Abstract
Cells achieve highly efficient and accurate communication through cellular projections such as neurites and filopodia, yet there is a lack of genetically encoded tools that can selectively manipulate their composition and dynamics. Here, we present a versatile optogenetic toolbox of artificial multi-headed myosin motors that can move bidirectionally within long cellular extensions and allow for the selective transport of GFP-tagged cargo with light. Utilizing these engineered motors, we could transport bulky transmembrane receptors and organelles as well as actin remodellers to control the dynamics of both filopodia and neurites. Using an optimized in vivo imaging scheme, we further demonstrate that, upon limb amputation in axolotls, a complex array of filopodial extensions is formed. We selectively modulated these filopodial extensions and showed that they re-establish a Sonic Hedgehog signalling gradient during regeneration. Considering the ubiquitous existence of actin-based extensions, this toolbox shows the potential to manipulate cellular communication with unprecedented accuracy.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
Supporting data are available upon request to the corresponding author. Source data are provided with this paper.
Change history
19 February 2021
A Correction to this paper has been published: https://doi.org/10.1038/s41556-021-00650-9
08 April 2021
A Correction to this paper has been published: https://doi.org/10.1038/s41556-021-00675-0
References
Jacquemet, G., Hamidi, H. & Ivaska, J. Filopodia in cell adhesion, 3D migration and cancer cell invasion. Curr. Opin. Cell Biol. 36, 23–31 (2015).
Sanders, T. A., Llagostera, E. & Barna, M. Specialized filopodia direct long-range transport of SHH during vertebrate tissue patterning. Nature 497, 628–632 (2013).
Roy, S., Hsiung, F. & Kornberg, T. B. Specificity of Drosophila cytonemes for distinct signaling pathways. Science 332, 354–358 (2011).
Möller, J., Lühmann, T., Chabria, M., Hall, H. & Vogel, V. Macrophages lift off surface-bound bacteria using a filopodium–lamellipodium hook-and-shovel mechanism. Sci. Rep. 3, 2884 (2013).
Eddy, R. J. et al. Tumor cell invadopodia: invasive protrusions that orchestrate metastasis. Trends Cell Biol. 27, 595–607 (2017).
Spector, I. et al. Latrunculins: novel marine toxins that disrupt microfilament organization in cultured cells. Science 219, 493–495 (1983).
MacLean-Fletcher, S. & Thomas, D. P. Mechanism of action of cytochalasin B on actin. Cell 20, 329–341 (1980).
Heimsath, E. G. et al. Myosin-X knockout is semi-lethal and demonstrates that myosin-X functions in neural tube closure, pigmentation, hyaloid vasculature regression and filopodia formation. Sci. Rep. 7, 17354 (2017).
Davis, D. M. & Sowinski, S. Membrane nanotubes: dynamic long-distance connections between animal cells. Nat. Rev. Mol. Cell Biol. 9, 431–436 (2008).
Matus, A. Actin-based plasticity in dendritic spines. Science 290, 754–758 (2000).
Small, J. V., Isenberg, G. & Celis, J. E. Polarity of actin at the leading edge of cultured cells. Nature 272, 638–639 (1978).
Urban, Edit. et al. Electron tomography reveals unbranched networks of actin filaments in lamellipodia. Nat. Cell Biol. 12, 429–435 (2010).
Vale, R. D. The molecular motor toolbox for intracellular transport. Cell 112, 467–480 (2003).
Howard, J. Molecular motors: structural adaptations to cellular functions. Nature 389, 561–567 (1997).
Schindler, T. D. et al. Engineering myosins for long-range transport on actin filaments. Nat. Nanotechnol. 9, 33–38 (2014).
Harbury, P. B. et al. A switch between two-, three- and four-stranded coiled coils in GCN4 leucine zipper mutants. Science 262, 1401–1407 (1993).
Rothbauer, U. et al. Targeting and tracing antigens in live cells with fluorescent nanobodies. Nat. Methods 3, 887–889 (2006).
Sarov, M. et al. A genome-wide resource for the analysis of protein localisation in Drosophila. eLife 5, e12068 (2016).
Harikumar, A. et al. An endogenously tagged fluorescent fusion protein library in mouse embryonic stem cells. Stem Cell Rep. 9, 1304–1314 (2017).
Reznikoff, C. A., David, W. B. & Heidelberger, C. Establishment and characterization of a cloned line of C3H mouse embryo cells sensitive to postconfluence inhibition of division. Cancer Res. 33, 3231–3238 (1973).
Kerber, M. L. et al. A novel form of motility in filopodia revealed by imaging myosin-X at the single-molecule level. Curr. Biol. 19, 967–973 (2009).
Bischoff, M. et al. Cytonemes are required for the establishment of a normal hedgehog morphogen gradient in Drosophila epithelia. Nat. Cell Biol. 15, 1269–1281 (2013).
di Magliano, M. P. & Hebrok, M. Hedgehog signalling in cancer formation and maintenance. Nat. Rev. Cancer 3, 903–911 (2003).
Rohatgi, R., Milenkovic, L. & Scott, M. P. Patched1 regulates hedgehog signaling at the primary cilium. Science 317, 372–376 (2007).
Gong, X. et al. Structural basis for the recognition of sonic hedgehog by human Patched1. Science 361, eaas8935 (2018).
Yue, S. et al. Requirement of Smurf-mediated endocytosis of Patched1 in sonic hedgehog signal reception. eLife 3, e02555 (2014).
Callejo, A. et al. Dispatched mediates hedgehog basolateral release to form the long-range morphogenetic gradient in the Drosophila wing disk epithelium. Proc. Natl Acad. Sci. USA 108, 12591–12598 (2011).
Salles, F. T. et al. Myosin IIIa boosts elongation of stereocilia by transporting espin 1 to the plus ends of actin filaments. Nat. Cell Biol. 11, 443–450 (2009).
Berg, J. S. & Cheney, R. E. Myosin-X is an unconventional myosin that undergoes intrafilopodial motility. Nat. Cell Biol. 4, 246–250 (2002).
Bohil, A. B., Robertson, B. W. & Cheney, R. E. Myosin-X is a molecular motor that functions in filopodia formation. Proc. Natl Acad. Sci. USA 103, 12411–12416 (2006).
Barzik, M. et al. Ena/VASP regulates mDia2-initiated filopodial length, dynamics and function. Mol. Biol. Cell 25, 2604–2619 (2014).
Kennedy, M. J. et al. Rapid blue-light–mediated induction of protein interactions in living cells. Nat. Methods 7, 973–975 (2010).
Taslimi, A. et al. An optimized optogenetic clustering tool for probing protein interaction and function. Nat. Commun. 5, 4925 (2014).
Prigozhina, N. L. & Waterman-Storer, C. M. Protein kinase D-mediated anterograde membrane trafficking is required for fibroblast motility. Curr. Biol. 14, 88–98 (2004).
Okabe, S. & Hirokawa, N. Actin dynamics in growth cones. J. Neurosci. 11, 1918–1929 (1991).
Sheng, Z.-H. & Cai, Q. Mitochondrial transport in neurons: impact on synaptic homeostasis and neurodegeneration. Nat. Rev. Neurosci. 13, 77–93 (2012).
Hollenbeck, P. J. & Saxton, W. M. The axonal transport of mitochondria. J. Cell Sci. 118, 5411–5419 (2005).
Abe, Y. et al. Structural basis of presequence recognition by the mitochondrial protein import receptor Tom20. Cell 100, 551–560 (2000).
Imokawa, Y. & Yoshizato, K. Expression of Sonic Hedgehog gene in regenerating newt limb blastemas recapitulates that in developing limb buds. Proc. Natl Acad. Sci. USA 94, 9159–9164 (1997).
McCusker, C., Bryant, S. V. & Gardiner, D. M. The axolotl limb blastema: cellular and molecular mechanisms driving blastema formation and limb regeneration in tetrapods. Regeneration 2, 54–71 (2015).
Litingtung, Y. et al. Shh and Gli3 are dispensable for limb skeleton formation but regulate digit number and identity. Nature 418, 979–983 (2002).
Roy, S. & Gardiner, D. M. Cyclopamine induces digit loss in regenerating axolotl limbs. J. Exp. Zool. 293, 186–190 (2002).
Van Bergeijk, P. et al. Optogenetic control of organelle transport and positioning. Nature 518, 111–114 (2015).
Gutnick, A. et al. The light-sensitive dimerizer zapalog reveals distinct modes of immobilization for axonal mitochondria. Nat. Cell Biol. 21, 768–777 (2019).
Verhey, K. J. & Hammond, J. W. Traffic control: regulation of kinesin motors. Nat. Rev. Mol. Cell Biol. 10, 765–777 (2009).
Bird, J. E. et al. Harnessing molecular motors for nanoscale pulldown in live cells. Mol. Biol. Cell 28, 463–475 (2017).
Fatehullah, A., Tan, S. H. & Barker, N. Organoids as an in vitro model of human development and disease. Nat. Cell Biol. 18, 246–254 (2016).
Ariotti, N. et al. Modular detection of GFP-labeled proteins for rapid screening by electron microscopy in cells and organisms. Dev. Cell 35, 513–525 (2015).
Pédelacq, J.-D. et al. Engineering and characterization of a superfolder green fluorescent protein. Nat. Biotechnol. 24, 79–88 (2006).
Zhang, Y. et al. Rapid single-step induction of functional neurons from human pluripotent stem cells. Neuron 78, 785–798 (2013).
Ang, C. E. et al. The novel lncRNA lnc-NR2F1 is pro-neurogenic and mutated in human neurodevelopmental disorders. eLife 8, e41770 (2019).
Zhang, Z. et al. Optogenetic manipulation of cellular communication in axolotls. Protoc. Exch. https://doi.org/10.21203/rs.3.pex-1279/v1 (2020).
Nacu, E. et al. FGF8 and SHH substitute for anterior–posterior tissue interactions to induce limb regeneration. Nature 533, 407–410 (2016).
Acknowledgements
We thank Z. Bryant and P. Ruijgrok for discussion on the design and application of the motors and useful comments on the manuscript, the Cell Sciences Imaging Facility (CSIF) at Stanford University for technical assistance and the whole Barna Lab for helpful discussion and critical reading of the manuscript. The project was supported by awards R01HD088597 (M.B.) and U19MH104172 (M.W.) from the National Institutes of Health (NIH) and, in part, by award 1S10OD01227601 from the National Center for Research Resources (NCRR). The content of this Report is solely the responsibility of the authors and does not necessarily represent the official views of NIH or NCRR. Z.Z. is supported in part by an NIH P50 training grant. O.Z. is a Simons Fellow of the Helen Hay Whitney Foundation (HHWF) and in part supported by the Canadian Institutes for Health Research (CIHR) Fellowship. M.B. is a New York Stem Cell Foundation (NYSCF) Robertson Investigator and M.W. is a Howard Hughes Medical Institute (HHMI) Faculty Scholar.
Author information
Authors and Affiliations
Contributions
Z.Z., N.D. and M.B. conceived the project. N.D., Z.Z. and M.B. designed and constructed the expression vectors. Z.Z. and M.B. planned the experiments. Z.Z. and N.D. executed and analysed the experiments. Y.L., O.Z., H.D.R., M.W. and M.B. contributed to essential cell cultures, reagents and expertise. Z.Z. and M.B. wrote the manuscript with input from all authors. M.B. supervised all aspects of the work.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Cell Biology thanks the anonymous reviewers for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Characterization of ATV+ and ATV- in live cells.
a, Representative images of randomly chosen mouse fibroblasts expressing either (upper) ATV+ or (lower) ATV- with pmBFP chosen from 3 independent experiments. Scale, 20 µm. b, c, Mouse fibroblasts co-expressing sfGFP-ATV+, mKate2-ATV- and pmiRFP in the same cells. ATV+ and ATV- display significantly different localization patterns within the same cells. Representative images chosen from 3 independent experiments. Scale, 20 µm. Mouse fibroblasts expressing (d) sfGFP-ATV+ or (e) sfGFP-optoATV- were imaged under a total internal reflection (TIRF) microscope. d, Left: A kymograph of a cellular protrusion showing single molecule traces of sfGFP-ATV+ moving down the tip. Scale, 1 µm. Right: Quantification of the velocity of ATV+ moving within cellular protrusions. Single dots represent single molecule traces chosen from 3 independent experiments (N = 23 traces). e, Left: A kymograph of a cellular protrusion where single molecule traces of sfGFP-optoATV- retracting back to the cell body upon light stimulation. Scale, 100 nm. Right: Quantification of the velocity of activated optoATV- moving within cellular protrusions. Single dots represent single molecule traces chosen from 3 independent experiments (N = 16 traces). The graphs are shown as mean ± SEM.
Extended Data Fig. 2 Bidirectional transport of bulky transmembrane SHH receptors within filopodia using ATVs in live cells.
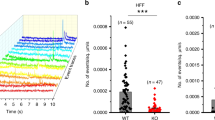
a, Expression of sPtch1-eGFP with or without co-expression of mKate2-ATV+ in C3H/10T1/2 cells. Representative images chosen from 3 independent experiments. Scale, 20 µm. b, Quantification of the relative concentration of sPtch1-eGFP at the tip compared to the cell body (N = 33 and 41 filopodia each), based on images from (a). c, Expression of Disp1-eGFP with or without co-expression of mKate2-ATV- in C3H/10T1/2 cells. Representative images chosen from 3 independent experiments. Scale, 20 µm. d, Quantification of the relative concentration of Disp1-eGFP at the tip compared to the cell body (N = 86 and 96 filopodia each), based on images from (c). Statistics determined by two-tailed Mann-Whitney test. Both bar graphs displayed as mean ± SEM.
Extended Data Fig. 3 Inducing filopodia growth by doxycycline in a stable Cos7 cell line.
a, A Cos7 cell line stably expressing mKate2-ATV+ and Espin1-sfGFP (linked with a T2A linker) under a TetOn-3G system was generated. b, The cells were imaged with or without 24 hours of 2 µg/ml doxycycline induction. Filopodia growth was only observed when doxycycline was applied. Representative images chosen from 3 independent experiments. Scale, 20 µm. pmBFP was used to visualize full cell morphology. c, Quantification of the filopodia density of each individual cell (N = 10 cells each). Statistics determined by two-tailed Mann-Whitney test. The bar graph displayed as mean ± SEM.
Extended Data Fig. 4 Preventing the outgrowth of mDia2M/A-mediated artificial filopodia using ATV- in live cells.
a, Expression of sfGFP-mDia2M/A with or without co-expression of mKate2-ATV- in Cos7 cells. The mutant actin nucleating protein mDia2M/A could generate artificial filopodia, while mDia2M/A trapped inside the cell body by mKate2-ATV- could not. Representative images chosen from 3 independent experiments. Scale, 20 µm. b, Quantification of the filopodia density of each individual cell (N = 10 cells each). Statistics determined by two-tailed Mann-Whitney test. The bar graph displayed as mean ± SEM.
Extended Data Fig. 5 GBP-GFP interaction is necessary for preserving native function of the transported cargo in live cells.
a, Expression of mKate2-ATV+ & sfGFP-mDia2M/A independently or the covalently linked cargo-motor in Cos7 cells. The prior is based on the binding interaction of GBP-GFP while the latter is based on covalent linkage. GBP-GFP interaction preserved the function of mDia2M/A in generating filopodia, while covalently linkage disrupted cargo function and cells formed fewer filopodia. The white dashed line delineates the smooth cell border. Representative images chosen from 3 independent experiments. Scale, 20 µm. b, Quantification of the filopodia density of each individual cell (N = 10 cells each). Statistics determined by two-tailed Mann-Whitney test. The bar graph displayed as mean ± SEM.
Extended Data Fig. 6 OptoATV+ could generate filopodia de novo by transporting MyoXtail to the cell border upon blue light stimulation in live cells.
a, Expression of mKate2-optoATV+ and eGFP-MyoXtail in Cos7 cells, before and after one pulse of 200 ms blue light every 10 s. de novo growth of filopodia could be readily observed after 2 minutes. The white dashed line delineates the cell border where the filopodium was grown. Representative images chosen from 3 independent experiments. Scale, 5 µm. b, Quantification of filopodia density of each cell before and after light activation (N = 10 cells each). Statistics determined by two-tailed Mann-Whitney test. The bar graph displayed as mean ± SEM.
Extended Data Fig. 7 Mouse induced neuronal (iN) cells mature within 14 days of induction.
Immunostaining of maturing iN cells at day 5 (upper lane) or mature iN cells at day 14 (lower lane). Representative images chosen from 3 independent experiments. iN cells readily express the pan-neuronal marker MAP2 at day 5, but only express SYN1 until fully mature at day 14. Scale, 100 µm (upper) or 50 µm (lower).
Extended Data Fig. 8 Electroporation is efficient in the axolotl blastema.
Electroporated axolotl blastema tissue was harvested and stained in Hoechst solution prior to mounting and confocal imaging. a, The cell nucleus is clearly stained by Hoechst and used to indicate total cell number inside the field of view. Representative image chosen from 3 independent experiments. Scale, 20 µm. b, Quantification of cell density based on either Hoechst-stained nucleus number or pmKate2-labelled cell number (N = 10 fields of view each). Approximately 65% of blastema cells were efficiently electroporated. The bar graph displayed as mean ± SEM.
Extended Data Fig. 9 The actin-based filopodial extensions in the axolotl blastema are gradually developed during regeneration.
Electroporation of the high-affinity F-actin probe utrophin calponin homology domain fused to eGFP (UCHD-eGFP) and LifeAct-iRFP into axolotl blastema. a, Live imaging of UCHD-eGFP inside axolotl blastema at 3 dpa. Representative images chosen from 3 independent experiments. The z-stack of the images are pseudocolor-coded to show the morphology at different depths. Scale, 20 µm. b, Histogram plotting the distribution of filopodia lengths at 3 dpa and 7 dpa based on the UCHD-eGFP fluorescence images (N = 63 filopodia each).
Extended Data Fig. 10 The selective defects in the regenerated posterior digits are not caused by differences in the axolotl background or the light/dark cycle.
(a) Quantification of the non-amputated right forelimb digit length of the axolotls at 35 dpa (N = 6 animals each). (b) Quantification of the regenerated left forelimb digit length ratio of the axolotls that were electroporated with pmKate2 only, with either 0 hour or 24 hours of light provided for 35 days (N = 4 animals each). Statistics determined by two-tailed Welch’s ANOVA test or two-tailed t-test. Both bar graphs displayed as mean ± SEM.
Supplementary information
Supplementary Video 1
Single molecule imaging of ATV+ in cellular extensions. Total internal reflection (TIRF) microscopy was used to record single molecule traces of sfGFP-ATV+ in a C3H/10T1/2 cell. Single particles could be observed moving towards filopodia tips, where sfGFP-ATV+ accumulated and formed bright puncta. Images were acquired every 200 ms, for 20 s. Scale, 10 µm.
Supplementary Video 2
Activation of optoATV+ in live cells. mKate2-optoATV+ was expressed in C3H/10T1/2 cells and remained inactive and evenly distributed at dark. Pulses of 200 ms blue light were applied when indicated in the video to activate optoATV+. Bright puncta of optoATV+ represented the accumulation of optoATV+ at filopodia tips post activation. The bright puncta gradually dissolved in the dark as oligomerization became reversed. Images were acquired every 1 min, for 60 min. Scale, 20 µm.
Supplementary Video 3
Activation of optoATV+ could transport sPtch1 to filopodia tips upon blue light stimulation in live cells. mKate2-optoATV+ and sPtch1-eGFP were expressed in C3H/10T1/2 cells. OptoATV+ remained inactive and sPtch1-eGFP was dominantly distributed in the cell body at dark. Pulses of 200 ms blue light were applied after every image acquisition to activate the transportation of sPtch1-eGFP by optoATV+. Bright puncta of green fluorescence represented the accumulation of transported sPtch1-eGFP at filopodia tips post activation. Images were acquired every 2 min, for 10 min. Scale, 5 µm. pmiRFP was used to visualize full cell morphology.
Supplementary Video 4
Activation of optoATV+ could generate filopodial extensions in Cos7 cells by transporting MyoXtail. mKate2-optoATV+ and MyoXtail-eGFP were expressed in Cos7 cells. Acquisition of images and activation of optoATV+ by pulses of 200 ms of blue light were both set at once every 30 s. The outgrowth of filopodial extensions could be observed after 1 min. Scale, 5 µm. pmiRFP was used to visualize full cell morphology.
Supplementary Data 1
Full nucleotide sequences for the plasmids generated in this study.
Source data
Source Data Fig. 1
Source data
Source Data Fig. 2
Source data
Source Data Fig. 3
Source data
Source Data Fig. 4
Source data
Source Data Fig. 5
Source data
Source Data Fig. 6
Source data
Source Data Fig. 7
Source data
Source Data Extended Data Fig. 1
Source data
Source Data Extended Data Fig. 2
Source data
Source Data Extended Data Fig. 3
Source data
Source Data Extended Data Fig. 4
Source data
Source Data Extended Data Fig. 5
Source data
Source Data Extended Data Fig. 6
Source data
Source Data Extended Data Fig. 8
Source data
Source Data Extended Data Fig. 9
Source data
Source Data Extended Data Fig. 10
Source data
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, Z., Denans, N., Liu, Y. et al. Optogenetic manipulation of cellular communication using engineered myosin motors. Nat Cell Biol 23, 198–208 (2021). https://doi.org/10.1038/s41556-020-00625-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-020-00625-2
This article is cited by
-
Cellular and molecular mechanisms of Hedgehog signalling
Nature Reviews Molecular Cell Biology (2023)
-
Efficient generation of functional neurons from mouse embryonic stem cells via neurogenin-2 expression
Nature Protocols (2023)
-
Optogenetic control of apical constriction induces synthetic morphogenesis in mammalian tissues
Nature Communications (2022)
-
Regulatory mechanisms of cytoneme-based morphogen transport
Cellular and Molecular Life Sciences (2022)
-
Generation of extracellular morphogen gradients: the case for diffusion
Nature Reviews Genetics (2021)