Abstract
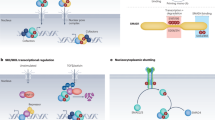
Wnt signalling drives many processes in development, homeostasis and disease; however, the role and mechanism of individual ligand–receptor (Wnt–Frizzled (Fzd)) interactions in specific biological processes remain poorly understood. Wnt9a is specifically required for the amplification of blood progenitor cells during development. Using genetic studies in zebrafish and human embryonic stem cells, paired with in vitro cell biology and biochemistry, we determined that Wnt9a signals specifically through Fzd9b to elicit β-catenin-dependent Wnt signalling that regulates haematopoietic stem and progenitor cell emergence. We demonstrate that the epidermal growth factor receptor (EGFR) is required as a cofactor for Wnt9a–Fzd9b signalling. EGFR-mediated phosphorylation of one tyrosine residue on the Fzd9b intracellular tail in response to Wnt9a promotes internalization of the Wnt9a–Fzd9b–LRP signalosome and subsequent signal transduction. These findings provide mechanistic insights for specific Wnt–Fzd signals, which will be crucial for specific therapeutic targeting and regenerative medicine.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
MS data have been deposited in ProteomeXchange with the primary accession code PXD010649 through MassIVE (MSV000082677). Summary data are seen in Supplementary Figs. 6e,f and 7d,e. Source data for the main and supplementary figures have been provided as Supplementary Table 3. Previously published sequencing data that were re-analysed here are available from the European Nucleotide Archive under the accession number PRJEB4197. All other data supporting the findings of this study are available from the corresponding authors on reasonable request.
References
Nusse, R. & Clevers, H. Wnt/β-catenin signaling, disease, and emerging therapeutic modalities. Cell 169, 985–999 (2017).
Richter, J. et al. WNT9A is a conserved regulator of hematopoietic stem and progenitor cell development. Genes (Basel) 9, E66 (2018).
Grainger, S. et al. Wnt9a is required for the aortic amplification of nascent hematopoietic stem cells. Cell Rep. 17, 1595–1606 (2016).
Ring, L., Neth, P., Weber, C., Steffens, S. & Faussner, A. β-Catenin-dependent pathway activation by both promiscuous ‘canonical’ WNT3a–, and specific ‘noncanonical’ WNT4– and WNT5a–FZD receptor combinations with strong differences in LRP5 and LRP6 dependency. Cell. Signal. 26, 260–267 (2014).
Voloshanenko, O., Gmach, P., Winter, J., Kranz, D. & Boutros, M. Mapping of Wnt–Frizzled interactions by multiplex CRISPR targeting of receptor gene families. FASEB J. 31, 4832–4844 (2017).
Perlin, J. R., Robertson, A. L. & Zon, L. I. Efforts to enhance blood stem cell engraftment: recent insights from zebrafish hematopoiesis. J. Exp. Med. 214, 2817–2827 (2017).
Clements, W. K. & Traver, D. Signalling pathways that control vertebrate haematopoietic stem cell specification. Nat. Rev. Immunol. 13, 336–348 (2013).
Zhen, F., Lan, Y., Yan, B., Zhang, W. & Wen, Z. Hemogenic endothelium specification and hematopoietic stem cell maintenance employ distinct Scl isoforms. Development 140, 3977–3985 (2013).
Leung, A. et al. Uncoupling VEGFA functions in arteriogenesis and hematopoietic stem cell specification. Dev. Cell 24, 144–158 (2013).
Butko, E. et al. Gata2b is a restricted early regulator of hemogenic endothelium in the zebrafish embryo. Development 142, 1050–1061 (2015).
Kobayashi, I. et al. Jam1a–Jam2a interactions regulate haematopoietic stem cell fate through Notch signalling. Nature 512, 319–323 (2014).
Damm, E. W. & Clements, W. K. Pdgf signalling guides neural crest contribution to the haematopoietic stem cell specification niche. Nat. Cell Biol. 19, 457–467 (2017).
Kissa, K. & Herbomel, P. Blood stem cells emerge from aortic endothelium by a novel type of cell transition. Nature 464, 112–115 (2010).
Bertrand, J. Y. et al. Haematopoietic stem cells derive directly from aortic endothelium during development. Nature 464, 108–111 (2010).
Luis, T. C. et al. Wnt3a deficiency irreversibly impairs hematopoietic stem cell self-renewal and leads to defects in progenitor cell differentiation. Blood 113, 546–554 (2009).
Zhao, C. et al. Loss of β-catenin impairs the renewal of normal and cml stem cells in vivo. Cancer Cell 12, 528–541 (2007).
Fleming, H. E. et al. Wnt signaling in the niche enforces hematopoietic stem cell quiescence and is necessary to preserve self-renewal in vivo. Cell Stem Cell 2, 274–283 (2008).
Reya, T. et al. A role for Wnt signaling in self-renewal of haematopoietic stem cells. Nature. 423, 409–414 (2003).
Willert, K. et al. Wnt proteins are lipid-modified and can act as stem cell growth factors. Nature 423, 448–452 (2003).
Malhotra, S. et al. Contrasting responses of lymphoid progenitors to canonical and noncanonical Wnt signals. J. Immunol. 181, 3955–3964 (2008).
Baba, Y. et al. Constitutively active β-catenin promotes expansion of multipotent hematopoietic progenitors in culture. J. Immunol. 177, 2294–2303 (2006).
Kirstetter, P., Anderson, K., Porse, B. T., Jacobsen, S. E. & Nerlov, C. Activation of the canonical Wnt pathway leads to loss of hematopoietic stem cell repopulation and multilineage differentiation block. Nat. Immunol. 7, 1048–1056 (2006).
Scheller, M. et al. Hematopoietic stem cell and multilineage defects generated by constitutive β-catenin activation. Nat. Immunol. 7, 1037–1047 (2006).
Luis, T. C., Ichii, M., Brugman, M. H., Kincade, P. & Staal, F. J. Wnt signaling strength regulates normal hematopoiesis and its deregulation is involved in leukemia development. Leukemia 26, 414–421 (2012).
Luis, T. C. et al. Canonical Wnt signaling regulates hematopoiesis in a dosage-dependent fashion. Cell Stem Cell 9, 345–356 (2011).
Goessling, W. et al. Genetic interaction of PGE2 and Wnt signaling regulates developmental specification of stem cells and regeneration. Cell 136, 1136–1147 (2009).
Lam, S. S. et al. Directed evolution of APEX2 for electron microscopy and proximity labeling. Nat. Methods 12, 51–54 (2015).
Veeman, M. T., Slusarski, D. C., Kaykas, A., Louie, S. H. & Moon, R. T. Zebrafish prickle, a modulator of noncanonical Wnt/Fz signaling, regulates gastrulation movements. Curr. Biol. 13, 680–685 (2003).
Ablain, J., Durand, E. M., Yang, S., Zhou, Y. & Zon, L. I. A CRISPR/Cas9 vector system for tissue-specific gene disruption in zebrafish. Dev. Cell 32, 756–764 (2015).
Ng, E. S., Davis, R., Stanley, E. G. & Elefanty, A. G. A protocol describing the use of a recombinant protein-based, animal product-free medium (APEL) for human embryonic stem cell differentiation as spin embryoid bodies. Nat. Protoc. 3, 768–776 (2008).
Thomson, J. A. et al. Embryonic stem cell lines derived from human blastocysts. Science 282, 1145–1147 (1998).
Janda, C. Y., Waghray, D., Levin, A. M., Thomas, C. & Garcia, K. C. Structural basis of Wnt recognition by Frizzled. Science 337, 59–64 (2012).
Dijksterhuis, J. P. et al. Systematic mapping of WNT–FZD protein interactions reveals functional selectivity by distinct WNT–FZD pairs. J. Biol. Chem. 290, 6789–6798 (2015).
Rulifson, E. J., Wu, C. H. & Nusse, R. Pathway specificity by the bifunctional receptor Frizzled is determined by affinity for Wingless. Mol. Cell 6, 117–126 (2000).
Tamai, K. et al. LDL-receptor-related proteins in Wnt signal transduction. Nature 407, 530–535 (2000).
Lawson, N. D. & Weinstein, B. M. In vivo imaging of embryonic vascular development using transgenic Zebrafish. Dev. Biol. 248, 307–318 (2002).
Lewis, J. L. et al. Reiterated Wnt signaling during zebrafish neural crest development. Development 131, 1299–1308 (2004).
Mishra, R., Hanker, A. B. & Garrett, J. T. Genomic alterations of ERBB receptors in cancer: clinical implications. Oncotarget 8, 114371–114392 (2017).
Doody, J. F. et al. Inhibitory activity of cetuximab on epidermal growth factor receptor mutations in non small cell lung cancers. Mol. Cancer Ther. 6, 2642–2651 (2007).
Zhao, Y. & Lin, S. Essential role of SH3-domain GRB2-like 3 for vascular lumen maintenance in zebrafish. Arterioscler. Thromb. Vasc. Biol. 33, 1280–1286 (2013).
Osherov, N. & Levitzki, A. Epidermal-growth-factor-dependent activation of the src-family kinases. Eur. J. Biochem. 225, 1047–1053 (1994).
Sultan, M. et al. Influence of RNA extraction methods and library selection schemes on RNA-seq data. BMC Genomics 15, 675 (2014).
Blom, N., Gammeltoft, S. & Brunak, S. Sequence and structure-based prediction of eukaryotic protein phosphorylation sites. J. Mol. Biol. 294, 1351–1362 (1999).
Schafer, B., Gschwind, A. & Ullrich, A. Multiple G-protein-coupled receptor signals converge on the epidermal growth factor receptor to promote migration and invasion. Oncogene 23, 991–999 (2004).
Prenzel, N. et al. EGF receptor transactivation by G-protein-coupled receptors requires metalloproteinase cleavage of proHB-EGF. Nature 402, 884–888 (1999).
Tomlins, S. A., Bolllinger, N., Creim, J. & Rodland, K. D. Cross-talk between the calcium-sensing receptor and the epidermal growth factor receptor in Rat-1 fibroblasts. Exp. Cell Res. 308, 439–445 (2005).
Yamamoto, H., Komekado, H. & Kikuchi, A. Caveolin is necessary for Wnt-3a-dependent internalization of LRP6 and accumulation of β-catenin. Dev. Cell 11, 213–223 (2006).
Blitzer, J. T. & Nusse, R. A critical role for endocytosis in Wnt signaling. BMC Cell Biol. 7, 28 (2006).
Yu, H., Ye, X., Guo, N. & Nathans, J. Frizzled 2 and Frizzled 7 function redundantly in convergent extension and closure of the ventricular septum and palate: evidence for a network of interacting genes. Development 139, 4383–4394 (2012).
Hsieh, J. C., Rattner, A., Smallwood, P. M. & Nathans, J. Biochemical characterization of Wnt–Frizzled interactions using a soluble, biologically active vertebrate Wnt protein. Proc. Natl Acad. Sci. USA 96, 3546–3551 (1999).
Cho, C., Smallwood, P. M. & Nathans, J. Reck and Gpr124 are essential receptor cofactors for Wnt7a/Wnt7b-specific signaling in mammalian CNS angiogenesis and blood–brain barrier regulation. Neuron 95, 1221–1225 (2017).
Zhou, Y. & Nathans, J. Gpr124 controls CNS angiogenesis and blood–brain barrier integrity by promoting ligand-specific canonical Wnt signaling. Dev. Cell 31, 248–256 (2014).
Eubelen, M. et al. A molecular mechanism for Wnt ligand-specific signaling. Science 361, eaat1178 (2018).
Lai, M. B. et al. TSPAN12 is a norrin co-receptor that amplifies Frizzled4 ligand selectivity and signaling. Cell Rep. 19, 2809–2822 (2017).
Fernandez, A. et al. The WNT receptor FZD7 is required for maintenance of the pluripotent state in human embryonic stem cells. Proc. Natl Acad. Sci. USA 111, 1409–1414 (2014).
Grainger, S. et al. CRISPR guide RNA validation in vitro. Zebrafish 14, 383–386 (2017).
Kwan, K. M. et al. The Tol2kit: a multisite gateway-based construction kit for Tol2 transposon transgenesis constructs. Dev. Dyn. 236, 3088–3099 (2007).
Huggins, I. J. et al. The WNT target SP5 negatively regulates WNT transcriptional programs in human pluripotent stem cells. Nat. Commun. 8, 1034 (2017).
Moya, N., Cutts, J., Gaasterland, T., Willert, K. & Brafman, D. A. Endogenous WNT signaling regulates hPSC-derived neural progenitor cell heterogeneity and specifies their regional identity. Stem Cell Rep. 3, 1015–1028 (2014).
Haas, J. P. Measurement of infection control department performance: state of the science. Am. J. Infect. Control 34, 543–549 (2006).
Lapek, J. D. Jr, Lewinski, M. K., Wozniak, J. M., Guatelli, J. & Gonzalez, D. J. Quantitative temporal viromics of an inducible HIV-1 model yields insight to global host targets and phospho-dynamics associated with protein Vpr. Mol. Cell. Proteomics 16, 1447–1461 (2017).
Lapek, J. D. Jr et al. Defining host responses during systemic bacterial infection through construction of a murine organ proteome atlas. Cell Syst. 6, 579–592.e4 (2018).
McAlister, G. C. et al. MultiNotch MS3 enables accurate, sensitive, and multiplexed detection of differential expression across cancer cell line proteomes. Anal. Chem. 86, 7150–7158 (2014).
Eng, J. K., McCormack, A. L. & Yates, J. R. An approach to correlate tandem mass spectral data of peptides with amino acid sequences in a protein database. J. Am. Soc. Mass Spectrom. 5, 976–989 (1994).
Elias, J. E. & Gygi, S. P. Target-decoy search strategy for increased confidence in large-scale protein identifications by mass spectrometry. Nat. Methods 4, 207–214 (2007).
Elias, J. E., Haas, W., Faherty, B. K. & Gygi, S. P. Comparative evaluation of mass spectrometry platforms used in large-scale proteomics investigations. Nat. Methods 2, 667–675 (2005).
Peng, J., Elias, J. E., Thoreen, C. C., Licklider, L. J. & Gygi, S. P. Evaluation of multidimensional chromatography coupled with tandem mass spectrometry (LC/LC–MS/MS) for large-scale protein analysis: the yeast proteome. J. Proteome Res. 2, 43–50 (2003).
Xiao, Y. et al. A novel significance score for gene selection and ranking. Bioinformatics 30, 801–807 (2014).
Huang da, W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Acknowledgements
We thank C. Fine and J. Olvera for FACS assistance; R. Espin-Palazon, J. Nussbacher and I. J. Huggins for manuscript reading; I. J. Huggins for providing cetuximab; N. Gohad for microscopy assistance; and M. Boutros for cell lines. S.G. was supported by awards from the American Heart Association (14POST18340021) and the Leukemia and Lymphoma Society (5431-15). Research reported in this publication was supported by the National Heart, Lung, and Blood Institute of the US National Institutes of Health (NIH) under award number K99HL133458 (awarded to S.G.). The content is solely the responsibility of the authors and does not necessarily represent the official views of the NIH. J.R. was supported in part by the UCSD Interdisciplinary Stem Cell Training Program (CIRM TG2-01154) and by the American Heart Association Predoctoral Fellowship (16PRE27340012). This work was supported in part by funding to K.W. from the UCSD Stem Cell Program and was made possible in part by the CIRM Major Facilities grant (FA1-00607) to the Sanford Consortium for Regenerative Medicine. D.T. was supported by a Scholar Award (1657-13) from The Leukemia and Lymphoma Society and CIRM (RB4-06158). This project was also supported by the NIH/National Heart, Lung, and Blood Institute grant R01HL135205 (awarded to D.T. and K.W.) and by the NIH/National Institute of General Medical Sciences grant R01GM110304 (awarded to K.W.). J.M.W. was supported by the Graduate Training in Cellular and Molecular Pharmacology Training Grant NIH T32 GM007752. I.A.D. was supported by the Harvard Stem Cell Institute and NIDDK grant UH2/3DK107372.
Author information
Authors and Affiliations
Contributions
S.G. conceived, designed and conducted the experiments and analysis, and wrote the manuscript. N.N. and J.R. designed and conducted the experiments and analysis. J.S., B.L., R.B., C.H.O. and J.H. conducted the experiments and analysis. J.M.W., M.C.-T. and D.G. performed MS and conducted the analysis. C.N.K. and I.A.D. provided the zebrafish lines. D.T. and K.W. supervised the experiments and edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Fig. 1 Fzd screening.
a, Schematic of strategy to screen for fzd expression in endothelial cells as marked by fli1a-GFP at 16.5 hpf. b, Expression of all 14 fzd transcripts in Fli1a+ cells at 16.5 hpf; n = 1 experiment shown as representative data from qPCR on pooled sorted cells; trend reproduced in 3 independent experiments. Bars represent the mean of three technical replicates. c, In the Wnt-off state, the STF reporter is silent and luciferase is not expressed. d, In the presence of a Wnt signal, β-catenin is translocated to the nucleus, where it activates the STF reporter and luciferase is expressed. e, Quantification of the HEK293T cell STF assay with Wnt9a and Fzd8a or Fzd9b in the context of STF or FOP:FLASH reporters; n = 3 biological replicates each. f, Quantification of the HEK293T (FZD1, FZD2 and FZD7 KO) cell STF assay with zebrafish Wnt3a and zebrafish Fzd8a, Fzd9a, Fzd9b or human FZD7; n = 3 biological replicates each. In all STF assays, the bars represent the mean, the error bars represent the standard deviation and the dots represent individual biological replicates; one-way ANOVA with Dunnett’s test. All STF assays were repeated independently with a similar trend.
Supplementary Fig. 2 Fzd9b is required for HSPC development.
a, Endothelial cell emerging from the aorta is labelled by fzd9b:Gal4; UAS:Cre; loxP-BFP-loxP-dsRed (pseudo-coloured green) at 3 dpf, during HSPC emergence (left). Scale bar, 10 µm. Cells in the thymus are labelled by fzd9b:Gal4; UAS:Cre; loxP-BFP-loxP-dsRed at 7 dpf (right). Scale bar, 25 µm. Images are representative of ten embryos from three experiments. b, Expression of Fzd9b-mKate fusion protein in fzd9b MO injected samples and no MO control at 3 dpf. Scale bar, 200 µm. Representative from n = 10 biological samples. c, Quantification of Runx1+ cells in the aorta of uninjected (n = 15) and fzd9b MO injected (n = 12) fish at 30 hpf. d, Quantification of runx1 transcripts by qPCR in uninjected and fzd9b MO injected fish at 30 hpf; n = 3 zebrafish each. e, Nascent HSPCs were labelled by gata2b and kdrl fluorescent transgenes at 46 hpf in uninjected and fzd9b MO injected fish. Scale bar, 10 µm. f, Quantification of Gata2b+ cells per millimetre of the aortic floor from e; n = 5 biological replicates each. g, WISH for the T cell marker rag1 at 5 dpf in uninjected and fzd9b MO injected fish. Scale bar, 200 µm. Representative from 30 biological samples as indicated. h, WISH for aortic (dlc, dll4 and notch1b), vascular (kdrl) and pronephros (cdh17) markers in uninjected and fzd9b MO-injected fish at 24 hpf. Scale bar, 500 µm. The numbers in the upper right corners represent the number of animals with the shown phenotype, over the total examined. i, WISH for cmyb in uninjected, fzd9b MO injected and fzd9b MO + fzd9b mRNA injected fish at 40 hpf. Scale bar, 30 µm. j, Quantification of i; n = 10, 7 and 10 biological replicates from left to right. k, Quantification of Cmyb+ cells in the aorta of fzd9b mutants and siblings injected with fzd9b MO or uninjected controls; n = 16, 6, 16 and 9 biological replicates from left to right. l, WISH for cmyb at 40 hpf in endothelial-specific fzd9b mutants. Scale bar, 30 µm. m, Quantification of l; n = 13 and 8 from left to right. In all graphs, each dot represents a biological replicate, the bars represent the mean and the error bars represent the standard deviation. Statistical analyses were done by ANOVA compared to controls as indicated. All STF assays were repeated independently with a similar trend.
Supplementary Fig. 3 WNT9A and FZD9 are required for HSPC development.
Expression of WNT9A (n = 4 biological replicates each) (a), FZD9 (n = 3 biological replicates each) (b), CD34 (n = 8 biological replicates each) (c), CD31 (n = 8 biological replicates each) (d), CD45 (n = 7, 8 and 8 biological replicates from left to right) (e) and CMYB (n = 7, 4 and 8 biological replicates from left to right) (f), after 14 d of haematopoietic-directed differentiation in shControl, shWNT9A and shFZD9 cells as measured by qPCR. The bars represent the mean and the error bars represent the standard deviation of biological replicates from three experiments, which are represented by individual dots. Statistical analyses were done by an unpaired, two-tailed t-test in a and b, and by ANOVA in c–f, compared to control shRNA. Representative FACS lots of TRA1-81 and SSEA4 staining in undifferentiated control (g), WNT9A (h) and FZD9 (i) shRNA-expressing hES cells; representative plots from two independent experiments. All STF assays were repeated independently with a similar trend.
Supplementary Fig. 4 LRP6-KO HEK293T STF cell line validation.
a, WISH for cmyb at 40 hpf in zebrafish injected with suboptimal MO dosages (0.1 ng wnt9a and 0.5 ng fzd9b). For quantification, see Fig. 2g. Scale bar, 30 µm. b, Quantification of the HEK293T cell STF assay with zebrafish Wnt9a, Fzd9b and Lrp6; n = 3 biological replicates each. Statistical analyses are compared to Wnt9a alone. c, Quantification of the HEK293T cell STF assay screen with mouse Wnt3a in WT and LRP6-KO cells; n = 3 biological replicates each. Statistical analyses were done by ANOVA as indicated. d, Quantification of the HEK293T cell STF assay with 500 nM CHIR in WT and LRP6-KO cells; n = 3 biological replicates each. In all graphs, the dots represent biological replicates from one experiment, the bars represent the mean and the error bars represent the standard deviation. Statistical analyses were done by ANOVA compared to controls as indicated. All STF assays were repeated independently with a similar trend.
Supplementary Fig. 5 Fzd9b intracellular domains are required for signalling.
a,b, Immunoblots of Fzd9b-V5 chimaeric proteins. Representative of two independent experiments. See also Supplementary Fig. 7. c, Quantification of the HEK293T cell STF assay with zebrafish Wnt9a, Fzd9b and Fzd4 chimaeras; n = 3 biological replicates each. d, Quantification of the HEK293T cell STF assay with human WNT9A, FZD9 and FZD4 chimaeras; n = 3 biological replicates each. e, Schematic of Fzd9b indicating the immunogen region used for antibody generation. f, Non-permeabilized immunofluorescence for Fzd9b chimaeras. Scale bar, 30 µm. Data are representative of ten fields of view from two independent experiments. g, Histograms of flow cytometry for Fzd9b chimaeras. Data are representative of two independent experiments using 10,000 cells for each condition. All STF assays were repeated independently with a similar trend. In c,d, the dots represent biological replicates from a single experiment, the bars represent the mean and the error bars represent the standard deviation; statistical analyses were by ANOVA compared to controls as indicated.
Supplementary Fig. 6 Fzd9b-5GS-APEX2 cell line validation and EGFR mediates Wnt9a–Fzd9b signalling.
a, Immunoblot for streptavidin after treatment of cells with vehicle, biotin-phenol, H2O2 or both, representative of five independent experiments. See also Supplementary Fig. 7. b, Immunoblots of cells for Fzd9b, FLAG (APEX2) and actin; representative of three independent experiments. See also Supplementary Fig. 7. c, Immunofluorescence for FLAG showing APEX2 localization on the membrane; representative of three independent experiments. Scale bar, 30 µm. d, Quantification of the HEK293T cell STF assay with zebrafish Wnt9a conditioned medium, WT Fzd9b and Fzd9b-5GS-APEX2; n = 3 biological replicates each; statistical analysis was done by ANOVA compared to Wnt9a conditioned medium alone. e, Fold enrichment of GO terms for biological processes identified from the top 5% of changed proteins in Fzd9b-APEX2 HEK293T cells treated with Wnt9a. The P values are listed next to each bar; n = 3 biological replicates; see the Methods for analysis and statistics. f, Normalized intensity of the top three transmembrane proteins detected in the APEX assay; n = 3 biological replicates. g, Immunoblot for EGFR in siControl-treated or siEGFR-treated HEK293T cells; representative of three independent experiments. See also Supplementary Fig. 7. h, Quantification of the HEK293T cell STF assay with human WNT9A and FZD9 with siEGFR or siControl; n = 3 biological replicates. i, WISH for cmyb at 40 hpf; quantified in j; n = 10 biological replicates each. Scale bar, 30 µm. k,l, STF assay with zebrafish Wnt9a and Fzd9b (k; n = 8 biological replicates each) or human WNT9A and FZD9 (l; n = 3 biological replicates each), treated with the selective EGFR inhibitor AG1478. m, Expression of runx1 and gata2b in vehicle and EGFR inhibitor zebrafish treated from 15–36 hpf; n = 4 biological replicates each. n, Expression of known EGFR ligands as transcripts per million (TPM) in HEK293T cells as reported from RNA-sequencing data sets42. The red dashed line indicates the commonly accepted cut-off for expression levels by TPM; n = 6 biological replicates from 2 experiments. o, Quantification of the HEK293T cell STF assay in Fzd9b-expressing cells cultured in serum-free conditions with neutralizing antibody for HBEGF as indicated; n = 3 biological replicates each. In all graphs, the dots represent biological replicates from a single experiment (except for panel n, which is pooled from two experiments), the bars represent the mean and the error bars represent the standard deviation. Statistical analyses were done by ANOVA compared to controls as indicated. All STF assays were repeated independently with a similar trend.
Supplementary Fig. 7 Immunoblots.
The original immunoblot images used in this study. Boxes indicate the cropped region shown in respective figures as indicated.
Supplementary information
Supplementary Information
Supplementary Figures 1–7, Supplementary Table titles and legends
Supplementary Table 1
Primers used
Supplementary Table 2
Source reagents
Supplementary Table 3
Statistics source data
Rights and permissions
About this article
Cite this article
Grainger, S., Nguyen, N., Richter, J. et al. EGFR is required for Wnt9a–Fzd9b signalling specificity in haematopoietic stem cells. Nat Cell Biol 21, 721–730 (2019). https://doi.org/10.1038/s41556-019-0330-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-019-0330-5
This article is cited by
-
Non-canonical non-genomic morphogen signaling in anucleate platelets: a critical determinant of prothrombotic function in circulation
Cell Communication and Signaling (2024)
-
Cross-linking of the endolysosomal system reveals potential flotillin structures and cargo
Nature Communications (2022)
-
Cardiac Wnt5a and Wnt11 promote fibrosis by the crosstalk of FZD5 and EGFR signaling under pressure overload
Cell Death & Disease (2021)
-
Deciphering molecular interactions by proximity labeling
Nature Methods (2021)
-
RAC1B modulates intestinal tumourigenesis via modulation of WNT and EGFR signalling pathways
Nature Communications (2021)