Abstract
Tau is an abundant microtubule-associated protein in neurons. Tau aggregation into insoluble fibrils is a hallmark of Alzheimer’s disease and other types of dementia1, yet the physiological state of tau molecules within cells remains unclear. Using single-molecule imaging, we directly observe that the microtubule lattice regulates reversible tau self-association, leading to localized, dynamic condensation of tau molecules on the microtubule surface. Tau condensates form selectively permissible barriers, spatially regulating the activity of microtubule-severing enzymes and the movement of molecular motors through their boundaries. We propose that reversible self-association of tau molecules, gated by the microtubule lattice, is an important mechanism of the biological functions of tau, and that oligomerization of tau is a common property shared between the physiological and disease-associated forms of the molecule.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Source data for all statistical analyses can been found on Supplementary Table 1. All other data that support the findings of this study are available from the corresponding authors on reasonable request.
Code availability
The custom analysis code used in this study is available from the corresponding authors on reasonable request.
References
Goedert, M., Eisenberg, D. S. & Crowther, R. A. Propagation of tau aggregates and neurodegeneration. Annu. Rev. Neurosci. 40, 189–210 (2017).
Kapitein, L. C. & Hoogenraad, C. C. Building the neuronal microtubule cytoskeleton. Neuron 87, 492–506 (2015).
Li, X. H. & Rhoades, E. Heterogeneous tau-tubulin complexes accelerate microtubule polymerization. Biophys. J. 112, 2567–2574 (2017).
Makrides, V. et al. Microtubule-dependent oligomerization of tau. Implications for physiological tau function and tauopathies. J. Biol. Chem. 278, 33298–33304 (2003).
Wegmann, S., Bennett, R. E., Amaral, A. S. & Hyman, B. T. Studying tau protein propagation and pathology in the mouse brain using adeno-associated viruses. Methods Cell Biol. 141, 307–322 (2017).
Hernandez-Vega, A. et al. Local nucleation of microtubule bundles through tubulin concentration into a condensed tau phase. Cell Rep. 20, 2304–2312 (2017).
Ambadipudi, S., Biernat, J., Riedel, D., Mandelkow, E. & Zweckstetter, M. Liquid-liquid phase separation of the microtubule-binding repeats of the Alzheimer-related protein Tau. Nat. Commun. 8, 275 (2017).
McVicker, D. P., Hoeprich, G. J., Thompson, A. R. & Berger, C. L. Tau interconverts between diffusive and stable populations on the microtubule surface in an isoform and lattice specific manner. Cytoskeleton 71, 184–194 (2014).
Dixit, R., Ross, J. L., Goldman, Y. E. & Holzbaur, E. L. Differential regulation of dynein and kinesin motor proteins by tau. Science 319, 1086–1089 (2008).
Hinrichs, M. H. et al. Tau protein diffuses along the microtubule lattice. J. Biol. Chem. 287, 38559–38568 (2012).
Vershinin, M., Carter, B. C., Razafsky, D. S., King, S. J. & Gross, S. P. Multiple-motor based transport and its regulation by Tau. Proc. Natl Acad. Sci. USA 104, 87–92 (2007).
Monroy, B. Y. et al. Competition between microtubule-associated proteins directs motor transport. Nat. Commun. 9, 1487 (2018).
Seitz, A. et al. Single-molecule investigation of the interference between kinesin, tau and MAP2c. EMBO J. 21, 4896–4905 (2002).
Vershinin, M., Xu, J., Razafsky, D. S., King, S. J. & Gross, S. P. Tuning microtubule-based transport through filamentous MAPs: the problem of dynein. Traffic 9, 882–892 (2008).
Vale, R. D. Severing of stable microtubules by a mitotically activated protein in Xenopus egg extracts. Cell 64, 827–839 (1991).
Qiang, L., Yu, W., Andreadis, A., Luo, M. & Baas, P. W. Tau protects microtubules in the axon from severing by katanin. J. Neurosci. 26, 3120–3129 (2006).
Yu, W. et al. The microtubule-severing proteins spastin and katanin participate differently in the formation of axonal branches. Mol. Biol. Cell 19, 1485–1498 (2008).
Zempel, H. & Mandelkow, E. M. Tau missorting and spastin-induced microtubule disruption in neurodegeneration: Alzheimer disease and hereditary spastic paraplegia. Mol. Neurodegener. 10, 68 (2015).
Ettinger, A., van Haren, J., Ribeiro, S. A. & Wittmann, T. Doublecortin is excluded from growing microtubule ends and recognizes the GDP-microtubule lattice. Curr. Biol. 26, 1549–1555 (2016).
Duan, A. R. et al. Interactions between tau and different conformations of tubulin: implications for tau function and mechanism. J. Mol. Biol. 429, 1424–1438 (2017).
Zhang, R., LaFrance, B. & Nogales, E. Separating the effects of nucleotide and EB binding on microtubule structure. Proc. Natl Acad. Sci. USA 115, E6191–E6200 (2018).
Kellogg, E. H. et al. Near-atomic model of microtubule-tau interactions. Science 360, 1242–1246 (2018).
Hagiwara, H., Yorifuji, H., Sato-Yoshitake, R. & Hirokawa, N. Competition between motor molecules (kinesin and cytoplasmic dynein) and fibrous microtubule-associated proteins in binding to microtubules. J. Biol. Chem. 269, 3581–3589 (1994).
Wegmann, S. et al. Tau protein liquid-liquid phase separation can initiate tau aggregation. EMBO J. 37, e98049 (2018).
Siahaan, V. et al. Kinetically distinct phases of tau on microtubules regulate kinesin motors and severing enzymes. Nat. Cell Biol. https://doi.org/10.1038/s41556-019-0374-6 (2019).
Kremer, A. et al. Early improved and late defective cognition is reflected by dendritic spines in Tau.P301L mice. J. Neurosci. 31, 18036–18047 (2011).
Garcia-Leon, J. A. et al. Generation of a human induced pluripotent stem cell-based model for tauopathies combining three microtubule-associated protein tau mutations which displays several phenotypes linked to neurodegeneration. Alzheimers Dement. 14, 1261–1280 (2018).
Hubbard, K. S. et al. High yield derivation of enriched glutamatergic neurons from suspension-cultured mouse ESCs for neurotoxicology research. BMC Neurosci. 13, 127 (2012).
Konzack, S., Thies, E., Marx, A., Mandelkow, E. M. & Mandelkow, E. Swimming against the tide: mobility of the microtubule-associated protein tau in neurons. J. Neurosci. 27, 9916–9927 (2007).
Niewidok, B. et al. Presence of a carboxy-terminal pseudorepeat and disease-like pseudohyperphosphorylation critically influence tau’s interaction with microtubules in axon-like processes. Mol. Biol. Cell 27, 3537–3549 (2016).
McKenney, R. J., Huynh, W., Tanenbaum, M. E., Bhabha, G. & Vale, R. D. Activation of cytoplasmic dynein motility by dynactin-cargo adapter complexes. Science 345, 337–341 (2014).
McKenney, R. J., Huynh, W., Vale, R. D. & Sirajuddin, M. Tyrosination of α-tubulin controls the initiation of processive dynein-dynactin motility. EMBO J. 35, 1175–1185 (2016).
McVicker, D. P., Chrin, L. R. & Berger, C. L. The nucleotide-binding state of microtubules modulates kinesin processivity and the ability of Tau to inhibit kinesin-mediated transport. J. Biol. Chem. 286, 42873–42880 (2011).
Urnavicius, L. et al. Cryo-EM shows how dynactin recruits two dyneins for faster movement. Nature 554, 202–206 (2018).
Gutierrez, P. A., Ackermann, B. E., Vershinin, M. & McKenney, R. J. Differential effects of the dynein-regulatory factor Lissencephaly-1 on processive dynein-dynactin motility. J. Biol. Chem. 292, 12245–12255 (2017).
Baumbach, J. et al. Lissencephaly-1 is a context-dependent regulator of the human dynein complex. eLife 6, e21768 (2017).
Ori-McKenney, K. M., Xu, J., Gross, S. P. & Vallee, R. B. A cytoplasmic dynein tail mutation impairs motor processivity. Nat. Cell Biol. 12, 1228–1234 (2010).
Hoang, H. T., Schlager, M. A., Carter, A. P. & Bullock, S. L. DYNC1H1 mutations associated with neurological diseases compromise processivity of dynein-dynactin-cargo adaptor complexes. Proc. Natl Acad. Sci. USA 114, E1597–E1606 (2017).
White, S. R., Evans, K. J., Lary, J., Cole, J. L. & Lauring, B. Recognition of C-terminal amino acids in tubulin by pore loops in Spastin is important for microtubule severing. J. Cell Biol. 176, 995–1005 (2007).
Tepper, K. et al. Oligomer formation of tau protein hyperphosphorylated in cells. J. Biol. Chem. 289, 34389–34407 (2014).
Acknowledgements
We thank all of the members of the McKenney and Ori-McKenney laboratories for feedback and B. Monroy for initial tau observations; J. Al-Bassam for feedback and M. Braun and S. Diez for sharing unpublished data. R.J.M. is supported by grants from NINDS (R00NS089428) and from NIGMS (R35GM124889). K.M.O.-M. is supported by grants from NICHD (R00HD080981), and from the Pew Charitable Trusts (A19-0406). S.S. is supported by grant number R01NS109176. M.V. is supported by a grant from the NSF (ENG-1563280).
Author information
Authors and Affiliations
Contributions
R.J.M., R.T. and K.M.O.-M. conceived the project. R.T., A.J.L. and T.T. produced reagents. R.T. performed all in vitro experiments. J.H. and S.S. provided hippocampal neuron cultures, D.W.N. created molecular models and M.V. performed data analysis.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Further Characterization of Tau Condensates.
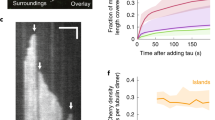
a, SDS-PAGE gel showing proteins used in this study. Protein constructs are listed to the right. N = 1 blot. b, Kymograph and images of the same MT as GFP-2N4R tau is added and subsequently washed out from the chamber. Scale bars: 2 μm (vertical), 2 min (horizontal). (left) and 10 min. (right). N = 2 chambers. c, Pixel intensity distribution of tau along MTs at various concentrations and MT types. WT (N) = 43, 57, 40, and 50. GMP-CPP (N) = 44, 96, and 82. Subtilisin (N) = 67, 52, 43, and 47 MTs, respectively. d, Distribution frequency of condensate lengths. N = 4, 52, 133, 233, 276, and 330 MTs for increasing tau concentrations respectively. e, Top: Intensity distribution of single TMR-SNAP-2N4R tau molecules. Fit to a Gaussian distribution is shown in red. N = 428 molecules from two independent trials. Bottom: Distribution of single- and two-step photobleaching observed for TMR-SNAP-2N4R tau molecules. Examples of single- and two-step photobleaching traces from the analysis. No molecules were observed to bleach in more than two steps. N = 49 molecules from 4 independent trials.
Supplementary Figure 2 Developmental Regulation of Tau Localization in Cultured Neurons.
a, Images of mouse hippocampal neurons at different days cultured in vitro. Neurons were immunostained with two different pan-tau antibodies, GTX49353 (Genetex) and T46 (Thermo Fisher). Scale bar: 25 μm. N = 4 preparations of neurons. b, Quantification of tau puncta frequency in neuron processes. Data represented the mean ± SD. N = 18, 26, and 23 images (from ≥ 3 different neuron preps) for DIV3, DIV4, and DIV7, respectively.
Supplementary Figure 3 The C-terminal Pseudo-Repeat Region of Tau Licenses the Rest of the Molecule Into Tau Condensates.
a, Schematic of tau isoforms and constructs. Orange boxes: alternatively spliced N-term. inserts. Blue: proline-rich domain. Green: MT binding repeats. Yellow: pseudo-repeat domain. b, Top: Images of tau isoforms incorporating into full-length tau (2N4R) condensates. Middle: intensity plots of both channels. Bottom: tau isoform intensities on the lattice and within 2N4R condensates. Scale bar: 2 μm. N = 176 and 211 continuous MT segments (3 chambers each) for 2N4R. N = 127 and 143 segments (3 chambers) for 2N3R. 309 and 158 segments (3 chambers) for 0N3R. c, Top: images of constructs incorporating in 2N4R-tau condensates. Middle: intensity plot of both channels. Bottom: isoform intensities on the lattice and in condensates. Scale bar: 2 μm. N = 152 and 119 segments (3 chambers) for MTBD, 189 and 208 segments (4 chambers) for Proj., 182 and 149 segments (4 chambers) for C-Term, 337 and 296 segments (6 chambers) for bonsai. d, Top: images of Mini-Tau and pseudo-repeat truncations incorporating into 2N4R-tau condensates. Middle: intensity plot of both channels. Bottom: tau isoform intensities on the lattice and in condensates. Scale bar: 2 μm. N = 144 and 239 segments (3 chambers) for Mini-Tau, 145 and 122 segments (3 chambers) for Mini-TauΔ9, 120 and 75 segments (4 chambers) for Mini-TauΔ28. Data represented the mean ± SD. For b, c, and d Student’s T-test (two sided). e, Sequence alignments showing identity conservation (top) and hydrophobicity (bottom) of the tau pseudo-repeat region. The pseudo-repeat is highlighted by green box above.
Supplementary Figure 4 Further Characterization of Tau Condensate Effects on Molecular Motors.
a, Cumulative frequency graph and table of pause times for DDX molecules. N = 7, 8, 4, and 10 chambers for DDB, DDH, DDF, and DDB-Lis1, respectively. b, Summary graph of all the behaviors for DDX complexes upon encountering tau condensates. Statistical significance calculated by both Pearson’s chi-squared test and Fisher’s exact test. c, Kymograph of a DDB molecule undergoing bidirectional movement when encountering a tau patch. Scale bar: 2 μm, 15 sec. N = 7 chambers. d, Model of MT-binding footprints of dynein, kinesin and tau. a, Left: Kinesin motor domain (KIF5B, pdb: 4HNA; green) footprint at the interface of the tubulin dimer and overlapping tau MT-binding repeats (R2x4, pdb: 6CVN; orange). Arrow highlights steric clash between kinesin motor domain and tau. Right: Dynein motor domain (DYNC1H1, pdb: 3JLT; yellow). e, End-on view of 4 tubulin dimers, shown from the minus (-) end. Arrow highlights steric clash between kinesin motor domain and tau. f, MT-lattice view. Arrow highlights steric clash between kinesin and tau.
Supplementary information
Supplementary Information
Supplementary Figs. 1–4, title and legend for Supplementary Table 1, titles and legends for Supplementary Videos 1–5.
Supplementary Table 1
Statistics source data.
Supplementary Video 1
Tau forms high density condensates on MTs.
Supplementary Video 2
Single-molecule behaviour of tau outside and within condensates.
Supplementary Video 3
Tau condensates form at specific locations on the MT lattice.
Supplementary Video 4
Behaviour of Dynein-Dynactin-BicD2 (DDB) complexes at tau condensates.
Supplementary Video 5
Tau condensates exclude spastin and protect MTs from severing.
Rights and permissions
About this article
Cite this article
Tan, R., Lam, A.J., Tan, T. et al. Microtubules gate tau condensation to spatially regulate microtubule functions. Nat Cell Biol 21, 1078–1085 (2019). https://doi.org/10.1038/s41556-019-0375-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-019-0375-5
This article is cited by
-
Enolase of Streptococcus suis serotype 2 promotes biomolecular condensation of ribosomal protein SA for HBMECs apoptosis
BMC Biology (2024)
-
Conserved roles for the dynein intermediate chain and Ndel1 in assembly and activation of dynein
Nature Communications (2023)
-
Initiation and modulation of Tau protein phase separation by the drug suramin
Scientific Reports (2023)
-
Multivalent interactions facilitate motor-dependent protein accumulation at growing microtubule plus-ends
Nature Cell Biology (2023)
-
Methylene blue accelerates liquid-to-gel transition of tau condensates impacting tau function and pathology
Nature Communications (2023)