Abstract
How microtubules (MTs) are generated in the cell is a major question in understanding how the cytoskeleton is assembled. For several decades, γ-tubulin has been accepted as the universal MT nucleator of the cell. Although there is evidence that γ-tubulin complexes are not the sole MT nucleators, identification of other nucleation factors has proven difficult. Here, we report that the well-characterized MT polymerase XMAP215 (chTOG/Msps/Stu2p/Alp14/Dis1 homologue) is essential for MT nucleation in Xenopus egg extracts. The concentration of XMAP215 determines the extent of MT nucleation. Even though XMAP215 and the γ-tubulin ring complex (γ-TuRC) possess minimal nucleation activity individually, together, these factors synergistically stimulate MT nucleation in vitro. The amino-terminal TOG domains 1–5 of XMAP215 bind to αβ-tubulin and promote MT polymerization, whereas the conserved carboxy terminus is required for efficient MT nucleation and directly binds to γ-tubulin. In summary, XMAP215 and γ-TuRC together function as the principal nucleation module that generates MTs in cells.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Voter, W. A. & Erickson, H. P. The kinetics of microtubule assembly. Evidence for a two-stage nucleation mechanism. J. Biol. Chem. 259, 10430–10438 (1984).
Petry, S. & Vale, R. Microtubule nucleation at the centrosome and beyond. Nat. Cell Biol. 17, 1089–1093 (2015).
Luders, J. & Stearns, T. Microtubule-organizing centres: a re-evaluation. Nat. Rev. Mol. Cell Biol. 8, 161–167 (2007).
Wiese, C. & Zheng, Y. Microtubule nucleation: γ-tubulin and beyond. J. Cell Sci. 119, 4143–4153 (2006).
Zheng, Y., Wong, M. L., Alberts, B. & Mitchison, T. Nucleation of microtubule assembly by a γ-tubulin-containing ring complex. Nature 378, 578–583 (1995).
Moritz, M., Braunfeld, M. B., Sedat, J. W., Alberts, B. & Agard, D. A. Microtubule nucleation by γ-tubulin-containing rings in the centrosome. Nature 378, 638–640 (1995).
Kollman, J. M., Merdes, A., Mourey, L. & Agard, D. A. Microtubule nucleation by γ-tubulin complexes. Nat. Rev. Mol. Cell Biol. 12, 709–721 (2011).
Moritz, M., Braunfeld, M. B., Guénebaut, V., Heuser, J. & Agard, D. A. Structure of the γ-tubulin ring complex: a template for microtubule nucleation. Nat. Cell Biol. 2, 365–370 (2000).
Kollman, J. M., Polka, J. K., Zelter, A., Davis, T. N. & Agard, D. A. Microtubule nucleating γ-TuSC assembles structures with 13-fold microtubule-like symmetry. Nature 466, 879–882 (2010).
Caudron, N. Microtubule nucleation from stable tubulin oligomers. J. Biol. Chem. 277, 50973–50979 (2002).
Groen, A. C., Maresca, T. J., Gatlin, J. C., Salmon, E. D. & Mitchison, T. J. Functional overlap of microtubule assembly factors in chromatin-promoted spindle assembly. Mol. Biol. Cell 20, 2766–2773 (2009).
Hannak, E. et al. The kinetically dominant assembly pathway for centrosomal asters in Caenorhabditis elegans is γ-tubulin dependent. J. Cell Biol. 157, 591–602 (2002).
Rogers, G. C., Rusan, N., Peifer, M. & Rogers, S. L. A multicomponent assembly pathway contributes to the formation of acentrosomal microtubule arrays in interphase Drosophila cells. Mol. Biol. Cell 19, 3163–3178 (2008).
Moritz, M., Zheng, Y., Alberts, B. M. & Oegema, K. Recruitment of the γ-tubulin ring complex to Drosophila salt-stripped centrosome scaffolds. J. Cell Biol. 142, 775–786 (1998).
Choi, Y., Liu, P., Sze, S., Dai, C. & Qi, R. CDK5RAP2 stimulates microtubule nucleation by the γ-tubulin ring complex. J. Cell Biol. 191, 1089–1095 (2010).
Liu, P., Choi, Y. & Qi, R. NME7 is a functional component of the γ-tubulin ring complex. Mol. Biol. Cell 25, 2017–2025 (2014).
Oegema, K. et al. Characterization of two related Drosophila γ-tubulin complexes that differ in their ability to nucleate microtubules. J. Cell Biol. 144, 721–733 (1999).
Kollman, J. M. et al. Ring closure activates yeast γTuRC for species-specific microtubule nucleation. Nat. Struct. Mol. Biol. 22, 132–137 (2015).
Gard, D. L. & Kirschner, M. W. A microtubule-associated protein from Xenopus eggs that specifically promotes assembly at the plus-end. J. Cell Biol. 105, 2203–2215 (1987).
Tournebize, R. et al. Control of microtubule dynamics by the antagonistic activities of XMAP215 and XKCM1 in Xenopus egg extracts. Nat. Cell Biol. 2, 13–19 (2000).
Popov, A. V. et al. XMAP215 regulates microtubule dynamics through two distinct domains. EMBO J. 20, 397–410 (2001).
Al-Bassam, J., Van Breugel, M., Harrison, S. C. & Hyman, A. Stu2p binds tubulin and undergoes an open-to-closed conformational change. J. Cell Biol. 172, 1009–1022 (2006).
Brouhard, G. J. et al. XMAP215 is a processive microtubule polymerase. Cell 132, 79–88 (2008).
Widlund, P. O. et al. XMAP215 polymerase activity is built by combining multiple tubulin-binding TOG domains and a basic lattice-binding region. Proc. Natl Acad. Sci. USA 108, 2741–2746 (2011).
Al-Bassam, J. et al. Fission yeast Alp14 is a dose-dependent plus end-tracking microtubule polymerase. Mol. Biol. Cell 23, 2878–2890 (2012).
Reber, S. B. et al. XMAP215 activity sets spindle length by controlling the total mass of spindle microtubules. Nat. Cell Biol. 15, 1116–1122 (2013).
Roostalu, J., Cade, N. & Surrey, T. Complementary activities of TPX2 and chTOG constitute an efficient importin-regulated microtubule nucleation module. Nat. Cell Biol. 17, 1422–1434 (2015).
Wilde, A. & Zheng, Y. Stimulation of microtubule aster formation and spindle assembly by the small GTPase Ran. Science 284, 1359–1362 (1999).
Slep, K. & Vale, R. Structural basis of microtubule plus end tracking by XMAP215, CLIP-170, and EB1. Mol. Cell 27, 976–991 (2007).
Ayaz, P., Ye, X., Huddleston, P., Brautigam, C. A. & Rice, L. M. A TOG:αβ-tubulin complex structure reveals conformation-based mechanisms for a microtubule polymerase. Science 337, 857–860 (2012).
Byrnes, A. E. & Slep, K. C. TOG-tubulin binding specificity promotes microtubule dynamics and mitotic spindle formation. J. Cell Biol. 216, 1641–1657 (2017).
Popov, A. V., Severin, F. & Karsenti, E. XMAP215 is required for the microtubule-nucleating activity of centrosomes. Curr. Biol. 12, 1326–1330 (2002).
Wieczorek, M., Bechstedt, S., Chaaban, S. & Brouhard, G. J. Microtubule-associated proteins control the kinetics of microtubule nucleation. Nat. Cell Biol. 17, 907–916 (2015).
Ohi, R. & Zanic, M. Ahead of the curve: new insights into microtubule dynamics. F1000Res. 5, 314 (2016).
Roostalu, J. & Surrey, T. Microtubule nucleation: beyond the template. Nat. Rev. Mol. Cell Biol. 18, 702–710 (2017).
Petry, S., Groen, A., Ishihara, K., Mitchison, T. & Vale, R. Branching microtubule nucleation in Xenopus egg extracts mediated by augmin and TPX2. Cell 152, 768–777 (2013).
Zanic, M., Widlund, P. O., Hyman, A. A. & Howard, J. Synergy between XMAP215 and EB1 increases microtubule growth rates to physiological levels. Nat. Cell Biol. 15, 688–693 (2013).
Bucciarelli, E. et al. Drosophila Dgt6 interacts with Ndc80, Msps/XMAP215, and γ-tubulin to promote kinetochore-driven MT formation. Curr. Biol. 19, 1839–1845 (2009).
Moritz, M., Rice, L. M. & Agard, D. A. Microtubule Nucleation (Wiley-VCH, Weinheim, 2004).
Wuhr, M. et al. Deep proteomics of the Xenopus laevis egg using an mRNA-derived reference database. Curr. Biol. 24, 1467–1475 (2014).
Oakley, C. E. & Oakley, B. R. Identification of γ-tubulin, a new member of the tubulin superfamily encoded by mipA gene of Aspergillus nidulans. Nature 338, 662–664 (1989).
Srayko, M., Kaya, A., Stamford, J. & Hyman, A. A. Identification and characterization of factors required for microtubule growth and nucleation in the early C. elegans embryo. Dev. Cell 9, 223–236 (2005).
Andersen, J. S. et al. Proteomic characterization of the human centrosome by protein correlation profiling. Nature 426, 570–574 (2003).
Gergely, F., Draviam, V. M. & Raff, J. W. The ch-TOG/XMAP215 protein is essential for spindle pole organization in human somatic cells. Genes Dev. 17, 336–341 (2003).
Hood, F. E. et al. Coordination of adjacent domains mediates TACC3-ch-TOG-clathrin assembly and mitotic spindle binding. J. Cell Biol. 202, 463–478 (2013).
Burgess, S. G., Bayliss, R. & Pfuhl, M. Solution NMR assignment of the cryptic sixth TOG domain of mini spindles. Biomol. NMR Assign. 9, 411–413 (2015).
Fox, J., Howard, A., Currie, J., Rogers, S. & Slep, K. The XMAP215 family drives microtubule polymerization using a structurally diverse TOG array. Mol. Biol. Cell 25, 2375–2392 (2014).
Lee, M. J., Gergely, F., Jeffers, K., Peak-Chew, S. Y. & Raff, J. W. Msps/XMAP215 interacts with the centrosomal protein D-TACC to regulate microtubule behaviour. Nat. Cell Biol. 3, 643–649 (2001).
Vasquez, R. J., Gard, D. L. & Cassimeris, L. Phosphorylation by CDK1 regulates XMAP215 function in vitro. Cell Motil. Cytoskeleton 43, 310–321 (1999).
Tan, S., Kern, R. C. & Selleck, W. The pST44 polycistronic expression system for producing protein complexes in Escherichia coli. Protein Expr. Purif. 40, 385–395 (2005).
Petry, S., Pugieux, C., Nedelec, F. & Vale, R. Augmin promotes meiotic spindle formation and bipolarity in Xenopus egg extracts. Proc. Natl Acad. Sci. USA 108, 14473–14478 (2011).
Alfaro-Aco, R., Thawani, A. & Petry, S. Structural analysis of the role of TPX2 in branching microtubule nucleation. J. Cell Biol. 216, 983–997 (2017).
Hannak, E. & Heald, R. Investigating mitotic spindle assembly and function in vitro using Xenopus laevis egg extracts. Nat. Protoc. 1, 2305–2314 (2006).
Murray, A. W. & Kirschner, M. W. Cyclin synthesis drives the early embryonic cell cycle. Nature 339, 275–280 (1989).
King, M. & Petry, S. in The Mitotic Spindle: Methods and Protocols Vol. 1413 (eds Chang, P. & Ohi, R.) 77–85 (Humana, New York, 2016).
Applegate, K. T. et al. plusTipTracker: quantitative image analysis software for the measurement of microtubule dynamics. J. Struct. Biol. 176, 168–184 (2011).
Jaqaman, K. et al. Robust single-particle tracking in live-cell time-lapse sequences. Nat. Methods 5, 695–702 (2008).
Zheng, Y., Wong, M. L., Alberts, B. & Mitchison, T. Purification and assay of gamma tubulin ring complex. Methods Enzymol. 298, 218–228 (1998).
Wiese, C. & Zheng, Y. A new function for the γ-tubulin ring complex as a microtubule minus-end cap. Nat. Cell Biol. 2, 358–364 (2000).
Choi, Y. & Qi, R. Z. Assaying microtubule nucleation by the γ-tubulin ring complex. Methods Enzymol. 540, 119–130 (2014).
Gell, C. et al. Microtubule dynamics reconstituted in vitro and imaged by single-molecule fluorescence microscopy. Methods Cell Biol. 95, 221–245 (2010).
Zanic, M. Measuring the effects of microtubule-associated proteins on microtubule dynamics in vitro. Methods Mol. Biol. 1413, 47–61 (2016).
Aitken, C. E., Marshall, R. A. & Puglisi, J. D. An oxygen scavenging system for improvement of dye stability in single-molecule fluorescence experiments. Biophys. J. 94, 1826–1835 (2008).
Le Maire, M., Aggerbeck, L. P., Monteilhet, C., Andersen, J. P. & Moller, J. V. The use of high-performance liquid chromatography for the determination of size and molecular weight of proteins: a caution and a list of membrane proteins suitable as standards. Anal. Biochem. 154, 525–535 (1986).
Acknowledgements
We thank J. Shaevitz for help with image analysis, H. A. Stone for insightful discussions, M. Moritz for providing the γ-tubulin protein, S. Reber and P. Widlund for providing plasmids of XMAP215, M. Moritz, F. Hughson and S. Reber for critical reading of the manuscript and Petry laboratory members for discussions. Proteomics experiments were carried out by the Mass Spectrometry Core at Princeton University. A.T. thanks the Image Analysis course at MBL for useful training. This work was supported by the NIH New Innovator Award, Pew Scholars Program in the Biomedical Sciences, David and Lucile Packard Foundation (all to S.P.), American Heart Association predoctoral fellowship 17PRE33660328 (to A.T.) and the NIH post-doctoral fellowship 1F32GM119195-01 (to R.S.K.).
Author information
Authors and Affiliations
Contributions
A.T. and S.P. conceived the project. A.T., R.S.K. and S.P. designed the experiments. A.T. and R.S.K. generated and characterized the reagents and tools. A.T. and R.S.K. performed and analysed all the experiments. A.T., R.S.K. and S.P. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 XMAP215 stimulates microtubule nucleation in Xenopus egg extracts, Related to Fig. 1.
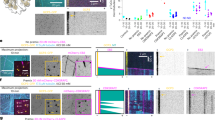
(A) Addition of excess wild-type XMAP215-GFP to Xenopus egg extracts in the presence of 5.5 μM RanQ69L. Representative images are displayed at 300 seconds of the reaction. Scale bar, 10 μm. See Supplementary Video 1. The experiment was repeated three times with independent extract preparations, with more than four additional supporting experiments. (B) EB1 comets in the entire field of view were counted as readout of microtubule (MT) nucleation. The number of MTs was plotted over time. The analysis was performed thrice with independent extract preparations. (C) Growth speed of MTs was obtained by tracking the EB1 comets over the image sequence. The number of growth speed measurements obtained from consecutive frames of all tracks (n): buffer (n = 17,798), 120 nM (n = 62,633), 240 nM (n = 75,532), 480 nM (n = 109,138). Mean growth speed (diamonds) is almost constant at 10 μm/min. Error bars display s.d. in growth speed across all measurements. The analysis was performed twice on experiments performed with independent extract preparations. (D) EB1 comets in the field of view were detected, counted and plotted over time for reactions from Fig. 1B. The nucleation measurements were repeated more than four times for experiments performed with independent extract preparations. Wild-type XMAP215-GFP was added back at 85 nM (equivalent to 120 nM in IgG-depleted extracts). MT lifetime (mean ± s.d.) was estimated to be 35 ± 15 seconds (IgG depletion), 43 ± 41 seconds (XMAP215 depletion + wt XMAP215). Lifetime of MTs was defined as the time from appearance of EB1 comet on the plus end (nucleation) to first disappearance of the EB1 comet from the plus end (catastrophe or pause) for n = 7 MTs analyzed. MT lifetime measurements were performed manually on one representative experiment displayed in Fig. 1B. (E) Related to Fig. 1A-B. Western blot analysis of α-tubulin, TACC3, TPX2, augmin subunits (HAUS1 and HAUS6), γ-TuRC subunits (γ-tubulin and GCP4) in XMAP215 immunodepletion, confirming that none of these proteins are co-depleted with XMAP215. The experiment was repeated identically at least twice with independent extracts and immunodepletions, and subsets of all proteins displayed were immunoblotted in more than three additional independent experiments. See Supplementary Fig. 9 for unprocessed scans.
Supplementary Figure 2 Microtubule polymerization in Xenopus egg extracts occurs with low XMAP215 concentration, Related to Fig. 2.
(A) De novo nucleation events were observed with increasing XMAP215 concentration (Fig. 2a, b). All EB1 comets in the field of view were tracked during the entire reaction, and growth speed of MTs was obtained from consecutive frames of all tracks. No MTs were observed when buffer (0 nM), 15 nM or 30 nM XMAP215 was added back; therefore, growth speed could not be measured. Mean growth speed for 60 nM and above XMAP215 concentrations is plotted. Error bars represent s.d. in all measurements. The number of growth speed measurements obtained from consecutive frames of all tracks (n). 60 nM (n = 87), 120 nM (n = 734), 240 nM (n = 1,889), 480 nM (n = 19,379), 720 nM (n = 43,995). Mean speed versus XMAP215 concentration was fitted to Michaelis-Menten kinetics (dashed curve). See Supplementary Video 3. The analysis was repeated twice with independent extract preparations. (B-C) Stabilized GMPCPP MTs (blue) were digested with subtilisin A and attached to passivated coverslips. Growth from MT seeds (red – tubulin, green – EB1-mCherry) was observed in XMAP215-depleted extracts supplemented with low XMAP215 concentration. Distinct MT growth events were observed starting from 7.5 nM XMAP215, while EB1 blinking events were seen at one end of seeds without any XMAP215 added. Scale bar, 10μm. Representative kymographs of these events are displayed in (C). This experiment was repeated thrice with independent extract preparations. (D) Growth speed was measured from kymograph analysis. All measurements versus XMAP215 concentration and fitted to Michaelis-Menten kinetics (red curve), and s.d. displayed in shaded red area. The number of kymographs analyzed (n): buffer (n = 10), 7.5 nM (n = 11), 15 nM (n = 16), 30 nM (n = 18), 60 nM (n = 17). The analysis was repeated on two independent experimental sets. (E) Centrally illuminated [104 × 104 μm] region was selected. All MT seeds were observed for a short period of 35–80 seconds. The number of MT seeds analyzed (n): buffer (n = 47), 7.5 nM (n = 100), additional reaction at 7.5 nM (n = 116), 15 nM (n = 31), 30 nM (n = 85), 60 nM (n = 65). Fraction of seeds that displayed MT growth on one end, based on composite tubulin and EB1 signal, was reported (blue diamonds). For buffer (0 nM XMAP215) addition, fraction of seeds displaying distinct EB1 blinking events on one end was reported (red square). The analysis was repeated on two independent experimental sets and one supporting set. See Supplementary Table 2 for source data.
Supplementary Figure 3 γ-TuRC is necessary for microtubule nucleation with XMAP215, Related to Fig. 3.
(A) Western blot analysis of γ-tubulin and GCP4 (core γ-TuRC subunit) in γ-tubulin depletion. XMAP215, augmin (HAUS6), TPX2 and TACC3 were verified to be present in γ-tubulin depleted extracts. The experiment was repeated with three independent extract preparations, including verification with a different γ-tubulin antibody (XenC) for γ-TuRC immunodepletion. See Supplementary Fig. 9 for unprocessed scans. (B-C) Total number of MTs and fraction of MTs generated via branching MT nucleation in control and γ-tubulin depleted extracts was plotted with time for increasing concentration of excess XMAP215. Few MTs that emerged in γ-tubulin depleted extracts were counted manually (MTs generated by both de novo nucleation and branching nucleation). For IgG depletion, EB1 comets were counted as a readout for total number of MTs in the field of view (see Methods), while MTs nucleated de novo were counted manually. The number of branched MTs was calculated by subtracting MTs nucleated de novo from total number of MTs. The fraction of branched MTs for IgG depletion was then calculated as the ratio of MTs generated by branching to total EB1 comets. See Fig. 3, and Supplementary Video 5. The analyses were repeated twice on experiments performed with independent extract preparations. (D) De novo nucleation events observed by adding 360 nM XMAP215-GFP in excess of the endogenous protein to control and γ-tubulin depleted extracts without RanQ69L. Representative images are displayed at 16 minutes of the reaction. Scale bar, 10 μm. The experiment was repeated three times with independent extract preparations. (E) EB1 comets were detected and tracked using the image analysis procedure described in Methods. The number of MT tracks was plotted over time. MT-independent nucleation was severely reduced in γ-tubulin depletion, in comparison to the control depletion, even after the addition of excess XMAP215. The analysis was repeated thrice on experiments performed with independent extract preparations.
Supplementary Figure 4 Characterization of purified γ-TuRC, Related to Fig. 4.
(A) Isolation of γ-TuRC from Xenopus egg extracts. Mass spectrometric analysis of prominent bands in the elution showed all γ-TuRC components: γ-tubulin, GCPs 2–6, Mzt2b and NEDD1. Some contaminating factors: PCM1, Clathrin heavy chain, SPAG5 and small kinetochore associated protein (KNSTRN) were still present. Representative image is displayed, and mass spectrometry was repeated twice with similar results. (B) Input, flow-through, and elution from the affinity purification were analyzed by immunoblot for XMAP215, TACC3, α-tubulin and TPX2. These components were not found in the elution, while γ-TuRC specific factors (γ-tubulin, GCP4, NEDD1) were present in purified γ-TuRC. These findings were consistent when repeated with two independent γ-TuRC preparations. (C) Purified γ-TuRC and XMAP215 were incubated to promote MT nucleation. For 5 nM XMAP215 experiments, representative images are shown in Fig. 4c. Additional experiments were performed with 30 nM XMAP215 and lower γ-TuRC concentrations. All individual MTs in the field of view were counted in these conditions. 6 to 10-fold increase in nucleation was observed with XMAP215 and γ-TuRC combined, as compared to the sum of MTs generated by either component. Data from two (set 1) and three (set 2) independent γ-TuRC preparations was pooled. The number of fields of view analyzed (n): γ-TuRC (set 1)n = 20; 5 nM XMAP215 n = 20; γ-TuRC + 5 nM XMAP215 n = 20; γ-TuRC (set 2) n = 66; 30 nM XMAP215 n = 62; γ-TuRC + 30 nM XMAP215 n = 61. Mean ± s.d. of number of MTs was reported. (D) Fluorescence intensity measurements (displayed in Fig. 4d) is plotted for individual data points and color-coded by 3 experiments performed with independent γ-TuRC preparations. n = 10 randomly chosen fields of view assessed for every reaction and plotted individually. Means±s.d. of pooled data (n = 30 fields of view), overlaid in blue circles, is displayed in Fig. 4d. (E) MT nucleation was evaluated with γ-TuRC sample that was further purified using sucrose gradient fractionation after affinity step. Identical synergy with XMAP215 is seen with γ-TuRC purified using this scheme (compare to Fig. 4d). n = 10 fields of view were analyzed for each condition, except buffer (n = 9) and 130 nM XMAP215 (n = 11). Mean ± s.d. is plotted. The experiment was performed once supporting Fig. 4c-d. See Supplementary Fig. 9 for unprocessed scans, and Supplementary Table 2 for source data.
Supplementary Figure 5 Microtubule nucleation by XMAP215 protein constructs, Related to Fig. 5.
(A) Branching MT nucleation activity of TOG3-CT construct of XMAP215, added back to XMAP215-depleted extracts at 120 nM final concentration in the presence of 5.5 μM RanQ69L and 1 μM GST-TPX2 α3-7. Representative image is displayed at 560 seconds of the reaction. Scale bar, 10 μm. The experiment was repeated three times with independent extract preparations. See Supplementary Video 6. (B) EB1 comets in the entire field of view were counted and plotted with time for addition of individual N-terminal deletion constructs of XMAP215. The analyses were repeated twice on experiments performed with independent extract preparations. See Fig. 5a. (C-D) Branching MT nucleation activity of full-length XMAP215 1-5AA construct was compared with wild-type XMAP215. All proteins were added back to XMAP215-depleted extracts at 120 nM concentration in the presence of 5.5 μM RanQ69L and 1 μM GST-TPX2 α3-7. Full-length protein with mutant TOG domains did not support any MT nucleation. Representative images are displayed at 722.5 seconds of the reaction. The number of EB1 comets was quantified as in (B). The experiment was repeated thrice with independent extract preparations, and the analysis was repeated on two independent experimental sets.
Supplementary Figure 6 C-terminus of XMAP215 is required for microtubule nucleation with γ-TuRC, Related to Fig. 6.
(A) Related to Fig. 6a. In vitro MT polymerization assay performed with several XMAP215 constructs: wild-type XMAP215, TOG1-5, TOG12-Kloop and TOG5-CT. Representative kymographs for each construct are displayed showing 15 minutes of growth for a representative MT in the field of view. Scale bar, 5 μm. Each experiment was repeated at least twice with freshly purified proteins for TOG1-5, TOG5-CT and TOG12-Kloop proteins. No difference in growth speed was observed using either fresh or frozen wild-type XMAP215, and reactions were performed once each with freshly purified protein and frozen-thawed wild-type protein. (B) Related to Fig. 6b. Branching MT nucleation activity of XMAP215 constructs was observed in Xenopus egg extracts. All proteins were added back to XMAP215-depleted extracts at varying final concentration in the presence of 5.5 μM RanQ69L and 1 μM GST-TPX2 α3-7. Representative images for each construct are displayed at 540 seconds of the reaction. A representative plot displays the nucleation curves generated with varying concentrations of TOG1-5 protein and wild-type XMAP215 in one Xenopus egg extracts. Scale bar, 10 μm. Number of times the experiments were repeated with independent extract preparations: wild-type XMAP215 (8), TOG1-5 (4), TOG12-Kloop (3), TOG5-CT (2). See Supplementary Video 9. (C) Related to Fig. 6c, d. Purified γ-TuRC was incubated with wild-type XMAP215 or TOG1-5 to promote MT nucleation in vitro. Individual MTs observed in the field of view were counted. TOG1-5 shows much lower synergistic MT generation than wild-type XMAP215 with purified γ-TuRC, although at the highest concentration of TOG1-5 tested (120 nM), the number of MTs approaches that seen with 30 nM full-length XMAP215. Roughly 20 fields of view for each reaction condition were analyzed, and data from two independent experiments performed with fresh γ-TuRC preparations was pooled. The number of fields analysed (n): γ-TuRC (n = 62); XMAP215 30 nM (n = 43); γ-TuRC + XMAP215 30nM (n = 41); TOG1-5 30 nM (n = 20); TOG1-5 60 nM (n = 39); TOG1-5 120 nM (n = 44); γ-TuRC + TOG1-5 30 nM (n = 20); γ-TuRC + TOG1-5 60 nM (n = 41); γ-TuRC + TOG1-5 120 nM (n = 41). Mean ± s.d. is plotted. See Supplementary Table 2 for source data.
Supplementary Figure 7 XMAP215 interacts with γ-TuRC subunits in Xenopus egg extracts and with αβ-tubulin in vitro in a salt dependent manner, Related to Figs. 7 and 8a.
(A) Proteins interacting with XMAP215 were analyzed using immunoprecipitation from Xenopus egg extracts. Input lane represents 125-fold dilution of egg extracts. Immunoblots against the C-terminus of XMAP215, γ-TuRC subunits (γ-tubulin, GCP4 and GCP5), augmin subunit (HAUS6), TPX2 and α-tubulin are shown. The experiment was repeated at least four times with independent extract preparations. (B) Reciprocal immunoprecipitation was performed by immunoprecipitating γ-TuRC using anti- γ-tubulin and anti-Mozart1 antibody and the interacting proteins analyzed using immunoblot. Input lane represents 25-fold dilution of CSF extracts. Immunoblots against the C-terminus of XMAP215, TPX2 and γ-TuRC subunits (γ-tubulin, GCP4 and GCP5) are shown, and were imaged using the iBright imager. The experiment was repeated at least two times with independent extract preparations. (C) 4.3 μM XMAP215 and 4.4 μM tubulin were mixed with 0.2 mM GTP and applied to size exclusion chromatography to verify the interaction between αβ-tubulin and wild-type XMAP215. The interaction study was performed at 75 mM salt concentration (left) and 150 mM salt concentration (near physiological salt concentration, right). Eluted fractions were analyzed with SDS-PAGE and stained with Coomassie blue dye. The 16-bit grey-scale gel image is displayed, where the contrast was adjusted to visualize the αβ-tubulin band clearly. The chromatogram displaying the elution profile and eluted fractions together show that while XMAP215 binds to αβ-tubulin at low salt concentration, significant binding was not observed near physiological salt concentration. The chromatography runs were performed and compared once at the exact buffer conditions specified, with three supporting runs performed at slightly different conditions. See Supplementary Fig. 9 for unprocessed scans.
Supplementary Figure 8 XMAP215 interacts with γ-tubulin and αβ-tubulin with distinct termini, Related to Figs. 7 and 8a.
(A) Size exclusion chromatography of 140 nM human γ-tubulin was performed in buffer containing 400 mM KCl to prevent γ-tubulin oligomerization. In high salt, γ-tubulin elutes in fractions L-M (marked [**], compare to first run in Fig. 7). This experiment was performed once at the specified condition, with three supporting runs at slightly different concentration and buffer conditions. (B) Immunoprecipitation of γ-tubulin by XMAP215 in vitro. GFP or XMAP215-GFP at 500 nM each, or buffer were mixed with 80 nM human γ-tubulin and precipitated using anti-GFP antibodies. γ-tubulin content co-precipitated was analyzed via immunoblot. The experiment was repeated independently three times. (C) Size exclusion chromatography was performed with 140 nM γ-tubulin and 1.2 μM TOG1212 or 0.7 μM XMAP215 FL 1-5AA. 500 μl sample volume was injected into the column, and 300 μl fractions were collected. Alternate fractions eluted between 8.5 ml to 16.6 ml were analyzed via SDS-PAGE followed by immunoblot with γ-tubulin antibodies and GFP antibodies to detect the elution profile of γ-tubulin and GFP labeled XMAP215 proteins, respectively. Shift in γ-tubulin signal was observed. Each chromatography run was repeated at least twice on different days at the specified concentration, and two supporting runs for TOG1212 at slightly different protein concentrations were performed. (D) Wild-type XMAP215 (4.3 μM) and bovine αβ-tubulin (4.4 μM) were mixed and applied to size exclusion chromatography. 100 μl sample volume was injected into the column, 200 μl fractions were collected and alternate fractions eluted between 8.5 ml to 13.9 ml were analyzed via SDS-PAGE. Elution fractions for full-length XMAP215, N-terminal TOG1-4 construct or C-terminal TOG5-CT construct with or without bovine αβ-tubulin were analyzed. Stoke’s radii of reference proteins are marked at their peak elution: Thyroglobulin (8.6 nm) and Aldolase (4.6 nm). Each chromatography run was performed once at the specified conditions. Two supporting runs for full-length XMAP215 and one for TOG5-CT performed under slightly different concentrations and buffers. Void volume of the column was measured as 8.7-8.8 ml. See Figs. 7 and 8a and Supplementary Fig. 9.
Supplementary Figure 9
Uncropped western blots and gels.
Supplementary information
Supplementary Information
Supplementary Figures 1–9, Supplementary Table and Supplementary Video legends.
Supplementary Table 1
Information on antibodies.
Supplementary Table 2
Source data.
Supplementary Video 1
Description: Addition of XMAP215 increases microtubule nucleation events; related to Supplementary Fig. 1a.
Supplementary Video 2
XMAP215’s concentration determines branching microtubule nucleation stimulated in XMAP215-immunodepleted extracts; related to Fig. 1c–f.
Supplementary Video 3
De novo microtubule nucleation events occur proportionately to XMAP215 concentration; related to Fig. 2a,b.
Supplementary Video 4
XMAP215 promotes microtubule nucleation in TPX2 and augmin immunodepleted extracts; related to Fig. 2c–e.
Supplementary Video 5
Addition of XMAP215 to γ-tubulin immunodepleted extracts; related to Fig. 3.
Supplementary Video 6
Microtubule nucleation by N-terminal deletion constructs of XMAP215; related to Fig. 5a and Supplementary Fig. 5a,b.
Supplementary Video 7
Microtubule nucleation by C-terminal deletion constructs of XMAP215; related to Fig. 5b,c.
Supplementary Video 8
Tubulin-binding mutations in TOG domains disrupt microtubule nucleation by XMAP215; related to Fig. 5d,e.
Supplementary Video 9
C-terminus is required for efficient microtubule nucleation by XMAP215 in Xenopus egg extracts; related to Fig. 6b and Supplementary Fig. 6b.
Rights and permissions
About this article
Cite this article
Thawani, A., Kadzik, R.S. & Petry, S. XMAP215 is a microtubule nucleation factor that functions synergistically with the γ-tubulin ring complex. Nat Cell Biol 20, 575–585 (2018). https://doi.org/10.1038/s41556-018-0091-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-018-0091-6
This article is cited by
-
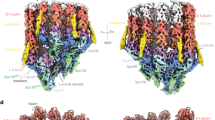
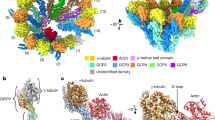
Structure of the γ-tubulin ring complex-capped microtubule
Nature Structural & Molecular Biology (2024)
-
CAMSAPs and nucleation-promoting factors control microtubule release from γ-TuRC
Nature Cell Biology (2024)
-
Structure of the native γ-tubulin ring complex capping spindle microtubules
Nature Structural & Molecular Biology (2024)
-
Microtubule nucleation and γTuRC centrosome localization in interphase cells require ch-TOG
Nature Communications (2023)
-
Mechanisms underlying spindle assembly and robustness
Nature Reviews Molecular Cell Biology (2023)