Abstract
Defining the ontogeny of the human adaptive immune system during embryogenesis has implications for understanding childhood diseases including leukaemias and autoimmune conditions. Using RAG1:GFP human pluripotent stem cell reporter lines, we examined human T-cell genesis from pluripotent-stem-cell-derived haematopoietic organoids. Under conditions favouring T-cell development, RAG1+ cells progressively upregulated a cohort of recognized T-cell-associated genes, arresting development at the CD4+CD8+ stage. Sort and re-culture experiments showed that early RAG1+ cells also possessed B-cell, myeloid and erythroid potential. Flow cytometry and single-cell-RNA-sequencing data showed that early RAG1+ cells co-expressed the endothelial/haematopoietic progenitor markers CD34, VECAD and CD90, whereas imaging studies identified RAG1+ cells within CD31+ endothelial structures that co-expressed SOX17+ or the endothelial marker CAV1. Collectively, these observations provide evidence for a wave of human T-cell development that originates directly from haemogenic endothelium via a RAG1+ intermediate with multilineage potential.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
All scRNA-seq and bulk RNA-seq data used in this study have been deposited in the Gene Expression Omnibus (GEO) database repository and are available under accession numbers GSE124172 (scRNA-seq) and GSE124173 (bulk RNAseq). All other data supporting the findings of this study are available from the corresponding author on reasonable request.
Code availability
All computational code used for the analysis in this manuscript is available from the corresponding author upon reasonable request.
References
Lin, Y., Yoder, M. C. & Yoshimoto, M. Lymphoid progenitor emergence in the murine embryo and yolk sac precedes stem cell detection. Stem Cells Dev. 23, 1168–1177 (2014).
Tian, Y. et al. The first wave of T lymphopoiesis in zebrafish arises from aorta endothelium independent of hematopoietic stem cells. J. Exp. Med. 214, 3347–3360 (2017).
Luis, T. C. et al. Initial seeding of the embryonic thymus by immune-restricted lympho-myeloid progenitors. Nat. Immunol. 17, 1424–1435 (2016).
Boiers, C. et al. Lymphomyeloid contribution of an immune-restricted progenitor emerging prior to definitive hematopoietic stem cells. Cell Stem Cell 13, 535–548 (2013).
Boiers, C. et al. A human IPS model implicates embryonic B-myeloid fate restriction as developmental susceptibility to B acute lymphoblastic leukemia-associated ETV6-RUNX1. Dev. Cell 44, 362–377 (2018).
Ivanovs, A. et al. Highly potent human hematopoietic stem cells first emerge in the intraembryonic aorta-gonad-mesonephros region. J. Exp. Med. 208, 2417–2427 (2011).
Bertrand, J. Y. & Traver, D. Hematopoietic cell development in the zebrafish embryo. Curr. Opin. Hematol. 16, 243–248 (2009).
Chen, A. T. & Zon, L. I. Zebrafish blood stem cells. J. Cell. Biochem. 108, 35–42 (2009).
Medvinsky, A. & Dzierzak, E. Definitive hematopoiesis is autonomously initiated by the AGM region. Cell 86, 897–906 (1996).
Yoshimoto, M. et al. Autonomous murine T-cell progenitor production in the extra-embryonic yolk sac before HSC emergence. Blood 119, 5706–5714 (2012).
Yoshimoto, M. et al. Embryonic day 9 yolk sac and intra-embryonic hemogenic endothelium independently generate a B-1 and marginal zone progenitor lacking B-2 potential. Proc. Natl Acad. Sci. USA 108, 1468–1473 (2011).
Godin, I., Dieterlen-Lièvre, F. & Cumano, A. Emergence of multipotent hemopoietic cells in the yolk sac and paraaortic splanchnopleura in mouse embryos, beginning at 8.5 days postcoitus. Proc. Natl Acad. Sci. USA 92, 773–777 (1995).
Nishikawa, S. I. et al. In vitro generation of lymphohematopoietic cells from endothelial cells purified from murine embryos. Immunity 8, 761–769 (1998).
Kawamoto, H., Ikawa, T., Ohmura, K., Fujimoto, S. & Katsura, Y. T cell progenitors emerge earlier than B cell progenitors in the murine fetal liver. Immunity 12, 441–450 (2000).
Ikawa, T. et al. Identification of the earliest prethymic T-cell progenitors in murine fetal blood. Blood 103, 530–537 (2004).
Benz, C. & Bleul, C. C. A multipotent precursor in the thymus maps to the branching point of the T versus B lineage decision. J. Exp. Med. 202, 21–31 (2005).
Yokota, T. et al. Tracing the first waves of lymphopoiesis in mice. Development 133, 2041–2051 (2006).
Mombaerts, P. et al. RAG-1-deficient mice have no mature B and T lymphocytes. Cell 68, 869–877 (1992).
Sobacchi, C., Marrella, V., Rucci, F., Vezzoni, P. & Villa, A. RAG-dependent primary immunodeficiencies. Hum. Mutat. 27, 1174–1184 (2006).
Ivanovs, A. et al. Human haematopoietic stem cell development: from the embryo to the dish. Development 144, 2323–2337 (2017).
Tavian, M., Biasch, K., Sinka, L., Vallet, J. & Peault, B. Embryonic origin of human hematopoiesis. Int. J. Dev. Biol. 54, 1061–1065 (2010).
Ditadi, A. et al. Human definitive haemogenic endothelium and arterial vascular endothelium represent distinct lineages. Nat. Cell Biol. 17, 580–591 (2015).
Sturgeon, C. M., Ditadi, A., Awong, G., Kennedy, M. & Keller, G. Wnt signaling controls the specification of definitive and primitive hematopoiesis from human pluripotent stem cells. Nat. Biotechnol. 32, 554–561 (2014).
Kennedy, M. et al. T lymphocyte potential marks the emergence of definitive hematopoietic progenitors in human pluripotent stem cell differentiation cultures. Cell Rep. 2, 1722–1735 (2012).
Mohtashami, M. et al. Direct comparison of Dll1- and Dll4-mediated Notch activation levels shows differential lymphomyeloid lineage commitment outcomes. J. Immunol. 185, 867–876 (2010).
Carpenter, L. et al. Human induced pluripotent stem cells are capable of B-cell lymphopoiesis. Blood 117, 4008–4011 (2011).
Ng, E. S. et al. Differentiation of human embryonic stem cells to HOXA+ hemogenic vasculature that resembles the aorta-gonad-mesonephros. Nat. Biotechnol. 34, 1168–1179 (2016).
Bertrand, J. Y., Cisson, J. L., Stachura, D. L. & Traver, D. Notch signaling distinguishes 2 waves of definitive hematopoiesis in the zebrafish embryo. Blood 115, 2777–2783 (2010).
Hadland, B. K. et al. A requirement for Notch1 distinguishes 2 phases of definitive hematopoiesis during development. Blood 104, 3097–3105 (2004).
Kumano, K. et al. Notch1 but not Notch2 is essential for generating hematopoietic stem cells from endothelial cells. Immunity 18, 699–711 (2003).
Schulz, T. C. et al. A scalable system for production of functional pancreatic progenitors from human embryonic stem cells. PLoS ONE 7, e37004 (2012).
Taoudi, S. et al. Extensive hematopoietic stem cell generation in the AGM region via maturation of VE-cadherin+CD45+ pre-definitive HSCs. Cell Stem Cell 3, 99–108 (2008).
Skelton, R. J. et al. CD13 and ROR2 permit isolation of highly enriched cardiac mesoderm from differentiating human embryonic stem cells. Stem Cell Rep. 6, 95–108 (2016).
Gama-Norton, L. et al. Notch signal strength controls cell fate in the haemogenic endothelium. Nat. Commun. 6, 8510 (2015).
Park, S. H. et al. HLA-DR expression in human fetal thymocytes. Hum. Immunol. 33, 294–298 (1992).
Melichar, H. J., Ross, J. O., Taylor, K. T. & Robey, E. A. Stable interactions and sustained TCR signaling characterize thymocyte–thymocyte interactions that support negative selection. J. Immunol. 194, 1057–1061 (2015).
Lancaster, J. N., Li, Y. & Ehrlich, L. I. R. Chemokine-mediated choreography of thymocyte development and selection. Trends Immunol. 39, 86–98 (2018).
Zlotoff, D. A. et al. CCR7 and CCR9 together recruit hematopoietic progenitors to the adult thymus. Blood 115, 1897–1905 (2010).
Plotkin, J., Prockop, S. E., Lepique, A. & Petrie, H. T. Critical role for CXCR4 signaling in progenitor localization and T cell differentiation in the postnatal thymus. J. Immunol. 171, 4521–4527 (2003).
Koenen, P. et al. Mutually exclusive regulation of T cell survival by IL-7R and antigen receptor-induced signals. Nat. Commun. 4, 1735 (2013).
Yokota, T. et al. Unique properties of fetal lymphoid progenitors identified according to RAG1 gene expression. Immunity 19, 365–375 (2003).
van Dijk, D. et al. Recovering gene interactions from single-cell data using data diffusion. Cell 174, 716–729 (2018).
Trapnell, C. et al. The dynamics and regulators of cell fate decisions are revealed by pseudotemporal ordering of single cells. Nat. Biotechnol. 32, 381–386 (2014).
Kuwata, N., Igarashi, H., Ohmura, T., Aizawa, S. & Sakaguchi, N. Cutting edge: absence of expression of RAG1 in peritoneal B-1 cells detected by knocking into RAG1 locus with green fluorescent protein gene. J. Immunol. 163, 6355–6359 (1999).
Igarashi, H. et al. Localization of recombination activating gene 1/green fluorescent protein (RAG1/GFP) expression in secondary lymphoid organs after immunization with T-dependent antigens in rag1/gfp knockin mice. Blood 97, 2680–2687 (2001).
Tavian, M., Robin, C., Coulombel, L. & Peault, B. The human embryo, but not its yolk sac, generates lympho-myeloid stem cells: mapping multipotent hematopoietic cell fate in intraembryonic mesoderm. Immunity 15, 487–495 (2001).
Klug, D. B. et al. Interdependence of cortical thymic epithelial cell differentiation and T-lineage commitment. Proc. Natl Acad. Sci. USA 95, 11822–11827 (1998).
Hess, I. & Boehm, T. Intravital imaging of thymopoiesis reveals dynamic lympho-epithelial interactions. Immunity 36, 298–309 (2012).
Costa, M. et al. A method for genetic modification of human embryonic stem cells using electroporation. Nat. Protoc. 2, 792–796 (2007).
Kao, T. et al. GAPTrap: a simple expression system for pluripotent stem cells and their derivatives. Stem Cell Rep. 7, 518–526 (2016).
Thomson, J. A. et al. Embryonic stem cell lines derived from human blastocysts. Science 282, 1145–1147 (1998).
Davis, R. P. et al. A protocol for removal of antibiotic resistance cassettes from human embryonic stem cells genetically modified by homologous recombination or transgenesis. Nat. Protoc. 3, 1550–1558 (2008).
Mohtashami, M., Shah, D. K., Kianizad, K., Awong, G. & Zúñiga-Pflücker, J. C. Induction of T-cell development by Delta-like 4-expressing fibroblasts. Int. Immunol. 25, 601–611 (2013).
McCarthy, D. J., Chen, Y. & Smyth, G. K. Differential expression analysis of multifactor RNA-Seq experiments with respect to biological variation. Nucleic Acids Res. 40, 4288–4297 (2012).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Acknowledgements
We would like to thank the members of the MCRI Flow cytometry support team for their assistance with cell sorting and microscopy and S. Chen for her assistance with the preparation of the art work. This study was funded by the National Health and Medical Research Council of Australia through research fellowships awarded to A.G.E. (grant no. GNT1117596) and E.G.S (grant no. GNT1079004), project grants awarded to A.G.E. and E.G.S. (grant nos GNT1068866, GNT1129861 and GNT1138717) as well as A.P.C. and E.G.S (grant no. GNT1123277), the Australian Research Council Special Research Initiative in Stem Cells (Stem Cells Australia), The Children’s Cancer Research Foundation (Australia) and the Stafford Fox Medical Research Foundation. Additional infrastructure funding to the Murdoch Children’s Research Institute was provided by the Australian Government National Health and Medical Research Council Independent Research Institute Infrastructure Support Scheme and the Victorian Government’s Operational Infrastructure Support Program.
Author information
Authors and Affiliations
Contributions
Conceptualization: A.M., A.G.E., E.G.S.; data curation: T.C.; formal analysis: A.M., F.F.B., T.C.; funding acquisition: A.P.C., C.A.W., A.G.E., E.G.S.; investigation: A.M., F.F.B., S.V.K., J.V.S., C.A.W.; methodology: A.M., F.F.B., E.S.N.; project administration: A.M., A.G.E., E.G.S.; resources: A.P.C.; supervision: A.G.E., E.G.S.; visualization: A.M., F.F.B., A.G.E., E.G.S.; writing—original draft: A.M., E.G.S.; writing—review and editing: A.M., A.P.C., C.A.W., A.G.E., E.G.S.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Characterization of RAG1GFP/w PSC reporter lines.
a, Schematic of the RAG1 locus showing the non-coding exon 1 and the coding exon 2. The cut site of RAG1 specific TALENs used to enhance gene targeting efficiency is indicated by a vertical blue arrowhead. The position of primer set 1 used to screen for clones that retained an intact RAG1 locus is indicated. The region of the native RAG1 locus in RAG1GFP/w PSC lines that was sequenced is indicated by the red dashed line. b, Schematic view of the RAG1 locus after insertion of a cassette encoding GFP and a loxP (red triangles) flanked selectable marker (PGK-Puro). Grey dashed lines denote sequences corresponding to those included in the targeting vector. c, Schematic view of the final targeted RAG1 locus. Primer set 3 was used to verify excision of sequences flanked by LoxP sites. d, Agarose gels showing the results of PCR analysis of genomic DNA representing individual H9 hESC and iPSC RM3.5 clones. As depicted in (a) and (b), primer set 1 amplified a DNA fragment of 2.2 Kb whereas primer set 2 amplified a DNA fragment of 2.3 Kb. Clone numbers corresponding to each PSC line are indicated. e, The loxP flanked selectable marker cassette was removed from the two heterozygous clones used in this study, H9 #36 and iPSC #13. Primer set 3 amplified a DNA fragment of 600 base pairs. control DNA derived from a targeted clone prior to the removal of the PGK-Puro cassette. f, Bright field-GFP fluorescence image showing individual GFP + human T-lymphoid progenitors. g, Histogram showing the results of real-time PCR indicating that RAG1 expression is restricted to the GFP + population. The Y axis shows the relative expression of RAG1 in arbitrary units from a single experiment. Results are derived from a single cell sorting experiment (n = 1). Scale bar = 20 μM.
Extended Data Fig. 2 Characterization of the haematopoietic organoid differentiation system.
a, Flow cytometry data from iPSC and hESC lines showing the expression of the key indicated marker genes between differentiation days 4 and 16. b, Bright-field image of embryoid bodies at differentiation day 8 and fluorescent images of RAG1+(GFP + ) cells within CD31+vascular networks of differentiation day 24 organoids. c, Flow cytometry plots showing the relationship between RAG1 expression levels and progression to the CD4+CD8+ stage. d, Flow cytometry analysis showing CD10+CD19 + RAG1+B-cell progenitors in organoid cultures treated with the NOTCH signalling inhibitor, DAPT. e, Flow cytometry analysis quantifying the proportion of CD56 + CD7+ cells (presumptive NK cells) in cultures supplemented with IL2 and IL15. f, Gating strategy used for isolation of cell populations profiled in Fig. 3. g, Relationship between GFP expression levels and specific T-cell differentiation stages. The frequency of cells with a given level of GFP fluorescence representing specific fractions is shown on the y-axis. Coloured lines with within histogram plots correspond to colours within the summary data shown in (h). h, Summary of data derived of 8 separate cell sorting experiments that contributed to the data presented in Fig. 3. Graphs show mean + /- SD for n = 8 differentiation experiments. i, Heat map representation of the expression levels of genes encoding markers used for isolation of specific differentiation stages of differentiation by FACS (see Fig. 3): All negative (AN), RAG1 negative (RN), Double negative (DN), Immature Single Positive (ISP) and Double Positive (DP). The row z-score is an indication of the degree of deviation from the mean for each row. j, The relationship between the number of reads mapping to the targeted RAG1 locus containing the GFP gene, the unmodified RAG1 locus and the RAG2 locus for the indicated fractions. RNAseq data corresponds to that analysed in Fig. 3. n = 3 or 4, as indicated, samples derived from independent cell sorting experiments (see Source Data for individual values. Because the y-axis is a log scale, samples with no reads are shown as yellow circles below each bar. Note that samples designated as RAG1- generally contained fewer that 10 (101) reads. For all flow cytometry plots, except for panels (d) and (f), numbers on the x and y axis are the exponent of log10 fluorescence. For panel d, the numbers for both x and y axis are the exponent of log10 Biexponential scaling from FlowJoⓇ flow cytometry software, where the (-) indicates -10. The percentage of cells in each quadrant or associated with each gate is indicated. All cells displayed as green are RAG1+. Scale bars for b, from left to right are 200, 600 and 500 μm respectively.
Extended Data Fig. 3 Characterization of FACS purified subfractions of RAG1+Cellfy.
a, Representative gating strategy used for isolation of cell populations profiled in Figs. 4, 5 and Extended Data Fig. 3. b, Flow cytometry plots showing the 2 most disparate examples of CD34 and CD90 expression in cells isolated for experiments that form part of the analysis presented in Fig. 4. In these examples the frequency of CD34+CD90+ cells varied from 0 to 10% of the RAG1+ population. c, d, Relationship between RNA sequencing reads for GFP, RAG1 and RAG2 across cell populations purified on the basis of GFP expression. c, Histograms summarizing the number of reads for the indicated genes derived from cell sorting experiments that contributed to data in Fig. 4. For iPSCs, n = 3 or 4, as indicated, For hESCs, n = 1 or as indicated. See source data for details. Because the y-axis is a log scale, samples with no reads are shown underneath each bar as a yellow point. d, Trend line representation of the relationship between the of number GFP and RAG1 reads for each sample across all experiments depicted in C above and supplementary figure 2j. n = 38 independent samples derived from cell sorting experiments The equation representing the line of best fit is shown, as is the correlation coefficient for this line (R2 = 0.69). e, Heat map representation of mean expression levels of 3′ HOXA genes (A9-A13) across all samples from the indicated fractions for experiments described in Fig. 4.
Extended Data Fig. 4 TCR gene expression of PSC organoid derived T-cell progenitors.
a, Graphical summary of the level of gene expression associated with specific TCR loci (see colour key) as indicated. The sorting parameters for individual populations are indicated across the X-axis, and the level of expression for each gene is given as RPKM on the Y-axis. Plots for both the iPSC and hESC based RAG1 reporter cells are shown. The mean expression level for each gene is shown, and where three or more independent samples were collected, error bars represent the SEM. Dots represent data from independent experiments and interconnecting the lines intersect the mean value for the indicated markers for each sorting parameter. For the iPSC and hESC based lines, the number of samples assayed for each fraction were as follows: CD34+CD90+RAG1- (n = 3, n = 1), RAG1+(n = 4, n = 2), CD5 + CD7+CD4-CD8-RAG1+(n = 4, n = 4), CD4+CD8-RAG1+(n = 4, n = 4), CD4+CD8+RAG1+(n = 4, n = 4). b, Heat map showing the expression levels of specific TCR genes representing the alpha, beta, gamma and delta loci. Genes included had more than 1 RPKM in 2 different samples. The samples generated using either the iPSC or hESC based reporter lines are indicated with open and filled circles respectively. As indicated by the Row Z-score colour key, genes more highly expressed are shown in red.
Extended Data Fig. 5 Potential of RAG1+ cells derived from the hESC RAG1:GFP reporter line.
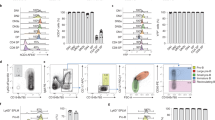
a, Methylcellulose colony forming assays for each of the indicated fractions. The number of samples contributing to data for each fraction was RAG1+(n = 3), CD90+CD34+RAG1- (n = 3) and CD90-CD34+RAG1- (n = 2). The mean number of colonies is shown, with error bars representing the s.e.m. The open circles indicate the data values that contributed to the calculation of the mean for each fraction. b, Flow cytometry analysis of methylcellulose cultures from the indicated fractions showing the presence of CD235a + (Glycophorin A) erythroid progenitors and CD14+and/or CD16 + myeloid lineage cells. c, Flow cytometry analysis of CD45 + blood cells derived from OP9 stromal cultures seeded with organoid derived progenitors for the fractions indicated. Robust numbers of the CD14+myeloid cells are observed in cultures derived from either the RAG1+or CD34+CD90+RAG- fractions. A small number of CD19 + B-lymphoid cells are also observed in these cultures, but not in those seeded with CD34+CD90-RAG1- cells. For all flow cytometry plots using antibody stains, numbers on the x and y axis are the exponent of log10 fluorescence. The percentage of cells in each quadrant or associated with each gate is indicated. All cells displayed as green are RAG1+.
Extended Data Fig. 6 Single cell sequencing analysis of emergent RAG1+ cells.
Heat map showing gene expression levels for key genes in individual cells used in the construction of the tSNE plots shown in Fig. 5a, b, with nominal assignment of lineage associations for each set indicated on the right of the map. The 9 subpopulations (0–8) are indicated across the top of the heat map (Clusters).
Extended Data Fig. 7 Gene expression analysis of early RAG1+ cells using single cell RNAseq.
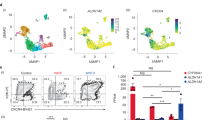
a, tSNE plot of the data shown in Fig. 5a without prior transformation using MAGIC42 (van Dijk, D et al 2018), showing that similar populations are present in both forms of the plot. b, tSNE plot summarizing the results of a second independent single cell RNAseq experiment (without prior transformation of the data using MAGIC). For both (a) and (b), an assignment of the identity of each group is suggested, along with a single exemplary gene for that cluster which formed part of the set used to assign that cluster. Note that numbering of clusters is based on their relative size, with the cluster designated as “0” containing the largest number of cells. The colouring of clusters was not specified such that similar clusters in each experiment share similar colours. However, the relative position of clusters representing the endothelial/haematopoietic progenitors and T-lineage cells are shared between the two experiments, as indicated by the transparent overlaid transparent grey arrow. c, tSNE plots of four genes illustrating the transition of cells from CDH5 + (VE-CAD + ) CLDN5 + endothelial/haematopoietic progenitor cells to GATA3 + IL7R + T-cell progenitors in Experiment 2. d, Violin plots indicating the relative level of endothelial/haematopoietic progenitor and T-cell progenitor gene expression associated with each cell in each of the fractions as indicated for Experiment 2. The y-axis shows the expression level (lnTPM + 1). e, Violin plots displaying the relative levels of expression of the haematopoietic progenitor and myeloid marker G-CSFR and the myeloid marker SPI1 across the CD45 + (PTPRC) blood-cell fractions. f, Violin plots showing the expression of the stromal marker (COL1A1) and erythroid genes (GATA1 and HBD) in the minor contaminating populations, 9 and 10. For d, e and f, dots represent individual data points (single cells), with the position on the y-axis representing the relative expression of the indicated gene in that cell. N numbers in d-f represent the individual cells (data points) with a given expression value for the indicated gene and are shown as individual data points in the plots.
Extended Data Fig. 8 Flow cytometry and immunofluorescence of early RAG1+ cells.
a, b, Flow cytometry plots showing that a proportion of emergent hESC-derived RAG1+ cells co-express CD90 and CD34 (a) but, in contrast to iPSC RAG1+ cells, only a small number of VE-CAD + cells retain CD90 expression (b). c, Flow cytometry plot showing a proportion of early RAG1+ cells are CD5-CD7-. For a, b and c, the number (a:136, b:141, c:59) and position of cells falling within the RAG1+gate that were used to generate accompanying contour plots is indicated. The scale numbers for both x and y axis are the exponent of log10 Biexponential scaling from FlowJoⓇ flow cytometry software, where the (-) indicates -10. d, e, Confocal immunofluorescence analysis showing the expression of RAG1, VE-CAD and CAV1 (d), and RAG1 and GATA3 (e) on cells localized or confined to defined structures. The presence of rare RAG1+ cells that co-express CAV1 and VE-CAD (marked by the arrow in d) is consistent with the initiation of RAG1 expression within cells that form part of the vascular structure. Nuclei were identified by staining with DAPI. Scale bars = 20 μm.
Supplementary information
Supplementary Video 1
Z-stack of confocal sections of haematopoietic organoids labelled with antibodies against GFP (RAG1, green), VE-CAD (white) and CAV1 (red) showing a group of GFP+ cells within a VE−CAD+CAV1+ structure. In this Z-stack, no co-localization of GFP with the other markers was discernable. Nuclei were visualized by labelling with DAPI.
Supplementary Video 2
Z-stack of confocal sections of haematopoietic organoids labelled with antibodies against GFP (RAG1, green), VE-CAD (white) and CAV1 (red) showing two cells in which expression of these markers seems to co-localize. Nuclei were visualized by labelling with DAPI. This Z-stack covers the area presented in Fig. 7c.
Supplementary Video 3
Z-stack of confocal sections of haematopoietic organoids labelled with antibodies against SOX17 (red) and GFP (RAG1, green) showing the co-localization of GFP expression with SOX17 in a subset of cells. Nuclei were visualized by labelling with DAPI. This Z-stack covers the area presented in Fig. 7d.
Supplementary Video 4
Time-lapse video of iPSC RAG1GFP/w reporter line-derived haematopoietic organoids showing the migration of RAG1+ (GFP, green) cells within CD31+ (red) vascular-like structures.
Supplementary Video 5
Time-lapse video of hESC RAG1GFP/w reporter line-derived haematopoietic organoids showing the migration of RAG1+ (GFP, green) cells within CD31+ (red) vascular-like structures.
Supplementary Video 6
Time-lapse video of iPSC RAG1GFP/w reporter line-derived haematopoietic organoids showing the migration of RAG1+ (GFP, green) cells within CD31+ (red) vascular-like structures
Supplementary Table
Supplementary Table 1. List of primers relevant to the Extended Data Fig. 1 schematics. Supplementary Table 2. List of antibodies, including those used for flow cytometry and immunofluorescence, referred to in the Methods section
Source data
Source Data Fig. 1
Source Data used construct histogram plots and line graphs shown in Main Figure 1
Source Data Fig. 2
Source Data used construct histogram plots and line graphs shown in Main Figure 2
Source Data Fig. 3
Source Data used construct histogram plots and line graphs shown in Main Figure 3
Source Data Fig. 4
Source Data used construct histogram plots and line graphs shown in Main Figure 4
Source Data Fig. 5
Source Data used construct histogram plots and line graphs shown in Main Figure 5
Source Data Fig. 7
Source Data used construct histogram plots and line graphs shown in Main Figure 7
Source Data Fig. 8
Source Data used construct histogram plots and line graphs shown in Main Figure 8
Source Data Extended Data Fig. 1
Source Data used construct histogram plots and line graphs shown in ED Figure 1
Source Data Extended Data Fig. 1
Uncropped versions of the gel images shown in ED figure 1, constructed from the source file from a digital camera.
Source Data Extended Data Fig. 2
Source Data used construct histogram plots and line graphs shown in ED Figure 2
Source Data Extended Data Fig. 3
Source Data used construct histogram plots and line graphs shown in ED Figure 3
Source Data Extended Data Fig. 4
Source Data used construct histogram plots and line graphs shown in ED Figure 4
Source Data Extended Data Fig. 5
Source Data used construct histogram plots and line graphs shown in ED Figure 5
Rights and permissions
About this article
Cite this article
Motazedian, A., Bruveris, F.F., Kumar, S.V. et al. Multipotent RAG1+ progenitors emerge directly from haemogenic endothelium in human pluripotent stem cell-derived haematopoietic organoids. Nat Cell Biol 22, 60–73 (2020). https://doi.org/10.1038/s41556-019-0445-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-019-0445-8
This article is cited by
-
Using stem cells to model the human bone marrow in a dish
Nature Methods (2024)
-
Induced pluripotent stem cells (iPSCs): molecular mechanisms of induction and applications
Signal Transduction and Targeted Therapy (2024)
-
Generation of complex bone marrow organoids from human induced pluripotent stem cells
Nature Methods (2024)
-
Gene expression in organoids: an expanding horizon
Biology Direct (2023)
-
Unwinding the Role of the CMG Helicase in Inborn Errors of Immunity
Journal of Clinical Immunology (2023)