Abstract
Tumour growth and invasiveness require extracellular matrix (ECM) degradation and are stimulated by the GALA pathway, which induces protein O-glycosylation in the endoplasmic reticulum (ER). ECM degradation requires metalloproteases, but whether other enzymes are required is unclear. Here, we show that GALA induces the glycosylation of the ER-resident calnexin (Cnx) in breast and liver cancer. Glycosylated Cnx and its partner ERp57 are trafficked to invadosomes, which are sites of ECM degradation. We find that disulfide bridges are abundant in connective and liver ECM. Cell surface Cnx–ERp57 complexes reduce these extracellular disulfide bonds and are essential for ECM degradation. In vivo, liver cancer cells but not hepatocytes display cell surface Cnx. Liver tumour growth and lung metastasis of breast and liver cancer cells are inhibited by anti-Cnx antibodies. These findings uncover a moonlighting function of Cnx–ERp57 at the cell surface that is essential for ECM breakdown and tumour development.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All data supporting the findings of this study are available from the corresponding author upon reasonable request. The mass spectrometry data are available via ProteomeXchange with the identifier PXD021202. Source data are provided with this paper.
References
Nissen, R., Cardinale, G. J. & Udenfriend, S. Increased turnover of arterial collagen in hypertensive rats. Proc. Natl Acad. Sci. USA 75, 451–453 (1978).
Jabłońska-Trypuć, A., Matejczyk, M. & Rosochacki, S. Matrix metalloproteinases (MMPs), the main extracellular matrix (ECM) enzymes in collagen degradation, as a target for anticancer drugs. J. Enzyme Inhib. Med. Chem. 31, 177–183 (2016).
Hotary, K., Allen, E., Punturieri, A., Yana, I. & Weiss, S. J. Regulation of cell invasion and morphogenesis in a three-dimensional type I collagen matrix by membrane-type matrix metalloproteinases 1, 2, and 3. J. Cell Biol. 149, 1309–1323 (2000).
Hotary, K. B. et al. Membrane type I matrix metalloproteinase usurps tumor growth control imposed by the three-dimensional extracellular matrix. Cell 114, 33–45 (2003).
Nguyen, A. T. et al. Organelle specific O-glycosylation drives MMP14 activation, tumor growth, and metastasis. Cancer Cell 32, 639–653.e6 (2017).
Lu, P., Takai, K., Weaver, V. M. & Werb, Z. Extracellular matrix degradation and remodeling in development and disease. Cold Spring Harb. Perspect. Biol. 3, a005058 (2011).
Bonnans, C., Chou, J. & Werb, Z. Remodelling the extracellular matrix in development and disease. Nat. Rev. Mol. Cell Biol. 15, 786–801 (2014).
Poincloux, R., Lizárraga, F. & Chavrier, P. Matrix invasion by tumour cells: a focus on MT1-MMP trafficking to invadopodia. J. Cell Sci. 122, 3015–3024 (2009).
Linder, S., Wiesner, C. & Himmel, M. Degrading devices: invadosomes in proteolytic cell invasion. Annu. Rev. Cell Dev. Biol. 27, 185–211 (2011).
Paterson, E. K. & Courtneidge, S. A. Invadosomes are coming: new insights into function and disease relevance. FEBS J. 285, 8–27 (2018).
Murphy, D. A. & Courtneidge, S. A. The ‘ins’ and ‘outs’ of podosomes and invadopodia: characteristics, formation and function. Nat. Rev. Mol. Cell Biol. 12, 413–426 (2011).
Destaing, O., Saltel, F., Géminard, J.-C., Jurdic, P. & Bard, F. Podosomes display actin turnover and dynamic self-organization in osteoclasts expressing actin–green fluorescent protein. Mol. Biol. Cell 14, 407–416 (2003).
Gill, D. J., Chia, J., Senewiratne, J. & Bard, F. Regulation of O-glycosylation through Golgi-to-ER relocation of initiation enzymes. J. Cell Biol. 189, 843–858 (2010).
Gill, D. J. et al. Initiation of GalNAc-type O-glycosylation in the endoplasmic reticulum promotes cancer cell invasiveness. Proc. Natl Acad. Sci. USA 110, E3152–E3161 (2013).
Bard, F. & Chia, J. Cracking the glycome encoder: signaling, trafficking, and glycosylation. Trends Cell Biol. 26, 379–388 (2016).
Chia, J., Tay, F. & Bard, F. The GalNAc-T activation (GALA) pathway: drivers and markers. PLoS ONE 14, e0214118 (2019).
Springer, G. F. T and Tn, general carcinoma autoantigens. Science 224, 1198–1206 (1984).
Ju, T., Otto, V. I. & Cummings, R. D. The Tn antigen-structural simplicity and biological complexity. Angew. Chem. Int. Ed. 50, 1770–1791 (2011).
Ju, T. et al. Tn and sialyl-Tn antigens, aberrant O-glycomics as human disease markers. Proteomics Clin. Appl. 7, 618–631 (2013).
Chia, J., Goh, G. & Bard, F. Short O-GalNAc glycans: regulation and role in tumor development and clinical perspectives. Biochim. Biophys. Acta 1860, 1623–1639 (2016).
Chia, J., Tham, K. M., Gill, D. J., Bard-Chapeau, E. A. & Bard, F. A. ERK8 is a negative regulator of O-GalNAc glycosylation and cell migration. eLife 3, e01828 (2014).
Naba, A. et al. The extracellular matrix: tools and insights for the ‘omics’ era. Matrix Biol. 49, 10–24 (2016).
Herchenhan, A. et al. Lysyl oxidase activity is required for ordered collagen fibrillogenesis by tendon cells. J. Biol. Chem. 290, 16440–16450 (2015).
Weadock, K. S., Miller, E. J., Keuffel, E. L. & Dunn, M. G. Effect of physical crosslinking methods on collagen-fiber durability in proteolytic solutions. J. Biomed. Mater. Res. 32, 221–226 (1996).
Boudko, S. P. & Bachinger, H. P. The von Willebrand Factor A3 domain binding region of type III collagen stabilized by the cysteine knot. RCSB Protein Data Bank https://doi.org/10.2210/pdb4gyx/pdb (2012).
Barth, D. et al. The role of cystine knots in collagen folding and stability, part I. Conformational properties of (Pro-Hyp-Gly)5 and (Pro-(4S)-FPro-Gly)5 model trimers with an artificial cystine knot. Chemistry 9, 3692–3702 (2003).
Khoshnoodi, J., Pedchenko, V. & Hudson, B. G. Mammalian collagen IV. Microsc. Res. Tech. 71, 357–370 (2008).
Engel, J. et al. Structure and macromolecular organization of type VI collagen. Ann. NY Acad. Sci. 460, 25–37 (1985).
Birk, D. E. Type V collagen: heterotypic type I/V collagen interactions in the regulation of fibril assembly. Micron 32, 223–237 (2001).
Ilani, T. et al. A secreted disulfide catalyst controls extracellular matrix composition and function. Science 341, 74–76 (2013).
Feige, M. J. & Hendershot, L. M. Disulfide bonds in ER protein folding and homeostasis. Curr. Opin. Cell Biol. 23, 167–175 (2011).
Bulleid, N. J. Disulfide bond formation in the mammalian endoplasmic reticulum. Cold Spring Harb. Perspect. Biol. 4, a013219 (2012).
Oka, O. B. V. & Bulleid, N. J. Forming disulfides in the endoplasmic reticulum. Biochim. Biophys. Acta 1833, 2425–2429 (2013).
Oliver, J. D., Roderick, H. L., Llewellyn, D. H. & High, S. ERp57 functions as a subunit of specific complexes formed with the ER lectins calreticulin and calnexin. Mol. Biol. Cell 10, 2573–2582 (1999).
Frickel, E.-M. et al. ERp57 is a multifunctional thiol-disulfide oxidoreductase. J. Biol. Chem. 279, 18277–18287 (2004).
Jessop, C. E. & Bulleid, N. J. Glutathione directly reduces oxidoreductasesin the endoplasmic reticulum of mammalian cells. J. Biol. Chem. 279, 55341–55347 (2004).
Sefried, S., Häring, H.-U., Weigert, C. & Eckstein, S. S. Suitability of hepatocyte cell lines HepG2, AML12 and THLE-2 for investigation of insulin signalling and hepatokine gene expression. Open Biol. 8, 180147 (2018).
Nakabayashi, H., Taketa, K., Miyano, K., Yamane, T. & Sato, J. Growth of human hepatoma cells lines with differentiated functions in chemically defined medium. Cancer Res. 42, 3858–3863 (1982).
Juin, A. et al. Discoidin domain receptor 1 controls linear invadosome formation via a Cdc42–Tuba pathway. J. Cell Biol. 207, 517–533 (2014).
Steentoft, C. et al. Precision mapping of the human O-GalNAc glycoproteome through SimpleCell technology. EMBO J. 32, 1478–1488 (2013).
Jessop, C. E. et al. ERp57 is essential for efficient folding of glycoproteins sharing common structural domains. EMBO J. 26, 28–40 (2007).
Holbrook, L.-M. et al. OX133, a monoclonal antibody recognizing protein-bound N-ethylmaleimide for the identification of reduced disulfide bonds in proteins. mAbs 8, 672–677 (2016).
Barth, D., Kyrieleis, O., Frank, S., Renner, C. & Moroder, L. The role of cystine knots in collagen folding and stability, part II. Conformational properties of (Pro-Hyp-Gly)n model trimers with N- and C-terminal collagen type III cystine knots. Chemistry 9, 3703–3714 (2003).
Pankov, R. & Yamada, K. M. Fibronectin at a glance. J. Cell Sci. 115, 3861–3863 (2002).
Williams, D. B. Beyond lectins: the calnexin/calreticulin chaperone system of the endoplasmic reticulum. J. Cell Sci. 119, 615–623 (2006).
Fregno, I. & Molinari, M. Proteasomal and lysosomal clearance of faulty secretory proteins: ER-associated degradation (ERAD) and ER-to-lysosome-associated degradation (ERLAD) pathways. Crit. Rev. Biochem. Mol. Biol. 54, 153–163 (2019).
Lakkaraju, A. K. K. & van der Goot, F. G. Calnexin controls the STAT3-mediated transcriptional response to EGF. Mol. Cell 51, 386–396 (2013).
Van Duyn Graham, L., Sweetwyne, M. T., Pallero, M. A. & Murphy-Ullrich, J. E. Intracellular calreticulin regulates multiple steps in fibrillar collagen expression, trafficking, and processing into the extracellular matrix. J. Biol. Chem. 285, 7067–7078 (2010).
Fregno, I. et al. ER-to-lysosome-associated degradation of proteasome-resistant ATZ polymers occurs via receptor-mediated vesicular transport. EMBO J. 37, e99259 (2018).
Chakravarthi, S., Jessop, C. E. & Bulleid, N. J. The role of glutathione in disulphide bond formation and endoplasmic‐reticulum‐generated oxidative stress. EMBO Rep. 7, 271–275 (2006).
Chakravarthi, S. & Bulleid, N. J. Glutathione is required to regulate the formation of native disulfide bonds within proteins entering the secretory pathway. J. Biol. Chem. 279, 39872–39879 (2004).
Franco, R., Schoneveld, O. J., Pappa, A. & Panayiotidis, M. I. The central role of glutathione in the pathophysiology of human diseases. Arch. Physiol. Biochem. 113, 234–258 (2007).
Okazaki, Y., Ohno, H., Takase, K., Ochiai, T. & Saito, T. Cell surface expression of calnexin, a molecular chaperone in the endoplasmic reticulum. J. Biol. Chem. 275, 35751–35758 (2000).
Holbrook, L.-M. et al. The platelet-surface thiol isomerase enzyme ERp57 modulates platelet function. J. Thromb. Haemost. 10, 278–288 (2012).
Obeid, M. ERP57 membrane translocation dictates the immunogenicity of tumor cell death by controlling the membrane translocation of calreticulin. J. Immunol. 181, 2533–2543 (2008).
Dihazi, H. et al. Secretion of ERP57 is important for extracellular matrix accumulation and progression of renal fibrosis, and is an early sign of disease onset. J. Cell Sci. 126, 3649–3663 (2013).
Kessenbrock, K., Plaks, V. & Werb, Z. Matrix metalloproteinases: regulators of the tumor microenvironment. Cell 141, 52–67 (2010).
Courtneidge, S. A., Azucena, E. F., Pass, I., Seals, D. F. & Tesfay, L. The Src substrate Tks5, podosomes (invadopodia), and cancer cell invasion. Cold Spring Harbor Symp. Quant. Biol. 70, 167–171 (2005).
Boersema, P. J., Raijmakers, R., Lemeer, S., Mohammed, S. & Heck, A. J. R. Multiplex peptide stable isotope dimethyl labeling for quantitative proteomics. Nat. Protoc. 4, 484–494 (2009).
Rappsilber, J., Mann, M. & Ishihama, Y. Protocol for micro-purification, enrichment, pre-fractionation and storage of peptides for proteomics using StageTips. Nat. Protoc. 2, 1896–1906 (2007).
Steentoft, C. et al. Mining the O-glycoproteome using zinc-finger nuclease-glycoengineered SimpleCell lines. Nat. Methods 8, 977–982 (2011).
Carlson, C. M., Frandsen, J. L., Kirchhof, N., McIvor, R. S. & Largaespada, D. A. Somatic integration of an oncogene-harboring Sleeping Beauty transposon models liver tumor development in the mouse. Proc. Natl Acad. Sci. USA 102, 17059–17064 (2005).
Acknowledgements
We thank D. Fass and M. Molinari for insightful discussions about disulfide bonds, Cnx and ERp57, and G. Van der Goot for plasmids and advice. We also thank the Advanced Molecular Pathology Laboratory at the IMCB for mouse histology services and the Bordeaux Imaging Center (BIC) for their support in this study. This work was supported by A*STAR, Cancer Research UK (grant C13329/A21671 to M.J.H.), La Fondation pour la Recherche Médicale (grant number DEQ20180839586) and Integrated Cancer Research (SIRIC, BRIO).
Author information
Authors and Affiliations
Contributions
M.R. designed and performed experiments. J.C., S.V. and H.C. contributed to the analyses of Cnx glycosylation and glycosylation sites. A.T.N. performed most of the mouse experiments, and S.L.T. and X.L.G. performed the ECM characterization and degradation experiments. R.M. and M.J.H. participated in the initial characterization of the function of Cnx glycosylation. M.J.H. contributed to the manuscript and mentoring of R.M., and F.S. contributed to discussion, experimental design and mentoring of M.R. F.A.B. designed experiments and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Cnx glycosylation and translocation.
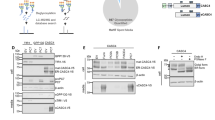
a, Immunoblot analysis of the level of Tn-modified Cnx in human liver tumours (T) versus the corresponding normal liver tissues (NT) from 20 liver cancer patients. b, Quantification of calnexin staining colocalizing with cortactin staining in NIH3T3vSrc cells. c, Immunofluorescence of MDA ER-G2 cells stained with calnexin and cortactin. Scale bar, 20μm. Source data and unmodified blots of Extended Data Fig. 1 can be found in files named Source Data Extended Data Fig. 1.
Extended Data Fig. 2 Cnx and ERp57 are essential for ECM degradation.
a, Schematic of matrix degradation assay. b, Relative intensity of degraded collagen antibody staining after incubation with NIH 3T3vSrc control cells and NIH 3T3vSrc Cnx KD cells. Values indicate the mean ± SEM for 3 replicates and were analyzed using a two-tailed Student’s t-test. Representatives images can be found on the right. Scale bar, 20 μm. siRNA efficacy and its quantification can be found below. c, Workflow on ImageJ software to quantify degradation per nuclei. d, Quantification of ECM degradation assay of NIH3T3vSrc cells incubated with control antibody (IgG control) or 2 different anti-Cnx antibodies. Values indicate the mean ± SEM of normalized fold change for 3 replicates and were analyzed using a two-tailed Student’s t-test. e, Western blot analysis and quantification of cnx siRNA associated to Fig. 2d. Values were analyzed using a two-tailed Student’s t-test. f, Quantification of ECM degradation assay of NIH3T3vSrc cells transfected with control siRNA (siNT5) or siRNA against Cnx (siCnx). Values indicate the mean ± SEM of normalized fold changes for 3 replicates and were analyzed using a two-tailed Student’s t-test. Western blot and quantification of siRNA efficacy can be found on the right. g, Immunoblot analysis of MMPs protein level in NIH3T3vSrc transfected with control siRNA (siNT5) or siRNA against Cnx (siCnx). siRNA efficacy can be found on the right (this quantification is for Fig S2G, S2H and S2I, the experiments were done with the same samples). Values indicate the mean ± SEM of 3 replicates and were analyzed using a two-tailed Student’s t-test. h, Quantification of MMP activity by FRET assay of MDA ER-G2 cells transfected with control siRNA (siNT5) or siRNA against cnx (siCnx). Quantification of siRNA efficacy can be found on Supp Fig. 2g. i, Immunofluorescence of NIH3T3vSrc cells transfected with control siRNA (siNT5) or siRNA against Cnx (siCnx) and stained with MMP14 and the invadosome marker cortactin. Scale bar, 5 μm. Quantification of siRNA efficacy can be found on Supp Fig. 2g. j, Western blot and quantification of siRNA in Fig. 2h. Values indicate the mean ± SEM of 3 replicates and were analyzed using a two-tailed Student’s t-test. Source data and unmodified blots of Extended Data Fig. 2 can be found in files named Source Data Extended Data 2.
Extended Data Fig. 3 Disulfide bonds are abundant in liver ECM and reduced by Cnx/ERp57.
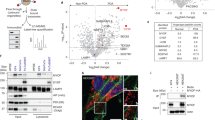
a, Immunofluorescence of collagens I and III and OX133 on decellularized liver, untreated or treated with TCEP and NEM. Scale bar, 50 μm. b, Immunofluorescence of collagen I and fibronectin on decellularized liver, untreated or treated with TCEP and NEM. Scale bar, 50 μm. c, Immunofluorescence of rat tail ECM, treated or non-treated with TCEP and NEM and stained with OX133 antibody. Scale bar, 20 μm. d, Western blot and quantification of siRNA efficacy associated to Fig. 4d. Values represent the mean ± SEM of 2 replicates. e, Immunofluorescence of collagen incubated with NIH3T3vSrc (not shown), with or without NEM, and stained with ERp57 antibody. Collagen was previously coupled to 5-carboxy-X-rhodamin succinimidyl ester before cell seeding. Scale bar, 10 μm. f, Quantification of ECM degradation assay of NIH3T3vSrc cells incubated with GSH (+GSH) or with GPx and H2O2 (+GPx). Values indicate the mean ± SEM of normalized fold changes for 3 and replicates respectively and were analyzed using a two-tailed Student’s t-test. g, Western blot and quantification of siRNA efficacy associated to Fig. 4f. Values represent the mean ± SEM of 3 replicates and were analyzed using a two-tailed Student’s t-test. h, Quantification of ECM degradation assay of MDA ER-G2 transfected with control siRNA or 2 different ERp57 siRNA (siERp57 #1 and #2). Collagen disulfide bonds were chemically reduced using TCEP (+TCEP) or left untreated (- TCEP) before cell seeding. Values indicate the mean ± SEM of normalized fold changes for 3 replicates and were analyzed using a two-tailed Student’s t-test. Western blot and quantification of siRNA efficacy can be found on the right. Source data and unmodified blots of Extended Data Fig. 3 can be found in files named Source Data Extended Data Fig. 3.
Extended Data Fig. 4 Cnx is essential for tumour growth.
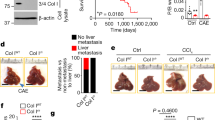
a, Schematic diagram of experiments using a mouse model of breast cancer metastasis to the lung to test anti-calnexin antibody. b, Schematic diagram of the workflow of the calnexin antibody treatment in a mouse model for liver cancer and its lung metastasis.
Supplementary information
Supplementary Table 1
Table containing patient data.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 1
Unprocessed western blots.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 3
Unprocessed western blots.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 1
Unprocessed western blots.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 2
Unprocessed western blots.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 3
Unprocessed western blots.
Rights and permissions
About this article
Cite this article
Ros, M., Nguyen, A.T., Chia, J. et al. ER-resident oxidoreductases are glycosylated and trafficked to the cell surface to promote matrix degradation by tumour cells. Nat Cell Biol 22, 1371–1381 (2020). https://doi.org/10.1038/s41556-020-00590-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-020-00590-w
This article is cited by
-
Crosstalk between KDEL receptor and EGF receptor mediates cell proliferation and migration via STAT3 signaling
Cell Communication and Signaling (2024)
-
Mechanisms and roles of podosomes and invadopodia
Nature Reviews Molecular Cell Biology (2023)