Abstract
Totipotency refers to the ability of a cell to generate all of the cell types of an organism. Unlike pluripotency, the establishment of totipotency is poorly understood. In mouse embryonic stem cells, Dux drives a small percentage of cells into a totipotent state by expressing 2-cell-embryo-specific transcripts. To understand how this transition takes place, we performed single-cell RNA-seq, which revealed a two-step transcriptional reprogramming process characterized by downregulation of pluripotent genes in the first step and upregulation of the 2-cell-embryo-specific elements in the second step. To identify factors controlling the transition, we performed a CRISPR–Cas9-mediated screen, which revealed Myc and Dnmt1 as two factors preventing the transition. Mechanistic studies demonstrate that Myc prevents downregulation of pluripotent genes in the first step, while Dnmt1 impedes 2-cell-embryo-specific gene activation in the second step. Collectively, the findings of our study reveal insights into the establishment and regulation of the totipotent state in mouse embryonic stem cells.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
RNA-seq, Drop-seq, RRBS and CRISPR screen-related data, including the sgRNA read counts, that support the findings of this study have been deposited in the Gene Expression Omnibus (GEO) under the accession code GSE121459. Previously published sequencing data that were reanalysed here are available in the GEO under the accession codes GSE85766 (Promoter methylation in ESCs and 2C-like cells), GSE95517 (Dux ChIP-seq) and GSM1171648 (Myc ChIP-seq), and the samples GSM1966767, GSM1966768 and GSM1966769 (Zscan+ and MuERVL+ spontaneous 2C-like cell transcriptome)16,18,28. Source data for all figures have been provided as Supplementary Table 10. All other data supporting the findings of this study are available from the corresponding author on reasonable request.
Code availability
All of the codes used in this study are available from the corresponding author upon reasonable request.
References
Lee, M. T., Bonneau, A. R. & Giraldez, A. J. Zygotic genome activation during the maternal-to-zygotic transition. Annu. Rev. Cell Dev. Biol. 30, 581–613 (2014).
Lu, F. & Zhang, Y. Cell totipotency: molecular features, induction, and maintenance. Natl Sci. Rev. 2, 217–225 (2015).
Falco, G. et al. Zscan4: a novel gene expressed exclusively in late 2-cell embryos and embryonic stem cells. Dev. Biol. 307, 539–550 (2007).
Macfarlan, T. S. et al. Embryonic stem cell potency fluctuates with endogenous retrovirus activity. Nature 487, 57–63 (2012).
Yang, Y. et al. Derivation of pluripotent stem cells with in vivo embryonic and extraembryonic potency. Cell 169, 243–257.e25 (2017).
Yang, J. et al. Establishment of mouse expanded potential stem cells. Nature 550, 393–397 (2017).
Bao, S. et al. Derivation of hypermethylated pluripotent embryonic stem cells with high potency. Cell Res. 28, 22–34 (2018).
Li, M. & Izpisua Belmonte, J. C. Deconstructing the pluripotency gene regulatory network. Nat. Cell Biol. 20, 382–392 (2018).
Boskovic, A. et al. Higher chromatin mobility supports totipotency and precedes pluripotency in vivo. Genes Dev. 28, 1042–1047 (2014).
Ishiuchi, T. et al. Early embryonic-like cells are induced by downregulating replication-dependent chromatin assembly. Nat. Struct. Mol. Biol. 22, 662–671 (2015).
Baker, C. L. & Pera, M. F. Capturing totipotent stem cells. Cell Stem Cell 22, 25–34 (2018).
Lu, F., Liu, Y., Jiang, L., Yamaguchi, S. & Zhang, Y. Role of Tet proteins in enhancer activity and telomere elongation. Genes Dev. 28, 2103–2119 (2014).
Rodriguez-Terrones, D. et al. A molecular roadmap for the emergence of early-embryonic-like cells in culture. Nat. Genet. 50, 106–119 (2018).
Choi, Y. J. et al. Deficiency of microRNA miR-34a expands cell fate potential in pluripotent stem cells. Science 355, eaag1927 (2017).
Whiddon, J. L., Langford, A. T., Wong, C. J., Zhong, J. W. & Tapscott, S. J. Conservation and innovation in the DUX4-family gene network. Nat. Genet. 49, 935–940 (2017).
Hendrickson, P. G. et al. Conserved roles of mouse DUX and human DUX4 in activating cleavage-stage genes and MERVL/HERVL retrotransposons. Nat. Genet. 49, 925–934 (2017).
De Iaco, A. et al. DUX-family transcription factors regulate zygotic genome activation in placental mammals. Nat. Genet. 49, 941–945 (2017).
Eckersley-Maslin, M. A. et al. MERVL/Zscan4 network activation results in transient genome-wide DNA demethylation of mESCs. Cell Rep. 17, 179–192 (2016).
Eidahl, J. O. et al. Mouse Dux is myotoxic and shares partial functional homology with its human paralog DUX4. Hum. Mol. Genet. 25, 4577–4589 (2016).
Kumar, R. M. et al. Deconstructing transcriptional heterogeneity in pluripotent stem cells. Nature 516, 56–61 (2014).
Kolodziejczyk, A. A. et al. Single cell RNA-sequencing of pluripotent states unlocks modular transcriptional variation. Cell Stem Cell 17, 471–485 (2015).
Smith, A. Formative pluripotency: the executive phase in a developmental continuum. Development 144, 365–373 (2017).
Doench, J. G. et al. Optimized sgRNA design to maximize activity and minimize off-target effects of CRISPR-Cas9. Nat. Biotechnol. 34, 184–191 (2016).
Li, W. et al. MAGeCK enables robust identification of essential genes from genome-scale CRISPR/Cas9 knockout screens. Genome Biol. 15, 554 (2014).
Chappell, J. & Dalton, S. Roles for MYC in the establishment and maintenance of pluripotency. Cold Spring Harb. Perspect. Med. 3, a014381 (2013).
Kim, J. et al. A Myc rather than core pluripotency module accounts for the shared signatures of embryonic stem and cancer cells. Cell 143, 313–324 (2010).
Nie, Z. et al. c-Myc is a universal amplifier of expressed genes in lymphocytes and embryonic stem cells. Cell 151, 68–79 (2012).
Krepelova, A., Neri, F., Maldotti, M., Rapelli, S. & Oliviero, S. Myc and max genome-wide binding sites analysis links the Myc regulatory network with the polycomb and the core pluripotency networks in mouse embryonic stem cells. PLoS One 9, e88933 (2014).
Percharde, M., Bulut-Karslioglu, A. & Ramalho-Santos, M. Hypertranscription in development, stem cells, and regeneration. Dev. Cell 40, 9–21 (2017).
Lin, C. Y. et al. Transcriptional amplification in tumor cells with elevated c-Myc. Cell 151, 56–67 (2012).
Jones, P. A. & Liang, G. Rethinking how DNA methylation patterns are maintained. Nat. Rev. Genet. 10, 805–811 (2009).
Dan, J. et al. Zscan4 inhibits maintenance DNA methylation to facilitate telomere elongation in mouse embryonic stem cells. Cell Rep. 20, 1936–1949 (2017).
De Iaco, A., Coudray, A., Duc, J. & Trono, D. DPPA2 and DPPA4 are necessary to establish a 2C‐like state in mouse embryonic stem cells. EMBO Rep. 20, e47382 (2019).
Eckersley-Maslin, M. et al. Dppa2 and Dppa4 directly regulate the Dux-driven zygotic transcriptional program. Genes Dev. 33, 194–208 (2019).
Campbell, A. E. et al. NuRD and CAF-1-mediated silencing of the D4Z4 array is modulated by DUX4-induced MBD3L proteins. eLife 7, e31023 (2018).
Percharde, M. et al. A LINE1-nucleolin partnership regulates early development and ESC identity. Cell 174, 391–405.e19 (2018).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
The ENCODE Project Consortium An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Shalem, O. et al. Genome-scale CRISPR-Cas9 knockout screening in human cells. Science 343, 84–87 (2014).
Chen, R., Wu, X., Jiang, L. & Zhang, Y. Single-cell RNA-seq reveals hypothalamic cell diversity. Cell Rep. 18, 3227–3241 (2017).
Criscione, S. W., Zhang, Y., Thompson, W., Sedivy, J. M. & Neretti, N. Transcriptional landscape of repetitive elements in normal and cancer human cells. BMC Genomics 15, 583 (2014).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Macosko, E. Z. et al. Highly parallel genome-wide expression profiling of individual cells using nanoliter droplets. Cell 161, 1202–1214 (2015).
Robinson, M. D. & Oshlack, A. A scaling normalization method for differential expression analysis of RNA-seq data. Genome Biol. 11, R25 (2010).
McCarthy, D. J., Chen, Y. & Smyth, G. K. Differential expression analysis of multifactor RNA-Seq experiments with respect to biological variation. Nucleic Acids Res. 40, 4288–4297 (2012).
Kramer, A., Green, J., Pollard, J. Jr. & Tugendreich, S. Causal analysis approaches in Ingenuity Pathway Analysis. Bioinformatics 30, 523–530 (2014).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411 (2018).
Lun, A. T., Bach, K. & Marioni, J. C. Pooling across cells to normalize single-cell RNA sequencing data with many zero counts. Genome Biol. 17, 75 (2016).
McDavid, A. et al. Data exploration, quality control and testing in single-cell qPCR-based gene expression experiments. Bioinformatics 29, 461–467 (2013).
McInnes, L., Healy, J., Saul, N. & Großberger, L. UMAP: uniform manifold approximation and projection. J. Open Source Softw. 3, 861 (2018).
Trapnell, C. et al. The dynamics and regulators of cell fate decisions are revealed by pseudotemporal ordering of single cells. Nat. Biotechnol. 32, 381–386 (2014).
Leek, J. T., Johnson, W. E., Parker, H. S., Jaffe, A. E. & Storey, J. D. The sva package for removing batch effects and other unwanted variation in high-throughput experiments. Bioinformatics 28, 882–883 (2012).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Acknowledgements
We thank S. L. Pfaff for providing the MERVL-tdTomato reporter, F. Lu for assistance with the establishment of the reporter cell line and Z. Chen for critical reading of the manuscript. This project was supported by the NIH (R01HD092465) and HHMI. Y.Z. is an Investigator of the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
Y.Z. conceived the project; X.F., X.W. and Y.Z. designed the experiments; X.F. and X.W. performed the experiments. M.N.D. performed bioinformatics analyses. All authors were involved in the interpretation of data. X.F., X.W. and Y.Z. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Transcriptomic anlysis indicated the existance of an intermediate cell state during ESC to 2C-like cell transition.
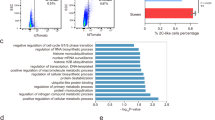
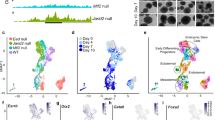
a, Images showing Dux induction can gradually induce 2C-like cells generation in cultured ESCs. Scale bar: 0.1 mm. Experiment was repeated independently twice with similar results. b, The 2C-like cell percentage of two independent ESC clones after one day of Dux induction. c, Dux transgene mRNA level relative to Gapdh in two independent ESC clones. b-c, Shown are mean ± SD, n=3 biologically independent cell cultures. P values (indicated as numbers in the graphs) are calculated by unpaired t-test, two-tailed, two-sample unequal variance. Experiments were repeated independently twice with similar results. d, Bar plots showing the -log10(q value) (right-tailed Fisher-exact test) of the GO terms of upregulated and downregulated genes in 2C-like cells. e, Box plot showing the distance of the nearest gene TSS (n=Stable: 24,739, Down: 2,466, Up: 2,146 genes) to MERVL repeats (median: Stable: ~213kb, Down: ~242.5kb, Up: ~93kb). f, Scatter plot comparing 2-cell-embryo-specific genes expression profile between spontaneous 2C-like cells and Dux-induced 2C-like cells (Pearson correlation, r=0.9, n=1,400 genes). g, The percentage of 2C-like cells after 2-hour and 24-hour culture of D1 2C+ cells. h, Scatter plot comparing gene expression profile between cells which exit from Dux-induced 2C-like state (D2 2C− cells) and D1 2C− cells, (Pearson correlation, r=0.95, n=27,221 genes/repeats), supporting that Dux-induced 2C-like cells can exit from 2C-like state spontanesouly. i, Relative percentage of genes and repeats in each category. j, Box plot showing the log2 expression level of 2C+-upregulated genes/repeats (n=2,976 genes/repeats) and 2C+-downregulated genes/repeats (n=2,726 genes/repeats) in each of the D0 2C−, D1 2C−, and D1 2C+ cell populations. e, j, P values were calculated by two-tailed mann-whitney U-test. The black central line is the median, boxes limits indicate the upper and lower quartiles, whiskers indicate the 1.5 interquartile range, dots represent outliers. k, Pie chart reporting the percentage of Group1 and Group2 genes in 2C+-upregulated genes and Dux bound 2C+-upregulated genes. d-k were based on two independent biological replicates of RNA-seq experiments. The effect-size of each statistical analysis and statistical source data can be found in Supplementary Table 10.
Supplementary Figure 2 Single-cell transcriptomic analyses revealed an intermediate state from ESC to 2C-like cell transition.
a, Violin plot showing the log2 expression of representative genes or repeats in each time point (n= 0h: 738, 12h: 456 cells, 24h: 568, 36h: 871). Violin plots show the kernel density estimation for the distribution of the gene expression in each cluster. The width of the plot represents the proportion of data with the corresponding expression value. b, UMAP plot showing the expression of representative genes in each sequenced cells (n=2,633 cells). c, UMAP plot showing the subpopulations of of cluster 1 cells (n=1,780 cells). Around 7% pluripotent cells (n=115 cells) belong to the minor subcluster b. d, Box plots comparing the gene expression between subcluster a (n=1,665 cells) and subcluster b cells (n=115 cells). The black central line is the median, boxes limits indicate the upper and lower quartiles, whiskers indicate the 1.5 interquartile range, dots represent outliers. P values (shown as p) are calculated by two-tailed Mann-Whitney U-test and effect-sizes (shown as r) are calculated as \(Z/\sqrt N\) where Z is the Z value of the Mann–Whitney U-test and N is the number of samples. e, FACS histogram plot showing the expression of Rex1 in mESC culture. About 5% of cells are Rex1-negative. Experiment was repeated independently three times with similar results. f, Bar plot showing the proportion of the different cell clusters at different time point of Dux induction. g, The partial 2-cell embyro-specfic genes and elements activated in D1 2C− cells leads to less dramatic transcriptional variability between D1 2C− cells and cluster 2 cells compared to that of cluster 1 cells (n=470 genes/repeats). The red dots show the top 30 genes/repeats that define the variance between D1 2C− and cluster 1 cells. Correlation was calculated using spearman correlation. Statistical source data can be found in Supplementary Table 10.
Supplementary Figure 3 Single-cell transcriptomic analyses revealed the transcriptional dynamics from ESC to 2C-like cell transition.
a, Scatter plots showing cells (n=2,633 cells) along the projected pseudo-time following the dynamics of the ESC to 2C-like cell transition. b, Cell distribution along with the pseudotime in each time point. Cells collected at the early hours of Dux-induction (0 h and 12 h) are located at the beginning of the timeline while cells collected at the late hours (24 h and 36 h) are gradually located towards the end, supporting that the projected timeline captures the transcriptional changes during Dux-induced 2C-like transition. c, Mean expression of representative genes in each cell cluster. d, Violin plot showing the expression of representative transcripts in each cell cluster (n=1,780 cells, 512 cells and 341 cells respectively for cluster 1, 2, and 3). Violin plots show the kernel density estimation of the distribution for the expression of the genes in each cluster. The width of the plot represents the proportion of data with the corresponding expression value. e, Box plot showing the expression of the detected 2C+-downregulated pluripotent genes (n=135 genes), 2C+-upregulated 2-cell-embyro-specific genes (n=75 genes) and activated repeats (n=501 repeats) in each cell cluster when analyzed after subcluster b cells are excluded. The black central line is the median, boxes limits indicate the upper and lower quartiles, whiskers indicate the 1.5 interquartile range, dots represent outliers. P values (shown as p) are calculated by two-tailed Mann-Whitney U-test and effect-sizes (shown as r) are calculated as \(Z/{\sqrt{N}}\) where Z is the Z value of the Mann–Whitney U-test and N is the number of samples. Statistical source data can be found in Supplementary Table 10.
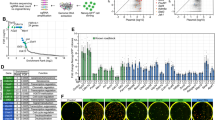
Supplementary Figure 4 CRISPR-Cas9 screen identified novel regulators mediating ESC to 2C-like transition.
a,b, RRA score as calculated by MAGeCK tool of negative effect (a) and positive effect (b) of genes on the ESC to 2C-like cell transition from the screen. The results were based on two biologically independent screens.
Supplementary Figure 5 Myc perturbation facilitats ESC to 2C-like cell transition.
a, Western-blot showing the sgRNA efficiency. Unprocessed raw blots can be found in Supplementary Fig. 7. b, Relative percentage of 2C-like cells after 24-hour culture of D1 2C+ cells isolated from ESC infected with sgGFP or sgMyc. Shown are mean ± SD, n=3 biologically independent cell cultures. c, Relative percentage of spontaneous 2C-like cells and representative FACS plot of ESC culture. Shown are mean ± SD, n=3 biologically independent cell cultures. d, Myc knockdown decreases total RNA amount per cell in ESCs (left panel) and D1 2C+ cells (right panel), confirming the transcrition amplification effect of Myc in mESCs. Shown are mean ± SD, n=3 biologically independent cell cultures. b-d, P values (indicated as numbers in the graphs) are calculated by unpaired t-test, two-tailed, two-sample unequal variance. Statistical source data can be found in Supplementary Table 10. a-d, experiments were repeated independently twice with similar results.
Supplementary Figure 6 Dnmt1 and Myc perturbation facilitates ESC to 2C-like cell transition.
a, Western-blot showing the sgRNA efficiency. Unprocessed raw blots can be found in Supplementary Fig. 7. b, Relative percentage of 2C-like cells after 24-hour culture of D1 2C+ cells isolated from ESC infected with sgGFP or sgDnmt1. Shown are mean ± SD, n=3 biologically independent cell cultures. c, Relative percentage of spontaneous 2C-like cells and representative FACS plot of ESC culture. Shown are mean ± SD, n=3 biologically independent cell cultures. b-c, P values (indicated as numbers in the graphs) are calculated by unpaired t-test, two-tailed, two-sample unequal variance. d, Mean promoter methylation change from ESC to 2C-like cell of 2C+-upregulated (n=1,611 genes) and 2C+-downregulated genes (n=1,398 genes). The black central line is the median, boxes limits indicate the upper and lower quartiles, whiskers indicate the 1.5 interquartile range, dots represent outliers. P values (shown as p) are calculated by two-tailed Mann-Whitney U-test and effect-sizes (shown as r) are calculated as \(Z/{\sqrt{N}}\) where Z is the Z value of the Mann–Whitney U-test and N is the number of samples. e, The percentage of 2C-like cells with prolonged Dux induction. Prolonged Dux induction did not induce complete 2C-like transition as Dux induction initiates an unsynchronized 2C-like transition (Fig. 1b) and cannot maintain 2C-like state (Supplementary Fig. 1g). f, Bootstrapped correlation analysis showing that the Zscan4+ intermediate cells reported by Rodriguez-Terrones et al. are assigned to clusters 1 and 3 of our single-cell analysis but cannot be firmly assigned to cluster 2, the intermediate state. g, Density plot showing the distribution of cells from the different clusters generated in our study and study from Rodriguez-Terrones et al. along the pseudotime. Cells were ordered using the 93 genes selected by Rodriguez-Terrones et al. after technical batch removal using the MNN method. We note that the date from Rodriguez-Terrones et al. did to capture the intermediate state (Cluster 2) identified in our study. h-i, FACS histogram plot showing the expression of Myc (h) or Dnmt1 (i) in mESCs. a-c, e, h-i, experiments were repeated independently twice with similar results. Statistical source data can be found in Supplementary Table 10.
Supplementary information
Supplementary Information
Supplementary Figures 1–7 and their legends and legends for Supplementary Tables 1–10.
Rights and permissions
About this article
Cite this article
Fu, X., Wu, X., Djekidel, M.N. et al. Myc and Dnmt1 impede the pluripotent to totipotent state transition in embryonic stem cells. Nat Cell Biol 21, 835–844 (2019). https://doi.org/10.1038/s41556-019-0343-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-019-0343-0
This article is cited by
-
A CRISPR/Cas9 screen in embryonic stem cells reveals that Mdm2 regulates totipotency exit
Communications Biology (2024)
-
Maternal factor Trim75 contributes to zygotic genome activation program in mouse early embryos
Molecular Biology Reports (2024)
-
ZINBMM: a general mixture model for simultaneous clustering and gene selection using single-cell transcriptomic data
Genome Biology (2023)
-
Selective binding of retrotransposons by ZFP352 facilitates the timely dissolution of totipotency network
Nature Communications (2023)
-
A genome-wide screen reveals new regulators of the 2-cell-like cell state
Nature Structural & Molecular Biology (2023)