Rights and permissions
About this article
Cite this article
Davies, A., Nouruzi, S., Ganguli, D. et al. An androgen receptor switch underlies lineage infidelity in treatment-resistant prostate cancer. Nat Cell Biol 23, 1023–1034 (2021). https://doi.org/10.1038/s41556-021-00743-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-021-00743-5
This article is cited by
-
A novel exosome based therapeutic intervention against neuroendocrine prostate cancer
Scientific Reports (2024)
-
Chromatin accessibility landscape of relapsed pediatric B-lineage acute lymphoblastic leukemia
Nature Communications (2023)
-
Epigenetic mechanisms underlying subtype heterogeneity and tumor recurrence in prostate cancer
Nature Communications (2023)
-
Nucleophosmin 1 cooperates with BRD4 to facilitate c-Myc transcription to promote prostate cancer progression
Cell Death Discovery (2023)
-
Repurposing ketotifen as a therapeutic strategy for neuroendocrine prostate cancer by targeting the IL-6/STAT3 pathway
Cellular Oncology (2023)