Abstract
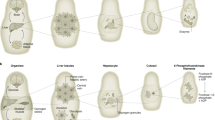
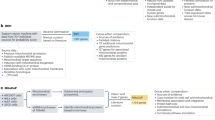
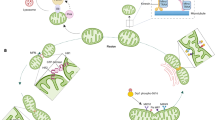
Although classically appreciated for their role as the powerhouse of the cell, the metabolic functions of mitochondria reach far beyond bioenergetics. In this Review, we discuss how mitochondria catabolize nutrients for energy, generate biosynthetic precursors for macromolecules, compartmentalize metabolites for the maintenance of redox homeostasis and function as hubs for metabolic waste management. We address the importance of these roles in both normal physiology and in disease.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Castresana, J. & Saraste, M. Evolution of energetic metabolism: the respiration-early hypothesis. Trends Biochem. Sci. 20, 443–448 (1995).
Lyons, T. W., Reinhard, C. T. & Planavsky, N. J. The rise of oxygen in Earth’s early ocean and atmosphere. Nature 506, 307–315 (2014).
Sagan, L. On the origin of mitosing cells. J. Theoret. Biol 14, 225–274 (1966).
Timmis, J. N., Ayliffe, M. A., Huang, C. Y. & Martin, W. Endosymbiotic gene transfer: organelle genomes forge eukaryotic chromosomes. Nat. Rev. Genet. 5, 123–135 (2004).
Vyas, S., Zaganjor, E. & Haigis, M. C. Mitochondria and cancer. Cell 166, 555–566 (2016).
Walsh, C. T., Tu, B. P. & Tang, Y. Eight kinetically stable but thermodynamically activated molecules that power cell metabolism. Chem. Rev. 118, 1460–1494 (2017).
Sazanov, L. A. A giant molecular proton pump: structure and mechanism of respiratory complex I. Nat. Rev. Mol Cell Biol. 16, 375–388 (2015).
Watt, I. N., Montgomery, M. G., Runswick, M. J., Leslie, A. G. & Walker, J. E. Bioenergetic cost of making an adenosine triphosphate molecule in animal mitochondria. Proc. Natl Acad Sci. USA 107, 16823–16827 (2010).
DiMauro, S. & Schon, E. A. Mitochondrial respiratory-chain diseases. New Eng. J. Med. 348, 2656–2668 (2003).
Dimauro, S. & Rustin, P. A critical approach to the therapy of mitochondrial respiratory chain and oxidative phosphorylation diseases. Biochim. Biophys. acta 1159–1167, 2009 (1792).
Nelson, D. L. & Cox, M. M. Lehninger Principles of Biochemistry. (W.H. Freeman and company, New York, 2005).
Hui, S. et al. Glucose feeds the TCA cycle via circulating lactate. Nature 551, 115–118 (2017).
Faubert, B. et al. Lactate metabolism in human lung tumors. Cell 171, 358–371 (2017).
Vander Heiden, M. G., Cantley, L. C. & Thompson, C. B. Understanding the Warburg effect: the metabolic requirements of cell proliferation. Science 324, 1029–1033 (2009).
Herzig, S. et al. Identification and functional expression of the mitochondrial pyruvate carrier. Science 337, 93–96 (2012).
Bricker, D. K. et al. A mitochondrial pyruvate carrier required for pyruvate uptake in yeast, Drosophila, and humans. Science 337, 96–100 (2012).
Papandreou, I., Cairns, R. A., Fontana, L., Lim, A. L. & Denko, N. C. HIF-1 mediates adaptation to hypoxia by actively downregulating mitochondrial oxygen consumption. Cell Metab. 3, 187–197 (2006).
Connett, R. J., Honig, C. R., Gayeski, T. E. J. & Brooks, G. A. Defining hypoxia: a systems view of VO2, glycolysis, energetics and intracellular PO2. J. Appl. Physiol. 68, 833–842 (1990).
Sherratt, H. S. A. The metabolism of the small intestine: oxygen uptake and L-lactate production along the length of the small intestine of rat and guinea pig. Comp. Biochem. Physiol 24, 745–761 (1968).
Gerich, J. E. Role of the kidney in normal glucose homeostasis and in the hyperglycaemia of diabetes mellitus: therapeutic implications. Diabetic Med. 27, 136–142 (2010).
Diaz-Ruiz, R., Rigoulet, M. & Devin, A. The Warburg and Crabtree effects: on the origin of cancer cell energy metabolism and of yeast glucose repression. Biochim. Biophys. acta 1807, 568–576 (2011).
Schell, J. C. et al. A role for the mitochondrial pyruvate carrier as a repressor of the Warburg effect and colon cancer cell growth. Mol. Cell 56, 400–413 (2014).
Divakaruni, A. S. et al. Thiazolidinediones are acute, specific inhibitors of the mitochondrial pyruvate carrier. Proc. Natl Acad. Sci. USA 110, 5422–5427 (2013).
Schell, J. C. et al. Control of intestinal stem cell function and proliferation by mitochondrial pyruvate metabolism. Nat. Cell Biol. 19, 1027–1036 (2017).
Olson, K. A., Schell, J. C. & Rutter, J. Pyruvate and metabolic flexibility: illuminating a path toward selective cancer therapies. Trends Biochem. Sci. 41, 219–230 (2016).
Metallo, C. M., Walther, J. L. & Stephanopoulos, G. Evaluation of 13C isotopic tracers for metabolic flux analysis in mammalian cells. J. Biotech. 144, 167–174 (2009).
Sellers, K. et al. Pyruvate carboxylase is critical for non-small-cell lung cancer proliferation. J. Clin. Invest. 125, 687–698 (2015).
Patel, K. P., O’Brien, T. W., Subramony, S. H., Shuster, J. & Stacpoole, P. W. The spectrum of pyruvate dehydrogenase complex deficiency: clinical, biochemical and genetic features in 371 patients. Mol. Genet. Metab. 105, 34–43 (2012).
Christen, S. et al. Breast cancer-derived lung metastases show increased pyruvate carboxylase-dependent anaplerosis. Cell Rep. 17, 837–848 (2016).
Davidson, S. M. et al. Environment impacts the metabolic dependencies of Ras-driven non-small cell lung cancer. Cell Metab. 23, 517–528 (2016).
DeBerardinis, R. J. & Cheng, T. Q’s next: the diverse functions of glutamine in metabolism, cell biology and cancer. Oncogene 29, 313–324 (2010).
DeBerardinis, R. J. et al. Beyond aerobic glycolysis: transformed cells can engage in glutamine metabolism that exceeds the requirement for protein and nucleotide synthesis. Proc. Natl Acad. Sci. USA 104, 19345–19350 (2007).
Haigis, M. C. et al. SIRT4 inhibits glutamate dehydrogenase and opposes the effects of calorie restriction in pancreatic beta cells. Cell 126, 941–954 (2006).
Yang, C. et al. Glutamine oxidation maintains the TCA cycle and cell survival during impaired mitochondrial pyruvate transport. Mol. Cell 56, 414–424 (2014).
Vacanti, N. M. et al. Regulation of substrate utilization by the mitochondrial pyruvate carrier. Mol. Cell 56, 425–435 (2014).
Wang, R. et al. The transcription factor Myc controls metabolic reprogramming upon T lymphocyte activation. Immunity 35, 871–882 (2011).
Wise, D. R. et al. Myc regulates a transcriptional program that stimulates mitochondrial glutaminolysis and leads to glutamine addiction. Proc. Natl Acad. Sci. 105, 18782–18787 (2008).
Wang, J. B. et al. Targeting mitochondrial glutaminase activity inhibits oncogenic transformation. Cancer Cell 18, 207–219 (2010).
Jin, L., Alesi, G. N. & Kang, S. Glutaminolysis as a target for cancer therapy. Oncogene 35, 3619–3625 (2016).
Muir, A. et al. Environmental cystine drives glutamine anaplerosis and sensitizes cancer cells to glutaminase inhibition. eLife 6, e27713 (2017).
Bhutia, Y. D. & Ganapathy, V. Glutamine transporters in mammalian cells and their functions in physiology and cancer. Biochim. Biophys. acta 1863, 2531–2539 (2016).
Sastrasinh, A. & Sastrasinh, M. Glutamine transport in submitochondrial particles. Am. J. Physiol. 257, 1050–1058 (1989).
Indiveri, C., Abruzzo, G., Stipani, I. & Palmieri, F. Identification and purification of the reconstitutively active glutamine carrier from rat kidney mitochondria. Biochem. J. 333, 285–290 (1998).
Brosnan, J. T. & Brosnan, M. E. Branched-chain amino acids: metabolism, physiological function and application. J. Nutr. 136, 207–211 (2006).
Mayers, J. R. et al. Tissue of origin dictates branched-chain amino acid metabolism in mutant Kras-driven cancers. Science 353, 1161–1165 (2016).
Shimomura, Y., Murakami, T., Nakai, N., Nagasaki, M. & Harris, R. A. Exercise promotes BCAA catabolism: effects of BCAA supplementation on skeletal muscle during exercise. J. Nutr. 134, 1583–1587 (2004).
Green, C. R. et al. Branched-chain amino acid catabolism fuels adipocyte differentiation and lipogenesis. Nat. Chem. Biol. 12, 15–21 (2016).
Blackburn, P. R. et al. Maple syrup urine disease: mechanisms and management. Appl. Clin. Genet. 10, 57–66 (2017).
Rohrig, F. & Schulze, A. The multifaceted roles of fatty acid synthesis in cancer. Nat. Rev. Cancer 16, 732–749 (2016).
McGarry, J. D. & Brown, N. F. The mitochondrial carnitine palmitoyltransferase system from concept to molecular analysis. Eur. J. Biochem. 244, 1–14 (1997).
Houten, S. M. & Wanders, R. J. A. A general introduction to the biochemistry of mitochondrial fatty acid β-oxidation. J. Inherit. Metab. Dis. 33, 469–477 (2010).
Schoors, S. et al. Fatty acid carbon is essential for dNTP synthesis in endothelial cells. Nature 520, 192–197 (2015).
Ruderman, N. & Prentki, M. AMP kinase and malonyl-CoA: targets for therapy of the metabolic syndrome. Nat. Rev. Drug Discov. 3, 340–351 (2004).
Munday, M. R. Regulation of mammalian acetyl-CoA carboxylase. Biochem. Soc. Trans. 30, 1059–1063 (2002).
German, N. J. et al. PHD3 loss in cancer enables metabolic reliance on fatty acid oxidation via deactivation of ACC2. Mol. Cell 63, 1006–1020 (2016).
Currie, E., Schulze, A., Zechner, R., Walther, T. C. & Farese, R. V. Jr. Cellular fatty acid metabolism and cancer. Cell Metab. 18, 153–161 (2013).
Svensson, R. U. et al. Inhibition of acetyl-CoA carboxylase suppresses fatty acid synthesis and tumor growth of non-small-cell lung cancer in preclinical models. Nat. Med. 22, 1108–1119 (2016).
Garcia, D. & Shaw, R. J. AMPK: mechanisms of cellular energy sensing and restoration of metabolic balance. Mol. Cell 66, 789–800 (2017).
Carracedo, A., Cantley, L. C. & Pandolfi, P. P. Cancer metabolism: fatty acid oxidation in the limelight. Nat. Rev. Cancer 13, 227–232 (2013).
Pearce, E. L. et al. Enhancing CD8 T-cell memory by modulating fatty acid metabolism. Nature 460, 103–107 (2009).
O’Sullivan, D. et al. Memory CD8+ T cells use cell-intrinsic lipolysis to support the metabolic programming necessary for development. Immunity 41, 75–88 (2014).
Stern, J. H., Rutkowski, J. M. & Scherer, P. E. Adiponectin, leptin, and fatty acids in the maintenance of metabolic homeostasis through adipose tissue crosstalk. Cell Metab. 23, 770–784 (2016).
Serra, D., Mera, P., Malandrino, M. I., Mir, J. F. & Herrero, L. Mitochondrial fatty acid oxidation in obesity. Antioxid. Redox Sign. 19, 269–284 (2013).
Ahn, C. S. & Metallo, C. M. Mitochondria as biosynthetic factories for cancer proliferation. Cancer Metab. 3, 1 (2015).
Walker, M. A., Volpi, S., Sims, K. B., Walter, J. E. & Traggiai, E. Powering the immune system: mitochondria in immune function and deficiency. J. Immunol. Res. 2014, 164309 (2014).
Ducker, G. S. & Rabinowitz, J. D. One-carbon metabolism in health and disease. Cell Metab. 25, 27–42 (2017).
Tibbetts, A. S. & Appling, D. R. Compartmentalization of mammalian folate-mediated one-carbon metabolism. Annu. Rev. Nutrit. 30, 57–81 (2010).
Chasin, L. A., Feldman, A., Konstam, M. & Urlaub, G. Reversion of a chinese hamster cell auxotrophic mutant. Proc. Natl Acad. Sci. USA 71, 718–722 (1974).
Ducker, G. S. et al. Reversal of cytosolic one-carbon flux compensates for loss of the mitochondrial folate pathway. Cell Metab. 23, 1140–1153 (2016).
Ben-Sahra, I., Hoxhaj, G., Ricoult, S. J. H., Asara, J. M. & Manning, B. D. mTORC1 induces purine synthesis through control of the mitochondrial tetrahydrofolate cycle. Science 351, 728–733 (2016).
Nilsson, R. et al. Metabolic enzyme expression highlights a key role for MTHFD2 and the mitochondrial folate pathway in cancer. Nat. Commun. 5, 3128 (2014).
Ron-Harel, N. et al. Mitochondrial biogenesis and proteome remodeling promote one-carbon metabolism for T cell activation. Cell Metab. 24, 104–117 (2016).
Munier-Lehmann, H., Vidalain, P. O., Tangy, F. & Janin, Y. L. On dihydroorotate dehydrogenases and their inhibitors and uses. J. Med. Chem. 56, 3148–3167 (2013).
Herrmann, M. L., Schleyerbach, R. & Kirschbaum, B. J. Leflunomide: an immunomodulatory drug for the treatment of rheumatoid arthritis and other autoimmune diseases. Immunopharmacology 47, 273–289 (2000).
Sykes, D. B. et al. Inhibition of dihydroorotate dehydrogenase overcomes differentiation blockade in acute myeloid leukemia. Cell 167, 171–186 (2016).
Jeong, S. M. et al. SIRT4 has tumor-suppressive activity and regulates the cellular metabolic response to DNA damage by inhibiting mitochondrial glutamine metabolism. Cancer Cell 23, 450–463 (2013).
Brown, K. K., Spinelli, J. B., Asara, J. M. & Toker, A. Adaptive reprogramming of de novo pyrimidine synthesis is a metabolic vulnerability in triple-negative breast cancer. Cancer Discov. 7, 391–399 (2017).
Hatzivassiliou, G. et al. ATP citrate lyase inhibition can suppress tumor cell growth. Cancer Cell 8, 311–321 (2005).
Metallo, C. M. et al. Reductive glutamine metabolism by IDH1 mediates lipogenesis under hypoxia. Nature 481, 380–384 (2012).
Mullen, A. R. et al. Reductive carboxilation supports growth in tumor cells with defective mitochondria. Nature 481, 385–388 (2012).
Jiang, L. et al. Reductive carboxylation supports redox homeostasis during anchorage-independent growth. Nature 532, 255–258 (2016).
Sun, J., Aluvila, S., Kotaria, R., Mayor, J. A., Walters, E. D. & Kaplan, R. Mitochondrial and plasma membrane citrate transporters: discovery of selective inhibitors and application to structure/function analysis. Mol. Cell Pharmacol. 2, 101–110 (2010).
Kim, C. W. et al. Induced polymerization of mammalian acetyl-CoA carboxylase by MIG12 provides a tertiary level of regulation of fatty acid synthesis. Proc. Natl Acad. Sci. USA 107, 9626–9631 (2010).
Glerum, D. M., Claeys, D., Mertens, W. SpringerAmpamp; Azzi, A. The tricarboxylate carrier from rat liver mitochondria. Euro. J. Biochem. 194, 681–684 (1990).
Zara, V. & Gnoni, G. V. V. Effect of starvation on the activity of the mitochondrial tricarboxylate carrier. Biochim. Biophys. acta 1239, 33–38 (1995).
Lee, J. V. et al. Akt-dependent metabolic reprogramming regulates tumor cell histone acetylation. Cell Metab. 20, 306–319 (2014).
Londono Gentile, T. et al. DNMT1 is regulated by ATP-citrate lyase and maintains methylation patterns during adipocyte differentiation. Mol. Cell. Biol. 33, 3864–3878 (2013).
Wellen, K. E. et al. ATP-citrate lyase links cellular metabolism to histone acetylation. Science 324, 1076–1080 (2009).
Xiao, M. et al. Inhibition of alpha-KG-dependent histone and DNA demethylases by fumarate and succinate that are accumulated in mutations of FH and SDH tumor suppressors. Genes Dev. 26, 1326–1338 (2012).
Yang, L. et al. Targeting stromal glutamine synthetase in tumors disrupts tumor microenvironment-regulated cancer cell growth. Cell Metab. 24, 685–700 (2016).
Matthews, G. D., Gur, N., Koopman, W. J., Pines, O. & Vardimon, L. Weak mitochondrial targeting sequence determines tissue-specific subcellular localization of glutamine synthetase in liver and brain cells. J. Cell Sci. 123, 351–359 (2010).
Tardito, S. et al. Glutamine synthetase activity fuels nucleotide biosynthesis and supports growth of glutamine-restricted glioblastoma. Nat. Cell Biol. 17, 1556–1568 (2015).
Spinelli, J. B. et al. Metabolic recycling of ammonia via glutamate dehydrogenase supports breast cancer biomass. Science 358, 941–946 (2017).
Frigerio, F., Casimir, M., Carobbio, S. & Maechler, P. Tissue specificity of mitochondrial glutamate pathways and the control of metabolic homeostasis. Biochim. Biophys. acta 1777, 965–972 (2008).
Coloff, J. L. et al. Differential glutamate metabolism in proliferating and quiescent mammary epithelial cells. Cell Metab. 23, 867–880 (2016).
Perez-Arellano, I., Carmona-Alvarez, F., Martinez, A. I., Rodriguez-Diaz, J. & Cervera, J. Pyrroline-5-carboxylate synthase and proline biosynthesis: from osmotolerance to rare metabolic disease. Protein Sci. 19, 372–382 (2010).
Palmieri, F. The mitochondrial transporter family SLC25: identification, properties and physiopathology. Mol. Med. 34, 465–484 (2013).
Bahl, J. J., Matsuda, M., DeFronzo, R. A. & Bressler, R. In vitro and in vivo suppression of gluconeogenesis by inhibition of pyruvate carboxylase. Biochem. Pharmacol. 53, 67–74 (1997).
Kumashiro, N. et al. Targeting pyruvate carboxylase reduces gluconeogenesis and adiposity and improves insulin resistance. Diabetes 62, 2183–2194 (2013).
Monne, M. & Palmieri, F. Antiporters of the mitochondrial carrier family. Curr. Topics Membranes 73, 289–320 (2014).
Yang, J., Kalhan, S. C. & Hanson, R. W. What is the metabolic role of phosphoenolpyruvate carboxykinase? J. Biol. Chem. 284, 27025–27029 (2009).
Dailey, H. A. & Meissner, P. N. Erythroid heme biosynthesis and its disorders. Cold Spring Harb. Persp. Med. 3, a011676 (2013).
Ulrich, D. L. et al. ATP-dependent mitochondrial porphyrin importer ABCB6 protects against phenylhydrazine toxicity. J. Biol. Chem. 287, 12679–12690 (2012).
Bonkowsky, H. L., Bloomer, J. R., Ebert, P. S. & Mahoney, M. J. Heme synthetase deficiency in human protoporphyria: demonstration of the defect in liver and cultured skin fibroblasts. J. Clin. Investig. 56, 1139–1148 (1975).
Yang, H. et al. Nutrient-sensitive mitochondrial NAD+ levels dictate cell survival. Cell 130, 1095–1107 (2007).
Titov, D. V. et al. Complementation of mitochondrial electron transport chain by manipulation of the NAD+/NADH ratio. Science 352, 231–235 (2016).
van de Ven, R. A. H., Santos, D. & Haigis, M. C. Mitochondrial sirtuins and molecular mechanisms of aging. Trends Mol. Med. 23, 320–331 (2017).
Stein, L. R. & Imai, S. The dynamic regulation of NAD metabolism in mitochondria. Trends Endocrinol. Metab. 23, 420–428 (2012).
Todisco, S., Agrimi, G., Castegna, A. & Palmieri, F. Identification of the mitochondrial NAD+ transporter in Saccharomyces cerevisiae. J. Biol. Chem. 281, 1524–1531 (2006).
Schantz, P. G., Sjoberg, B. & Svedenhag, J. Malate-aspartate and alpha-glycerophosphate shuttle enzyme levels in human skeletal muscle: methodological considerations and effect of endurance training. Acta Physiol. Scand. 128, 397–407 (1986).
Wallace, D. C. Mitochondria and cancer. Nat. Rev. Cancer 12, 685–698 (2012).
Hayes, D. J. et al. An unusual metabolic myopathy: a malate—aspartate shuttle defect. J. Neurol. Sci. 82, 27–39 (1987).
Son, J. et al. Glutamine supports pancreatic cancer growth through a KRAS-regulated metabolic pathway. Nature 496, 101–105 (2013).
Sullivan, L. B. et al. Supporting aspartate biosynthesis is an essential function of respiration in proliferating cells. Cell 162, 552–563 (2015).
Birsoy, K. et al. An essential role of the mitochondrial electron transport chain in cell proliferation is to enable aspartate synthesis. Cell 162, 540–551 (2015).
Gnoni, G. V., Priore, P., Geelen, M. J. & Siculella, L. The mitochondrial citrate carrier: metabolic role and regulation of its activity and expression. IUBMB Life 61, 987–994 (2009).
Palmieri, F. Diseases caused by defects of mitochondrial carriers: a review. Biochim. Biophys. acta 1777, 564–578 (2008).
Mracek, T., Drahota, Z. & Houstek, J. The function and the role of the mitochondrial glycerol-3-phosphate dehydrogenase in mammalian tissues. Biochim. Biophys. acta 1827, 401–410 (2013).
Fan, J. et al. Quantitative flux analysis reveals folate-dependent NADPH production. Nature 510, 298–302 (2014).
Lewis, C. A. et al. Tracing compartmentalized NADPH metabolism in the cytosol and mitochondria of mammalian cells. Mol. Cell 55, 253–263 (2014).
Bao, X. R. et al. Mitochondrial dysfunction remodels one-carbon metabolism in human cells. eLife 5, e10575 (2016).
Tedeschi, P. M. et al. Contribution of serine, folate and glycine metabolism to the ATP, NADPH and purine requirements of cancer cells. Cell Death Dis. 4, e877 (2013).
Schieber, M. & Chandel, N. S. ROS function in redox signaling and oxidative stress. Curr. Biol. 24, 453–462 (2014).
Wallace, J. L. & Wang, R. Hydrogen sulfide-based therapeutics: exploiting a unique but ubiquitous gasotransmitter. Nat. Rev. Drug Discov. 14, 329–345 (2015).
Lyssiotis, C. A. & Kimmelman, A. C. Metabolic interactions in the tumor microenvironment. Trends Cell Biol. 27, 863–875 (2017).
Rai, R., Saraswat, V. A. & Dhiman, R. K. Gut microbiota: its role in hepatic encephalopathy. J. Clin. Exp. Hepatol. 5, 29–36 (2015).
Eng, C. H., Yu, K., Lucas, J., White, E. & Abraham, R. T. Ammonia derived from glutaminolysis is a diffusible regulator of autophagy. Sci. Signal. 3, ra31 (2010).
Braissant, O., McLin, V. A. & Cudalbu, C. Ammonia toxicity to the brain. J. Inherit. Metab. Dis. 36, 595–612 (2013).
Morris, S. M. Jr. Regulation of enzymes of the urea cycle and arginine metabolism. Annu. Rev. Nutrit. 22, 87–105 (2002).
McCudden, C. R. & Powers-Lee, S. G. Required allosteric effector site for N-acetylglutamate on carbamoyl-phosphate synthetase I. J. Biol. Chem. 271, 18285–18294 (1996).
Kim, J. et al. CPS1 maintains pyrimidine pools and DNA synthesis in KRAS/LKB1-mutant lung cancer cells. Nature 546, 168–172 (2017).
Jones, E. A., Smallwood, R. A., Craigie, A. & Rosenoer, V. M. The enterohepatic circulation of urea nitrogen. Clin. Sci. 37, 825–836 (1969).
Brusilow, S. W. Urea cycle disorders: clinical paradigm of hyperammonemic encephalopathy. Prog. Liver Dis. 13, 293–309 (1995).
Kenny, D. J. & Balskus, E. P. Engineering chemical interactions in microbial communities. Chem. Soc. Rev. 47, 1705–1729 (2017).
Treberg, J. R., Brosnan, M. E., Watford, M. & Brosnan, J. T. On the reversibility of glutamate dehydrogenase and the source of hyperammonemia in the hyperinsulinism/hyperammonemia syndrome. Advanc. Enzyme Reg. 50, 34–43 (2010).
Sena, L. A. & Chandel, N. S. Physiological roles of mitochondrial reactive oxygen species. Mol. Cell 48, 158–167 (2012).
Block, K., Gorin, Y. & Abboud, H. E. Subcellular localization of Nox4 and regulation in diabetes. Proc. Natl Acad. Sci. USA 106, 14385–14390 (2009).
Balaban, R. S., Nemoto, S. & Finkel, T. Mitochondria, oxidants, and aging. Cell 120, 483–495 (2005).
Chang, T. S. et al. Peroxiredoxin III, a mitochondrion-specific peroxidase, regulates apoptotic signaling by mitochondria. J. Biol. Chem. 279, 41975–41984 (2004).
Schneider, M. et al. Mitochondrial glutathione peroxidase 4 disruption causes male infertility. FASEB 23, 3233–3242 (2009).
Cox, A. G., Winterbourn, C. C. & Hampton, M. B. Mitochondrial peroxiredoxin involvement in antioxidant defence and redox signalling. Biochem. J. 425, 313–325 (2009).
Rhee, S. G. Cell signaling: H2O2, a necessary evil for cell signaling. Science 312, 1882–1883 (2006).
Bae, Y. S. et al. Epidermal growth factor (EGF)-induced generation of hydrogen peroxide. J. Biol. Chem. 272, 217–221 (1997).
Leslie, N. R. et al. Redox regulation of PI 3-kinase signalling via inactivation of PTEN. EMBO J. 22, 5501–5510 (2003).
Chandel, N. S. et al. Reactive oxygen species generated at mitochondrial complex III stabilize hypoxia-inducible factor-1alpha during hypoxia: a mechanism of O2 sensing. J. Biol. Chem. 275, 25130–25138 (2000).
Finley, L. W. et al. SIRT3 opposes reprogramming of cancer cell metabolism through HIF1alpha destabilization. Cancer Cell 19, 416–428 (2011).
West, A. P., Shadel, G. S. & Ghosh, S. Mitochondria in innate immune responses. Nat. Rev. Immunol. 11, 389–402 (2011).
Sena, L. A. et al. Mitochondria are required for antigen-specific T cell activation through reactive oxygen species signaling. Immunity 38, 225–236 (2013).
Wheeler, M. L. & Defranco, A. L. Prolonged production of reactive oxygen species in response to B cell receptor stimulation promotes B cell activation and proliferation. J. Immunol. 189, 4405–4416 (2012).
Linden, D. R., Levitt, M. D., Farrugia, G. & Szurszewski, J. H. Endogenous production of H2S in the gastrointestinal tract: still in search of a physiologic function. Antioxid. Redox Signal. 12, 1135–1146 (2010).
Levitt, M. D., Abdel-Rehim, M. S. & Furne, J. Free and acid-labile hydrogen sulfide concentrations in mouse tissues: anomalously high free hydrogen sulfide in aortic tissue. Antioxid. Redox Signal. 15, 373–378 (2011).
Luna-Sanchez, M. et al. CoQ deficiency causes disruption of mitochondrial sulfide oxidation, a new pathomechanism associated with this syndrome. EMBO Mol. Med. 9, 78–95 (2017).
Lash, L. H. Mitochondrial glutathione transport: physiological, pathological and toxicological implications. Chemico-Biol. Interact. 163, 54–67 (2006).
Szabo, C. et al. Tumor-derived hydrogen sulfide, produced by cystathionine-beta-synthase, stimulates bioenergetics, cell proliferation, and angiogenesis in colon cancer. Proc. Natl Acad. Sci. USA 110, 12474–12479 (2013).
Rigottier-Gois, L. Dysbiosis in inflammatory bowel diseases: the oxygen hypothesis. ISME J. 7, 1256–1261 (2013).
Calvo, S. E., Clauser, K. R. & Mootha, V. K. MitoCarta2.0: an updated inventory of mammalian mitochondrial proteins. Nucleic Acids Res. 44, 1251–1257 (2016).
Szendroedi, J., Phielix, E. & Roden, M. The role of mitochondria in insulin resistance and type 2 diabetes mellitus. Nat. Rev. Endocrinol. 8, 92–103 (2011).
Bratic, A. & Larsson, N. G. The role of mitochondria in aging. J. Clin. Investig. 123, 951–957 (2013).
Weinberg, S. E. & Chandel, N. S. Targeting mitochondria metabolism for cancer therapy. Nat. Chem. Biol. 11, 9–15 (2015).
Chen, W. W., Freinkman, E., Wang, T., Birsoy, K. & Sabatini, D. M. Absolute quantification of matrix metabolites reveals the dynamics of mitochondrial metabolism. Cell 166, 1324–1337 (2016).
Fatehullah, A., Tan, S. H. & Barker, N. Organoids as an in vitro model of human development and disease. Nat. Cell Biol. 18, 246–254 (2016).
Debnath, J., Muthuswamy, S. K. & Brugge, J. S. Morphogenesis and oncogenesis of MCF-10A mammary epithelial acini grown in three-dimensional basement membrane cultures. Methods 30, 256–268 (2003).
Cantor, J. R. et al. Physiologic medium rewires cellular metabolism and reveals uric acid as an endogenous inhibitor of UMP synthase. Cell 169, 258–272 (2017).
Acknowledgements
We would like to thank Sarah Tucker and Liam Kelley for editing this article. J.B.S is supported by the National Science Foundation Graduate Research Fellowship DGE1144152. M.C.H. is supported by the Ludwig Center at Harvard and NIH grant R01CA213062.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Spinelli, J.B., Haigis, M.C. The multifaceted contributions of mitochondria to cellular metabolism. Nat Cell Biol 20, 745–754 (2018). https://doi.org/10.1038/s41556-018-0124-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-018-0124-1
This article is cited by
-
Glutaminolysis regulates endometrial fibrosis in intrauterine adhesion via modulating mitochondrial function
Biological Research (2024)
-
Micropeptides: potential treatment strategies for cancer
Cancer Cell International (2024)
-
Focusing on mitochondria in the brain: from biology to therapeutics
Translational Neurodegeneration (2024)
-
Spike structure of gold nanobranches induces hepatotoxicity in mouse hepatocyte organoid models
Journal of Nanobiotechnology (2024)
-
SIRT5-mediated ME2 desuccinylation promotes cancer growth by enhancing mitochondrial respiration
Cell Death & Differentiation (2024)