Abstract
Organs consist of multiple cell types that ensure proper architecture and function. How different cell types coexist and interact to maintain their homeostasis in vivo remains elusive. The skin epidermis comprises mostly epithelial cells, but also harbours Langerhans cells (LCs) and dendritic epidermal T cells (DETCs). Whether and how distributions of LCs and DETCs are regulated during homeostasis is unclear. Here, by tracking individual cells in the skin of live adult mice over time, we show that LCs and DETCs actively maintain a non-random spatial distribution despite continuous turnover of neighbouring basal epithelial cells. Moreover, the density of epithelial cells regulates the composition of LCs and DETCs in the epidermis. Finally, LCs require the GTPase Rac1 to maintain their positional stability, density and tiling pattern reminiscent of neuronal self-avoidance. We propose that these cellular mechanisms provide the epidermis with an optimal response to environmental insults.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Additional source data for Figs. 4, 5, and Extended Data Figs. 1, 2, 4, 6, 8 have been deposited at Figshare and are accessible from https://doi.org/10.6084/m9.figshare.14199746, https://figshare.com/s/6855f86218f9026b0250, https://figshare.com/s/016423de8cf0ce293c1e, https://figshare.com/s/5c97c32da91f8810fed7, https://figshare.com/s/78598f01a1438569eb9b, https://figshare.com/s/03291104075fbb3492e7, https://figshare.com/s/f4ccb37bd6bd4557dbe2, https://figshare.com/s/c199699135ff4b446dc8 and https://figshare.com/s/336af7d8ebf313fcb2a5. Source data are provided with this paper. All other data that support the conclusions are available from the authors upon reasonable request.
Code availability
Positions of immune cells in the x–y plane were identified using Fiji. The minimum distance and displacement analyses were performed using the Matlab function squareform. The Voronoi tessellation used to determine nuclei neighbour relationships was performed using Matlab function voronoi. To make an artificially generated random pattern, random x–y positions were generated by Matlab function randi. All code is available from the corresponding author on request.
References
Gonzales, K. A. U. & Fuchs, E. Skin and its regenerative powers: an alliance between stem cells and their niche. Dev. Cell 43, 387–401 (2017).
Park, S., Greco, V. & Cockburn, K. Live imaging of stem cells: answering old questions and raising new ones. Curr. Opin. Cell Biol. 43, 30–37 (2016).
Simpson, C. L., Patel, D. M. & Green, K. J. Deconstructing the skin: cytoarchitectural determinants of epidermal morphogenesis. Nat. Rev. Mol. Cell Biol. 12, 565–580 (2011).
Solanas, G. & Benitah, S. A. Regenerating the skin: a task for the heterogeneous stem cell pool and surrounding niche. Nat. Rev. Mol. Cell Biol. 14, 737–748 (2013).
Tai, K., Cockburn, K. & Greco, V. Flexibility sustains epithelial tissue homeostasis. Curr. Opin. Cell Biol. 60, 84–91 (2019).
Xin, T., Greco, V. & Myung, P. Hardwiring stem cell communication through tissue structure. Cell 164, 1212–1225 (2016).
Ho, A. W. & Kupper, T. S. T cells and the skin: from protective immunity to inflammatory skin disorders. Nat. Rev. Immunol. 19, 490–502 (2019).
Pasparakis, M., Haase, I. & Nestle, F. O. Mechanisms regulating skin immunity and inflammation. Nat. Rev. Immunol. 14, 289–301 (2014).
Tay, S. S. et al. The skin-resident immune network. Curr. Dermatol. Rep. 3, 13–22 (2014).
Eming, S. A., Krieg, T. & Davidson, J. M. Inflammation in wound repair: molecular and cellular mechanisms. J. Invest. Dermatol. 127, 514–525 (2007).
Jameson, J. et al. A role for skin γδ T cells in wound repair. Science 296, 747–749 (2002).
Kaplan, D. H. Ontogeny and function of murine epidermal Langerhans cells. Nat. Immunol. 18, 1068–1075 (2017).
Leoni, G. et al. Wound repair: role of immune-epithelial interactions. Mucosal Immunol. 8, 959–968 (2015).
Merad, M., Ginhoux, F. & Collin, M. Origin, homeostasis and function of Langerhans cells and other langerin-expressing dendritic cells. Nat. Rev. Immunol. 8, 935–947 (2008).
Nielsen, M. M., Witherden, D. A. & Havran, W. L. γδ T cells in homeostasis and host defence of epithelial barrier tissues. Nat. Rev. Immunol. 17, 733–745 (2017).
Strbo, N., Yin, N. & Stojadinovic, O. Innate and adaptive immune responses in wound epithelialization. Adv. Wound Care 3, 492–501 (2014).
Takashima, A. & Bergstresser, P. R. Cytokine-mediated communication by keratinocytes and Langerhans cells with dendritic epidermal T cells. Semin. Immunol. 8, 333–339 (1996).
Sugita, K. et al. Innate immunity mediated by epidermal keratinocytes promotes acquired immunity involving Langerhans cells and T cells in the skin. Clin. Exp. Immunol. 147, 176–183 (2007).
Bernard, J. J., Gallo, R. L. & Krutmann, J. Photoimmunology: how ultraviolet radiation affects the immune system. Nat. Rev. Immunol. 19, 688–701 (2019).
Lewis, J. M. et al. Mechanisms of chemical cooperative carcinogenesis by epidermal Langerhans cells. J. Invest. Dermatol. 135, 1405–1414 (2015).
Chodaczek, G. et al. Body-barrier surveillance by epidermal γδ TCRs. Nat. Immunol. 13, 272–282 (2012).
Kitashima, D. Y. et al. Langerhans cells prevent autoimmunity via expansion of keratinocyte antigen-specific regulatory T cells. EBioMedicine 27, 293–303 (2018).
Kubo, A., Nagao, K. & Amagai, M. 3D visualization of epidermal Langerhans cells. Methods Mol. Biol. 961, 119–127 (2013).
Kubo, A. et al. External antigen uptake by Langerhans cells with reorganization of epidermal tight junction barriers. J. Exp. Med. 206, 2937–2946 (2009).
Matsui, T. & Amagai, M. Dissecting the formation, structure and barrier function of the stratum corneum. Int. Immunol. 27, 269–280 (2015).
Nagao, K. et al. Stress-induced production of chemokines by hair follicles regulates the trafficking of dendritic cells in skin. Nat. Immunol. 13, 744–752 (2012).
Yoshida, K. et al. Distinct behavior of human Langerhans cells and inflammatory dendritic epidermal cells at tight junctions in patients with atopic dermatitis. J. Allergy Clin. Immunol. 134, 856–864 (2014).
Bobr, A. et al. Autocrine/paracrine TGF-β1 inhibits Langerhans cell migration. Proc. Natl Acad. Sci. USA 109, 10492–10497 (2012).
Madisen, L. et al. A robust and high-throughput Cre reporting and characterization system for the whole mouse brain. Nat. Neurosci. 13, 133–140 (2010).
Jung, S. et al. Analysis of fractalkine receptor CX3CR1 function by targeted deletion and green fluorescent protein reporter gene insertion. Mol. Cell. Biol. 20, 4106–4114 (2000).
Mesa, K. R. et al. Homeostatic epidermal stem cell self-renewal is driven by local differentiation. Cell Stem Cell 23, 677–686 e4 (2018).
Xin, T. et al. Flexible fate determination ensures robust differentiation in the hair follicle. Nat. Cell Biol. 20, 1361–1369 (2018).
Rompolas, P. et al. Live imaging of stem cell and progeny behaviour in physiological hair-follicle regeneration. Nature 487, 496–499 (2012).
Rompolas, P. et al. Spatiotemporal coordination of stem cell commitment during epidermal homeostasis. Science 352, 1471–1474 (2016).
Pineda, C. M. et al. Intravital imaging of hair follicle regeneration in the mouse. Nat. Protoc. 10, 1116–1130 (2015).
Rompolas, P., Mesa, K. R. & Greco, V. Spatial organization within a niche as a determinant of stem-cell fate. Nature 502, 513–518 (2013).
Kissenpfennig, A. et al. Dynamics and function of Langerhans cells in vivo: dermal dendritic cells colonize lymph node areas distinct from slower migrating Langerhans cells. Immunity 22, 643–654 (2005).
Itohara, S. et al. T cell receptor δ gene mutant mice: independent generation of αβ T cells and programmed rearrangements of γδ TCR genes. Cell 72, 337–348 (1993).
Bobr, A. et al. Acute ablation of Langerhans cells enhances skin immune responses. J. Immunol. 185, 4724–4728 (2010).
De Creus, A. et al. Langerhans cells that have matured in vivo in the absence of T cells are fully capable of inducing a helper CD4 as well as a cytotoxic CD8 response. J. Immunol. 165, 645–653 (2000).
Taveirne, S. et al. Langerhans cells are not required for epidermal Vγ3 T cell homeostasis and function. J. Leukoc. Biol. 90, 61–68 (2011).
Zhang, B. et al. Differential requirements of TCR signaling in homeostatic maintenance and function of dendritic epidermal T cells. J. Immunol. 195, 4282–4291 (2015).
Ivanova, A. et al. In vivo genetic ablation by Cre-mediated expression of diphtheria toxin fragment A. Genesis 43, 129–135 (2005).
Brown, S. et al. Correction of aberrant growth preserves tissue homeostasis. Nature 548, 334–337 (2017).
Pineda, C. M. et al. Hair follicle regeneration suppresses Ras-driven oncogenic growth. J. Cell Biol. 218, 3212–3222 (2019).
Pruitt, S. C. et al. Cdkn1b overexpression in adult mice alters the balance between genome and tissue ageing. Nat. Commun. 4, 2626 (2013).
Xie, W. et al. Conditional expression of the ErbB2 oncogene elicits reversible hyperplasia in stratified epithelia and up-regulation of TGFα expression in transgenic mice. Oncogene 18, 3593–3607 (1999).
Park, S. et al. Tissue-scale coordination of cellular behaviour promotes epidermal wound repair in live mice. Nat. Cell Biol. 19, 155–163 (2017).
Bauer, J. et al. A strikingly constant ratio exists between Langerhans cells and other epidermal cells in human skin. A stereologic study using the optical disector method and the confocal laser scanning microscope. J. Invest. Dermatol. 116, 313–318 (2001).
Numahara, T. et al. Spatial data analysis by epidermal Langerhans cells reveals an elegant system. J. Dermatol. Sci. 25, 219–228 (2001).
Sandrock, I. et al. Genetic models reveal origin, persistence and non-redundant functions of IL-17-producing γδ T cells. J. Exp. Med. 215, 3006–3018 (2018).
Gentek, R. et al. Epidermal γδ T cells originate from yolk sac hematopoiesis and clonally self-renew in the adult. J. Exp. Med. 215, 2994–3005 (2018).
Ghigo, C. et al. Multicolor fate mapping of Langerhans cell homeostasis. J. Exp. Med. 210, 1657–1664 (2013).
Merad, M. et al. Langerhans cells renew in the skin throughout life under steady-state conditions. Nat. Immunol. 3, 1135–1141 (2002).
Marsh, E. et al. Positional stability and membrane occupancy define skin fibroblast homeostasis in vivo. Cell 175, 1620–1633 e13 (2018).
Swetman, C. A. et al. Extension, retraction and contraction in the formation of a dendritic cell dendrite: distinct roles for Rho GTPases. Eur. J. Immunol. 32, 2074–2083 (2002).
Grueber, W. B. & Sagasti, A. Self-avoidance and tiling: mechanisms of dendrite and axon spacing. Cold Spring Harb. Perspect. Biol. 2, a001750 (2010).
Zipursky, S. L. & Grueber, W. B. The molecular basis of self-avoidance. Annu Rev. Neurosci. 36, 547–568 (2013).
Glogauer, M. et al. Rac1 deletion in mouse neutrophils has selective effects on neutrophil functions. J. Immunol. 170, 5652–5657 (2003).
Hotulainen, P. & Hoogenraad, C. C. Actin in dendritic spines: connecting dynamics to function. J. Cell Biol. 189, 619–629 (2010).
Dogterom, M. & Koenderink, G. H. Actin-microtubule crosstalk in cell biology. Nat. Rev. Mol. Cell Biol. 20, 38–54 (2019).
Luckashenak, N. et al. Rho-family GTPase Cdc42 controls migration of Langerhans cells in vivo. J. Immunol. 190, 27–35 (2013).
Nishibu, A. et al. Behavioral responses of epidermal Langerhans cells in situ to local pathological stimuli. J. Invest. Dermatol. 126, 787–796 (2006).
Mohammed, J. et al. Stromal cells control the epithelial residence of DCs and memory T cells by regulated activation of TGF-β. Nat. Immunol. 17, 414–421 (2016).
Van den Bossche, J. et al. Regulation and function of the E-cadherin/catenin complex in cells of the monocyte-macrophage lineage and DCs. Blood 119, 1623–1633 (2012).
Mayumi, N. et al. E-cadherin interactions are required for Langerhans cell differentiation. Eur. J. Immunol. 43, 270–280 (2013).
Vasioukhin, V. et al. The magical touch: genome targeting in epidermal stem cells induced by tamoxifen application to mouse skin. Proc. Natl Acad. Sci. USA 96, 8551–8556 (1999).
Peron, S. P. et al. A cellular resolution map of barrel cortex activity during tactile behavior. Neuron 86, 783–799 (2015).
Chen, X. et al. Endogenous expression of HrasG12V induces developmental defects and neoplasms with copy number imbalances of the oncogene. Proc. Natl Acad. Sci. USA 106, 7979–7984 (2009).
Jung, S. et al. In vivo depletion of CD11c+ dendritic cells abrogates priming of CD8+ T cells by exogenous cell-associated antigens. Immunity 17, 211–220 (2002).
Acknowledgements
We thank A. Anderson of Life Science Editors for critical feedback on the manuscript; N. Anandasabapathy for the Lang-eGFP mice; S. Beronja for the HrasG12V mice; and Akiko Iwasaki for the Tcrd KO mice. This work is supported by the Howards Hughes Medical Institute Scholar award, National Institutes if Health grants no. 1R01AR072668-01. S.P. was supported by The New York Stem Cell Foundation (NYSCF–D–F58). Y.B. is supported by the Centre National de la recherche Scientifique, The Institut Curie and the Institut National de la santé et de la recherche médicale (INSERM). E.M. was supported in part by the National Institute of Health (T32 GM007499). D.P.M. was supported by The National Insitutes of Health (T32-GM007223-44). J.L.M. was supported by Lo Graduate Fellowship for Excellence in Stem Cell Research and The National Institutes of Health (T32 GM 7499-41 A1 and T32 HD 7149-40).
Author information
Authors and Affiliations
Contributions
S.P., C.M.-M. and V.G. designed experiments and analysed data. S.P. performed two-photon imaging, laser ablations, Matlab and IMARIS analysis, mouse genetics and toxin injections. C.M.-M. performed two-photon imaging, whole-mount staining, FACS preparatory work and analysis and mouse genetics. D.G.G. assisted with two-photon imaging, Matlab and IMARIS analysis, and experimental discussions throughout the project. E.A.L. assisted with whole mounts and mouse genetics. J.D.B. assisted with mouse genetics. C.M.P. assisted with the HRAS model development. E.M. assisted with laser ablations and Matlab analysis. J.L.M. assisted with analysis and critical feedback on the manuscript. D.P.M. assisted with Airyscan microscopy and critical feedback on the manuscript. A.S.-S. assisted with quantifications of epithelial density and cell–cell contact. Y.B., K.C. and O.M. assisted with critical feedback on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Cell Biology thanks the anonymous reviewers for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 The depletion of LCs or DETCs does not impact epithelial basal density or epidermal architecture of the ear.
a, X-y view of the basal layer of the epidermis with LCs in red (anti-MHC II) and DETCs in green (anti-TCRγδ) comparing Lang-DTR control mice without the addition of diphtheria toxin (or – DT) to Lang-DTR mice given one dose of 1 µg/body weight diphtheria toxin for the acute depletion of LCs 5 days before harvest (+ DT) (left panels) or comparing TCRδ WT to TCRδ KO mice (right panels) (representative images from 3 mice each). Scale bar, 50 μm. b, X-y view of the basal layer and x-z view of the epidermis with epithelial nuclei in white and collagen in magenta, comparing Lang-DTR mice either - DT or + DT (left panels) or comparing TCRδ WT to TCRδ KO mice (right panels) (representative images from 3 mice each). Scale bar, 30 μm. c,e, Quantification of epithelial basal cell number comparing Lang-DTR mice either - DT or + DT mice (c) and TCRδ WT to TCRδ KO mice (e). Unpaired, two-tailed Student’s t-test (n=3 mice respectively). Data presented as absolute values +/- SD. d,f, Quantification of LC and DETC cell numbers comparing Lang-DTR mice either - DT or + DT, (d) and TCRδ WT to TCRδ KO mice (f). Unpaired, two-tailed Student’s t-test (n=3 mice respectively). Data presented as absolute values +/- SD. g,h, Ratio between epithelial basal and DETCs in Lang-DTR + DT (g) and LCs in TCRδ KO (h) compared to their respective control mice (n=3 mice respectively). Area quantified for cell number 0.0625 mm2. Unpaired, two-tailed Student’s t-test. Data presented as absolute values +/- SD.
Extended Data Fig. 2 Full depletion of LCs and DETCs does not alter epidermal architecture.
a, X-y view of basal layer of epidermis in paw with LCs (Lang-EGFP) in green comparing Lang-DTR control mice without diphtheria toxin (or – DT) to Lang-DTR mice given 1 dose of 1 µg/body weight diphtheria toxin for the acute depletion of LCs 5 days before harvest (+ DT) (representative images from 2 mice each). Scale bar, 50 μm. b, X-y view of basal layer and x-z view of epidermis with epithelial nuclei in white (K14-H2B-mCherry), comparing Lang-DTR mice either – DT or + DT (representative images from 3 mice each). Scale bar, 50 μm. c, Quantifications of epithelial basal cell number comparing Lang-DTR mice - DT or + DT pre depletion (D0) and 5 days post depletion (D5). Area quantified 0.0625 mm2. (n=2 mice respectively). d, Quantification of epidermal thickness comparing Lang-DTR mice -DT or +DT. Area quantified 0.0625 mm2. (n=2 mice respectively). e, X-y view of basal layer and x-z view of epidermis in the ear with all nuclei (Hoechst) in gray and immune cells (CD45+) in white comparing control huLangerin-CreER; TCR δ-CreER; Rosa-GFP-stop-DTA mice without tamoxifen (DTA) to huLangerin-CreER; TCR δ-CreER; Rosa-GFP-stop-DTA mice given 2 mgs of tamoxifen on 2 consecutive days 8 days before harvest (DTA + tamoxifen) (representative images from 2 mice each). Scale bar, 50 μm. f, Quantification of epithelial basal cell number comparing huLangerin-CreER; TCR δ-CreER; Rosa-GFP-stop-DTA either – Tamoxifen or + Tamoxifen at day 8. Area quantified 0.0625 mm2. (n=2 mice respectively). g, Quantification of epidermal thickness comparing huLangerin-CreER; TCR δ-CreER; Rosa-GFP-stop-DTA either – Tamoxifen or + Tamoxifen at day 8. Area quantified 0.0625 mm2. (n=2 mice respectively). All data in figure presented as absolute values.
Extended Data Fig. 3 LCs and DETCs remain embedded in the basal layer of the ear epidermis upon changes in epithelial basal cell density.
(a-c) 3D reconstruction with Imaris surface rendering shows that LCs (red), DETCs (green), and epithelial cells (blue) are complicatedly intermixed in the epidermis. a, X-y and x-z views show that LCs and DETCs embed in the basal layer of the epidermis in HrasG12V mice (representative of 2 mice) 6 weeks after 1 dose of 2 mgs tamoxifen similar to (b) WT LCs and DETCs in homeostasis (representative of 3 mice) and (c) Cdkn1b;rtTA mice 3 days after induction with 1 mg/ml doxycycline (representative of 2 mice). All images: Scale bar, 10 μm.
Extended Data Fig. 4 Proliferation in LCs and DETCs is enhanced in a model of increased epithelial and immune cell density.
a,b, Staining of the epidermal basal layer of the ear in HrasG12V/+ compared to control Hras +/+ mice 6 weeks after induction with 1 dose of 2 mgs of tamoxifen for the proliferation marker ki67 (representative of 2 mice respectively). Scale bar, 50µm. c-e, Quantifications of cycling immune cells (CD45+ki67+) in the HrasG12V/+ mice compared to control Hras+/+mice. Area quantified 0.0625 mm2 X 3 regions per mouse. (n=2 mice respectively). Data presented as absolute values. f-j, Stainings of the epidermal basal layer of the ear in 6 week old CD1 mice for proliferation markers shown in red (f) ki67 and (g) phosphohistone H3 to enumerate proliferating immune cells (CD45+) shown in green (representative of 3 mice for ki67 and 2 mice for pH3). Scale bar, 50µm. h-j, Quantifications of cycling immune cells (CD45+ki67+) and of immune cells that are actively dividing (CD45+pH3+) during homeostasis. Area quantified 0.0625 mm2 X 3 regions per mouse. Unpaired, two-tailed Student’s t-test (n=3 mice for ki67 and n=2 mice for pH3). Data presented as absolute values.
Extended Data Fig. 5 Immune cell density gradually changes along with changes in epithelial basal density.
a, The densities of LCs and DETCs in mice with blocked proliferation of epithelial stem cells (Cdkn1b; K14-rtTA) during a 3-day revisit experiment post induction with 1 mg/ml doxyclycline. Epithelial nuclei are white (top panel). LCs (huLangerin-CreER; Rosa-stop-tdTomato) are red and DETCs (CX3CR1-GFP+/−) are green (bottom panel). Yellow asterisk (hair follicle). (3 mice respectively). Scale bar, 50 μm. b, Voronoi diagrams showing spatial distribution of LCs (top panel, red) and DETCs (bottom panel, green) (c) Quantification of epithelial basal cell number over the timecourse * p=0.0282 (Day 1 vs. Day 0), ** p=0.0076 (Day 2 vs. Day 0), and * p=0.0253 (Day 3 vs. Day 0), paired, two-tailed Student’s t-test (n=3 mice respectively). Data presented as percentage values +/- SD. d, Quantification of LC and DETC number for both LCs and DETCs over the timecourse. Area quantified 0.25 mm2. * p= 0.0101 (Day 0 LC vs. Day 2 LC) and * p=0.0113 (Day 0 LC vs. Day 3 LC), paired, two-tailed Student’s t-test (n=3 mice respectively). Data presented as percentage values +/- SD (e) Fiji software was used to count nucleated cells in the epidermis based on arbitrary distance from the SHG of the collagen in 260 X 260 ROIs of 40X images (0.27µm/pixel, 1µm step size). Basal cell counts comparing Cdkn1b;rtTA mice to Cdkn1b controls at day 3 post induction with 1 mg/ml doxycycline. Quantification of suprabasal cell counts which include spinous and granular layers, comparing Cdkn1b;rtTA mice to Cdkn1b controls at day 3 post induction with 1 mg/ml doxycycline. Area quantified 0.0625 mm2. * p=0.0318 and ** p=0.0063, unpaired, two-tailed Student’s t-test (n=5 separate regions from 3 Cdkn1b mice and n=4 separate regions from 2 Cdkn1b;rtTA mice). Data presented as absolute values +/- SD.
Extended Data Fig. 6 Cell apoptosis and activation do not contribute to the density drop of LCs and DETCs in Cdkn1b;rtTA mice.
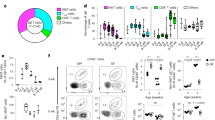
(a) Ears were taken from Cdkn1b and Cdkn1b;rtTA mice 3 days post induction with 1 mg/ml of doxycycline. Staining of epithelial basal layer of the ear for the apoptotic marker cleaved-caspase 3 (CC3) shown in white of Cdkn1b;rtTA (right panels) and control Cdkn1b mice (left panels) (representative images of 3 mice respectively). LCs and DETCs are shown in red (anti-CD45) and epithelial basal cell nuclei (K14-H2B-Cerulean) are shown in green. Inset in bottom right panel shows an epithelial cell positive for CC3. Scale bar, 40µm. b,c, Quantifications of immune cells undergoing apoptosis (CD45+CC3+) in Cdkn1b;rtTA compared to controls. Area quantified 0.0625 mm2 X 3 regions per mouse. Unpaired, two-tailed Student’s t-test (n=3 mice respectively). Data presented as absolute values/percentages respectively +/- SD. d, Epidermal single cell suspensions were processed from ear of mice 3 days post induction with 1mg/ml of doxycycline for flow cytometry and gated for LCs using (anti-CD207). K14-H2B-Cerulean was used to gate out the epithelial cells. e,f, The expression levels of the known activation marker (MHCII) from LCs from Cdkn1b;rtTA mice (closed red) compared to those from Cdkn1b control mice (solid black) in the epidermis. Negative control (gray) gated on cells negative for epithelial cell marker K14-H2B-Cerulean and immune cell markers for LCs (CD207) and DETCs.Unpaired, two-tailed Student’s t-test (n=3 mice respectively). Data presented as absolute values +/- SD. g-h, The expression levels of E-cadherin, a mediator of cell-cell interaction known to be down-regulated in activated LCs, from LCs of Cdkn1b;rtTA mice (closed red) and those from Cdkn1b control mice (solid black) in the epidermis. Negative control (gray) gated on cells negative for the epithelial cell marker K14-H2B-Cerulean and immune cell markers for LCs (CD207) and DETCs. Unpaired, two-tailed Student’s t-test (n= 3 mice respectively). Data presented as absolute values +/- SD.
Extended Data Fig. 7 The distribution of one immune population is maintained upon the depletion of the other and surviving DETCs re-establish a regular pattern after local cell loss.
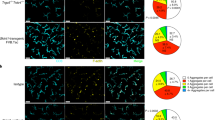
a,c, Voronoi diagrams were generated from images acquired for the quantifications in Extended Data Fig. 1 comparing (a) the distribution of DETCs in Lang-DTR mice with and without one dose of 1 ug/body weight of diphtheria toxin 5 days post depletion or (c) that of LCs in TCRδ KO compared to WT controls showing that the distribution of the remaining immune population is not disrupted in either models. b,d, Minimum distance quantifications showed no significant differences regardless of which immune population had been ablated when compared to controls showing that (b) DETC (n=198 cells Lang-DTR –DT, n=195 cells Lang-DTR +DT) and (d) LC (n=172 cells WT, n=176 cells TCRδ KO) patterns are maintained in the absence of the other population. Area quantified 0.625 mm2. Unpaired, two-tailed Student’s t-test (3 mice respectively). Data presented as absolute values +/- SD. e, Local laser ablation of DETCs. DETCs within the yellow box (100 μm × 100 μm) are ablated by multiphoton laser (810 nm) and the same region is revisited 3 days after the ablation. The upper panel shows actual images from a mouse from day 0 before ablation to day 3 post ablation. Scale bar, 10µm. The lower panel displays the Voronoi diagram for DETCs generated from the images at each timepoint and encompasses a larger area around the ablation site. Area size 0.25 mm2 (red, LCs (huLangerin-CreER; Rosa-stop-tdTomato); green, DETCs (CX3CR1-GFP+/−); white, nuclei of epithelial cells (K14-H2B-Cerulean); representative images from 4 mice). f, The DETC pattern within ablated region was quantified from day 1 to day 7 and compared to the initial number at day 0. * p=0.0102 (Day 1 vs. Day 0), ** p=0.0034 (Day 2 vs. Day 0), ** p=0.0087 (Day 3 vs. Day 0), paired, two-tailed Student’s t-test (n=4 mice respectively). Data presented as absolute values +/- SD.
Extended Data Fig. 8 LCRac1KO have increased mobility in the epidermis compared to models with similar LC density and cell apoptosis does not contribute to the density drop observed in the LCRac1KO phenotype.
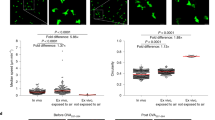
a, Quantification of cell number across models (n=3 mice respectively). b, Minimum distance across models. Note, the data used for Lang-DTR is the same data set as in Fig. 4d,e for Day 8 and the data used for LCRac1KO at 1 week and 3 weeks are the same data sets used in Fig. 5f (n= 811 Cdkn1b, n=480 Cdkn1b; rtTA, n=365 Lang-DTR, n=893 LCRac1KO(1week), n=797 LCRac1KO(3weeks) cells from 3 mice respectively (c) Displacement analysis of LCRac1KO at day 0 versus day 3 compared to LCs of Cdkn1b;rtTA mice during 72 h and LCs of Lang-DTR mice over the course of 8 days.. The data used for LCRac1KO at 1 week and 3 weeks are the same data sets used in Fig. 5b (n=705 Cdkn1b, n=858 Cdkn1b;rtTA, n=789 Lang-DTR, n=657 LCWT, n=498 LCRac1KO from cells 3 mice respectively). Data presented as absolute values +/- SD. d-e, Staining of epithelial basal layer of the ear for apoptotic marker cleaved-caspase (CC3) shown in white of (d) huLangerin-CreER; Rac1+/+ mice and (e) huLangerin-CreER; Rac1fl/fl mice days post induction with 2 mg dose of tamoxifen given on 5 consecutive days. LCs are shown in red (anti-CD207), all nuclei are shown in green (Hoechst). Inset in bottom right panel shows an LC cell positive for CC3. All images: scale bar, 40 μm. f,g, Quantifications show very few apoptotic events (< 1%) in the LCs of both huLangerin-CreER; Rac1+/+ mice and huLangerin-CreER; Rac1+/+ control mice (CC3+CD207+) (data is representative n=3 for huLangerin-CreER; Rac1+/+ mice, n=3 for huLangerin-CreER; Rac1fl/fl mice). Area quantified 0.0625 mm2 X 3 regions/mouse. Unpaired, two-tailed Student’s t-test (n=3 mice respectively). Data presented as absolute/percentage values respectively +/- SD.
Extended Data Fig. 9 Activation does not contribute to the drop in density observed in LCRac1KO.
a, Ears and lymph nodes were taken from mice 7 days post induction with 2mg dose of tamoxifen given on 5 consecutive days. Epidermal and lymph node single cell suspensions were processed for flow cytometry and gated for LCs using anti-langerin antibody (CD207). K14-H2B-Cerulean was used to gate out the epithelial cells. b-e, LCRac1KO (open red) show no significant difference in the expression levels of the known activation markers MHCII and CD86 compared to LCwt (solid black) in either the epidermis or draining lymph nodes showing that LCRac1KO are not activated in this model. Unpaired, two-tailed Student’s t-test (data is representative n=4 for huLangerin-CreER; Rac1+/+ mice, n=5 for huLangerin-CreER; Rac1fl/fl mice in c and n=4 for both groups in e). Data presented as absolute values +/- SD. f, Epidermal cell suspensions were processed for flow cytometry and gated for LCs using anti-langerin antibody (CD207). K14-H2B-Cerulean was used to gate out the epithelial cells. g-h, LCRac1KO (closed red, n=3) show no significant difference in the expression levels of the known mediator of cell-cell interaction E-cadherin compared to LCwt (solid black, n=3). Negative control (gray) is gated on cells negative for the epithelial cell marker K14-H2B-Cerulean and immune cell markers for LCs (CD207) and DETCs. Unpaired, two-tailed Student’s t-test (data is representative n=3 for huLangerin-CreER; Rac1+/+ mice, n=3 for huLangerin-CreER; Rac1fl/fl mice). Data presented as absolute values +/- SD.
Extended Data Fig. 10 Skin-resident immune cells actively coordinate their distribution with epithelial basal cells.
During homeostasis, LCs and DETCs actively maintain a non-random distribution. Although these immune cells continuously move within epithelial basal cell neighbors, they sustain their regular tiling pattern. Moreover, epithelial basal cells determine the density of LCs and DETCs in the skin epidermis. However, LCs and DETCs do not impact the density of epithelial basal cells nor the architecture of the epidermis signifying a unidirectional regulation of density. Finally, immune cell dendrites enable maximal coverage of the basal layer and do not come into contact with neighbouring immune cells in homeostasis and their dendritic behaviors, mediated by Rac1, coordinate their patterned organization.
Supplementary information
Supplementary Video 1
3D reconstruction of epidermis. 3D reconstruction with Imaris surface rendering shows that LCs (red), DETCs (green) and epithelial cells (white) are intermixed in a complicated pattern in the epidermis. LCs and DETCs extrude their dendrite vertically to the surface of the epidermis as well as laterally between the epithelial cells. Scale bar, 40 μm.
Supplementary Video 2
Dendritic movement of LCs and DETCs during homeostasis. Time-lapse recording over 4 h of LCs and DETCs shows stable cell bodies while their dendrites dynamically explore junctional spaces between neighbouring epithelial basal cells. LCs (red), DETCs (green) and nuclei of epithelial basal cells (white). Yellow arrows point to dendritic activity in the junctional space of epithelial basal cells. Scale bar, 50 μm.
Supplementary Video 3
Time-lapse movies of LCRac1KO and LCWT. Time-lapse recording over 4 h of LCRac1KO and LCWT shows that LCs are stable in position and protrude dendrites over time regardless of whether Rac1 is knocked out in the LCs. Yellow arrows point to dendritic activity. The magnified movies repeat 5 times. Scale bar, 50 μm (wide view), 20 μm (magnified view).
Source data
Source Data Fig. 1
Statistical source data
Source Data Fig. 2
Statistical source data
Source Data Fig. 3
Statistical source data
Source Data Fig. 4
Statistical source data
Source Data Fig. 5
Statistical source data
Source Data Extended Data Fig. 1
Statistical source data
Source Data Extended Data Fig. 2
Statistical source data
Source Data Extended Data Fig. 4
Statistical source data
Source Data Extended Data Fig. 5
Statistical source data
Source Data Extended Data Fig. 6
Statistical source data
Source Data Extended Data Fig. 7
Statistical source data
Source Data Extended Data Fig. 8
Statistical source data
Source Data Extended Data Fig. 9
Statistical source data
Rights and permissions
About this article
Cite this article
Park, S., Matte-Martone, C., Gonzalez, D.G. et al. Skin-resident immune cells actively coordinate their distribution with epidermal cells during homeostasis. Nat Cell Biol 23, 476–484 (2021). https://doi.org/10.1038/s41556-021-00670-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-021-00670-5
This article is cited by
-
Cellular and molecular mechanisms of skin wound healing
Nature Reviews Molecular Cell Biology (2024)
-
Polar microalgae extracts protect human HaCaT keratinocytes from damaging stimuli and ameliorate psoriatic skin inflammation in mice
Biological Research (2023)
-
Tissue-specific macrophages: how they develop and choreograph tissue biology
Nature Reviews Immunology (2023)
-
Tracking skin and immune cell interactions
Nature Cell Biology (2021)