Abstract
Dystrophin proteomic regulation in muscular dystrophies (MDs) remains unclear. We report that a long noncoding RNA (lncRNA), H19, associates with dystrophin and inhibits E3-ligase-dependent polyubiquitination at Lys 3584 (referred to as Ub-DMD) and its subsequent protein degradation. In-frame deletions in BMD and a DMD non-silent mutation (C3340Y) resulted in defects in the ability of the protein to interact with H19, which caused elevated Ub-DMD levels and dystrophin degradation. Dmd C3333Y mice exhibited progressive MD, elevated serum creatine kinase, heart dilation, blood vessel irregularity and respiratory failure with concurrently reduced dystrophin and increased Ub-DMD status. H19 RNA oligonucleotides conjugated with agrin (AGR–H19) and nifenazone competed with or inhibited TRIM63. Dmd C3333Y animals, induced-pluripotent-stem-cell-derived skeletal muscle cells from patients with Becker MD and mdx mice subjected to exon skipping exhibited inhibited dystrophin degradation, preserved skeletal and cardiac muscle histology, and improved strength and heart function following AGR–H19 or nifenazone treatment. Our study paves the way for meaningful targeted therapeutics for Becker MD and for certain patients with Duchenne MD.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
Mass spectrometry data to identify DMD-binding proteins and post-translational modifications have been deposited in ProteomeXchange with the primary accession code PXD020566 (ftp://ftp.pride.ebi.ac.uk/pride/data/archive/2020/08/PXD020566). All other data supporting the findings of this study are available from the corresponding authors upon reasonable request. Source data are provided with this paper.
References
Dalkilic, I. & Kunkel, L. M. Muscular dystrophies: genes to pathogenesis. Curr. Opin. Genet. Dev. 13, 231–238 (2003).
Nowak, K. J. & Davies, K. E. Duchenne muscular dystrophy and dystrophin: pathogenesis and opportunities for treatment. EMBO Rep. 5, 872–876 (2004).
Blake, D. J., Weir, A., Newey, S. E. & Davies, K. E. Function and genetics of dystrophin and dystrophin-related proteins in muscle. Physiol. Rev. 82, 291–329 (2002).
Brooke, M. H. et al. Duchenne muscular dystrophy: patterns of clinical progression and effects of supportive therapy. Neurology 39, 475–481 (1989).
McDonald, C. M. et al. Profiles of neuromuscular diseases. Duchenne muscular dystrophy. Am. J. Phys. Med. Rehabil. 74, S70–S92 (1995).
Walcher, T. et al. Cardiac involvement in a female carrier of Duchenne muscular dystrophy. Int. J. Cardiol. 138, 302–305 (2010).
Ceulemans, B. P., Storm, K., Reyniers, E. Jr, Callewaert, L. & Martin, J. J. Muscle pain as the only presenting symptom in a girl with dystrophinopathy. Pediatr. Neurol. 38, 64–66 (2008).
Hoffman, E. P., Arahata, K., Minetti, C., Bonilla, E. & Rowland, L. P. Dystrophinopathy in isolated cases of myopathy in females. Neurology 42, 967–975 (1992).
Song, T. J., Lee, K. A., Kang, S. W., Cho, H. & Choi, Y. C. Three cases of manifesting female carriers in patients with Duchenne muscular dystrophy. Yonsei Med. J. 52, 192–195 (2011).
Echigoya, Y., Lim, K. R. Q., Nakamura, A. & Yokota, T. Multiple exon skipping in the Duchenne muscular dystrophy hot spots: prospects and challenges. J. Pers. Med 8, 41 (2018).
Torella, A. et al. One hundred twenty-one dystrophin point mutations detected from stored DNA samples by combinatorial denaturing high-performance liquid chromatography. J. Mol. Diagn. 12, 65–73 (2010).
Ehmsen, J., Poon, E. & Davies, K. The dystrophin-associated protein complex. J. Cell Sci. 115, 2801–2803 (2002).
Lin, C. & Yang, L. Long noncoding RNA in cancer: wiring signaling circuitry. Trends Cell Biol. 28, 287–301 (2018).
Brown, R. S. Zinc finger proteins: getting a grip on RNA. Curr. Opin. Struct. Biol. 15, 94–98 (2005).
Dormoy-Raclet, V. et al. The RNA-binding protein HuR promotes cell migration and cell invasion by stabilizing the β-actin mRNA in a U-rich-element-dependent manner. Mol. Cell. Biol. 27, 5365–5380 (2007).
Hu, Q. et al. Oncogenic lncRNA downregulates cancer cell antigen presentation and intrinsic tumor suppression. Nat. Immunol. 20, 835–851 (2019).
Lenk, U. et al. Non-isotopic analysis of single strand conformation polymorphism (SSCP) in the exon 13 region of the human dystrophin gene. J. Med. Genet. 30, 951–954 (1993).
Feng, J., Yan, J., Buzin, C. H., Towbin, J. A. & Sommer, S. S. Mutations in the dystrophin gene are associated with sporadic dilated cardiomyopathy. Mol. Genet. Metab. 77, 119–126 (2002).
Flanigan, K. M. et al. Rapid direct sequence analysis of the dystrophin gene. Am. J. Hum. Genet. 72, 931–939 (2003).
Goldberg, L. R. et al. A dystrophin missense mutation showing persistence of dystrophin and dystrophin-associated proteins yet a severe phenotype. Ann. Neurol. 44, 971–976 (1998).
Lenk, U. et al. A cysteine 3340 substitution in the dystroglycan-binding domain of dystrophin associated with Duchenne muscular dystrophy, mental retardation and absence of the ERG b-wave. Hum. Mol. Genet. 5, 973–975 (1996).
Campbell, K. P. & Kahl, S. D. Association of dystrophin and an integral membrane glycoprotein. Nature 338, 259–262 (1989).
Ishikawa-Sakurai, M., Yoshida, M., Imamura, M., Davies, K. E. & Ozawa, E. ZZ domain is essentially required for the physiological binding of dystrophin and utrophin to β-dystroglycan. Hum. Mol. Genet. 13, 693–702 (2004).
Abdel-Salam, E., Abdel-Meguid, I. & Korraa, S. S. Markers of degeneration and regeneration in Duchenne muscular dystrophy. Acta Myol. 28, 94–100 (2009).
Ghafoor, T., Mahmood, A. & Shams, S. Duchenne muscular dystrophy with associated growth hormone deficiency. J. Coll. Physicians Surg. Pak. 13, 722–723 (2003).
McNally, E. M. et al. Contemporary cardiac issues in Duchenne muscular dystrophy. working group of the national heart, lung, and blood institute in collaboration with parent project muscular dystrophy. Circulation 131, 1590–1598 (2015).
Ho, R., Nguyen, M. L. & Mather, P. Cardiomyopathy in Becker muscular dystrophy: overview. World J. Cardiol. 8, 356–361 (2016).
Fang, S., Jensen, J. P., Ludwig, R. L., Vousden, K. H. & Weissman, A. M. Mdm2 is a RING finger-dependent ubiquitin protein ligase for itself and p53. J. Biol. Chem. 275, 8945–8951 (2000).
Liew, C. W., Sun, H., Hunter, T. & Day, C. L. RING domain dimerization is essential for RNF4 function. Biochemical J. 431, 23–29 (2010).
Kuniyoshi, K. et al. Pivotal role of RNA-binding E3 ubiquitin ligase MEX3C in RIG-I-mediated antiviral innate immunity. Proc. Natl Acad. Sci. USA 111, 5646–5651 (2014).
Choi, Y. E. et al. The E3 ubiquitin ligase cIAP1 binds and ubiquitinates caspase-3 and -7 via unique mechanisms at distinct steps in their processing. J. Biol. Chem. 284, 12772–12782 (2009).
Morreale, F. E. & Walden, H. Types of ubiquitin ligases. Cell 165, 248–248.e1 (2016).
Talis, A. L., Huibregtse, J. M. & Howley, P. M. The role of E6AP in the regulation of p53 protein levels in human papillomavirus (HPV)-positive and HPV-negative cells. J. Biol. Chem. 273, 6439–6445 (1998).
Percherancier, Y. et al. Role of SUMO in RNF4-mediated promyelocytic leukemia protein (PML) degradation: sumoylation of PML and phospho-switch control of its SUMO binding domain dissected in living cells. J. Biol. Chem. 284, 16595–16608 (2009).
Oda, H., Kumar, S. & Howley, P. M. Regulation of the Src family tyrosine kinase Blk through E6AP-mediated ubiquitination. Proc. Natl Acad. Sci. USA 96, 9557–9562 (1999).
Wei, W., Li, M., Wang, J., Nie, F. & Li, L. The E3 ubiquitin ligase ITCH negatively regulates canonical Wnt signaling by targeting dishevelled protein. Mol. Cell. Biol. 32, 3903–3912 (2012).
Wang, D. et al. Engineering a lysosomal enzyme with a derivative of receptor-binding domain of apoE enables delivery across the blood-brain barrier. Proc. Natl Acad. Sci. USA 110, 2999–3004 (2013).
Zong, Y. et al. Structural basis of agrin–LRP4–MuSK signaling. Genes Dev. 26, 247–258 (2012).
Barik, A. et al. LRP4 is critical for neuromuscular junction maintenance. J. Neurosci. 34, 13892–13905 (2014).
Allerson, C. R. et al. Fully 2′-modified oligonucleotide duplexes with improved in vitro potency and stability compared to unmodified small interfering RNA. J. Med. Chem. 48, 901–904 (2005).
Aslesh, T., Maruyama, R. & Yokota, T. Skipping multiple exons to treat DMD—promises and challenges. Biomedicines 6, 1 (2018).
Aartsma-Rus, A. & Krieg, A. M. FDA approves eteplirsen for duchenne muscular dystrophy: the next chapter in the eteplirsen saga. Nucleic Acid Ther. 27, 1–3 (2017).
Jearawiriyapaisarn, N. et al. Sustained dystrophin expression induced by peptide-conjugated morpholino oligomers in the muscles of mdx mice. Mol. Ther. 16, 1624–1629 (2008).
Wu, B. et al. Effective rescue of dystrophin improves cardiac function in dystrophin-deficient mice by a modified morpholino oligomer. Proc. Natl Acad. Sci. USA 105, 14814–14819 (2008).
Quinlan, J. G. et al. Evolution of the mdx mouse cardiomyopathy: physiological and morphological findings. Neuromuscul. Disord. 14, 491–496 (2004).
Judge, L. M., Arnett, A. L., Banks, G. B. & Chamberlain, J. S. Expression of the dystrophin isoform Dp116 preserves functional muscle mass and extends lifespan without preventing dystrophy in severely dystrophic mice. Hum. Mol. Genet. 20, 4978–4990 (2011).
Vulin, A. et al. The ZZ domain of dystrophin in DMD: making sense of missense mutations. Hum. Mutat. 35, 257–264 (2014).
Iuchi, S. Three classes of C2H2 zinc finger proteins. Cell. Mol. Life Sci. 58, 625–635 (2001).
Tuffery-Giraud, S. et al. Genotype–phenotype analysis in 2,405 patients with a dystrophinopathy using the UMD-DMD database: a model of nationwide knowledgebase. Hum. Mutat. 30, 934–945 (2009).
Gao, Q. Q. & McNally, E. M. The dystrophin complex: structure, function, and implications for therapy. Compr. Physiol. 5, 1223–1239 (2015).
Aartsma-Rus, A. et al. Development of exon skipping therapies for Duchenne muscular dystrophy: a critical review and a perspective on the outstanding issues. Nucleic Acid Ther. 27, 251–259 (2017).
Berteaux, N. et al. H19 mRNA-like noncoding RNA promotes breast cancer cell proliferation through positive control by E2F1. J. Biol. Chem. 280, 29625–29636 (2005).
Hart, F. D. & Boardman, P. L. Trial of nifenazone (‘thylin’). BMJ 1, 1553–1554 (1964).
Wu, J. et al. A myogenic double-reporter human pluripotent stem cell line allows prospective isolation of skeletal muscle progenitors. Cell Rep. 25, 1966–1981.e4 (2018).
Chakraborty, S., Christoforou, N., Fattahi, A., Herzog, R. W. & Leong, K. W. A robust strategy for negative selection of Cre-loxP recombination-based excision of transgenes in induced pluripotent stem cells. PLoS ONE 8, e64342 (2013).
Geng, L., Zhang, H. L. & Peng, H. B. The formation of acetylcholine receptor clusters visualized with quantum dots. BMC Neurosci. 10, 80 (2009).
Yang, L. et al. ncRNA- and Pc2 methylation-dependent gene relocation between nuclear structures mediates gene activation programs. Cell 147, 773–788 (2011).
Lin, A. et al. The LINK-A lncRNA interacts with PtdIns(3,4,5)P3 to hyperactivate AKT and confer resistance to AKT inhibitors. Nat. Cell Biol. 19, 238–251 (2017).
Yoon, J. H., Srikantan, S. & Gorospe, M. MS2-TRAP (MS2-tagged RNA affinity purification): tagging RNA to identify associated miRNAs. Methods 58, 81–87 (2012).
Aartsma-Rus, A. & van Putten, M. Assessing functional performance in the mdx mouse model. J. Vis. Exp. 27, 51303 (2014).
Hoffman, E. & Winder, S. J. A modified wire hanging apparatus for small animal muscle function testing. PLoS Curr. https://doi.org/10.1371/currents.md.1e2bec4e78697b7b0ff80ea25a1d38be (2016).
Conner, J. D., Wolden-Hanson, T. & Quinn, L. S. Assessment of murine exercise endurance without the use of a shock grid: an alternative to forced exercise. J. Vis. Exp. 14, e51846 (2014).
Hamer, P. W., McGeachie, J. M., Davies, M. J. & Grounds, M. D. Evans Blue dye as an in vivo marker of myofibre damage: optimising parameters for detecting initial myofibre membrane permeability. J. Anat. 200, 69–79 (2002).
Weyers, J. J., Carlson, D. D., Murry, C. E., Schwartz, S. M. & Mahoney, W. M. Jr. Retrograde perfusion and filling of mouse coronary vasculature as preparation for micro computed tomography imaging. J. Vis. Exp. 10, e3740 (2012).
Li, C. et al. A ROR1–HER3–lncRNA signalling axis modulates the Hippo–YAP pathway to regulate bone metastasis. Nat. Cell Biol. 19, 106–119 (2017).
Acknowledgements
We would like to thank the Baylor College of Medicine Human Stem Cell Core (HSCC) for technical help and consultation for iPSC line derivation, maintenance and differentiation. The HSCC is supported in part by Shared Resource funding from the NIH (CA125123, PI: Osborne). The generation of knock-in mice was supported by CCSG grant NCI number CA016672 (GEMF) of the MD Anderson Cancer Center. The Proteomics and Metabolomics Facility was supported in part by Cancer Prevention Research Institute of Texas (CPRIT) grant number RP130397 and NIH grant numbers 1S10OD012304-01 and P30CA016672. This work was partially supported by the Texas A&M University Chancellor’s Research Initiative. This work was partially supported by the National Institute of Arthritis and Musculoskeletal and Skin Diseases (NIAMS) of the NIH under award numbers 1R01AR068293 and 1R21AR071583 to R.D. This work was partially supported by the Cancer Prevention and Research Institute of Texas (RR150085) and CPRIT Scholar in Cancer Research (to L. Han). This work was supported by NIH R01 awards (1 R01 CA218025-01, 1R01CA231011-01), a CPRIT individual investigator research award (180259) and a DoD Breakthrough award (BC180196) to C.L., and a NIH R01 award (1 R01 CA218036-01) and DoD Breakthrough awards (BC151465, BC181384), a CPRIT individual investigator research award (200423) and a Andrew Sabin Family Foundation Fellows award to L.Y.
Author information
Authors and Affiliations
Contributions
L.Y. and C.L. conceived the project and designed the experiments. Y.Z. executed the primary studies. Y.Z. and Y.L. developed the genetic mouse models and related experiments with assistance from Q.H, Z.X. and T.K.N. Z. Zhang and L. Han performed bioinformatics analysis with assistance from J.L. and D.S. Histological staining and corresponding analyses were performed by Y.Z. and K.L. D.H.H. executed the mass spectrometry analysis. CLIP assays were performed by L.Y. The hiPSCs were generated by J.J.K. and P.Z., and hiPSC culture was performed by Y.Z. with assistance from L. Huang, Y.X., C.S., Z. Zhao, J.W. and R.D. J.C., M.-C.H., A.F. and R.D. contributed to experimental design and data interpretation. S.D.E. assisted with manuscript drafting. Y.Z., L.Y. and C.L. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 H19 interacts with dystrophin.
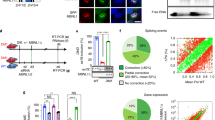
a, CLIP assays using mouse skeletal muscle tissues, followed by autoradiography. Protein-bound RNA (blue box) were extracted for Sanger sequencing. b, RIP assay using mouse skeletal muscle tissue. Mean values±SD, n = 3 independent experiments, two-way ANOVA. c, Representative images of immune RNA Fluorescence In Situ Hybridization in mouse skeletal and cardiac muscle tissues. Data represents three independent experiments. Scale bars, 50 μm. d, Streptavidin pull-down assay using indicated recombinant protein and biotinylated-H19, followed by immunoblotting (IB). Streptavidin (Strep.)-HRP indicated the presence of biotinylated-H19 transcripts. e, Detection of H19 depletion in C2C12 cells. f, RT-qPCR detection of indicated genes in C2C12 H19-proficient or -KO cell line. Mean values±SD, n = 4 independent experiments, one-way ANOVA. g and h, RT-qPCR detection (g) or IB detection (h) in Dmd-proficient or -KO C2C12 cell lines. Mean values±SD (g), n = 4 independent experiments, one-way ANOVA. i, H19 Copy number determination in iPS-derived cardiomyocytes (iPS-CMs) using qPCR. Mean values±SD, n = 5 independent experiments, one-way ANOVA. j, Representative image of immunofluorescence staining (IF) of iPS-SkMCs and cardiomyocytes (iPS-CMs) derived from healthy donor (GM09503), upon H19 depletion. Scale bars, 50 μm. k and l, IB detection of indicated proteins (k) or RIP assay (l) in H19-proficient or deficient iPS-SkMCs expressing indicated expression vectors. Mean values±SD (l), n = 3 independent experiments, two-way ANOVA. m, H19 RNA copy number determination in H19-proficient or deficient iPS-CMs expressing indicated expression vectors. Mean values±SD, n = 6 independent experiments, one-way ANOVA. n, Annotated MS/MS spectrum assigned to the dystrophin peptide with ubiquitin modification at Lys 3577. No significance [n.s.], p > 0.05, *, p < 0.05, **, p < 0.01, ***, p < 0.001. Autoradiography and immunoblots are representative of two independent experiments. Statistical source data and unprocessed immunoblots are provided as Source Data Extended Data Fig. 1.
Extended Data Fig. 2 H19 antagonizes poly-ubiquitination and protein degradation of DMD.
a, Blocking Peptide Competition Assay using cell lysates extracted from H19-proficient, or -deficient iPS-SkMCs and blocking peptides of Scramble, TPSN, Ub-TPSN, DMD and Ub-DMD as shown, followed with IB detection using indicated antibodies. MG132 was used as proteasome inhibitor. b, Autoradiography of immunoprecipitated dystrophin in H19-proficient or -deficient iPS-CMs stably expressing MS2-tagged H19 WT or indicated mutants followed by L-[35S]-Methionine pulse-chase. Input was subjected to IB detection using indicated antibody. c, IB detection of indicated proteins in H19-proficient or deficient hiPS-SkMCs expressing MS2-tagged H19 WT or indicated mutants. d, IB detection of indicated proteins in H19-proficient or deficient C2C12 differentiated myotubes with or without MG-132. e, IB detection of DMD in Dmd-proficient or deficient C2C12 differentiated myotubes expressing DMD WT or the indicated mutants with PS-341. f, RT-qPCR detection of H19 relative expression level in Dmd-proficient or -deficient C2C12-differentiated myotubes expressing DMD WT or indicated mutants. Mean values±SD, n = 5 independent experiments, one-way ANOVA. g, and h, RT-qPCR detection of Dmd (g) and H19 (h) relative expression level in Dmd-proficient or –deficient C2C12 expressing DMD or H19 WT or indicated mutants. Mean values±SD, n = 4 independent experiments, two-way ANOVA. i, IB detection using indicated antibodies of Ni-NTA pulldown using recombinant proteins as indicated. No significance [n.s.], p > 0.05, *, p < 0.05, **, p < 0.01, ***, p < 0.001, and ****, p < 0.0001. Immunoblots are representative of two independent experiments. Statistical source data and unprocessed immunoblots are provided as Source Data Extended Data Fig. 2.
Extended Data Fig. 3 H19 competes with TRIM63 in interacting with dystrophin.
a, Immunoprecipitation followed with IB detection using indicated antibodies of hiPS-SkMCs derived from healthy donor (GM09503) expressing DMD WT or indicated mutations. b, RIP assay and RT-QPCR detection of indicated genes of hiPS-SkMC cells derived from healthy donor (GM09503) expressing DMD WT or indicated mutations. Mean values±SD, n = 3 independent experiments, two-way ANOVA. c, IB detection using indicated antibodies of indicated recombinant proteins in the presence of biotinylated H19 RNA transcript. The LINK-A RNA transcript were included as negative control. d, Schematic illustration of predict secondary structure of Scr (scramble RNA) mimic, H19 RNA mimic and H19 mimic carrying mutation (H19 mut). e, IB detection using indicated antibodies of indicated recombinant proteins in the presence of biotinylated scramble (Scr), or H19 RNA mimics. f and g, Representative images of genotyping (f) and Sanger sequencing analysis (g) of Dmd in Dmd WT, Dmd C3333Y and mdx mice. h and i, Body weight measurement of female (h) or male (i) Dmd WT, Dmd WT/C3333Y and Dmd C3333Y/C3333Y mice. Mean values±SD, n = 5 mice in each group, two-way ANOVA. j, H&E staining representative images of the lung, liver, kidney and spleen tissues in Dmd WT and Dmd C3333Y mice. Scale bars, 100 μm. Data represent independent replicates using 8 animals per group. k, Creatine kinase (CK) concentration in female Dmd WT, Dmd WT/C3333Y and Dmd C3333Y/C3333Y mice serum was tested of 4, 12 and 24-week-old, respectively. Mean values±SD, n = 5 mice in each group, two-way ANOVA. No significance [n.s.], p > 0.05, *, p < 0.05, **, p < 0.01, ***, p < 0.001, and ****, p < 0.0001. Immunoblots are representative of two independent experiments. Statistical source data and unprocessed immunoblots are provided as Source Data Extended Data Fig. 3.
Extended Data Fig. 4 Dmd C3333Y mouse models muscular dystrophy.
a, RT-qPCR detection of indicated genes in indicated female mice. Mean values±SD, n = 5 mice per group, two-way ANOVA. b, Mouse cytokine array assay using the serum of indicated mice. Data represent independent replicates using 3 animals per group c and d, Serum concentration of indicated cytokines in indicated mice. Mean values±SEM, n = 3 (c) or 5 (d) mice per group, unpaired student’s t-test. e, H&E staining representative images of Gastrocnemius, Quadriceps, or Tibialis Anterior in indicated female mice at age of 12 weeks. Scale bars, 100 μm. Data represent independent replicates using 8 animals per group. f, Transmission electron microscope representative images of tibialis anterior (TA) and cardiac muscles. Scale bars, 500 nm. Data represent independent replicates using 5 animals per group. g, Running distance and exhaustion time of indicated mice. Mean values±SD, Female: n = 8, 15, 13; Male: n = 7, 6, one-way ANOVA. h, Representative images of micro CT of mouse spine (left) and kyphotic index (right) of male indicated mice. Mean values±SEM, n = 6 per group unpaired student’s t-test. i, H&E staining representative images of 46-week-old indicated lung. Scale bars, 1 mm or 100 μm. Data represent independent replicates using 8 animals per group. j, Food intake of male indicated mice. Mean values±SD, n = 4 animals in each group, unpaired student’s t-test. k, VO2 (top) and VCO2 (bottom) consumption of male indicated mice. Mean values±SEM, n = 6 animals in each group, two-way ANOVA. l and m, Urinary albumin to creatinine ratio (ACR), blood urea nitrogen (BUN) and serum creatinine measurement of female (left) and male (right) indicated mice. Mean values±SD, Female: n = 8, 9; Male: n = 8, 6 for WT and C3333Y, respectively, unpaired student’s t-test. No significance [n.s.], p > 0.05, *, p < 0.05, **, p < 0.01, ***, p < 0.001, and ****, p < 0.0001. Statistical source data are provided as Source Data Extended Data Fig. 4.
Extended Data Fig. 5 Dmd C3333Y mutation facilitates poly-ubiquitination and protein degradation of DMD.
a, IF staining representative images using indicated antibodies in QUA (quadriceps) of Dmd C3333Y and mdx mice at indicated age. Scale bars, 100 μm. Data represent independent replicates using 5 animals per group. b, c, IF staining representative images (b) and staining intensity statistics (c) in QUA using DMD and Ub-DMD (Lys3584) antibodies of Dmd WT or Dmd C3333Y mice at indicated age. Scale bars (left), 100 μm. Mean values±SD (right), n = 8 mice in each group, two-way ANOVA. d, Colonies karyotyping representative images of hiPS cells derived from human fibroblast cells of indicated donors. Healthy donor: GM09503; BMD patients: GM04981, GM02298, GM05089, GM04569. Data represent independent replicates using 3 clones per cell line. e, Flow cytometer verification of human fibroblast cells derived hiPS cells by using OCT4 antibody. Data represent independent replicates using 3 clones per cell line. ****, p < 0.0001. Statistical source data are provided as Source Data Extended Data Fig. 5.
Extended Data Fig. 6 H19 RNA mimics and Nifenazone promote the protein stability of dystrophin.
a, IP and IB detection using indicated antibodies in healthy or BMD iPS-SkMCs. b, IP and IB detection using indicated antibodies in healthy or BMD iPS-SkMCs treated with H19 mimics, wild type or mut. c, Log of TRIM63 expression profile in different human tissues. d, Heat map of sandwich ELISA using His6-TUBE and Ub-DMD (Lys3584) antibody for screening customized compound library and the generation of the Ub-DMD was detected by OD450. Log2 of relative fold change of polyUb chain formation activity was shown. e, Competition curve determination of IC50 value of NIF on the enzymatic activity of indicated E3 ligases. The IC50 values of NIF against each E3 ligase are shown. Mean values±SD; n = 3 independent experiments. f, Left: IB detection using indicated antibodies of hiPS-SkMCs upon vehicle (Veh.) or NIF treatment. Right: list of E3 ligase and known cellular substrates as shown. g, Autoradiography of immunoprecipitated dystrophin in iPS-CMs derived from healthy donor or BMD patient stably expressing MS2-tagged H19 or treated with vehicle or Nifenazon (10 µM) followed by L-[35S]-Methionine pulse-chase. Input was subjected to IB detection using indicated antibody. Immunoblots are representative of two independent experiments. Statistical source data and unprocessed immunoblots are provided as Source Data Extended Data Fig. 6.
Extended Data Fig. 7 H19 and Nifenazone extend the half-life of DMD protein.
a, Representative images of IF staining using indicated antibody (top) and statistical analysis of DMD staining intensities (bottom) of hiPS-CMs derived from healthy donor or BMD patients, upon transfection of Scramble (Scr) mimic, H19 mimic or treatment Nifenazone (10 µM). Scale bars, 50 µm. Mean values±SD, n = 5 independent experiments, two-way ANOVA. b, Representative images of IF staining using indicated antibody (top) and statistical analysis of Ub-DMD (Lys3584) staining intensities (bottom) of hiPS-CMs derived from healthy donor or BMD patients, upon transfection of Scramble (Scr) mimic, H19 mimic or treatment Nifenazone (10 µM). Scale bars, 50 µm. Mean values±SD, n = 5 independent experiments, two-way ANOVA. c, IP using DMD antibody followed by IB detection using indicated antibodies in hiPS-SkMCs from healthy donor or BMD patients with the indicated treatments. d–g, RT-QCPR detection of indicated genes of hiPS-SkMCs derived from healthy donor or BMD patients, upon transfection of Scramble (Scr) mimic, H19 mimic or treatment Nifenazone (10 µM). Mean values±SD, n = 5 independent experiments, two-way ANOVA. No significance [n.s.], p > 0.05, *, p < 0.05, **, p < 0.01, ***, p < 0.001, ****, p < 0.0001. Immunoblots are representative of two independent experiments. Statistical source data and unprocessed immunoblots are provided as Source Data Extended Data Fig. 7.
Extended Data Fig. 8 DMD K3584R resists TRIM63-dependent polyUb and protein degradation.
a, Graphic illustration of the strategies used to generate knockin cell lines derived from hiPS cells. b, EGFP expression of hiPS cells upon electroporation. Scale bars, 50 µm. Data represent 3 independent experiments. c, DNA agarose gel detection of single-cell colonies derived from 04981 or 25313 using indicated genotyping primers. Green forward: Puro-GT-R(LJL); Green reverse: PM20003-A-R-GT-R; Red forward: PM20003-A-L-GT-F; Red reverse: Puro-F. d, DNA agarose gel detection of single-cell colonies derived from 04981 or 25313 respectively, following removal of resistance cassettes, using indicated genotyping primers. Blue forward: PM20003-A-L-GT-F; Blue reverse: PM20003-A-L-GT-R3. *: mixed colonies containing cells with resistance cassettes not removed. Yellow boxes: single cell colonies used for following studies. e, Sanger sequencing validation of DMD K3584R mutant. Data represent 3 independent experiments. f, IB detection using indicated antibodies of GM04981 parental (Par) or K3584R mutant cells in the presence of indicated treatment. MG132 was used as proteasome inhibitor. g-i, Concentration measurement of H19 mimic or AGR-H19 per milligram of tissue at indicated time points post i.p. administration of H19 mimics (g), SubQ administration of H19 mimics (h), or SubQ administration of AGR-H19 (i) (10 mg/kg, single dose). Mean values±SD, n = 5 animals per time point per experimental condition. j, Left, representative images of α-bungarotoxin staining (top) or Duolink PLA (proximity ligation assay) using antibodies targeting LPR4 and Biotin respectively (bottom) of hiPS-SkMCs with indicated treatment. rAgrin, recombinant Agrin (Z+ form, aa. 1260–2045). Scale bars, 50 µm. Right: Percentage of cells with AchR clustering (blue bars) or Duolink PLA (red bars) of hiPS-SkMCs with indicated treatment. Mean values±SD, n = 6 independent experiments, two-way ANOVA, ***, p < 0.001. DNA agarose gel and immunoblots are representative of two independent experiments. Statistical source data and unprocessed immunoblots are provided as Source Data Extended Data Fig. 8.
Extended Data Fig. 9 AGR-H19 and Nifenazone restore skeletal muscle and heart function.
a, Immunohistological detection of Biotin-labeled mimics in quadriceps, heart, kidney, liver and lung tissues in the presence of indicated mimics. Scale bars, 100 μm. Data represent independent replicates using 5 animals per group. b, Body weight measurement of male Dmd WT treated with scramble (Scr) or Dmd C3333Y mice with indicated treatments. Mean values±SD, n = 5 mice per group, two-way ANOVA. c, d, Serum IGF1 (c) and IGF2 (d) level concentration of Dmd WT or Dmd C3333Y mice with the indicated treatments. Mean values±SD, n = 5 mice per group, one-way ANOVA. e, RT-qPCR detection of Dmd in Dmd WT or Dmd C3333Y QUA (quadriceps) with indicated treatments. Mean values±SD, n = 6 mice in each group, one-way ANOVA. f-i, RT-qPCR detection of indicated genes in Dmd C3333Y QUA with indicated treatments. Mean values±SD, n = 5 (f, g, h) or 6 (i) mice in each group, one-way ANOVA. j, Serum CK concentration in Dmd WT treated with AGR-Scr or Dmd C3333Y mice with indicated treatments. Mean values±SD, n = 5 mice per group, one-way ANOVA. k, Percentage of central nucleic of indicated muscle fiber of Dmd WT treated with AGR-Scr or Dmd C3333Y mice with indicated treatments. Mean values±SD, n = 5 mice per group, two-way ANOVA. l, Body weight measurement of male Dmd WT mice treated with vehicle or Dmd C3333Y mice with indicated treatments from 4 weeks old to age of 16 weeks. Mean values±SD, n = 5 mice per group, two-way ANOVA. m, Serum CK concentration in Dmd WT mice treated with vehicle or Dmd C3333Y mice with indicated treatments. Mean values±SD, n = 5 mice per group, one-way ANOVA. No significance [n.s.], p > 0.05, *, p < 0.05, **, p < 0.01, ***, p < 0.001, and ****, p < 0.0001. Statistical source data are provided as Source Data Extended Data Fig. 9.
Extended Data Fig. 10 AGR-H19 and Nifenazone improve the protein stability of dystrophin following exon-skipping.
a, IP and IB detection using indicated antibodies of hiPS-SkMCs derived from healthy or DMD donor in the presence of biotinylated Scr or H19 mimics. b, Representative images of IF staining of hiPS-CMs derived from healthy donor treated with scramble and vehicle, or DMD patient (GM25313) treated with h44AON1 alone or in combination with H19 mimics or Nifenazone respectively. Scale bars, 100 μm. c, Statistical analysis of DMD (top) or Ub-DMD (Lys3584) (bottom) of hiPS-CMs derived from healthy donor (GM09503) treated with scramble and vehicle, or DMD patient (GM25313) treated with h44AON1 alone or in combination with H19 mimics or Nifenazone respectively. Mean values±SD. n = 5 independent experiments in each group, one-way ANOVA. d, IB detection using indicated antibodies of hiPS-SkMCs derived from healthy donor (H. 09503) or DMD patient (25313) in the presence of h44AON1 and/or indicated treatment. e, IB detection using indicated antibodies of GM25313 parental (Par) or K3584R mutant cells in the presence of control (Ctrl.) or TRIM63 siRNA. hiPS-SkMCs derived from healthy donor (H., GM09503) were included as control. f, IB detection using indicated antibodies of GM25313 parental (Par) or K3584R mutant cells in the presence of control (Ctrl.) or TRIM63 siRNA. hiPS-SkMCs derived from healthy donor (H., GM09503) were included as control. MG132 was used as proteasome inhibitor. No significance [n.s.], p > 0.05, *, p < 0.05, **, p < 0.01, ***, p < 0.001. Immunoblots are representative of two independent experiments. Statistical source data and unprocessed immunoblots are provided as Source Data Extended Data Fig. 10.
Supplementary information
Supplementary Information
Supplementary Figs. 1–5.
Supplementary Tables
Supplementary Table 1 clinical information of hiPSCs from patients with BMD used in this study. Basic clinical parameters of human skeletal muscle tissues and hiPSCs derived from healthy donors, patients with BMD and patients with DMD. Supplementary Table 2: summary of CLIP assay. Summary of Sanger sequencing of CLIP assay results. CLIP assays of the dystrophin α-domain were performed in both fresh frozen human and mouse skeletal muscle tissues, followed by Sanger sequencing. This table contains two tabs: tab 1: Sanger sequencing of CLIP assay using human skeletal muscle tissue tab 2: Sanger sequencing of CLIP assay using mouse skeletal muscle tissue. Supplementary Table 3: summary of DMD-binding proteins and post-translational modifications. In vitro mouse DMD protein pulldown using H19 WT or H19 KO C2C12 cells, followed by liquid chromatography–mass spectrometry. Mouse IgG was used as the negative control. This table contains four tabs: tab 1: mouse IgG control in H19 WT C2C12 cells; tab 2: anti-DMD antibody in H19 WT C2C12 cells; tab 3: mouse IgG control in H19 KO C2C12 cells; tab 4: anti-DMD antibody in H19 KO C2C12 cells. Supplementary Table 4: characterization of Dmd C3333Y animal fertility. Comparison of animal fertility, C57BL/6N male mice with or without the C3333Y mutation on the X chromosome were crossed with heterozygous C3333Y female mice by natural mating at 6–8 weeks old. The fertility rate experiments were repeated three times. Each time we used eight males and eight females, and one male and one female were mated per cage. Data are presented as the mean ± s.d., unpaired Student’s t-test. Supplementary Table 5: characterization of the Dmd C3333Y animal genotype, gender numbers and ratios. Mendelian ratios and mice number comparison of the indicated genotypes and genders. One C57BL/6N heterozygous C3333Y male mouse and one female heterozygous mouse were mated per cage, and we used eight pairs in each experiment, and the experiment was repeated a total of three times. Data are presented as the mean ± s.d., χ2 test. Supplementary Table 6: small-molecule inhibitor screen. Supplementary Table 7: list of compounds. Supplementary Table 8: blood chemistry of Dmd C3333Y animals subjected to AGR–H19 or NIF treatment. The Clinical Pathology Workgroup at the MD Anderson Cancer Center measured all blood chemistry. Serum was collected from mice by cardiac puncture after fasting for 4–6 h. Mice serum albumin, alkaline phosphatase, BUN, calcium, chloride, globulin, potassium, sodium and total protein values are given as the mean ± s.d., n = 5 mice in each group. Unpaired Student’s t-test. This table contains two tabs: tab 1: blood chemistry analysis of Dmd C3333Y animal subjected to AGR–H19 treatment; tab 2: blood chemistry analysis of Dmd C3333Y animals subjected to NIF treatment. Supplementary Table 9: oligonucleotides used in this study. Supplementary Table 10: antibodies used in this study.
Supplementary Video 1
Video of muscle incoordination of Dmd C3333Y mice.
Supplementary Video 2
Video of muscle tremor of Dmd C3333Y mice.
Supplementary Video 3
Hanging test of Dmd WT mice.
Supplementary Video 4
Hanging test of Dmd C3333Y mice.
Supplementary Video 5
Video of in vivo ECG of Dmd WT and Dmd C3333Y mice. Surface lead II ECG recordings were performed in conscious and freely mobile mice that were 40 weeks old. The upper part of the video recorded the ECG of a Dmd WT mouse, and the lower part recorded the ECG of a Dmd C3333Y mouse.
Source data
Source Data Fig. 1
Source data for Fig. 1c,f,g,i.
Source Data Fig. 1
Full gels for Fig. 1a,d,e,j.
Source Data Fig. 2
Source data for Fig. 2a,c,d,h.
Source Data Fig. 2
Full gels for Fig. 2b,e,f.
Source Data Fig. 3
Source data for Fig. 3b,c,f,g,h.
Source Data Fig. 4
Source data for Fig. 4b,c,e,g–k.
Source Data Fig. 5
Source data for Fig. 5b,d,f,g.
Source Data Fig. 5
Full gels for Fig. 5a,c,e.
Source Data Fig. 6
Source data for Fig. 6c,d–i.
Source Data Fig. 7
Source data for Fig. 7b,c–g.
Source Data Fig. 8
Source data for Fig. 8b,c,e–g.
Source Data Extended Data Fig. 1
Source data for Extended Data Fig. 1b,f–i,l,m.
Source Data Extended Data Fig. 1
Full gels for Extended Data Fig. 1a,d,h,k.
Source Data Extended Data Fig. 2
Source data for Extended Data Fig. 2f,g,h.
Source Data Extended Data Fig. 2
Full gels for Extended Data Fig. 2a–e,i.
Source Data Extended Data Fig. 3
Source data for Extended Data Fig. 3b,h,i,k.
Source Data Extended Data Fig. 3
Full gels for Extended Data Fig. 3a,c,e.
Source Data Extended Data Fig. 4
Source data for Extended Data Fig. 4a,c,d,g,h,j–m.
Source Data Extended Data Fig. 5
Source data for Extended Data Fig. 5c.
Source Data Extended Data Fig. 6
Source data for Extended Data Fig. 6c–e.
Source Data Extended Data Fig. 6
Full gel for Extended Data Fig. 6a,b,f,g.
Source Data Extended Data Fig. 7
Source data for Extended Data Fig. 7a,b,d–g.
Source Data Extended Data Fig. 7
Full gel for Extended Data Fig. 7c.
Source Data Extended Data Fig. 8
Source data for Extended Data Fig. 8g–I.
Source Data Extended Data Fig. 8
Full gel for Extended Data Fig. 8f.
Source Data Extended Data Fig. 9
Source data for Extended Data Fig. 9b,9c–l.
Source Data Extended Data Fig. 10
Source data for Extended Data Fig. 10c.
Source Data Extended Data Fig. 10
Full gel for Extended Data Fig. 10a,d–f.
Rights and permissions
About this article
Cite this article
Zhang, Y., Li, Y., Hu, Q. et al. The lncRNA H19 alleviates muscular dystrophy by stabilizing dystrophin. Nat Cell Biol 22, 1332–1345 (2020). https://doi.org/10.1038/s41556-020-00595-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-020-00595-5